Structural basis for the recognition of spliceosomal SmN/B/B’ proteins by the RBM5 OCRE domain in splicing regulation
Figures
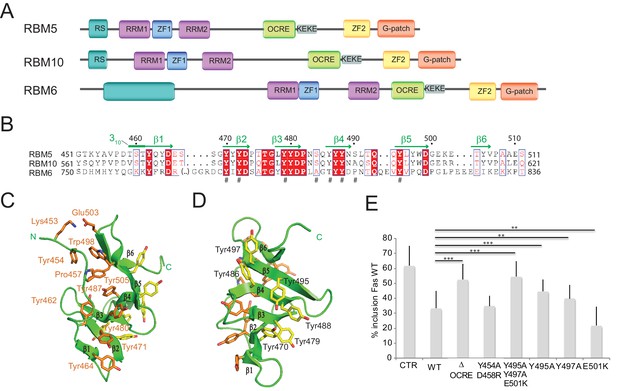
Structure and functional analysis of the RBM5 OCRE domain.
(A) Domain composition of the related RBM5, RBM6 and RBM10 family proteins. (B) Sequence alignment of OCRE domains of human RBM5, RBM10 and RBM6 proteins. (C,D) Two views in a cartoon presentation of the RBM5 OCRE domain, showing side chains of conserved and exposed tyrosine residues, some of which were probed by mutational analysis. (C) At one side of the β-sheet surface the N-terminal extension shields the tyrosines (orange) from solvent exposure, (D) on the opposite surface of the β-sheet, numerous tyrosine residues (yellow) are solvent accessible. (E) Mutational analysis of conserved residues of the RBM5 OCRE domain. Specific mutations of conserved and accessible residues of RBM5 OCRE domain impair the activity of the protein in FAS alternative splicing regulation ex vivo. HeLa cells were co-transfected with a Fas wild type alternative splicing reporter (harbouring sequences between the 5’ end of exon 5 and the first 47 nucleotides of exon 7) and T7-RBM5 expression plasmids. RNA and proteins were isolated 24 hr after transfection. Patterns of alternative splicing were studied by RT-PCR using specific primers (PT1, PT2) and the percentage of inclusion was calculated and is presented in the histogram for a minimum of 16 replicas of the experiment. T-test (two-tailed distribution, homoscedastic) results are mentioned (**<0,01; ***<0,001). Full quantification and T-test results are provided in Figure 5—figure supplement 2.
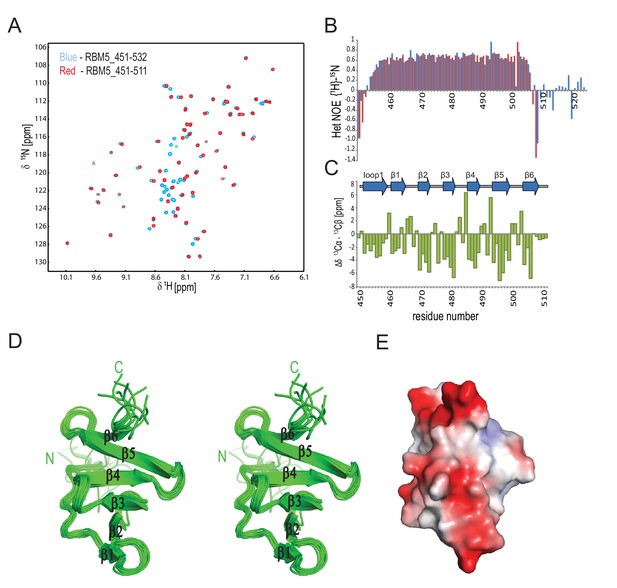
NMR analysis of the RBM5 OCRE domain.
(A) Superimposition of 1H,15N-HSQC spectra of two different constructs comprising the RBM5 OCRE domain, residues 451–511 (red) and residues 451–532 (blue). (B) {1H}-15N heteronuclear NOE relaxation analysis of the two constructs (residues 451–511, red) and residues 451–532, blue) indicate that the peptide backbone beyond residue 511 is flexible. (C) Secondary chemical shifts of OCRE 451–511 indicates that OCRE comprises β-strand and extended conformations only.(D) Stereo view of the ensemble of 10 lowest energy NMR structures of the RBM5 OCRE domain and (E) Electrostatic surface charge of the OCRE domain, same view as in Figure 1B.
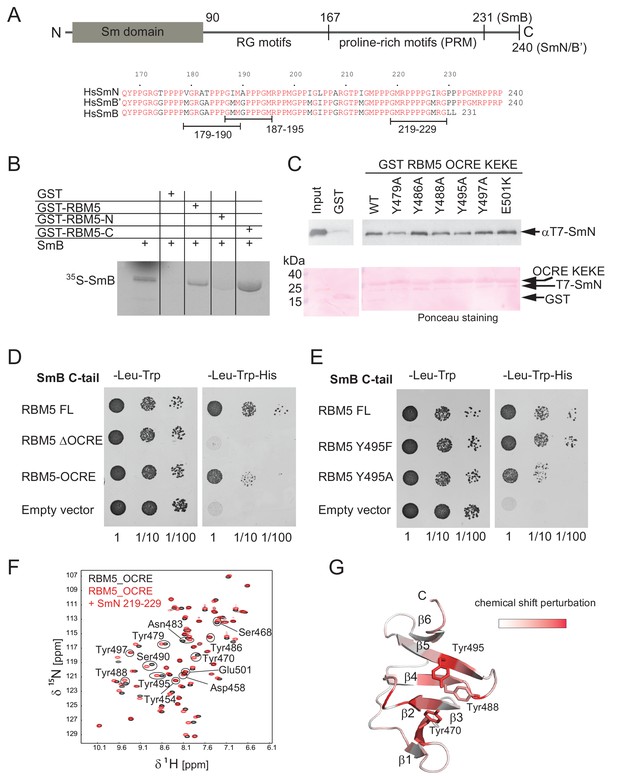
Interaction of RBM5 OCRE with SmN/B/B’ proteins.
