Nucleosomes impede Cas9 access to DNA in vivo and in vitro
Figures
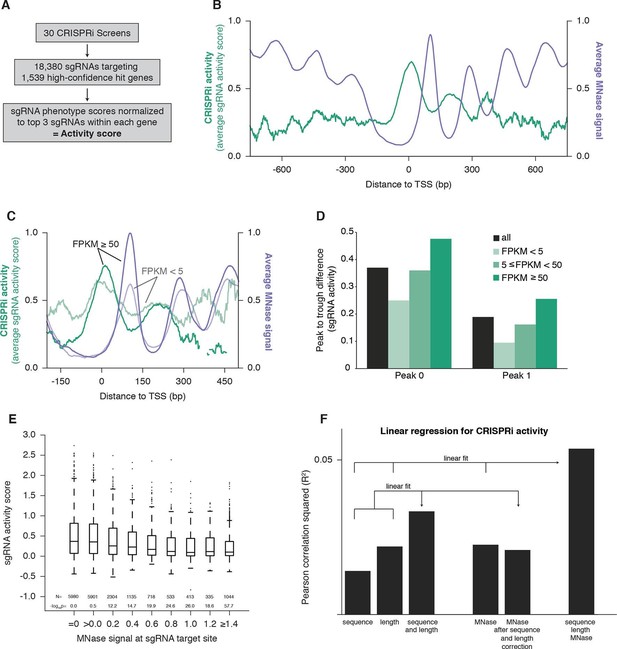
CRISPRi activity anti-correlates with nucleosome occupancy.
(A) Workflow for generating CRISPRi activity scores from pooled genetic screens. The resulting values are distributed around 0 for inactive sgRNAs and around 1 for highly active sgRNAs. (B) Average CRISPRi activity and MNase-seq signal relative to the TSS. Green line represents average CRISPRi activity score of all sgRNAs within a 50 bp window around each position. Purple line represents the K562 MNase-seq signal at each position averaged across all genes analyzed. (C) Average CRISPRi activity and MNase-seq signal for genes grouped by expression value. Genes were grouped into lower expression (light lines; N=240), higher expression (heavy lines; N=368), and medium expression (omitted for clarity; N=930), and analyzed as in (B). Expression values were obtained as fragments per kilobase million (FPKM) from ENCODE K562 RNA-seq data. Average activity at positions with fewer than 10 sgRNAs within the 50bp window was not calculated. (D) Quantification of the amplitude of periodic CRISPRi activity. Peak and trough coordinates were obtained by calculating the local maxima and minima of the activity traces from analyses in (B) and (C). Peak 0 was defined as the local maximum closest to the TSS, peak 1 was defined as the next maximum downstream of peak 0, and troughs were defined as the minima immediately downstream of the respective peaks. (E) CRISPRi activity and target site nucleosome occupancy for individual sgRNAs. Target site nucleosome occupancy was calculated from the average MNase-seq signal at all genomic coordinates across the length of the sgRNA protospacer and the protospacer adjacent motif (PAM). sgRNAs were then binned by the target site nucleosome occupancy, displayed as box-and-whisker plots, and labeled according to the minimum value within the bin except where indicated. P-values were calculated by a two-tailed Mann-Whitney test comparing each bin to the =0.0 bin. (F) Linear regression for CRISPRi activity. The squared Pearson correlation was calculated for the sgRNA activity scores compared to the indicated individual parameters (bars 1, 2, and 4) or linear fits of multiple parameters (bars 3 and 6). sgRNA activity scores were corrected for sequence and length features (bar 5) by subtracting the linear fit of those two features.
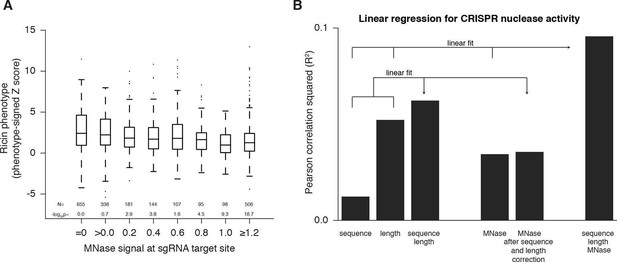
Cas9 nuclease activity anti-correlates with nucleosome occupancy.
