Structural basis for germline antibody recognition of HIV-1 immunogens
Figures
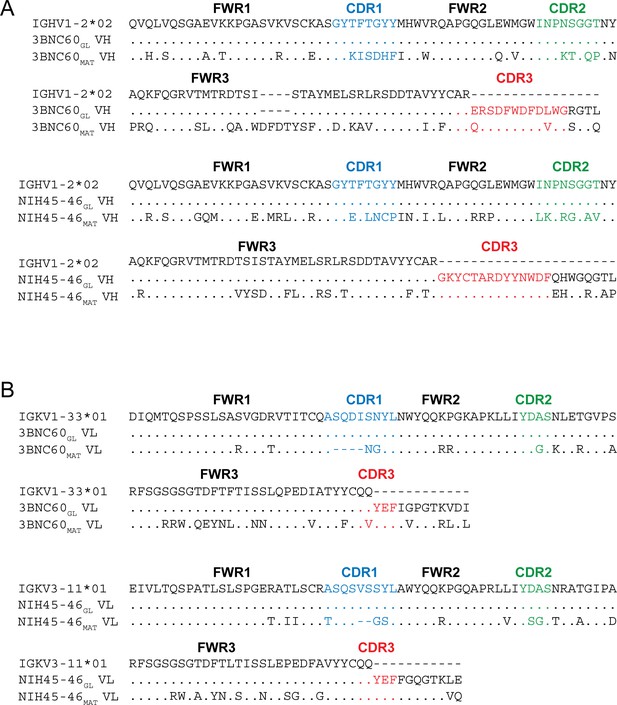
Sequence Alignments of Inferred Germline and Mature Forms of 3BNC60 and NIH45-46.
Alignments of (A) VH and (B) VL sequences of inferred germline progenitors (3BNC60GL and NIH45-46GL), mature 3BNC60 (3BNC60MAT), mature NIH45-46 (NIH45-46MAT), and the predicted germline V gene segments from which they were derived. Antibody framework regions (FWR) and CDR loops (CDR) are marked and CDR loops are colored blue (CDR1), green (CDR2), and red (CDR3). CDR3 sequences for germline Fabs were taken from mature antibody sequences as done in previous studies (Dosenovic et al., 2015; Hoot et al., 2013; McGuire et al., 2013; Scharf et al., 2013).
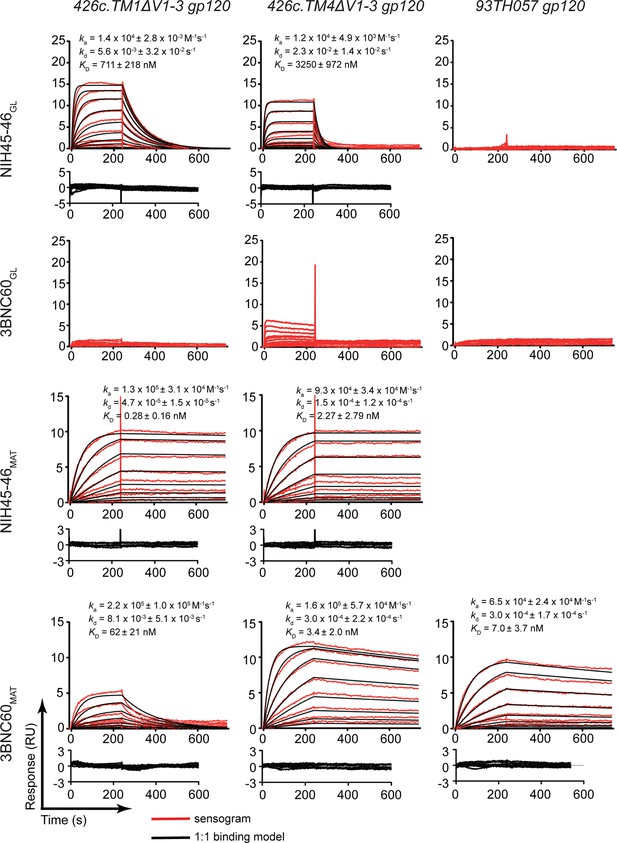
SPR binding assays.