(A) Domain structure and multiple sequence alignment of the C-terminal tails of human SmN/B/B’ proteins. (B) In vitro pull down assays of 35S-labeled SmB with GST-RBM5 full-length, N- and C-terminal halves of human RBM5. First lane represents 20% of the input used in the pull down. (C) Conserved tyrosine residues in the RBM5 OCRE domain are important for the interaction with SmN protein. GST-pull down assays were carried out with different variants of RBM5 comprising the OCRE and KEKE domains harboring the mutations indicated. The detection of the protein was carried out by western blot with T7 epitope antibody and Ponceau staining was performed as a loading control. Quantification is provided in Figure 2—figure supplement 1. (D) The C-terminal domain of human SmB binds to the OCRE domain of RBM5 in vivo. Yeast two hybrid plasmids encoding the C-terminal domain of human SmB (SmB C-tail) and the indicated RBM5 coding regions were transformed into yeast. Serial dilutions of equivalent amounts of exponentially growing yeast were plated on double and triple dropout media. Growth on -Leu -Trp -His is indicative of an interaction between the tested proteins: RBM5-FL: full length RBM5 protein; RBM5 ∆OCRE: deletion of the RBM5 OCRE domain (amino-acids 452–535); RBM5 OCRE: RBM5 OCRE domain (amino-acids 452–535). (E) Surface-exposed tyrosine residues are important for SmB C-tail binding in vivo. RBM5-FL: full length RBM5 protein; RBM5 Y495F: RBM5-FL carrying a tyrosine to phenylalanine substitution at position 495; RBM5-FL Y495A: RBM5 carrying a tyrosine to alanine substitution at position 495. The two-hybrid assay was performed as described in panel d. (F) NMR titrationof 15N-labeled RMB5 OCRE domain (0.2 mM- black) with SmN residues 219–229 (red) at two-fold molar excess. (G) Mapping of NMR chemical shift perturbations upon titration of the OCRE domain with SmN (residues 219–229) onto a surface representation of the RBM5 OCRE domain structure.
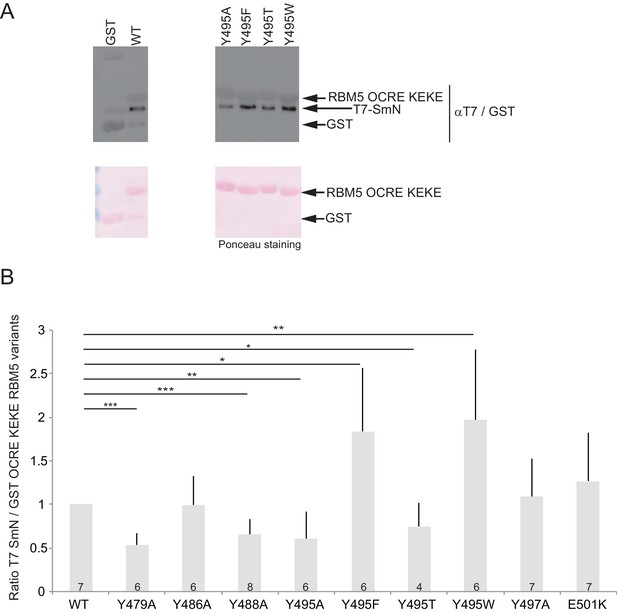
Aromatic residues in RBM5 OCRE are important for interaction with SmN.
(A) GST-pull down assays were carried out with T7 epitope-tagged-SmN (T7-SmN) and wildtype RBM5 fragment of the region comprising the OCRE and KEKE domain and different variants bearing mutations of Tyr495 to Ala, Thr, Phe and Trp (as in Figure 2C).The detection of the protein was carried out by Western blot using anti-T7 and GST-epitope antibodies and by Ponceau staining as loading control. (B) Quantification of the ratio between T7-SmN and GST-fusion proteins for 4 to 7 replicates of the experiment. The ratio was normalized to 1 for wild type GST-OCRE. T-test (two-tailed distribution, homoscedastic) results are indicated (*<0,05; **<0,01 and ***<0,001).
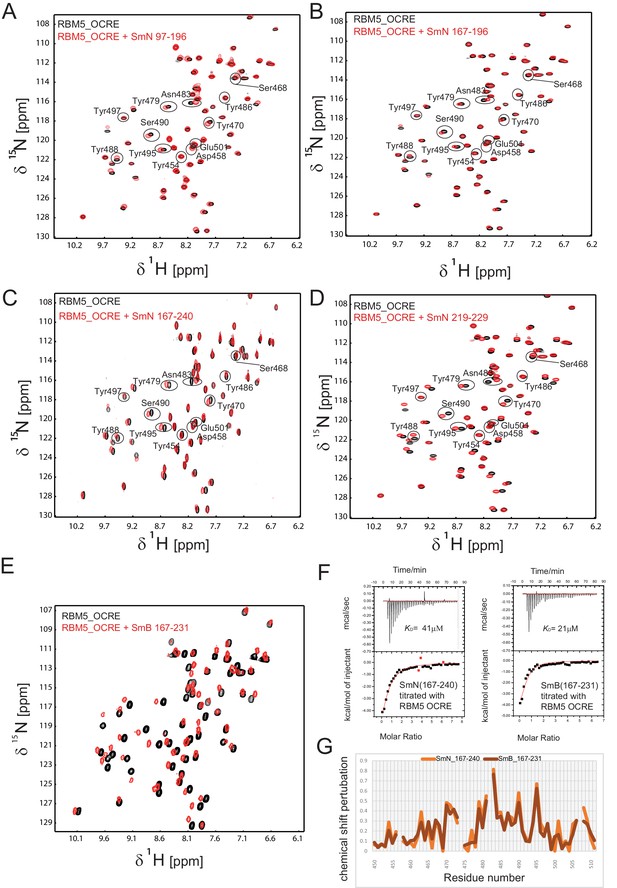
Interaction of RBM5 OCRE with C-terminal regions of SmN and SmB.