(A) Cas9 nuclease phenotypes and target site nucleosome occupancy for individual sgRNAs targeting CDS regions. Ricin susceptibility phenotypes for each sgRNA are expressed as a Z score and are positive if the phenotype matches the expected knockdown phenotype. Target site nucleosome occupancy was calculated as in Figure 1E. sgRNAs were then binned by the target site nucleosome occupancy, displayed as box-and-whisker plots, and labeled according to the minimum value within the bin except where indicated. P-values were calculated by a two-tailed Mann-Whitney test comparing each bin to the =0.0 bin. (B) Linear regression for Cas9 nuclease phenotypes. The squared Pearson correlation was calculated for the sgRNA activity scores compared to the indicated individual parameters (bars 1, 2, and 4) or linear fits of multiple parameters (bars 3 and 6). sgRNA activity scores were corrected for sequence and length features (bar 5) by subtracting the linear fit of those two features.
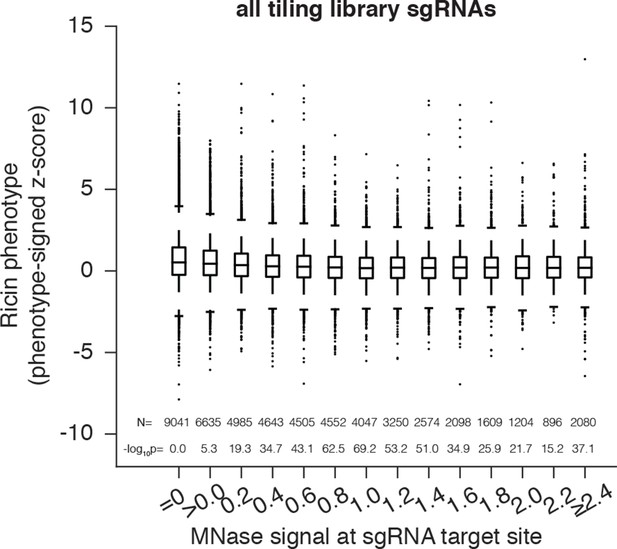
Cas9 nuclease activity anti-correlates with nucleosome occupancy at all target sites.
Cas9 nuclease phenotypes and target site nucleosome occupancy for individual sgRNAs. As in Figure 2A, but incorporating all CDS- and non-CDS-targeting sgRNAs in the ricin-susceptibility gene tiling library.
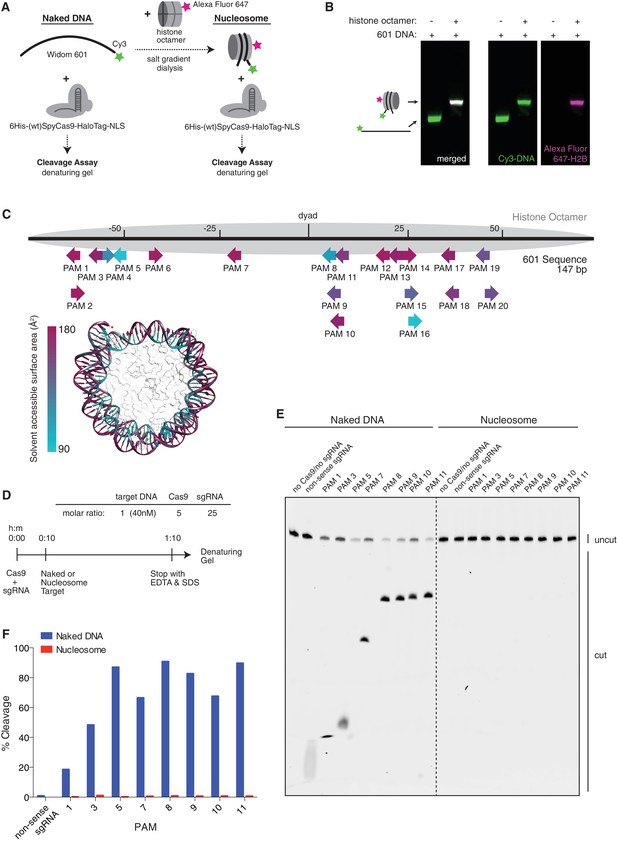
Cas9 nuclease activity is blocked by the presence of a nucleosome in vitro.
