An evolutionarily conserved mechanism for cAMP elicited axonal regeneration involves direct activation of the dual leucine zipper kinase DLK
Figures
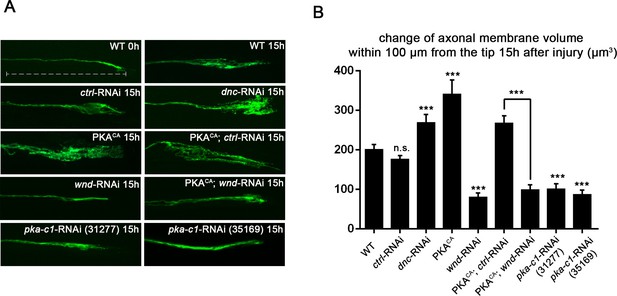
PKA stimulates and is required for axonal regeneration in Drosophila motoneurons.
Single motoneuron axons were labeled by expression of UAS-mCD8-GFP using the m12-Gal4 driver and imaged either 0 hr or 15 hr after nerve crush injury. Representative images are shown in (A), while (B) shows quantification of the increased volume in axonal membrane, which is measured within 100 µm of the proximal axon tip, indicated in dotted line. Genotypes used in (A): wild type (WT) (;;m12-Gal4, UAS-mCD8GFP/+); control-RNAi (UAS-dcr2;; UAS-moody-RNAi/m12-Gal4, UAS-mCD8GFP); dnc-RNAi (UAS-dcr2;; UAS-dnc-RNAi/m12-Gal4,UAS-mCD8GFP); wnd-RNAi (UAS-dcr2;; UAS-wnd-RNAi/m12-Gal4,UAS-mCD8GFP); PKACA(; UAS-PKACA/+; UAS-mCD8GFP/+); PKACA,control-RNAi (UAS-dcr2; UAS-PKACA/+; UAS-moody-RNAi (VDRC 100674)/m12-Gal4, UAS-mCD8GFP); PKACA,wnd-RNAi (UAS-dcr2; UAS-PKACA/+; UAS-wnd-RNAi/m12-Gal4, UAS-mCD8GFP); pka-c1-RNAi (UAS-dcr2; UAS-pka-c1-RNAi/+; m12-Gal4, UAS-mCD8GFP [using two different lines, Bloomington 31277 and 35169]). All data are represented as mean ± SEM; At least 10 animals (≥50 axons) are examined per genotype; ***p<0.001; ‘n.s.’ indicates non-significant; scale bar, 100 μm.
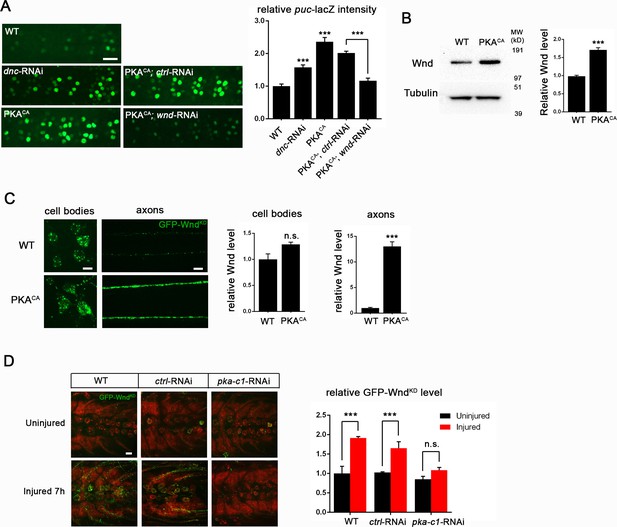
PKA modulates the levels of Wnd protein and downstream signaling in Drosophila neurons.
(A) The puc-lacZ transcriptional reporter for Wnd/JNK signaling indicates that activated PKA stimulates Wnd signaling. A pan-neuronal driver (BG380-Gal4) is used to express UAS-dnc-RNAi, UAS-PKACA or UAS-PKACA together with UAS-wnd-RNAi or a control-RNAi. Example images are shown of cell bodies in the dorsal midline of the ventral nerve cord; (all but two of these neurons are motoneurons). Quantification (described in methods) was carried out for 10 animals per genotype. (B) Endogenous Wnd protein levels are increased in PKACA expressing neurons. Ventral nerve cords were dissected from third instar larvae (BG380-Gal4 [WT control] and BG380-Gal4; UAS-PKACA/+) and processed for Western blotting with anti-Wnd and anti-tubulin antibodies. The quantification shows Wnd/tubulin ratios (normalized to WT control) averaged from 3 independent experiments (25 nerve cords per experiment). (C) PKA increases DLK levels via a posttranscriptional mechanism. GFP-tagged kinase dead Wnd (GFP-WndKD) was ectopically expressed using m12-Gal4 driver (WT control) or co-expressed with UAS-PKACA and imaged directly after fixation. Example images and quantification of GFP-WndKD intensity in cell bodies and axons within segmental nerves. n>10 animals for each condition. (D) PKA-C1 is required for induction of Wnd protein after axonal injury. Example images and quantification of GFP-WndKD in nerve cords and segmental nerves before and after (8 hr) injury. UAS-GFP-WndKD was expressed in motoneurons by OK6-Gal4. WT or together with UAS-pka-c1-RNAi or UAS-moody-RNAi (control) and imaged similarly to Figure 2C. The quantification method for GFP intensity is described in materials and methods. n>10 animals for each condition. All data are represented as mean ± SEM; ***p<0.001, **p<0.01, *p<0.05, ‘n.s.’ indicates non-significant; scale bars, 10 μm.
