Ensemble cryo-EM uncovers inchworm-like translocation of a viral IRES through the ribosome
Figures
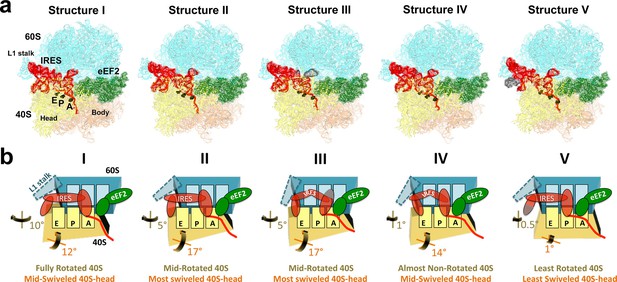
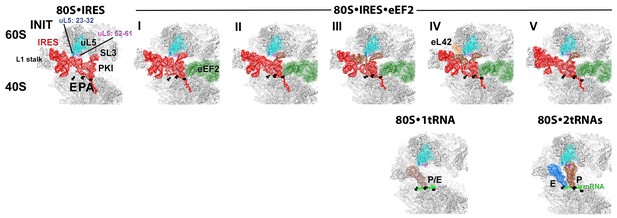
Cryo-EM structures of the 80S•TSV IRES bound with eEF2•GDP•sordarin.
(a) Structures I through V. In all panels, the large ribosomal subunit is shown in cyan; the small subunit in light yellow (head) and wheat-yellow (body); the TSV IRES in red, eEF2 in green. Nucleotides C1274, U1191 of the 40S head and G904 of the platform (C1054, G966 and G693 in E. coli 16S rRNA) are shown in black to denote the A, P and E sites, respectively. Unresolved regions of the IRES in densities for Structures III and V are shown in gray. (b) Schematic representation of the structures shown in panel a, denoting the conformations of the small subunit relative to the large subunit. A, P and E sites are shown as rectangles. All measurements are relative to the non-rotated 80S•2tRNA•mRNA structure (Svidritskiy et al., 2014). The colors are as in panel a.
-
Figure 1—source data 1
Structure refinement statistics for Structures I, II, III, IV, V.
- https://doi.org/10.7554/eLife.14874.003
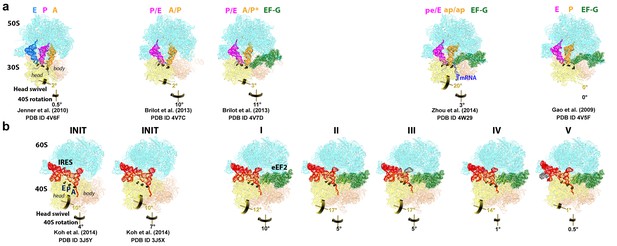
Comparison of 70S•2tRNA•mRNA and 80S•IRES translocation complexes.
(a) Structures of bacterial 70S•2tRNA•mRNA translocation complexes, ordered according to the position of the translocating A->P tRNA (orange). The large ribosomal subunit is shown in cyan; the small subunit in light yellow (head) and wheat-yellow (body), elongation factor G (EF-G) is shown in green. Nucleotides C1054, G966 and G693 of 16S rRNA are shown in black to denote the A, P and E sites, respectively. The extents of the 30S subunit rotation and head swivel relative to their positions in the post-translocation structure (Gao et al., 2009) are shown with arrows. References and PDB codes of the structures are shown. (b) Structures of the 80S•IRES complexes in the absence (Koh et al., 2014) and presence of eEF2 (this work). The large ribosomal subunit is shown in cyan; the small subunit in light yellow (head) and wheat-yellow (body); the TSV IRES in red, eEF2 in green. Nucleotides C1274, U1191 of the 40S head and G904 of the platform (corresponding to C1054, G966 and G693 in E. coli 16S rRNA) are shown in black to denote the A, P and E sites, respectively. Unresolved regions of the IRES in densities for Structures III and V are shown in gray. The extents of the 40S subunit rotation and head swivel relative to their positions in the post-translocation structure (Svidritskiy et al., 2014) are shown with arrows.
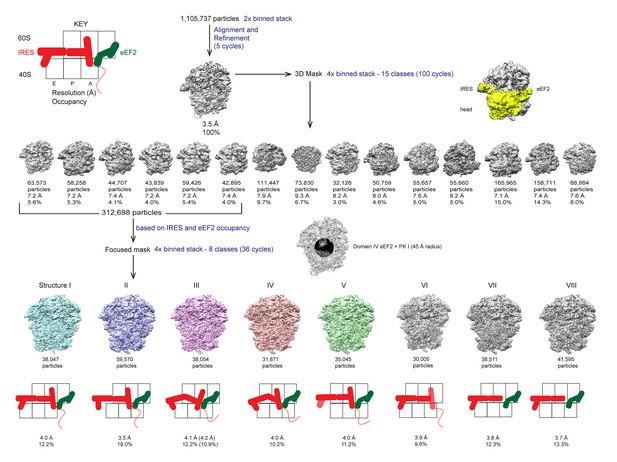
Schematic of cryo-EM refinement and classification procedures.