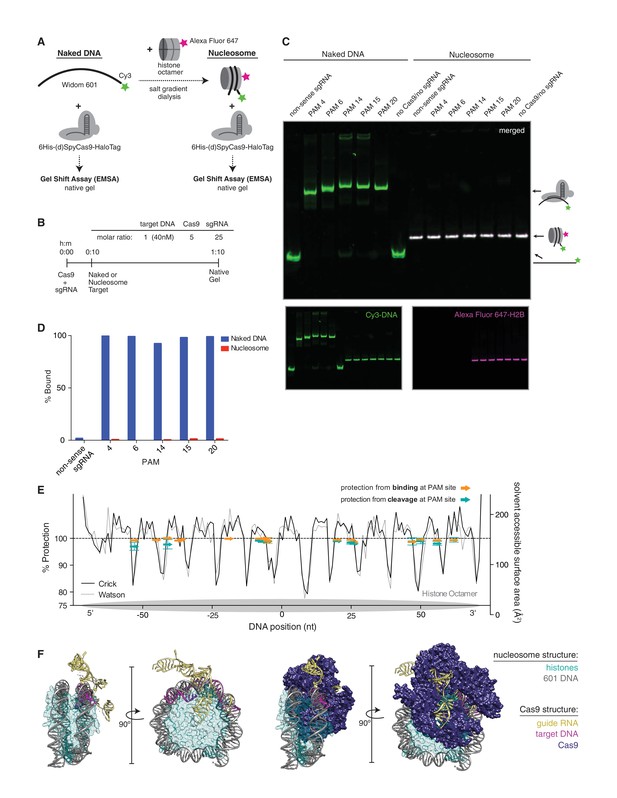
(A) Schematic of the experimental setup for in vitro cleavage assays. Mononucleosomes were assembled by salt gradient dialysis of purified mouse histone octamer with the minimal nucleosome positioning sequence, Widom 601 (147 bp). Prior to assembly, DNA was 5’-end-labeled with Cy3 and histone H2B was fluorescently labeled using an introduced cysteine (T115C) coupled to Alexa Fluor 647. Purified His-tagged and HaloTagged Cas9 pre-loaded with in vitro transcribed sgRNAs were added to naked DNA or assembled nucleosomes, and DNA cleavage products were visualized using a denaturing gel imaged for Cy3-DNA fluorescence. See also Figure 3—figure supplement 1A. (B) Confirmation that fully occupied, well-positioned nucleosomes were assembled. After assembly using salt gradient dialysis, the produced nucleosomes were visualized using a native PAGE gel imaged in the Cy3 and Alexa Fluor 647 channels. Full incorporation of the free DNA into a nucleosome occupying a single position on the DNA is indicated by the presence of a single-shifted band containing all of the detectable Cy3-DNA and Alexa Fluor 647-H2B signal. (C) Available PAMs and solvent accessibility of the 601 nucleosome positioning sequence. (Above) A schematic of the 601 sequence. The location of the histone octamer when assembled into a nucleosome is indicated by the gray oval. The location of PAMs within the double-stranded sequence are indicated with arrows spanning the 3 bp of the PAM, pointing in the 5’ to 3’ orientation of the NGG motif. The arrows are colored according to solvent accessibility at the center of the PAM as calculated from the crystal structure of the 601 nucleosome (PDBID 3LZ0, Vasudevan et al., 2010). (Below) Crystal structure of the 601 nucleosome. For clarity, the surface area of the histones in the crystal structure has been made transparent. The DNA in the crystal structure is colored according to solvent accessibility using the same color scale as the PAMs above. Residues colored teal are less accessible, while residues colored fuchsia are more accessible by solvent. (D) Experimental conditions and timeline for cleavage assays. (E) Denaturing PAGE gel showing the results of a cleavage assay targeting the indicated PAMs. Cleavage reactions containing naked DNA were loaded on the left half of the gel, while reactions containing nucleosomes were loaded on the right. The DNA was imaged via a Cy3 fluorophore attached to the 5’ end of the sgRNA-complimentary strand. A negative control was conducted with an sgRNA that had no sequence complementarity to the 601 sequence used (non-sense guide). See also Figure 3—figure supplement 1B,C for additional controls. (F) Quantification of the gel in (D). For each lane, percent cleavage was determined by calculating the percent of the total band signal corresponding to cleaved DNA.