Representative sensograms (red), fits (black, where applicable), residuals, and KD, ka, and kd values (mean ± standard deviation from 3 independent experiments) for binding of germline and mature Abs to gp120 cores. NIH45-46GL, 3BNC60GL, NIH45-46MAT and 3BNC60MAT IgG were captured on a protein A biosensor chip, and 426c.TM1△V1-3, 426c.TM4△V1-3 and 93TH057 gp120 cores were flowed over the chip as a 2-fold dilution series with top concentrations of 16 mM and 400 nM for germline and mature Abs, respectively.
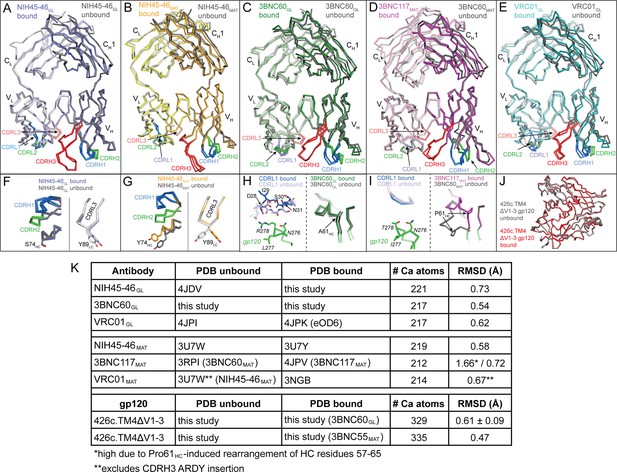
Overview of Bound and Unbound Structures of Germline and Mature Forms of NIH45-46 and 3BNC60.
Superposition of unbound (grey) and bound (colored) Fab structures of (A) NIH45-46GL (blue), (B) NIH45-46MAT (orange), (C) 3BNC60GL (green), (D) 3BNC60MAT (purple), and (E) VRC01GL (teal). The crystal structures were superimposed on their VHVL domains and are shown as wire representations with CDR loops colored blue (CDR1), green (CDR2), and red (CDR3). Panels (F–I) show detailed areas of interest for the corresponding structure comparisons shown in panels (A–D). Protein backbones are shown as wire diagrams, side chains are shown as stick representations (red, oxygen; blue, nitrogen). (J) Superposition of unbound (grey) and bound (red) structures of 426c.TM4△V1-3 shown as wire diagrams. (K) Table summarizing Cα rmsds for the indicated Ab and gp120 pairs. Since no unbound crystal structure of 3BNC117MAT was available, the structure of the close clonal relative, 3BNC60MAT (93%HC/96% LC sequence identity) was substituted. Similarly, no unbound structure of VRC01MAT was available, so that of NIH45-46MAT (88%HC/96% LC sequence identity) was substituted, omitting its four-residue insertion in CDRH3 from the rmsd calculation.
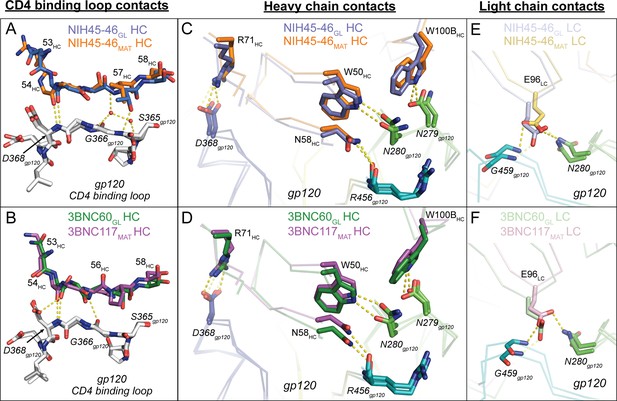
Comparison of Signature VRC01-class and CD4-mimickry Contacts in Germline Ab-gp120 Immunogen Complexes.