NMR titration of 0.2 mM 15N-labeled OCRE domain (black) with SmN and SmB peptides (red), (A) SmN 97–196, (B) SmN 167–196, (C) SmN 167–240 (A-C: 2-fold molar excess), (D) SmN 219–229 (6-fold molar excess), (E) SmB 167–231, 4-fold molar excess. (F) ITC analysis of the interaction of OCRE with C-terminal tails of SmN (residues 167 to 240) and SmB (residues 167 to 231). (G) Comparison of the chemical shift perturbations seen for the C-terminal tails of (C) SmN and SmB (E).
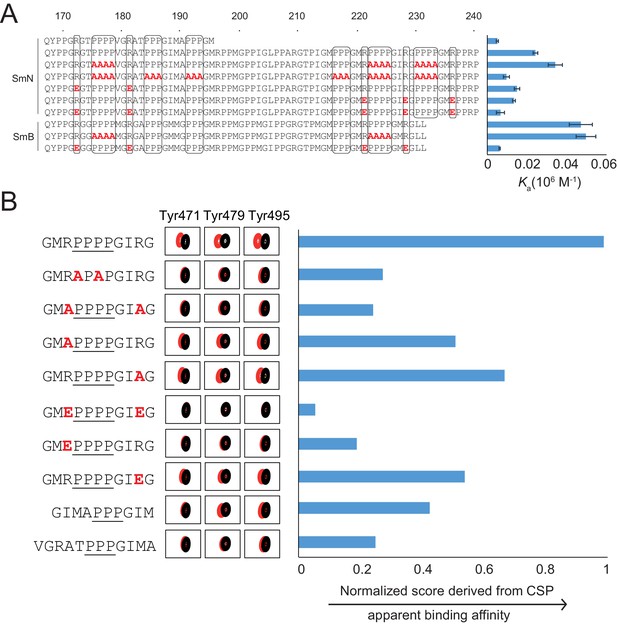
Proline-rich motif sequence requirements for OCRE binding.
(A) Sequence of the C-terminal tail of human SmN/B/B’ and mutations of proline-rich-motifs (PRMs) tested. Conserved three or four-proline motifs and flanking arginine residues are highlighted. The binding affinities of these tails to the RBM5 OCRE domain determined by ITC (Table 2) are indicated on the right. Note, that association constants are shown here. (B) Binding of various PRM peptides to RBM5 OCRE monitored by NMR titrations. Relative binding affinities of PRM peptides to RBM5 OCRE based on a normalized chemical shift perturbation score are shown.
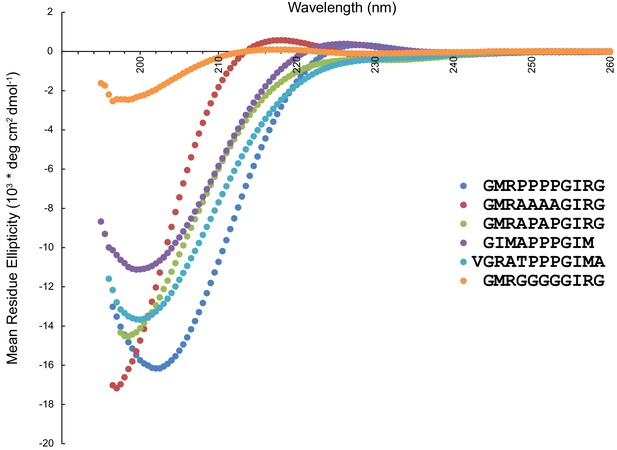
Circular dichroism spectra of peptides used in the OCRE binding study.
The presence of a strong negative band at 200 nm and a weak positive band at 217 nm are characteristic features of a PPII helix in CD spectra (Drake et al., 1988).In the presence of a higher number of proline residues, the positive CD band shifts towards 229 nm (Petrella et al., 1996). Therefore, wildtype (blue) and 4A mutant peptide (red) show PPII helical conformation, while the APAP mutant (green), which lacks the characteristic positive band at 217/229 nm does not exhibit a PPII helical conformation. The 4G mutant peptide (orange) also shows residual PPII helical conformation, consistent with previous reports (Kelly et al., 2001; Brown and Zondlo, 2012). Of the two peptides with three prolines, the one where the 3P motif is preceded by GIMA (purple) has PPII helical conformation (although the negative band at 199.5 nm is less intense as compared to the wildtype peptide), while the one preceded by VGRA (cyan) has a random coil conformation (lacking a positive band around ~229 nm).
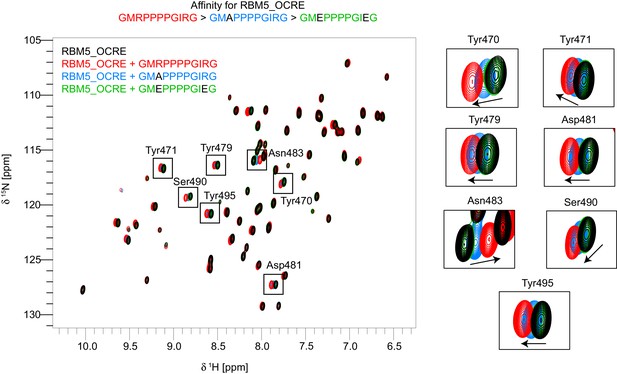
Residues used for normalized CSP score calculation.