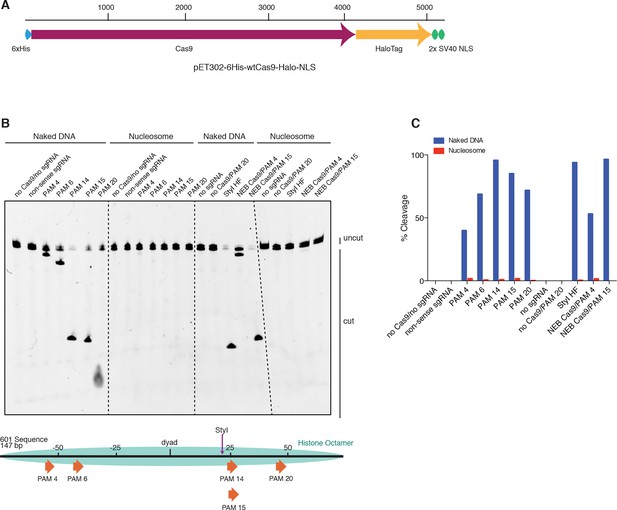
HaloTagged Cas9 activity is indistinguishable from untagged Cas9.
(A) Diagram of the CDS region in the Cas9 expression plasmid used in this study. (B) Cleavage assay comparing the HaloTagged Cas9 construct used in this study with an untagged Cas9 commercially purchased from New England Biolabs (NEB, Ipswich, MA). Both forms of Cas9 were incubated with either naked DNA or the same DNA assembled into a nucleosome (see Figure 3A). A positive control used the restriction enzyme, StyI-HF (from NEB), to target a sequence at a location within the DNA known to be fully protected upon assembly into a nucleosome. Unless explicitly labaled as NEB, all constructs of Cas9 (and dCas9) used were histidine tagged and HaloTagged.
dCas9 is unable to bind nucleosomal DNA in vitro.
(A) Schematic of the experimental setup for in vitro binding assays. Either naked DNA or assembled nucleosomes were incubated with catalytically dead Cas9 (dCas9, histidine tagged and HaloTagged), and binding was assessed by an electrophoretic mobility shift assay (EMSA). (B) Experimental conditions and timeline for binding assays. (C) A native PAGE gel showing the results of an EMSA in which dCas9 was targeted to the indicated PAMs on either naked or nucleosomal DNA. Gels were scanned for fluorescence from Cy3 on the DNA (green) and Alexa Fluor 647 on histone H2B (magenta). The two color channels were merged to identify the location of intact nucleosomes (white). See also Figure 4—figure supplement 1 and 3 for reagent preparation and experimental conditions, and Figure 4—figure supplement 2 for comparison with wtCas9 binding. (D) Quantification of the gel in (C). Percent bound was determined by calculating the percent of the total band signal in each lane corresponding to Cas9-bound target as determined by a shift in mobility within the gel. (E) Summary of binding and cleavage results for each PAM tested. The ability of Cas9 to bind or cleave nucleosomal DNA at a targeted PAM is displayed as percent protection by the nucleosome, and was calculated by taking the ratio of binding or cleavage on nucleosomal DNA to that on naked DNA. While only the largest error bars are visible, replicates were performed for 15 of the 30 data points and are displayed with error bars showing standard deviation from the mean. The DNA positions plotted correspond to the three nucleotides of the targeted PAM. In order to compare percent protection from binding and cleavage with solvent accessibility, the PAMs are overlaid with a plot of the solvent accessible surface area for each strand (Watson or Crick) of DNA in the 601 nucleosome structure. The percent protection at each PAM, as well as the solvent accessibility were plotted so that the 5’ end of each DNA strand begins at the left of the graph, where position 0 indicates the dyad. (F) Structural assessment of the ability of Cas9 to bind nucleosomal DNA. Superposition of the Cas9-guideRNA-DNA crystal structure (PDBID 4UN3, Anders et al., 2014) onto the 601 nucleosome crystal structure (PDBID 3LZ0, Vasudevan et al., 2010) was achieved by alignment of the DNA path in both structures. (Left) To better view the alignment of the Cas9 target DNA with the nucleosomal DNA, the Cas9 protein density has been removed. (Right) After alignment of the DNA, inclusion of the Cas9 protein density reveals extensive steric clashes with the histones. Histone surface area was made partially transparent to better reveal the overlapping densities with Cas9.
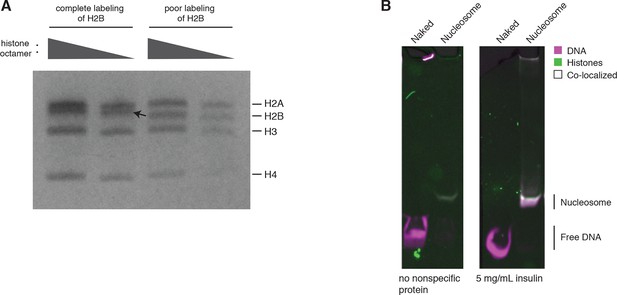
Quality controls.