Top panels show (A) CD4 binding loop, (C) HC, (E) LC contacts in superimposed NIH45-46GL/426c.TM1△V1-3 and NIH45-46MAT/93TH057 (PDB 3U7Y) complexes. Bottom panels show (B) CD4-binding loop, (D) HC, (F) LC contacts in superimposed 3BNC60GL/426c.TM4△V1-3 and 3BNC117MAT/93TH057 (PDB 4JPV) complexes. Protein backbones are shown as wire diagrams, interacting residues are shown as stick representations (red, oxygen; blue, nitrogen). Yellow dashed lines indicate putative hydrogen bonds (distance < 3.5 Å, A-H–D angle > 90°). Ab coloring: blue, NIH45-46GL HC; light blue, NIH45-46GL LC; orange, NIH45-46MAT HC; yellow, NIH45-46MAT LC; green, 3BNC60GL HC; light green, 3BNC60GL LC; purple, 3BNC60MAT HC; light pink 3BNC60MAT LC. gp120 coloring: blue, CD4-binding loop; green, loop D; teal, loop V5. (E, F) Interacting residues of the C” strand of Fab HCs and the CD4-binding loop of gp120 (grey) are shown as sticks. The Fab residue numbers are indicated since the amino acid sequence differs at some positions between germline and mature Abs.
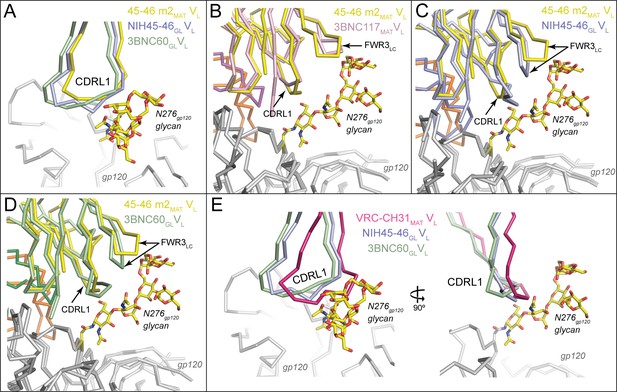
Accommodation of Asn276gp120 Glycan by Germline Ab Light Chains.
Superposition of gp120 (grey) complexes with germline and mature Abs. (A) CDRL1 from 45-46m2MAT (yellow), NIH45-46GL (blue) and 3BNC60GL (green) is positioned near the Asn276gp120 glycan. Two- and four-residue insertions in NIH45-46GL and 3BNC60GL, respectively, result in a widening of the tip of CDRL1 rather than a more extended loop, which would clash with gp120 protein residues and/or the Asn276gp120 glycan. gp120 (grey) complexes with (B) 3BNC117MAT (pink) and 45-46m2MAT (yellow) (C) NIH45-46GL (blue) and 45-46m2MAT (yellow), (D) 3BNC60GL (green) and 45–46 m2MAT (yellow). Protein backbones are shown as wire diagrams and the Asn276gp120 glycan from the 45-46m2MAT/93TH057 gp120 complex (PDB code 4JKP) is shown as sticks (yellow, carbon; red, oxygen; blue, nitrogen). The positions of CDRL1 and FWR3LC are indicated. (E) CDRL1 from VRC-CH31MAT (magenta), NIH45-46GL (blue) and 3BNC60GL (green) is positioned near the Asn276gp120 glycan. The CDRL1 loop of VRC-CH31MAT is of the same length as that of 3BNC60GL, and uses increased backbone conformational flexibility due to somatically mutated glycine residues to avoid clashes with gp120 protein residues and/or the Asn276gp120 glycan.