Overlay of 15N-HSQC spectra of free OCRE (black) vs bound to wild type peptide-GMRPPPPGIRG (red), mutant peptides -GMAPPPPGIRG (blue) and GMEPPPPGIEG (green). Zoomed regions on the right show the residues that were used for calculation of the normalized chemical shift perturbations (CSP) score to allow a comparison of relative binding affinities of the different peptides. Only spectra of three peptide bound states of OCRE are shown to illustrate the affinity range: largest CSPs are observed for GMRPPPPGIRG, intermediate for GMAPPPPGIRG, and lowest for GMEPPPPGIEG.
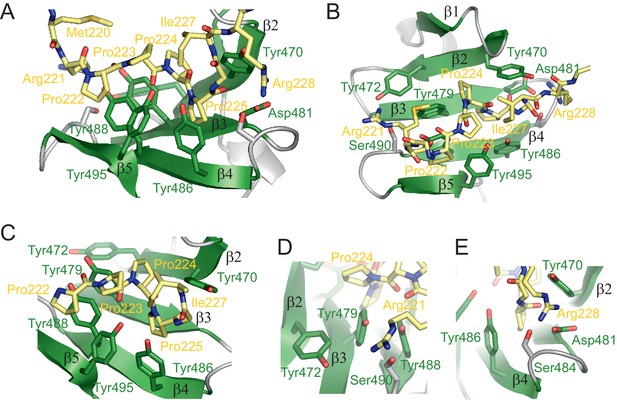
Structure of the RBM5 OCRE/SmN peptide complex.
(A,B) Side and top views in a cartoon presentation of the RBM5 OCRE/SmN peptide complex.The secondary structure and loops in the RBM5 OCRE domain are colored in green and grey, respectively, the proline-rich motif (PRM) peptide corresponding to SmN residues 221–229 is shown in yellow. (C–E) Zoomed views of key interaction sites. (C) The SmN PRM adopts a proline-type II helical conformation and is recognized by stacking with key tyrosine residues from the OCRE domain. The side chain of SmN Ile227 packs against the hydrophobic surface of the PPII helix. (D) Recognition of SmN Arg221 by interactions with hydroxyl groups of Tyr472, Tyr479 and Ser490 (E) Arg228 forms electrostatic contacts with the side chains of Asp481 and Ser484.
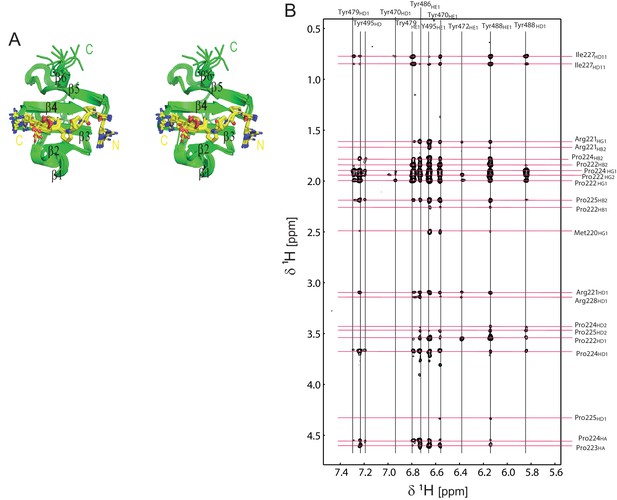
Structural analysis of the OCRE/SmN complex.
(A) Stereo view of the ensemble of 10 lowest energy NMR structures of the RBM5 OCRE domain in complex with SmN peptide, same view as in Figure 1D. (B) Intermolecular NOEs between the RBM5 OCRE domain and the SmN peptide. A zoomed view of a 2D ω1-filtered NOESY spectrum recorded on a 2 mM 15N,13C-labeled OCRE domain bound to 14 mM unlabelled SmN (residues 219–229). Assignments of the OCRE domain and the unlabelled SmN peptide are indicated on top and on the right, respectively.
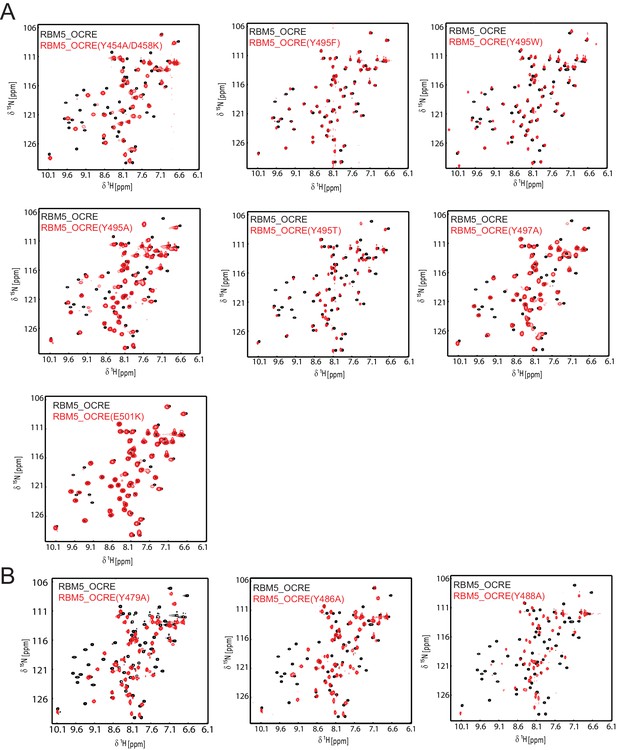
NMR spectra and ITC data for OCRE domain mutants.