All particles were initially aligned to a single model. 3D classification using a 3D mask around the 40S head, TSV IRES and eEF2, of the 4x binned stack was used to identify particles containing both the IRES and eEF2. Subsequent 3D classification using a 2D mask comprising PKI and domain IV of eEF2 yielded 5 'purified' classes representing Structures I through V. Sub-classification of each class did not yield additional classes, but helped improve density in the PKI region of class III (estimated resolution and percentage of particles in the sub-classified reconstruction are shown in parentheses).
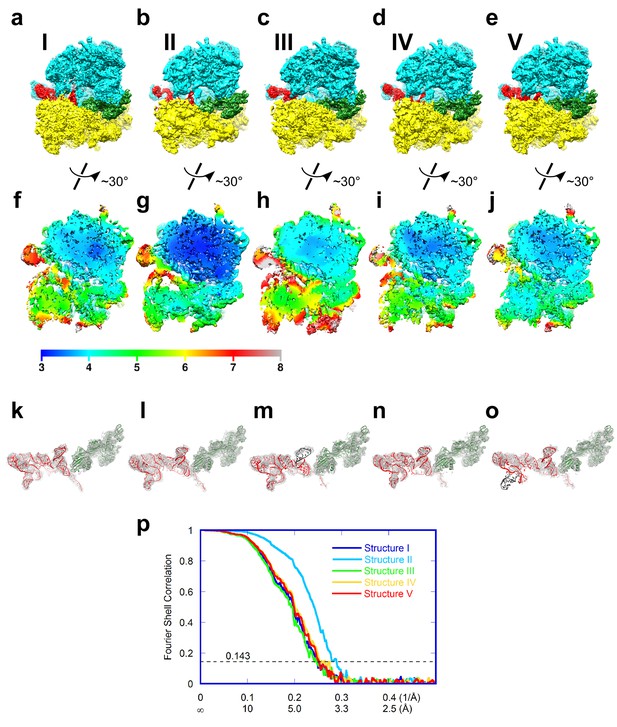
Cryo-EM density of Structures I-V.
In panels (a-e), the maps are segmented and colored as in Figure 1. The maps in all panels were B-softened by applying a B-factor of 30 Å2. (a-e) Cryo-EM map of Structures I, II, III, IV and V. (f-j) Local resolution of unfiltered and unmasked cryo-EM reconstructions, assessed using Blocres from the BSoft package (Cardone et al., 2013), for Structures I, II, III, IV and V. (k-o) Cryo-EM density for the TSV IRES (red model) and eEF2 (green model) in Structures I, II, III, IV and V. (p) Fourier shell correlation (FSC) curves for Structures I-V. The horizontal axis is labeled with spatial frequency Å-1 and with Å. The resolutions stated in the text correspond to an FSC threshold value of 0.143, shown as a dotted line, for the FREALIGN-derived FSC ('Part_FSC').
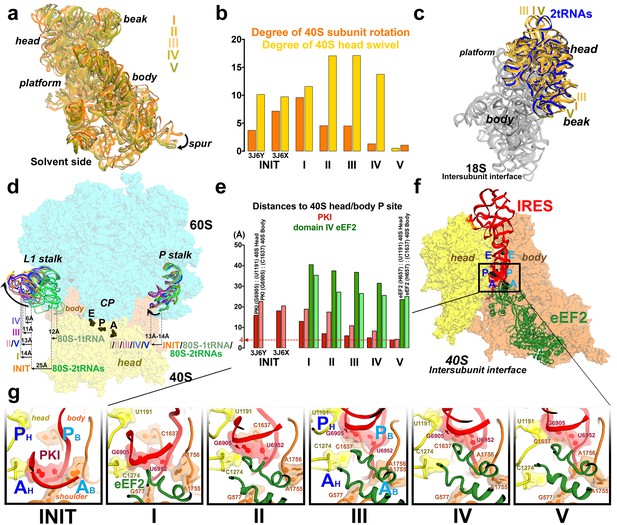
Large-scale rearrangements in Structures I through V, coupled with the movement of PKI from the A to P site and eEF2 entry into the A site.