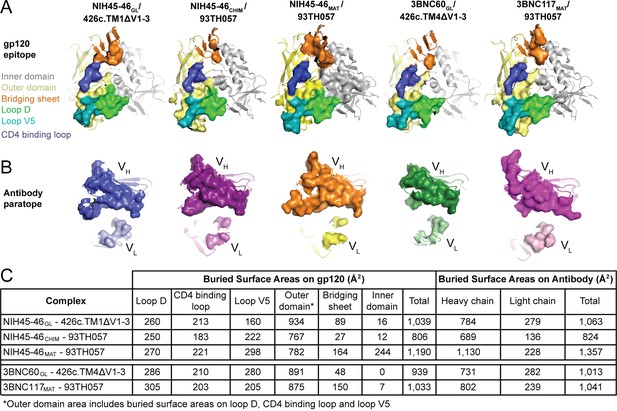
Comparison of the Binding Interfaces in gp120 Complexes with Germline and Mature Abs.
(A) gp120 residues contacted by Fabs and (B) Fab residues contacted by gp120s are shown as surfaces over ribbon diagrams. gp120 domains are colored as in Figure 3. Ab coloring: blue, NIH45-46GL HC; light blue, NIH45-46GL LC; purple, NIH45-46CHIM HC; light purple, NIH45-46CHIM LC; orange, NIH45-46MAT HC; yellow, NIH45-46MAT LC; green, 3BNC60GL HC; light green, 3BNC60GL LC; pink, 3BNC60MAT HC; light pink, 3BNC60MAT LC. (C) Quantitation of buried surface areas (Å2) depicted in (A) and (B). The columns labeled total are the sums of areas for outer domain, bridging sheet and inner domain for gp120, and of heavy chain and light chain for Abs. Surface areas buried due to complex formation were calculated using a 1.4 Å probe.
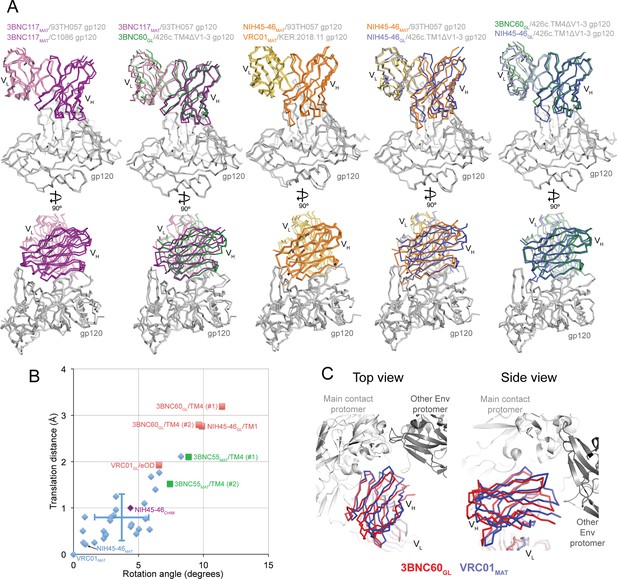
Comparisons of Binding Mode in Germline Ab-gp120 Immunogen Complexes.