Comparison of NMR spectra of the RBM5 OCRE domain wild type (black) and different mutants (red, as indicated).Spectra in (A) indicate that the overall fold is intact, with some potential local effects near the residue mutated. (B) These mutants suggest local unfolding of the β-strand, which comprises the altered residue, but are also consistent with an overall folded domain.
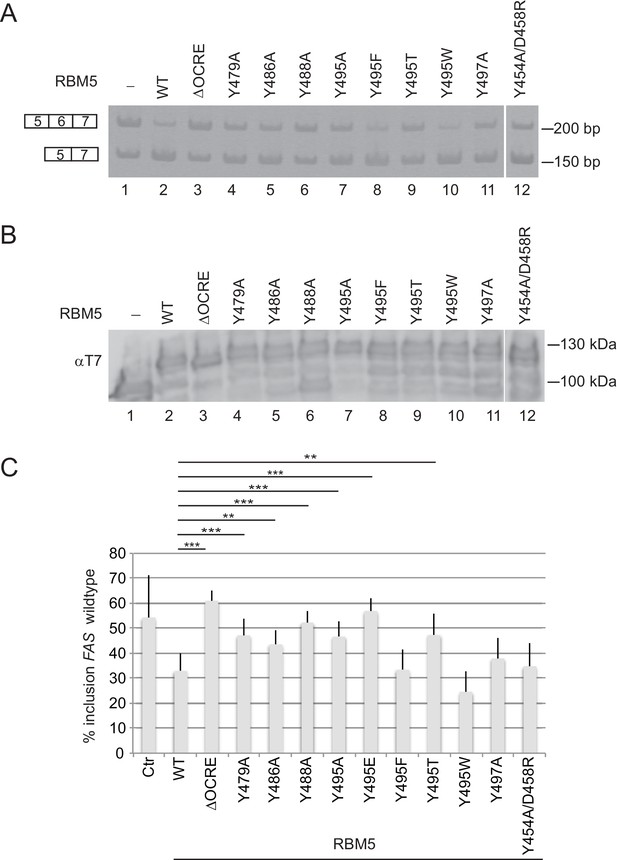
Mutational analysis of the RBM5 OCRE – SmN/B/B’ interaction.
Specific mutations of conserved residues in the RBM5 OCRE domain impair the activity of the protein in FAS alternative splicing regulation ex vivo. HeLa cells were co-transfected with a FAS alternative splicing reporter and T7-RBM5 expression plasmids (wild type or mutations of tyrosine residues as indicated). RNA and proteins were isolated 24 hr after transfection. Patterns of alternative splicing were studied by RT-PCR using specific primers. (A) Inclusion and skipping products are annotated. (B) Protein expression was detected by western blot with an anti-T7 epitope antibody. (C) Quantification of the activity of RBM5 OCRE domain mutants in FAS alternative splicing regulation of 3 to 10 replicates of the experiment. The percentage of inclusion is presented. T-test (two-tailed distribution, homoscedastic) results are mentioned (**<0,01; ***<0,001).
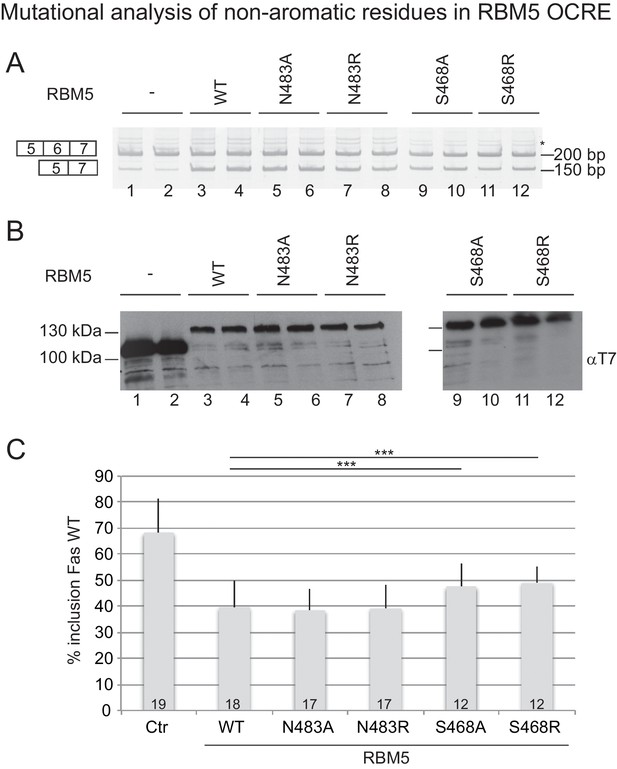
Effect of mutation of non-aromatic residues in RBM5 OCRE on FAS alternative splicing regulation.
(A) Mutations at position Ser468 of RBM5 OCRE domain impair the activity of the protein in FAS alternative splicing regulation ex vivo. HeLa cells were co-transfected with a FAS alternative splicing reporter and T7-RBM5 expression plasmids. RNA and proteins were isolated 24 hr after transfection. The pattern of alternative splicing was studied by RT-PCR using specific primers (PT1/PT2). Inclusion and skipping products are annotated. The asterisk indicates non-specific amplification. (B) Protein expression corresponding to the samples in (A), detected by western blot with an anti-T7 epitope antibody. (C) Quantification of the activity of RBM5 OCRE domain mutants on FAS alternative splicing regulation for 12 to 19 replicates, as indicated at the bottom of the histogram bars. Average and standard deviation of the percentage of inclusion is represented. T-test (two-tailed distribution, homoscedastic) results are indicated (***<0,001).