(A) H2B labeling is near complete. An SDS-PAGE gel after refolding and purifying the histone octamer containing fully labeled versus poorly labeled H2B. The arrow indicates a mobility shift of H2B corresponding to a fully labeled band. Each histone is present at equimolar ratios as indicated by PageBlue protein stain. (B) Nucleosome stability in Cas9 binding and cleavage assays is ensured by including a nonspecific protein in solution. On the left, a native PAGE gel showing naked DNA and nucleosomes under Cas9 binding and cleavage reaction conditions without a nonspecific protein in solution. On the right, the same conditions plus inclusion of a nonspecific competitor protein, insulin.
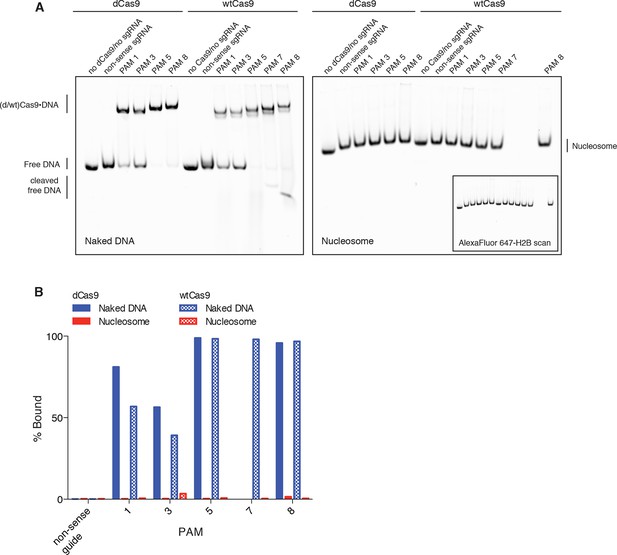
DNA binding by dCas9 is also representative of wtCas9 binding.
(A) Two native PAGE gels showing the results of an EMSA binding assay comparing dCas9 and wtCas9. Binding to naked DNA is shown in the gel on the left, while binding to nucleosomes is shown in the gel on the right. Both gels were imaged in the Cy3-DNA channel, while the gel on the right was also imaged in the Alexa Fluor 647 – H2B channel (inset image). Binding conditions for both dCas9 and wtCas9 were identical, and were as described in the methods. (B) Quantification of the Native PAGE gel in (A).
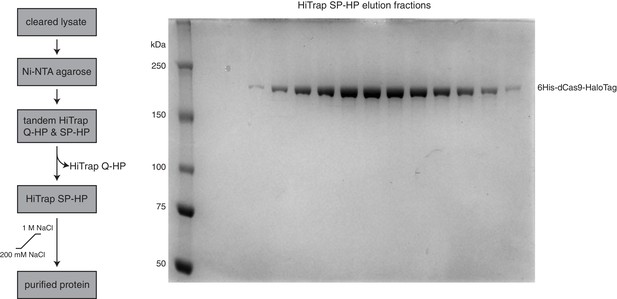
(wt/d)Cas9 purification strategy.
(A) Schematic showing the (wt/d)Cas9 purification strategy used in this study (left). SDS-PAGE gel stained with PageBlue (Life Technologies, Carlsbad, CA) showing the elution fractions from the HiTrap SP-HP column during purification of 6His-dCas9-HaloTag (right).
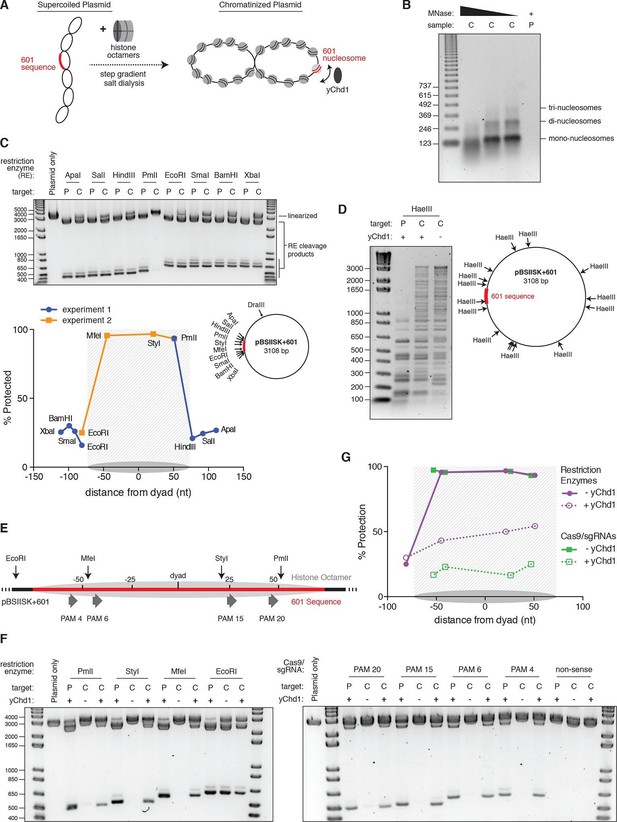
Nucleosomes within chromatinized DNA can block cleavage by Cas9, but a chromatin remodeling factor can restore Cas9 access.