(A) Superpositions of Fab-gp120 complexes depicted as wire diagrams. The following Fab/gp120 complexes were compared by alignment of their gp120s: 3BNC117MAT/93TH057 (PDB code 4JPV), 3BNC117MAT/C1086 (PDB code 4LSV), 3BNC117MAT/93TH057 (PDB code 4JPV), 3BNC60GL/426c.TM4△V1-3, NIH45-46MAT/93TH057 (PDB code 3U7Y), VRC01MAT/KER.2018.11 (PDB code 4LSS), NIH45-46GL/426c.TM1△V1-3. (B) Rotation angle and translation distance of VH domains of mature, chimeric and germline Fabs in complex with gp120s relative to VRC01MAT in complex with 93TH057 gp120 (PDB code 3NGB). Data points for complexes of mature Fabs bound to non-immunogen gp120s are shown as blue diamonds, complexes of germline Fabs bound to immunogen candidates are shown as red squares (TM1 = 426c.TM1△V1-3, TM4 = 426c.TM4△V1-3, eOD = eOD-GT6), the complex between the half mature, half germline NIH45-46CHIM and to a non-immunogen gp120 is shown as a purple diamond, and 3BNC55MAT bound to 426c.TM4△V1-3 is shown as a green square. When two complexes were found in the crystallographic asymmetric unit, rotation and translation parameters are shown for both complexes (denoted as #1 and #2). Standard deviations for the translation distance and rotation angle for mature VRC01-class bNAb–gp120 complexes shown as vertical and horizontal lines, respectively. (C) Alignment of the 3BNC60GL/426c.TM4△V1-3 (VHVL shown in red) and VRC01MAT Fab/gp120 (PDB code 3NGB) (VHVL shown in blue) structures onto the gp120 region of a native-like Env trimer structure (BG505 SOSIP.664; PDB code 5CEZ) (gray). Modeled structures are shown looking down the trimer three-fold axis (left panel) and from the side (right panel).
-
Figure 7—source data 1
Rotation angle and translation distance data of VH domains of mature, chimeric and germline Fabs in complex with gp120s relative to VRC01MAT in complex with 93TH057 gp120.
- https://doi.org/10.7554/eLife.13783.011
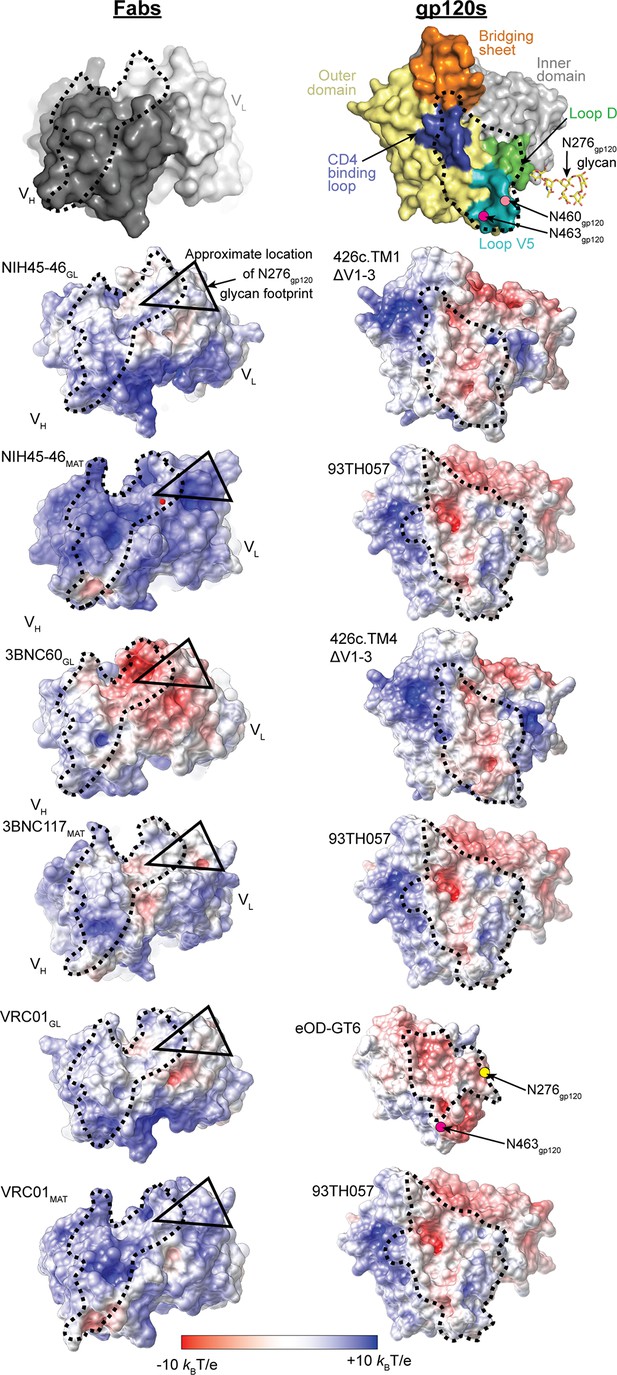
Comparison of Electrostatic Surface Characteristics of Fab-gp120 Complexes.