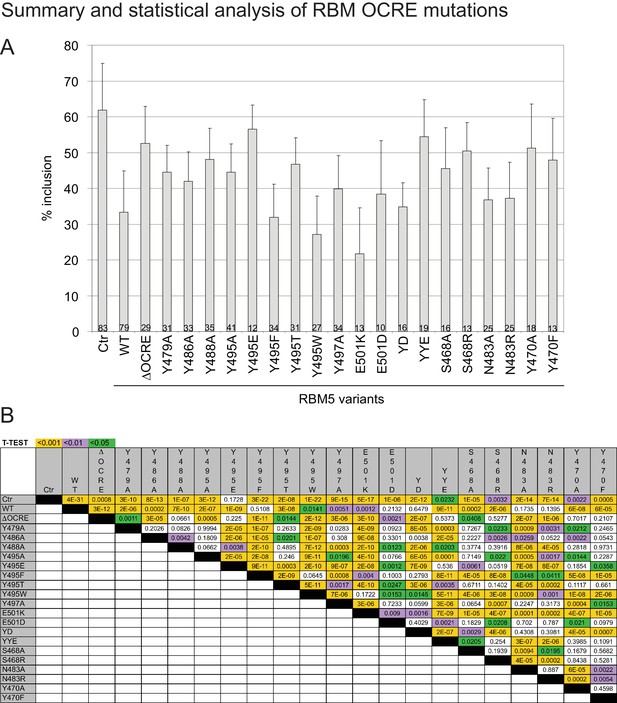
Effects and statistical analysis of RBM5 mutations on FAS alternative splicing.
(A) Quantification of the effect of RBM5 wild type and mutants on FAS wild type alternative splicing reporter regulation for all the experiments carried out in this study.Ctr: no RBM5 expressed. The number of replicates per mutant is annotated at the bottom of each histogram bar. (B) T-test (two-tailed distribution, homoscedastic) results for the data in (a), corresponding to each pairwise comparison (green<0,05; purple<0,01; orange<0,001).
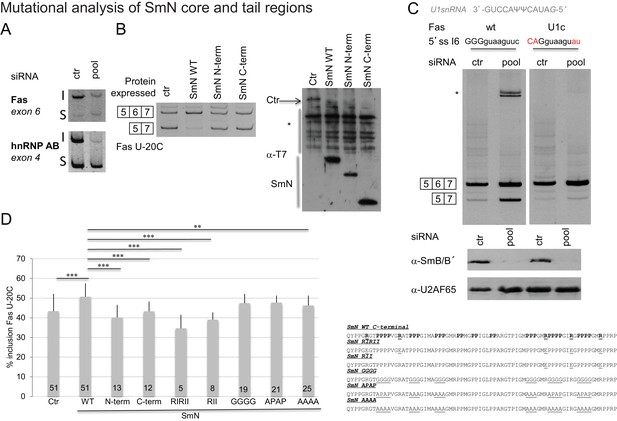
Effects of SmN wild type and mutants expression on FAS alternative splicing.
(A) Silencing of SmB/B’ affects alternative splicing of Fas and hnRNP AB genes.RNA was isolated from siRNA (on target smart pool siRNA, Dharmacon)-transfected HeLa cells. After reverse transcription using oligodT and random primers, PCR were performed using gene specific primers (Saltzman et al., 2011). The inclusion (I) and skipping (S) products of each gene are indicated. (B) SmN overexpression regulates FAS alternative splicing regulation. HeLa cells were co-transfected with a FAS alternative splicing reporter mutated at position -20 (U-20C) upstream of the 3’ splice site associated to exon 6 and with T7-SmN expression plasmids. RNA and proteins were isolated 24 hr after transfection. The pattern of alternative splicing was studied 24 hr later by RT-PCR using vector-specific primers (PT1/PT2). Inclusion and skipping products are indicated. Protein expression was detected by western blot using an anti-T7 epitope antibody. The asterisk indicates non-specific products. (C) Weakness of the 5’ splice site associated to FAS exon 6 is important for alternative splicing regulation by Sm proteins. RNA was isolated from siRNA HeLa transfected cells and the pattern of alternative splicing of FAS WT and FAS U1c mutant reporter (harbouring a mutation in the 5’ splice site of exon 6 that increases the base pairing potential with the 5’ end of U1 snRNA(Izquierdo et al., 2005) (sequence indicated) was detected by RT-PCR using vector-specific primers (PT1/PT2). Inclusion and skipping products are indicated. The asterisk indicates an additional amplification product. Western blot against SmB/B’ proteins (using the antibody 12F5, Santa Cruz, reference sc130670) was performed to evaluate the level of inactivation of the expression of the proteins and U2AF65 western blot analysis was performed as a control using the MC3 antibody. (D) Mutations in the C-terminal tail of SmN impair activity in FAS alternative splicing regulation. HeLa cells were co-transfected as in panel (B) using plasmids expressing T7-SmN variants bearing mutations indicated in the right part of the figure and containing an amino-terminal SV40 Large T antigen NLS (PKKKRKV). Average and standard deviation of the percentage of inclusion is indicated in the histogram. T-test (two-tailed distribution, homoscedastic) results are indicated (**<0,01; ***<0,001) as well as the number of replicates carried out for each mutant (from 5 to 51).