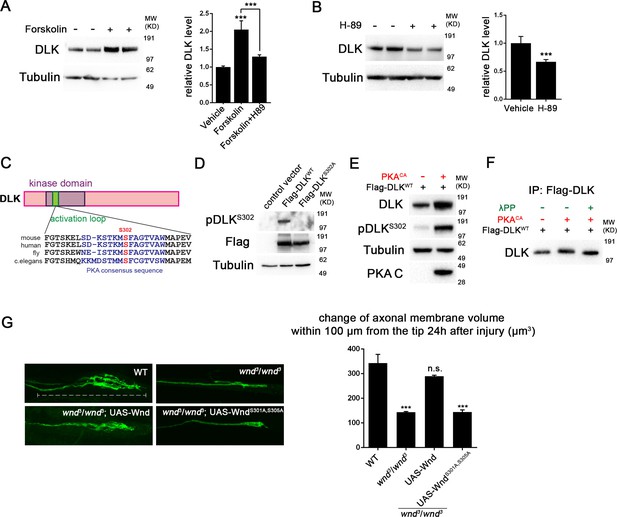
PKA activates DLK via phosphorylation of its activation loop.
(A-B) Changes in endogenous DLK abundance in response to treatment with forskolin (30 µM) (A) or the PKA inhibitor H-89 (5 µM) (B) for 6 hr in cultured rat embryonic cortical neurons. Quantification shows relative DLK/Tubulin levels in western blots. (C) Alignment of activation loop sequences in different species. (D) The anti-pDLKS302 antibodies recognize transfected Flag-DLKWT, but not activation loop mutation Flag-DLKS301A,S305A. Both proteins were transiently expressed in HEK293 cells. Western blots were probed with anti-pDLKS302 antibody, anti-Flag antibody to detect the total DLK expression levels, and anti-Tubulin (which remains similar in all manipulations) for normalization. (E) PKACA stimulates phosphorylation of DLK S302 in HEK293 cells. HEK293 cells were co-transfected with Flag-tagged DLKWT and an empty control plasmid or PKACA. Cell lysates were probed with anti-DLK antibody, anti-pDLKS302 antibody, anti-PKA C antibody and anti-tubulin antibody. (F) PKACA stimulates an increase in DLK molecular weight. Flag-DLK protein was immunoprecipitated from HEK293 cells co-transfected with DLK and either Flag-tagged DLKWT and an empty control plasmid or PKACA. The immunoprecipitated Flag-DLK was then incubated with either glycerol (control) or lambda protein phosphatase (λPP). PKACA induced a upward shift in DLK molecular weight, which was lost upon phosphatase treatment. (G) The activation loop is required for axonal regeneration in Drosophila neurons. Single axons in Drosophila third instar larva are labeled by mCD8RFP using eve-Gal4 driver. 24 hr after injury, these neurons in animals heterozygyous for wnd (wnd3/+) show robust axonal sprouting. However, sprouting fails to occur in wnd3/wnd3 animals. Expression of UAS-Wnd (WT) can restore axonal regeneration in wnd mutant background (UAS-Wnd, wnd3; wnd3,eve-Gal4, UAS-mCD8RFP). However, expression of activation loop mutant UAS-WndS301A,S305A failed to rescue the sprouting defect in wnd mutant animals (UAS-WndS301A,S305A, wnd3; wnd3, eve-Gal4,m12-mCD8RFP). Quantification of the volume of axonal membrane within 100 µm of the distal ending of the proximal stump. n> 50 axons for each genotype. Data are presented as mean ± SEM for 3 independent experiments; ***p<0.001; scale bar, 100 µm.
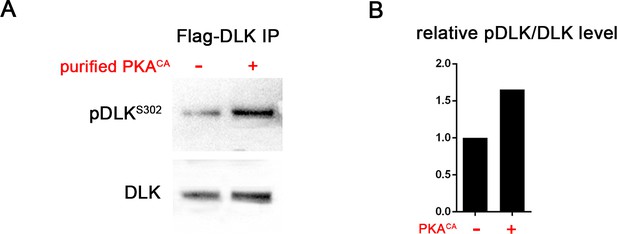
PKA can directly phosphorylate DLK at S302.