The binding surfaces of Fabs (left panels) and gp120s (right panels) are shown. Each binding partner is shown in an orientation looking into the binding interface; the corresponding complex would be obtained by rotating each binding partner by ~90˚ about the vertical axis. The top panel shows the locations of landmarks on surface representations of Fabs (dark grey, VH; light grey, VL) and gp120s (yellow, outer domain; grey, inner domain; orange, bridging sheet; blue, CD4-binding loop; green, loop D; teal, loop V5; ordered residues of the Asn276gp120 glycan shown as yellow sticks (normally a complex-type N-glycan, but a high mannose N-glycan in crystal structures); approximate locations of Asn460gp120 and Asn463gp120 shown as light pink and magenta dots, respectively). The lower panels show electrostatic potentials on surface representations of Fabs (left panels) and gp120s (right panels) colored blue (positive electrostatic potential) to red (negative electrostatic potential). The binding interfaces are outlined with a dotted black line. The approximate footprints of the complex-type Asn276gp120 glycan on Fab surfaces are indicated with a black triangle (the Asn463gp120 glycan, also complex-type, is not resolved in any Env structures, thus its footprint on Fab surfaces cannot be shown).
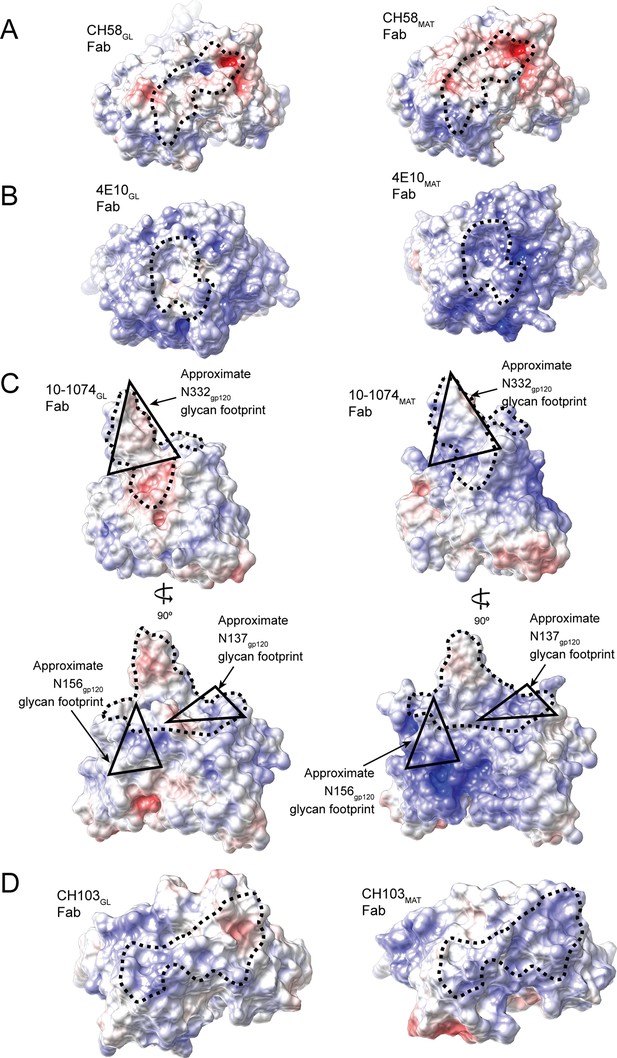
The combining sites of germline (left panels) and mature Fabs (right panels) are shown as surface representations and electrostatic potentials are indicated using blue for positive electrostatic potential and red for negative electrostatic potential.