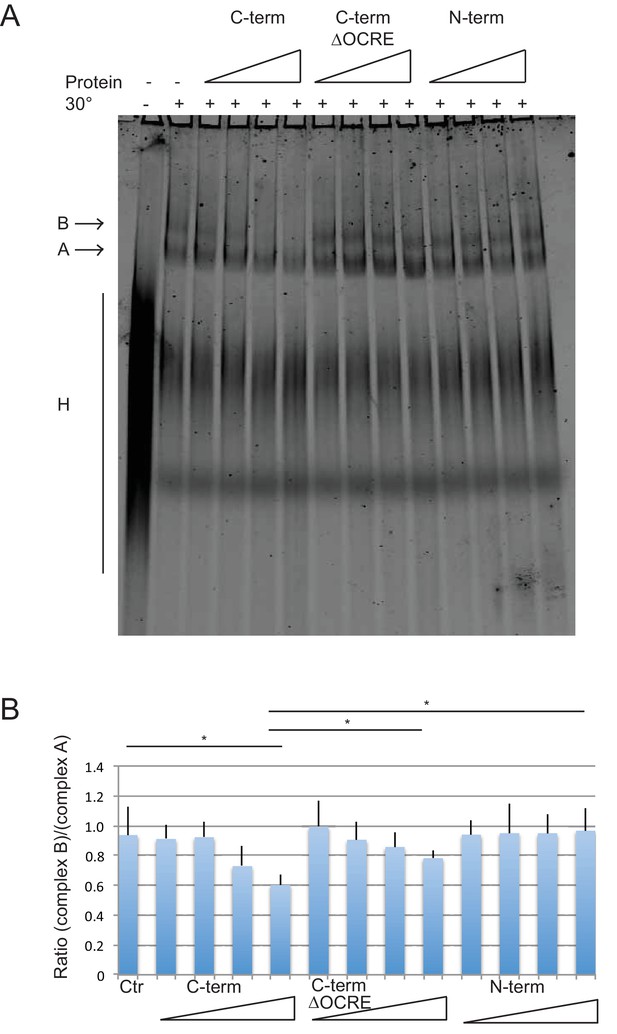
Spliceosome assembly assays with different regions of RBM5.
(A) RBM5 inhibits the full spliceosome assembly on AdML and blocks the transition from complexes A to B in an OCRE dependent manner.Splicing complexes assembled on AdML were resolved by electrophoresis on native gel in the presence or absence of 25, 50, 100 and 200 ng/ul of the indicated proteins. The position of the H, A and B complexes are indicated. (B) Quantification of the activity of spliceosome assembly of the C-term, N-term and ∆OCRE proteins. The experiment was performed three times and the results of a T-test (two-tailed distribution, homoscedastic) are indicated (*<0,05).
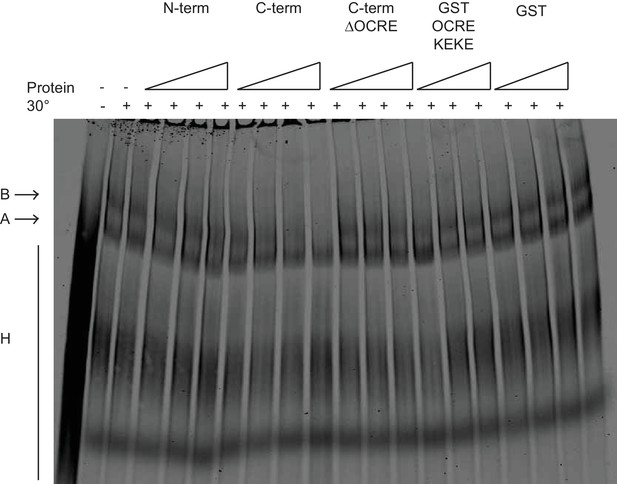
Effects of RBM5 full-length and fragments on spliceosome assembly.
The OCRE KEKE domains of RBM5 are not sufficient to trigger the block in transition from of A to B complexes. Splicing complexes assembled on AdML were resolved by electrophoresis on native gel in the presence or absence of 80, 400 and 2000 ng/ul of GST and GST OCRE KEKE proteins and 25, 50, 100 and 200 ng/ul of N-term, C-term and C-term ΔOCRE proteins. The presented result was reproduced twice under different conditions.
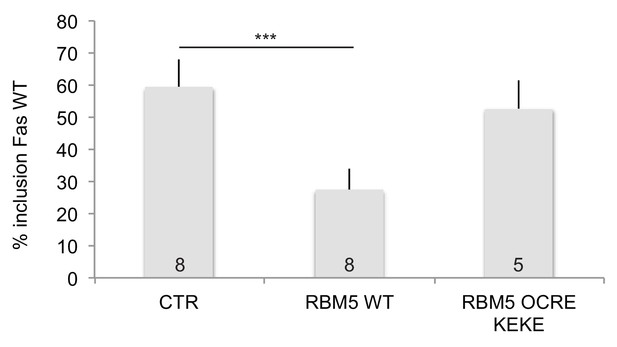
Effects of RBM5 full-length and OCRE on FAS alternative splicing.
The OCRE KEKE domains of RBM5 are not sufficient to trigger Fas exon 6 skipping.HeLa cells were co-transfected using plasmids expressing T7-RBM5 wild type protein and T7-NLS-RBM5 OCRE KEKE (T7 epitope fused to the SV40 Large T antigen SV40 NLS (PKKKRKV) and region from amino acid 452 to amino acid 535 of RBM5 protein). Average and standard deviation of the percentage of inclusion are indicated in the histogram. T-test (two-tailed distribution, homoscedastic) result is 8.4 E-07. The number of replicates carried out for each construct is indicated at the bottom.