(a) Comparison of the 40S-subunit rotational states in Structures I through V, sampling a ~10° range between Structure I (fully rotated) and Structure V (non-rotated). 18S ribosomal RNA is shown and ribosomal proteins are omitted for clarity. The superpositions of Structures I-V were performed by structural alignments of the 25S ribosomal RNAs. (b) Bar graph of the angles characterizing the 40S rotational and 40S head swiveling states in Structures I through V. Measurements for the two 80S•IRES (INIT) structures (Koh et al., 2014) are included for comparison. All measurements are relative to the non-rotated 80S•2tRNA•mRNA structure (Svidritskiy et al., 2014). (c) Comparison of the 40S conformations in Structures I through V shows distinct positions of the head relative to the body of the 40S subunit (head swivel). Conformation of the non-swiveled 40S subunit in the S. cerevisiae 80S ribosome bound with two tRNAs (Svidritskiy et al., 2014) is shown for reference (blue). (d) Comparison of conformations of the L1 and P stalks of the large subunit in Structures I through V with those in the 80S•IRES (Koh et al., 2014) and tRNA-bound 80S (Svidritskiy et al., 2014) structures. Superpositions were performed by structural alignments of 25S ribosomal RNAs. The central protuberance (CP) is labeled. (e) Bar graph of the positions of PKI and domain IV of eEF2 relative to the P site residues of the head (U1191) and body (C1637) in Structures I through V. (f and g) Close-up view of rearrangements in the A and P sites from the initiation state (INIT: PDB ID 3J6Y) to the post-translocation Structure V. The fragment shown within a rectangle in panel f is magnified in panel g. Nucleotides of the 40S body are shown in orange, 40S head in yellow. The superpositions of structures were performed by structural alignments of the 18S ribosomal RNAs excluding the head region (nt 1150–1620).
-
Figure 2—source data 1
Measurements for conformations and positions in Structures I through V.
- https://doi.org/10.7554/eLife.14874.008
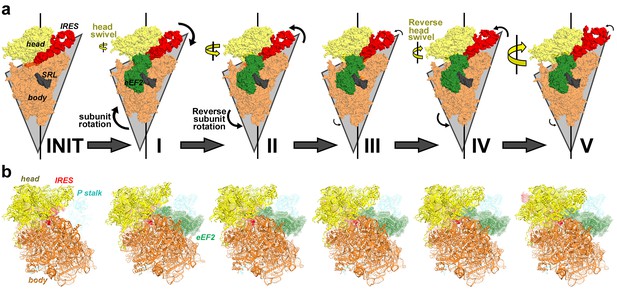
Large-scale rearrangements in Structures I through V, coupled with the movement of PKI from the A to P site and eEF2 entry into the A site.
(a) Rotational states of the 40S subunit in the 80S•IRES structure (INIT; PDB 3J6Y; Koh et al., 2014) and in 80S•IRES•eEF2 Structures I, II, III, IV and V (this work). For each structure, the triangle outlines the contours of the 40S body; the lower angle illustrates the extent of intersubunit (body) rotation. The sizes of the arrows correspond to the extent of the head swivel (yellow) and subunit rotation (black). The views were obtained by structural alignment of the 25S rRNAs; the sarcin-ricin loop (SRL) of 25S rRNA is shown in gray for reference. (b) Solvent view (opposite from that shown in (a)) of the 40S subunit in the 80S•IRES structure (INIT; PDB 3J6Y; Koh et al., 2014) and in 80S•IRES•eEF2 Structures I, II, III, IV and V (this work). The structures are colored as in Figure 1.
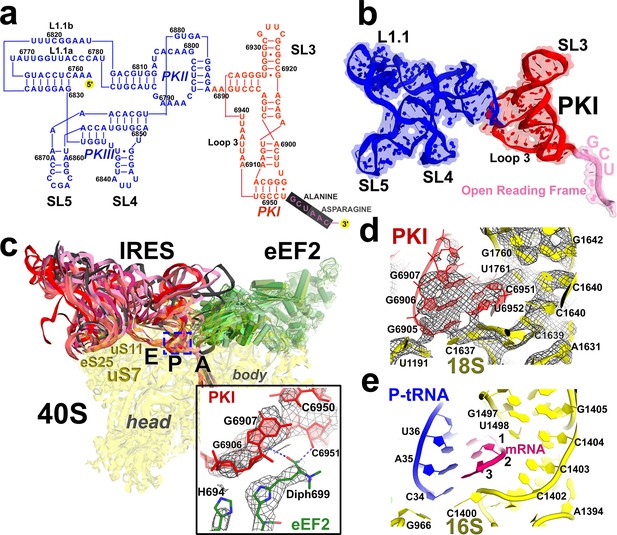
Positions of the IRES relative to eEF2 and elements of the ribosome in Structures I through V.