(A) Schematic of the experimental setup. Supercoiled plasmid containing the 601 sequence inserted into a pBlueScript II SK (+) backbone (pBSIISK+601) was chromatinized by step gradient salt dialysis in the presence of histone octamer. Purified yeast Chd1 (yChd1) remodeling factor was used to test the effect of ATP-dependent remodeling factors on Cas9 access to nucleosomal DNA. (B) Quality assessment of the chromatinized plasmid used in this study. Titrated amounts of Micrococcal Nuclease (MNase) were incubated with the chromatinized plasmid, and the resulting pattern of protection by assembled nucleosomes was visualized on a 1.3% agarose gel post-stained with ethidium bromide (EtBr). As a control, the supercoiled plasmid was also incubated with the lowest concentration of MNase. (C) A restriction enzyme accessibility assay (REAA) was used to assess the occupancy and position of the nucleosome assembled at the 601 sequence within the chromatinized plasmid. A panel of unique restriction enzyme sites spanning the 601 sequence were incubated with either the supercoiled plasmid, or the chromatinized plasmid. Cleavage was stopped, and protein was removed by incubation with proteinase K followed by Phenol:Chloroform:Isoamyl alcohol extraction and ethanol precipitation. (Top) The resulting DNA was then linearized using DraIII, and the level of cleavage by the restriction enzyme panel was visualized on a 1% agarose gel post-stained with EtBr. The label 'P' represents supercoiled plasmid, while 'C' represents chromatinized plasmid. (Bottom right) The location of the restriction sites used are indicated on a diagram of the plasmid. (Bottom left) After quantification of the gel, the percent protection from cleavage experienced in the chromatinized plasmid was plotted versus the location of the cleavage sites on the top strand of the 601 sequence. Experiment 1 refers to the REAA experiment shown in the gel above, while experiment 2 refers to the REAA experiment without remodeler shown in Figure 5F. The grey shading indicates the borders of the 601 sequence, and the grey oval represents the corresponding nucleosome. (D) REAA experiment using the frequent cutter, HaeIII, to assess the remodeling activity around the chromatinized plasmid by the purified yChd1 chromatin remodeler. The resulting banding patterns were visualized on a 1.5% agarose gel post-stained with EtBr. Low molecular weight fragments indicate a high degree of HaeIII accessibility, while higher weight bands indicate protection from digestion. (E) Diagram showing the location of the restriction enzyme cleavage sites and the PAMs targeted by Cas9/sgRNA in the experiment shown in (F) and (G). (F) An accessibility assay was performed essentially as in C using either restriction enzymes or Cas9/sgRNAs in the presence or absence of the remodeler yChd1. The level of cleavage by the restriction enzyme panel (left) or Cas9/sgRNAs (right) was visualized on a 1.3% agarose gel post-stained with EtBr. A negative control was conducted with an sgRNA that had no sequence complementarity to the plasmid used (non-sense guide). The concentration of yChd1 used was the same as in panel (D) (G) Quantification of the gels shown in F. Percent protection from cleavage of the chromatinized plasmid in the presence or absence of the chromatin remodeler was calculated relative to the percent cleavage in the corresponding supercoiled plasmid control, and plotted at the location of the restriction enzyme cleavage sites or the center of the PAMs with respect to the 601 dyad.
Additional files
-
Supplementary file 1
Table S1.
CRISPRi sgRNA annotations, activity scores, and target site MNase signal, related to Figure 1.
- https://doi.org/10.7554/eLife.12677.013
-
Supplementary file 2
Table S2.
Ricin tiling library sgRNA annotations, phenotype scores, and target site MNase signal, related to Figure 2.
- https://doi.org/10.7554/eLife.12677.014