(A) PKA can induce DLK S302 phosphorylation in vitro. Flag-DLK was purified from HEK293 cells by anti-Flag immunoprecipitation and used for an in vitro kinase activity with purified PKA catalytic subunit. 5 µg of purified DLK was incubated with or without 10,000 U PKA catalytic subunit. Equal amounts of DLK in both samples (as demonstrated by probing with anti-DLK antibody) were analyzed by western blotting with anti-pDLKS302 antibody. (B) Quantification shows relative pDLKS302/total DLK levels.

DLK activation by PKA does not require TORC1.
(A-B) The phosphorylation level of DLK S302 is not sensitive to treatments of TORC1 inhibitors. HEK293 cells were either untransfected or transfected with Flag-DLKWT + empty plasmid or Flag-DLKWT + PKACA. Cells were treated with torin1 (A) or rapamycin (B) for 2 hr. The efficiency of the drugs were demonstrated by probing with anti-phospho-S6K antibody.
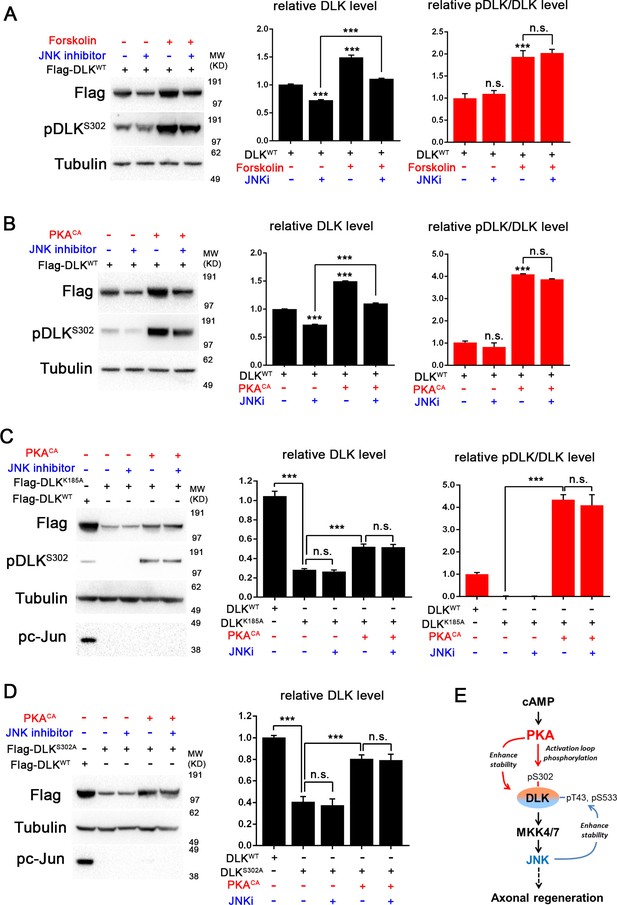
PKA promotes the stability of DLK independently of DLK downstream signaling.
(A-B) Activation of PKA promotes DLK stability independently of JNK. HEK293 cells were transiently transfected with Flag-DLKWT, and (A) treated with forskolin (6 hr, 30 µM) or (B) co-transfected with either PKACA or empty vector (control). In both cases, co-treatment with JNK inhibitor VIII (10 µM, 6 hr) led to a decrease in total Flag-DLK levels. However, both forskolin and PKACA induced an increase in DLK levels even in the presence of JNK inhibitor. Quantification shows average total DLK/Tubulin intensities and average pDLKS302/total DLK ratios (where total DLK is detected using anti-Flag antibody). All data are represented as mean ± SEM; quantifications of relative intensity from Western Blots were averaged from 3 independent experiments; ***p<0.001, **p<0.01, *p<0.05, ‘n.s.’ indicates non-significant. (C-D) Activation of PKA increases the stability of kinase dead DLK mutants, DLKK185A (C) and DLKS302A (D). HEK293 cells were transiently transfected with Flag-DLKK185A or Flag-DLKS302A together with PKACA or empty plasmid. Treatment JNK inhibitor VIII (10 µM) for 6 hr had no effect upon the PKACA induced levels of DLKK185A and DLKS302A mutant protein. Quantifications are similar to Figure 4A–B. Western bands intensity were averaged from 4 independent experiments; data are shown as mean ± SEM; ***p<0.001, **p<0.01, *p<0.05, ‘n.s.’ indicates non-significant. (E) Proposed model for the activation and stabilization of DLK by cAMP and PKA. cAMP elevation and PKA activation leads to the phosphorylation of S302 on DLK, which activates its kinase activity. Indicated in the blue arrow, downstream signaling via JNK leads to enhanced DLK stability and phosphorylation of DLK at other sites (Huntwork-Rodriguez et al., 2013). PKA also enhances DLK’s stability via an additional mechanism that is independent of S302 (red arrow).