(a) Secondary structure of the TSV IRES. The TSV IRES comprises two domains: the 5' domain (blue) and the PKI domain (red). The open reading frame (gray) is immediately following pseudoknot I (PKI). (b) Three-dimensional structure of the TSV IRES (Structure II). Pseudoknots and stem loops are labeled and colored as in (a). (c) Positions of the IRES and eEF2 on the small subunit in Structures I to V. The initiation-state IRES is shown in gray. The insert shows density for interaction of diphthamide 699 (eEF2; green) with the codon-anticodon-like helix (PKI; red) in Structure V. (d and e) Density of the P site in Structure V shows that interactions of PKI with the 18S rRNA nucleotides (c) are nearly identical to those in the P site of the 2tRNA•mRNA-bound 70S ribosome (d; Svidritskiy et al., 2013).
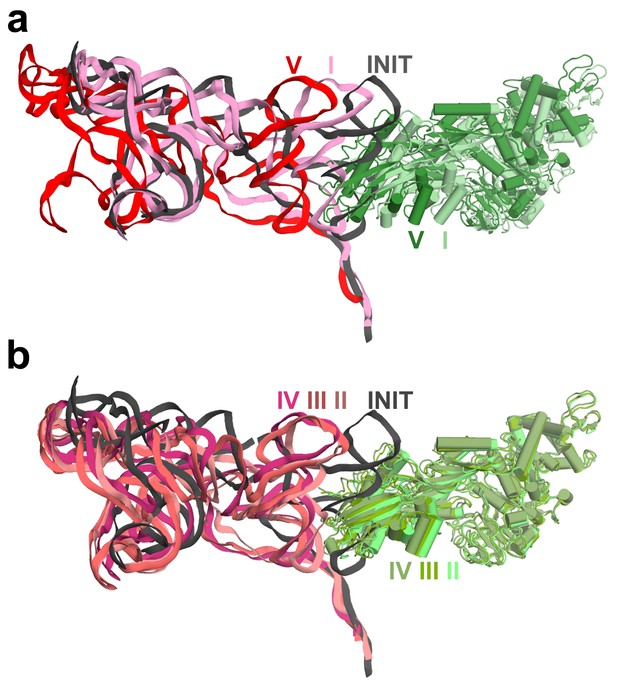
Comparison of the TSV IRES and eEF2 positions in Structures I through V.
(a) Positions of the IRES and eEF2 in the initiation, pre-translocation (I) and post-translocation (V) states, relative to the body of the 40S subunit (not shown) (b) Positions of the IRES and eEF2 in the initiation state (INIT) and intermediate steps of translocation (II, III and IV), relative to the body of the 40S subunit (not shown). Superpositions were obtained by structural alignments of the 18S rRNAs excluding the head domains (nt 1150–1620).
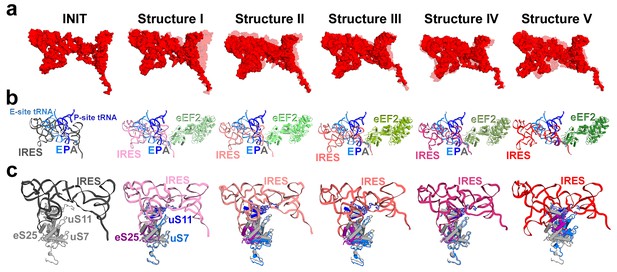
Positions of the IRES relative to proteins uS7, uS11 and eS25.
(a) Intra-IRES rearrangements from the 80S*IRES initiation structure (INIT; PDB 3J6Y, Koh et al., 2014) to Structures I through V. For each structure (shown in red), the conformation from a preceding structure is shown in light red for comparison. Superpositions were obtained by structural alignments of 18S rRNA. (b) Positions of the IRES and eEF2 relative to those of classical P- and E-site tRNAs in the 80S•tRNA complex (Svidritskiy et al, 2014). (c) Positions of the IRES relative to proteins uS11 (40S platform) and uS7 and eS25 (40S head), which interact with the 5′ domain of the IRES in the initiation state (left panel). In all panels, superpositions were obtained by structural alignments of the 18S rRNAs. Ribosomal proteins of the initiation state are shown in gray for comparison.
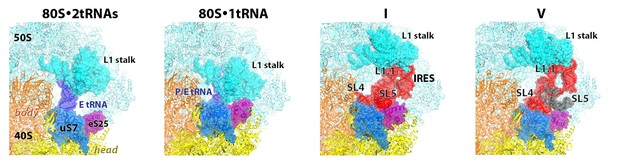
Positions of the L1stalk, tRNA and TSV IRES relative to proteins uS7 and eS25, in 80S•tRNA structures (Svidritskiy et al., 2014) and 80S•IRES structures I and V (this work).
The view shows the vicinity of the ribosomal E site. Loop 1.1 and stem loops 4 and 5 of the IRES are labeled.