Abs shown are CH58GL and CH58MAT (PDB codes 4RIR and 4HPO), 4E10GL and 4E10MAT (PDB codes 4ODX and 2FX7), 10-1074GL and 10-1074MAT (PDB codes 4FQQ and 4FQ2) (shown in two orientations to better illustrate electrostatic changes), and CH103GL and CH103MAT (PDB codes 4QHK and 4JAM). The antibody paratopes are outlined with a dotted black line. The approximate footprint of Asn137gp120, Asn156gp120 and Asn332gp120 glycans on 10-1074GL and 10-1074MAT Fab surfaces are indicated with black triangles.
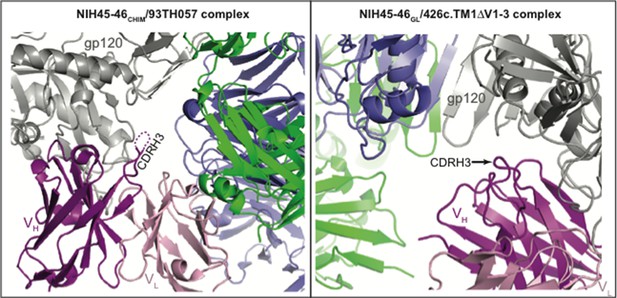
CDR3HC positions in NIH45-46CHIM/93TH057 gp120 (left) and NIH45-46GL/426c.TM1△V1-3 gp120 (right) complexes relative to symmetry mates (different views shown for clarity).
One complex of gp120 (grey) with antibody VH (pink) and VL (light pink) is shown with the nearest two symmetry mates (blue and green). The disordered region of NIH45-46CHIM CDRH3 is indicated by a dashed line. The CDRH3 loop is not engaged in crystal packing interactions in either complex.
Tables
Data collection and refinement statistics, molecular replacement.
| 3BNC60GL Fab (5F7E) | 426c.TM4ΔV1-3 (5FA2) | 3BNC60GL Fab-426c.TM4ΔV1-3 complex (5FEC) | NIH45-46GL Fab-426c.TM1ΔV1-3 complex (5IGX) | 3BNC55MAT Fab-426c.TM4ΔV1-3 complex (5I9Q) | |
|---|---|---|---|---|---|
| Resolution range | 34.13 - 1.9 (1.97 - 1.9) | 35.8 - 2.0 (2.072 - 2.0) | 37.44 - 3.1 (3.211 - 3.1) | 39.37 - 3.4 (3.521 - 3.4) | 37.54 - 2.8 (3.10 - 3.0) |
| Space group | P212121 | C121 | P212121 | F222 | P3121 |
| Unit cell dimensions | |||||
| a, b, c (Å) | 74.9, 74.9, 83.1 | 144.9, 85.9, 90.0 | 103.1, 134.1, 195.0 | 147.1, 169.3, 177.7 | 122.9, 122.9, 265.0 |
| α, β, γ (°) | 90.0, 90.0, 90.0 | 90.0, 104.8, 90.0 | 90.0, 90.0, 90.0 | 90.0, 90.0, 90.0 | 90.0, 90.0, 120.0 |
| Total reflections | 243498 (23377) | 416316 (24886) | 335536 (34539) | 207967 (20780) | 710649 (51477) |
| Unique reflections | 37256 (3651) | 81715 (7961) | 49653 (4885) | 15410 (1508) | 57299 (4353) |
| Multiplicity | 6.5 (6.5) | 4.6 (4.5) | 6.1 (6.4) | 13.5 (13.8) | 12.4 (11.8) |
| Completeness (%) | 0.95 (0.98) | 0.97 (0.95) | 1.00 (1.00) | 1.00 (1.00) | 0.71 (95.7) |
| Mean I/sigma(I) | 16.56 (2.36) | 13.35 (0.6) | 9.75 (1.47) | 21.76 (4.03) | 8.65 (1.3) |
| Wilson B-factor | 23.2 | 30.3 | 68.98 | 97.44 | 77.46 |