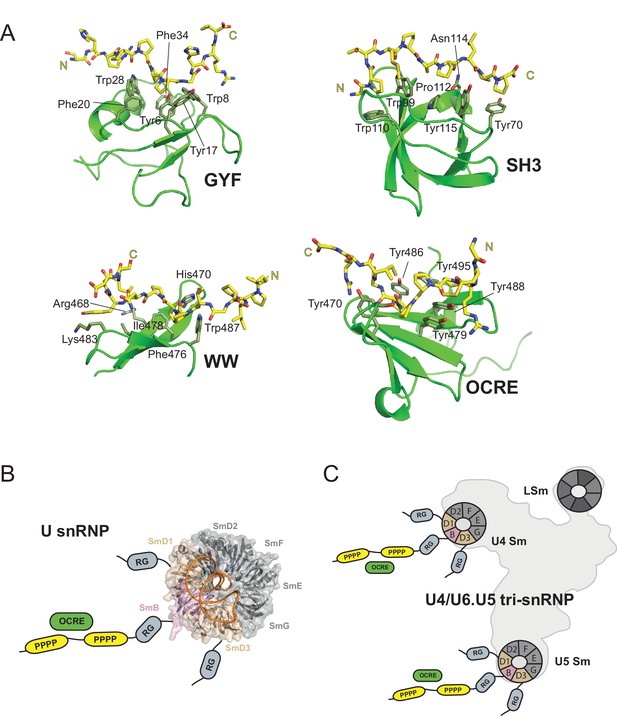
OCRE is a novel PRM-binding domain that interacts with snRNPs.
(A) Comparison of proline-rich motif (PRM) recognition by PRM binding domains.Cartoon representations of various PRM binding domains (green) with side chains of important residues shown as sticks and annotated. PRM ligands are shown by stick representation (yellow). PDB accession codes: GYF (PDB 1L2Z), SH3 (PDB accession 1ABO), WW (PDB 1I5H), and RBM5 OCRE. (B) The C-terminal tail of SmN/B/B’ comprising RG-rich regions and proline-rich motifs (PRM) extends far from the core Sm core fold in the seven-membered Sm ring of assembled U snRNPs. The crystal structure of U4 snRNP core (PDB 4WZJ) (Leung et al., 2011) is shown, the C-terminal tails of SmD1/D3, which harbour RG-rich regions and SmN/B/B’ comprising RG and PRM regions are not visible in the crystal structure and indicated schematically. (C) The seven-membered Sm core rings of U4 and U5 snRNPs are located at the outside of the assembled U4/U6.U5 tri-snRNP (Nguyen et al., 2015). Thus, also in the tri snRNP the SmN/B/B’ tails are accessible for interactions with the RBM5 OCRE domain.
Tables
Structural statistics RBM5 OCRE and OCRE/SmN complex.
https://doi.org/10.7554/eLife.14707.005| OCRE | OCRE + SmN | |
|---|---|---|
| NMR distance and dihedral restraints | ||
| Distance restraints | ||
| Total NOE | 1171 | 1127 |
| Intra-residue | 220 | 419 |
| Inter-residue | ||
| Sequential (|i-j| = 1) | 292 | 118 |
| Medium-range (|i-j| < 4) | 156 | 114 |
| Long-range (|i-j > 5) | 503 | 329 |
| Hydrogen bonds | 20 | 11 |
| Protein-peptide intermolecular | N/A | 109 |
| Dihedral angle restraints | ||
| ϕ | 47 | 51 |
| ψ | 51 | 51 |
| Structure statistics | ||
| Violations (mean and s.d.) | ||
| Distance restraints (>0.2 Å) | 0.685 ± 0.363 | 0.324 ± 0.111 |
| Dihedral angle restraints (>5 °) | 0 ± 0 | 0 ± 0 |
| Max. distance restraint violation (Å) | 0.657 ± 0.470 | 0.587 ± 0.142 |
| Max. dihedral angle restraint violation (°) | 0 | 0 |
| Deviations from idealized geometry | ||
| Bond lengths (Å) | 0.0038 ± 0.0002 | 0.0046 ± 0.0001 |
| Bond angles (°) | 0.444 ± 0.026 | 0.616 ± 0.015 |
| Impropers (°) | 1.230 ± 0.0681 | 1.737 ± 0.00889 |
| Average pairwise r.m.s.d.* (Å) | ||
| Heavy | 0.72 ± 0.05 | 0.71 ± 0.10 |
| Backbone | 0.38 ± 0.07 | 0.38 ± 0.09 |
-
∗Pairwise r.m.s.d. was calculated among 10 refined structures for residues 455-508 (RBM5 OCRE) and 221-229 (SmN) after water refinement.
Isothermal titration calorimetry data for the OCRE/SmN/B/B’ interaction.
https://doi.org/10.7554/eLife.14707.009| RBM5_OCRE | WT | Y495A | Y495T | Y495F | Y495W | Y488A | Y486A | Y479A | Y454A/D458K | E501K |
| SmN_167-240 | WT | WT | WT | WT | WT | WT | WT | WT | WT | |
| KD (µM) | 41 ± 2 | 172 ± 6 | 186 ± 12 | 87 ± 3 | 35 ± 3 | 220 ± 11 | 145 ± 5 | 111 ± 5 | 106 ± 15 | 48 ± 2 |
| KD values were determined from replicate measurements, with standard deviations as indicated. | ||||||||||
| RBM5_OCRE | WT | WT | WT | WT | WT | WT | ||||
| SmN_167-240 | 4P→4A | 4P/3P→4A/3A | 4P/3P→4G/3G | R1→E1 | R2→E2 | R1/R2→E1/E2 | ||||
| KD (µM) | 29 ± 3 | 103 ± 16 | 192 ± 66 | 66.5 ± 6 | 74 ± 4 | 149.5 ± 46 | ||||
| RBM5_OCRE | WT | |||||||||
| SmN_167-196 | WT | |||||||||
| KD (µM) | 195 ± 21 | |||||||||
| RBM5_OCRE | WT | WT | WT | |||||||
| SmB_167-231 | WT | 4P→4A | R1/R2→E1/E2 | |||||||
| KD (µM) | 21 ± 3 | 20 ± 2 | 159 ± 8 | |||||||