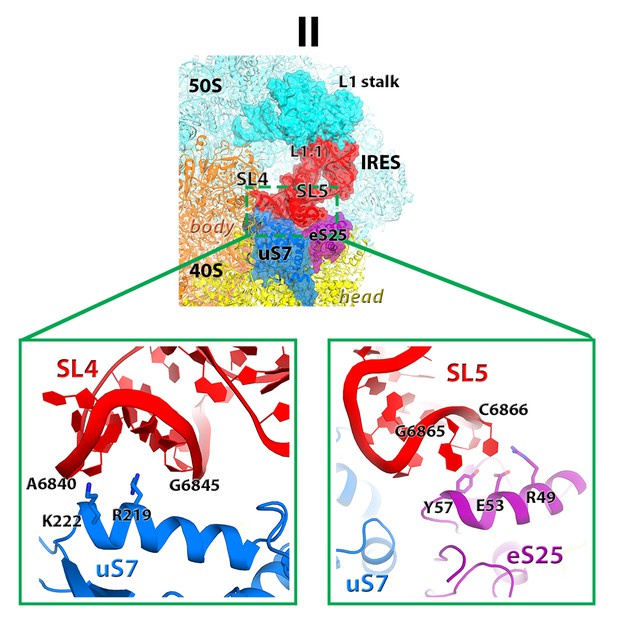
Interactions of the stem loops 4 and 5 of the TSV with proteins uS7 and eS25.
https://doi.org/10.7554/eLife.14874.014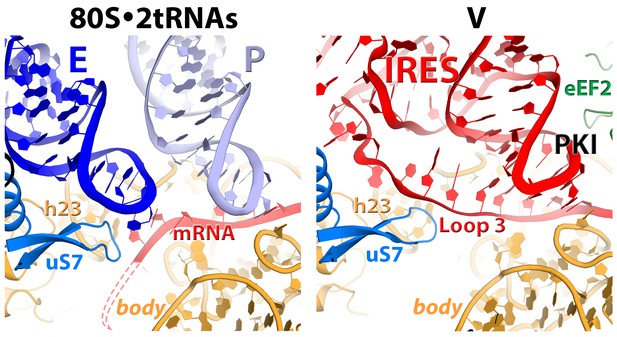
Position and interactions of loop 3 (variable loop region) of the PKI domain in Structure V (this work) resembles those of the anticodon stem loop of the E-site tRNA (blue) in the 80S•2tRNA•mRNA complex (Svidritskiy et al., 2014).
https://doi.org/10.7554/eLife.14874.015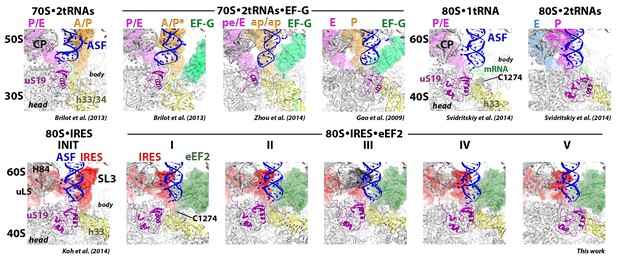
Positions of tRNAs and the TSV IRES relative to the A-site finger (blue, nt 1008–1043 of 25S rRNA) and the P site of the large subunit, comprising helix 84 of 25S rRNA (nt. 2668–2687) and protein uL5 (collectively labeled as central protuberance, CP, in the upper-row first figure, and individually labeled in the lower-row first figure).
Structures of translocation complexes of the bacterial 70S ribosome bound with two tRNAs and yeast 80S complexes with tRNAs are shown in the upper row and labeled. Structures of 80S•IRES complexes in the absence of eEF2 (INIT; PDB 3J6Y, Koh et al., 2014) and in the presence of eEF2 (this work) are shown in the lower row and labeled.
Interactions of the TSV IRES with uL5 and eL42.
Structures of 80S•IRES complexes in the absence of eEF2 (INIT; PDB 3J6Y, Koh et al., 2014) and in the presence of eEF2 (this work) are shown in the upper row and labeled. Structures of the 80S complexes with tRNAs (Svidritskiy et al., 2014) are shown in the lower row in a view similar to that for the 80S•IRES complex.