Summary of predicted PKA phosphorylation sites on DLK/Wnd in different species.
The sequence of DLK homologues in different species (human, mouse, Drosophila and C. elegans) were analyzed by Group-based Prediction System (GPS) to computationally predict PKA phosphorylation sites (Xue et al., 2005; 2008). Identified potential PKACA phosphorylation motifs in different species are shown, with the phosphorylation site highlighted in red. Numbering is shown for mouse DLK. Two sites, S302 and S389, are conserved among all the species. Other sites are conserved in mammals, while fly DLK has a similar number of sites but in distinct locations. Not shown, C. elegans DLK also has 11 additional predicted sites, but at distinct locations from mammalian and fly DLK.
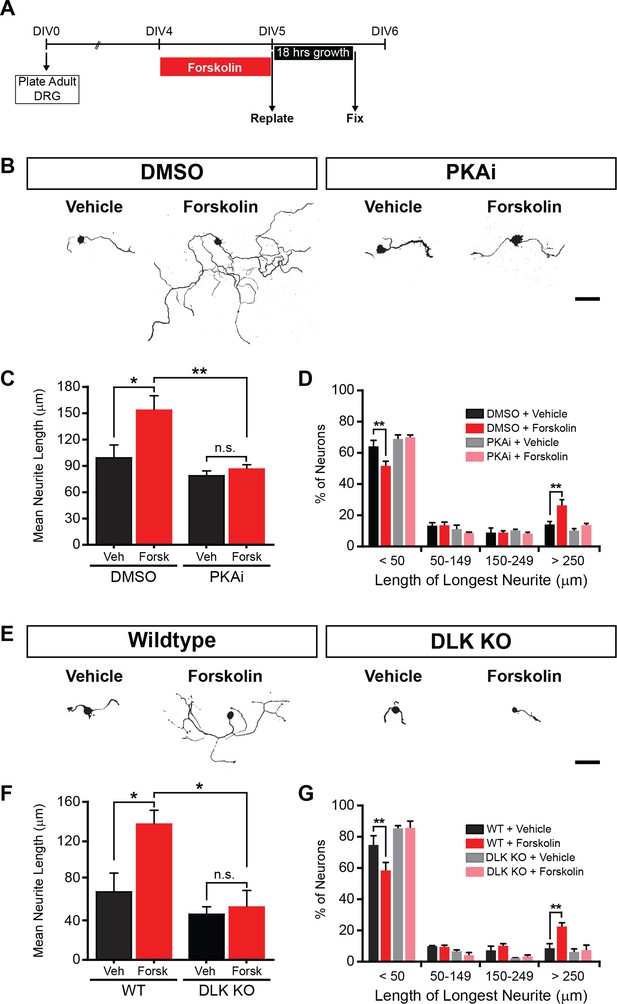
PKA stimulates axonal regeneration via DLK in adult DRG neurons.
(A-D) Induction of regeneration by forskolin requires PKA. Experimental design (A and also see Materials and Methods). To demonstrate that forskolin-induced neurite outgrowth is mediated by PKA, we assessed whether PKA signaling was required using the PKA inhibitor H-89 (PKAi, 5 µM). Representative neurons are shown in (B). Neurite outgrowth was assessed by quantifying mean neurite length (C) and distribution of longest neurite (D). Data are mean ± SEM for 4 independent experiments. (E-G) Induction of regeneration by forskolin requires DLK. WT and DLK KO neurons were treated with DMSO or forskolin (30 µM) as described in (A). Representative neurons are shown in (E). Neurite outgrowth was assessed by quantifying mean neurite length (F) and distribution of the longest neurite (G). Data are mean ± SEM for 3 independent experiments. ***p<0.001, **p<0.01, *p<0.05, ‘n.s.’ indicates non-significant; scale bars, 100 µm.
-
Figure 5—source data 1
Measurements of the longest neurite length for 100 neurons after replating in each condition.
The longest neurite length for 100 WT neurons after replating in four different conditions (DMSO+Vehicle, DMSO+forskolin, PKA inhibitor+Vehicle, PKA inhibitor+forskolin) are shown in Table ‘PKAi’. This table contains data from 4 independent experiments. The longest neurite length for 100 DLK KO neurons after replating in two different conditions (DMSO and forskolin) are shown in Table ‘DLK KO’. The table contains data from 3 independent experiments.
- https://doi.org/10.7554/eLife.14048.011