| R-merge | 0.09029 (0.8891) | 0.06624 (1.36) | 0.1729 (1.305) | 0.1087 (0.6832) | 0.327 (3.774) |
| CC1/2 | 0.999 (0.831) | 0.999 (0.437) | 0.995 (0.483) | 0.999 (0.909) | 0.993 (0.186) |
| CC* | 1 (0.953) | 1 (0.78) | 0.999 (0.807) | 1 (0.976) | 0.998 (0.56) |
| Rwork | 0.196 | 0.207 | 0.203 | 0.279 | 0.240 |
| Rfree | 0.210 | 0.232 | 0.267 | 0.286 | 0.279 |
| Number of atoms | 3502 | 5860 | 16611 | 5061 | 11572 |
| macromolecules | 3226 | 5124 | 16198 | 4956 | 11258 |
| ligands | 320 | 413 | 105 | 314 | |
| Protein residues | 429 | 672 | 2182 | 697 | 1537 |
| RMS (bonds) | 0.01 | 0.012 | 0.014 | 0.017 | 0.014 |
| RMS (angles) | 1.28 | 1.534 | 1.22 | 1.75 | 1.35 |
| Clashscore | 4.93 | 6.6 | 15.17 | 9.77 | 28.65 |
| Average B-factor | 33.0 | 48.66 | 84.92 | 95.14 | 86.29 |
| macromolecules | 31.14 | 47.27 | 83.94 | 93.81 | 86.98 |
| ligands | 70.79 | 123.27 | 165.32 | 61.48 | |
| solvent | 38.67 | 48.77 | |||
| Statistics for the highest-resolution shell are shown in parentheses. | |||||
Nominal net charges of VRC01-class bNAbs and of the human Ab repertoire. Nominal net charge is the total charge of the sequence calculated with Asp/Glu as -1 and Arg/Lys as +1. The human Ab repertoire average is calculated from 601,889 HC and 206,953 LC sequences reported in (Rubelt et al., 2012). Light chain Vgene assignments are from (Zhou et al., 2015). Standard deviations are indicated, except for human Ab repertoire VH + VL, which cannot be calculated with unpaired chains. *Two-tailed T test comparing the net charges of VRC01-class VHs with human repertoire VHs gives a p-value of 0.01.
| Nominal Net Charge | ||||
|---|---|---|---|---|
| Ab | VH + VL | VH | VL | LC V-gene |
| VRC01 | 5 | 4 | 1 | K3-20 |
| NIH45-46 | 9 | 8 | 1 | K3-20 |
| 3BNC117 | 5 | 2 | 3 | K1-33 |
| 12A12 | 5 | 1 | 4 | K1-33 |
| VRC-PG04 | 7 | 3 | 4 | K3-20 |
| VRC-CH31 | 2 | 2 | 0 | K1-33 |
| VRC-PG20 | 4 | 5 | -1 | L2-14 |
| VRC23 | 5 | 4 | 1 | K3-15 |
| VRC18 | 3 | 2 | 1 | K3-20 |
| VRC27 | 4 | 6 | -2 | K1-33 |
| Average of VRC01-class bNAbs | 4.9 ± 1.9 | 3.7* ± 2.1 | 1.2 ± 1.9 | |
| Human Ab repertoire average | 3.1 | 1.5 ± 2.4 | 1.6 ± 2.1 | |
Additional files
-
Supplementary file 1
Interfaces in NIH45-46GL-426c.TM1ΔV1-3 and 3BNC60GL-426c.TM4ΔV1-3 Complexes.
- https://doi.org/10.7554/eLife.13783.015