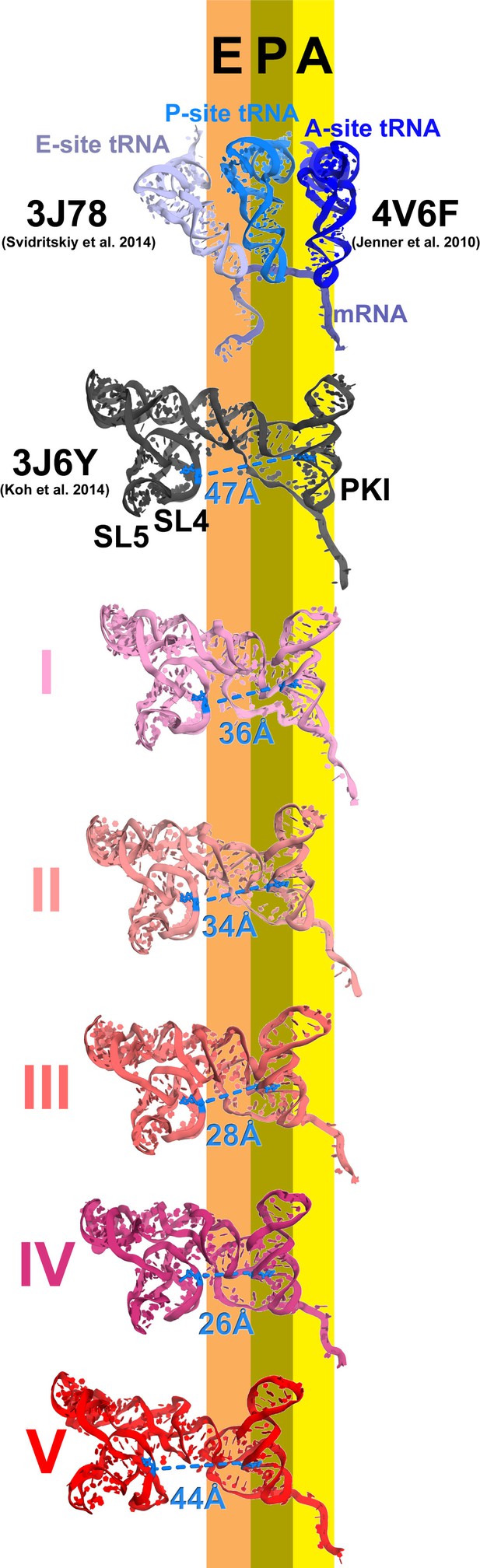
Inchworm-like translocation of the TSV IRES.
Conformations and positions of the IRES in the initiation state and in Structures I-V are shown relative to those of the A-, P- and E-site tRNAs. The view was obtained by structural alignment of the body domains of 18S rRNAs of the corresponding 80S structures. Distances between nucleotides 6848 and 6913 in SL4 and PKI, respectively, are shown (see also Figure 2—source data 1).
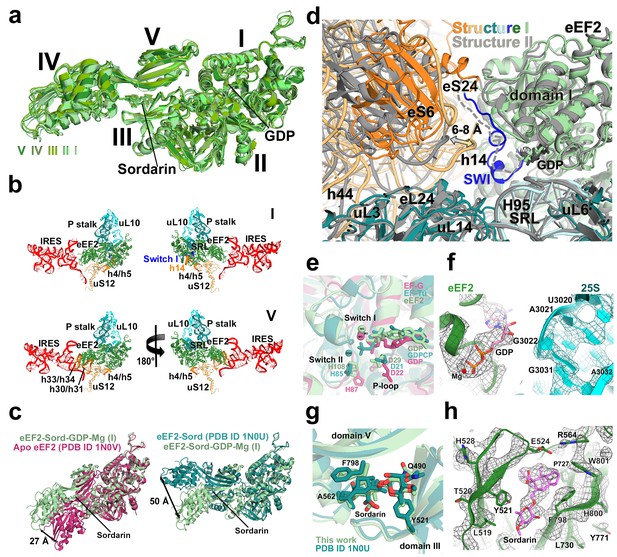
Conformations and interactions of eEF2.
(a) Conformations of eEF2 in Structures I-V and domain organization of eEF2 are shown. Roman numerals denote eEF2 domains. Superposition was obtained by structural alignment of domains I and II. (b) Elements of the 80S ribosome in Structures I and V that contact eEF2. eEF2 is shown in green, IRES RNA in red, 40S subunit elements in orange, 60S in cyan/teal. (c) Comparison of conformations of eEF2•sordarin in Structure I (light green) with those of free apo-eEF2 (magenta) and eEF2•sordarin (teal) (Jorgensen et al., 2003). (d) Interactions of the GTPase domains with the 40S and 60S subunits in Structure I (colored in green/blue, eEF2; orange, 40S; cyan/teal, 60S) and in Structure II (gray). Switch loop I (SWI) in Structure I is in blue; dashed line shows the putative location of unresolved switch loop I in Structure II. Superposition was obtained by structural alignment of the 25S rRNAs. (e) Comparison of the GTP-like conformation of eEF2•GDP in Structure I (light green) with those of 70S-bound elongation factors EF-Tu•GDPCP (teal; Voorhees et al. 2010) and EF-G•GDP•fusidic acid (magenta; fusidic acid not shown; Zhou et al., 2013). (f) Cryo-EM density showing guanosine diphosphate bound in the GTPase center (green) next to the sarcin-ricin loop of 25S rRNA (cyan) of Structure II. (g) Comparison of the sordarin-binding sites in the ribosome-bound (light green; Structure II) and isolated eEF2 (teal; Jorgensen et al., 2003). (h) Cryo-EM density showing the sordarin-binding pocket of eEF2 (Structure II). Sordarin is shown in pink with oxygen atoms in red.
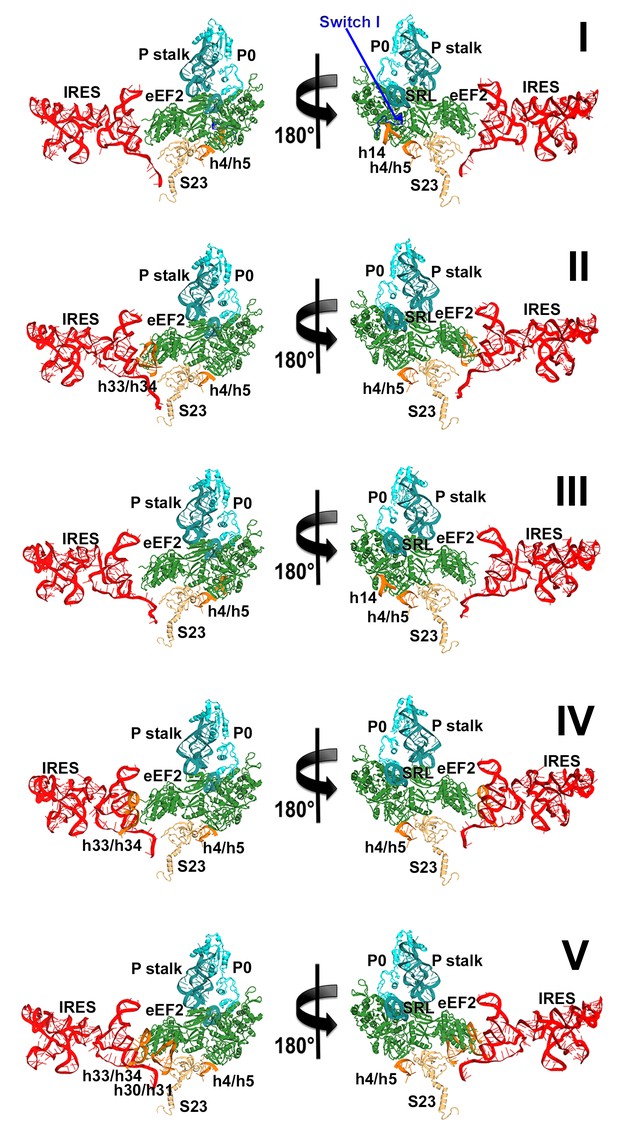
Elements of the 80S ribosome that contact eEF2 in Structures I through V.
The view and colors are as in Figure 5b: eEF2 is shown in green, IRES RNA in red, 40S subunit elements in orange, 60S in cyan/teal.
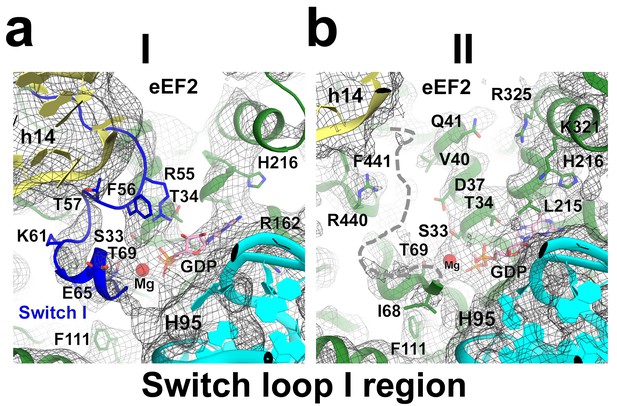
Cryo-EM density of the GTPase region in Structures I and II.
The switch loop I in Structure I is shown in blue. The putative position of the switch loop I, unresolved in the density of Structure II, is shown with a dashed line. Colors for the ribosome and eEF2 are as in Figure 1.
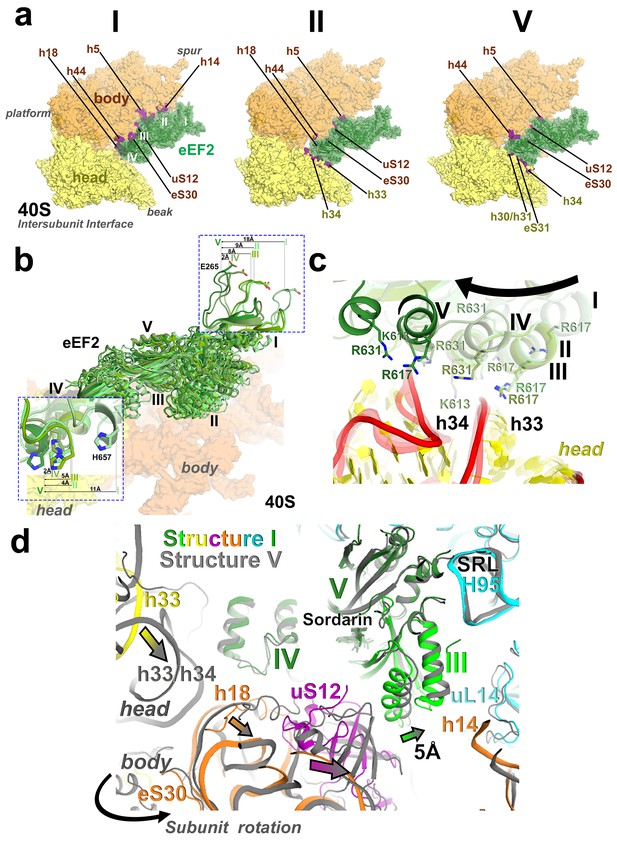
Interactions of eEF2 with the 40S subunit.
(a) eEF2 (green) interacts only with the body in Structure I (eEF2 domains are labeled with roman numerals in white), and with both the head and body in Structures II through V. Colors are as in Figure 1, except for the 40S structural elements that contact eEF2, which are labeled and shown in purple. (b) Entry of eEF2 into the 40S A site, from Structure I through V. Distances to the A-site accommodated eEF2 (Structure V) are shown. The view was obtained by superpositions of the body domains of 18S rRNAs. (c) Rearrangements, from Structure I through V, of a positively charged cluster of eEF2 (K613, R617 and R631) positioned over the phosphate backbone of 18S helices 33 and 34, suggesting a role of electrostatic interactions in eEF2 diffusion over the 40S surface. (d) Shift of the tip of domain III of eEF2, interacting with uS12 upon reverse subunit rotation from Structure I to Structure V. Structure I colored as in Figure 1, except uS12, which is in purple; Structure V is in gray.
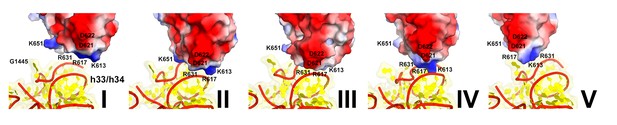
Repositioning (sliding) of the positively-charged cluster of domain IV of eEF2 over the phosphate backbone (red) of the 18S helices 33 and 34.
Structures I through V are shown. Electrostatic surface of eEF2 is shown; negatively and positively charged regions are shown in red and blue, respectively. The view was obtained by structural alignment of the 18S rRNAs.
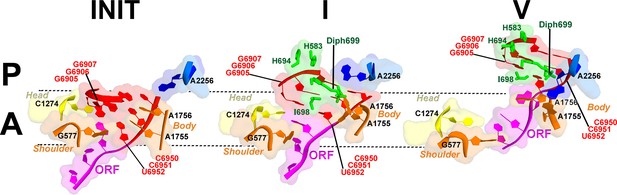
Interactions of the residues at the eEF2 tip with the decoding center of the IRES-bound ribosome.
Key elements of the decoding center of the 'locked' initiation structure (Koh et al., 2014), 'unlocked' Structure I, and post-translocation Structure V (this work) are shown. The histidine-diphthamide tip of eEF2 is shown in green. The codon-anticodon-like helix of PKI is shown in red, the downstream first codon of the ORF in magenta. Nucleotides of the 18S rRNA body are in orange and head in yellow; 25S rRNA nucleotide A2256 is blue. A and P sites are schematically demarcated by dotted lines.
Videos
Animation showing the transition from the initiation 80S•TSV IRES structures (Koh et al., 2014) to eEF2-bound Structures I through V (this work).
Four views (scenes) are shown: (1) A view down the intersubunit space, with the head of the 40S subunit oriented toward a viewer, as in Figure 1a; (2) A view at the solvent side of the 40S subunit, with the 40S head shown at the top, as in Figure 2—figure supplement 1; (3) A view down at the subunit interface of the 40S subunit; (4) A close-up view of the decoding center (A site) and the P site, as in Figure 2g. Each scene is shown twice. Colors are as in Figure 1. In scenes 1, 2 and 3, nucleotides C1274, U1191 of the 40S head and G904 of the 40S platform are shown in black to denote the A, P and E sites, respectively. In scene 4, C1274 and U1191 are labeled and shown in yellow; G577, A1755 and A1756 of the 40S body A site and C1637 of the body P site are labeled and shown in orange.





