Ancient origin of lubricated joints in bony vertebrates
Figures
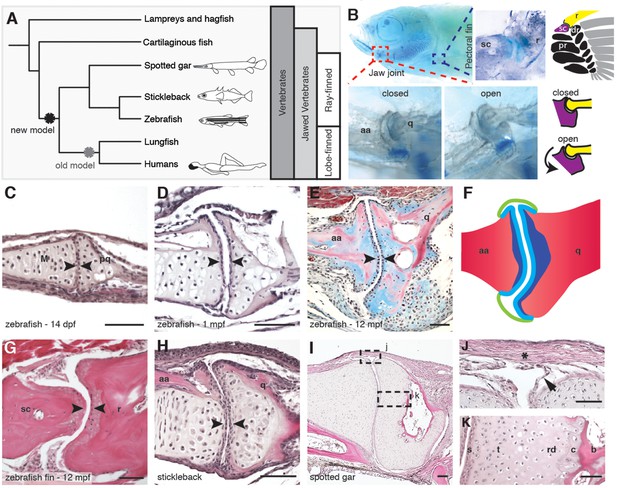
Synovial-like morphology of jaw and fin joints in ray-finned fish.
(A) Phylogenetic tree contrasts the old model of synovial joint evolution (grey asterisk) with the new model of synovial joints evolving in a common precursor of all bony vertebrates (black asterisk). (B) Alcian Blue-stained adult zebrafish and accompanying diagrams show the pectoral fin joints, and jaw joints in open and closed positions. (C–E, G–I) Sections of 14 dpf (n = 4), 1 mpf (n = 3), and adult (n = 6) zebrafish jaw joints; ray-scapula joint in the adult zebrafish pectoral fin (n = 4); and stickleback (1 mpf, n = 3) and spotted gar (10.2 cm, n = 3) jaw joints. Sections are stained by H&E (C, D, G–I) or trichrome (E). Articular chondrocytes (black arrowheads) line the cavity. (F) Schematic of adult jaw joint shows bone (red), cartilage (blue, lighter shade indicates articular), and synovial membrane (green). (J, K) Magnifications of (I) show the synovial membrane (arrow), fibrous capsule (asterisk) and multilayered articular cartilage (K). Scale bar in h, 100 μm; all other panels, 50 μm. aa: anguloarticular; q: quadrate; sc: scapula; r: ray; pr: proximal radial; dr: distal radial; M: Meckel’s; pq: palatoquadrate; s: superficial; t: transitional; rd: radial layer; c: calcified cartilage; b: bone.
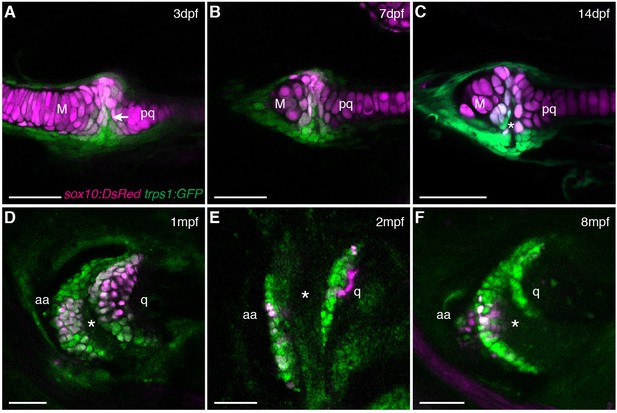
Live imaging of jaw joint cavitation.
(A–F) In double transgenic zebrafish, sox10:DsRed marks all chondrocytes and trps1:GFP labels nascent joint chondrocytes (arrow) and perichondrial cells at 3 and 7 dpf. By 14 dpf, a partial cavity (asterisk) was evident in 1/6 animals. In 3/3 animals at 1 mpf, 4/4 at 2 mpf, and 4/4 at 8 mpf, a fully formed cavity is evident and sox10:DsRed is expressed in a subset of trps1:GFP+ articular chondrocytes. Scale bar, 50 μm.
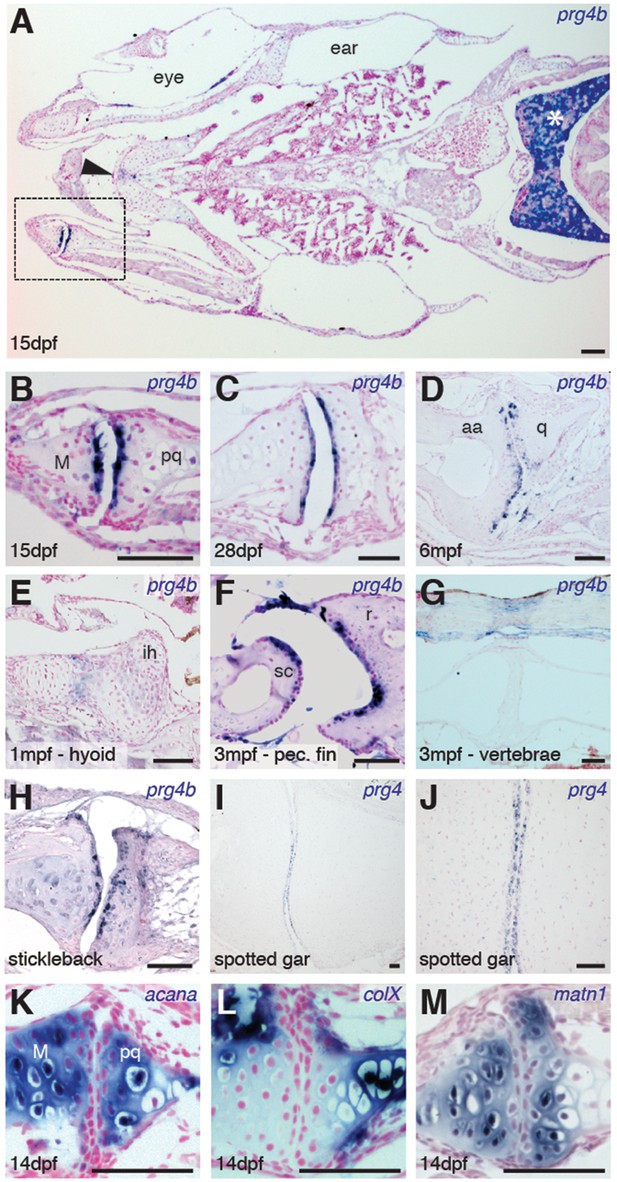
Expression of Prg4 genes in articular chondrocytes of ray-finned fish.
(A–G) prg4b expression in articular chondrocytes of the zebrafish jaw joint (boxed region in A, B–D), hyoid joint (E); ray-scapula joint (F); and vertebral column (G). prg4b is also expressed in the liver (asterisk), possibly in ligaments above the vertebrae, and weakly at the ceratohyal-ceratohyal joint (arrowhead). n = 3 each. (H–J) Expression of stickleback prg4b (1 mpf, n = 3) and gar prg4 (10.2 cm, n = 3) in jaw joint articular chondrocytes (J, magnification of I). (K–M) Exclusion of acana, col10a1, and matn1 expression from articular chondrocytes of the zebrafish jaw. n = 3 each. Scale bar, 50 μm. ih: interhyal. See also Figure 3—figure supplement 1 and 2.
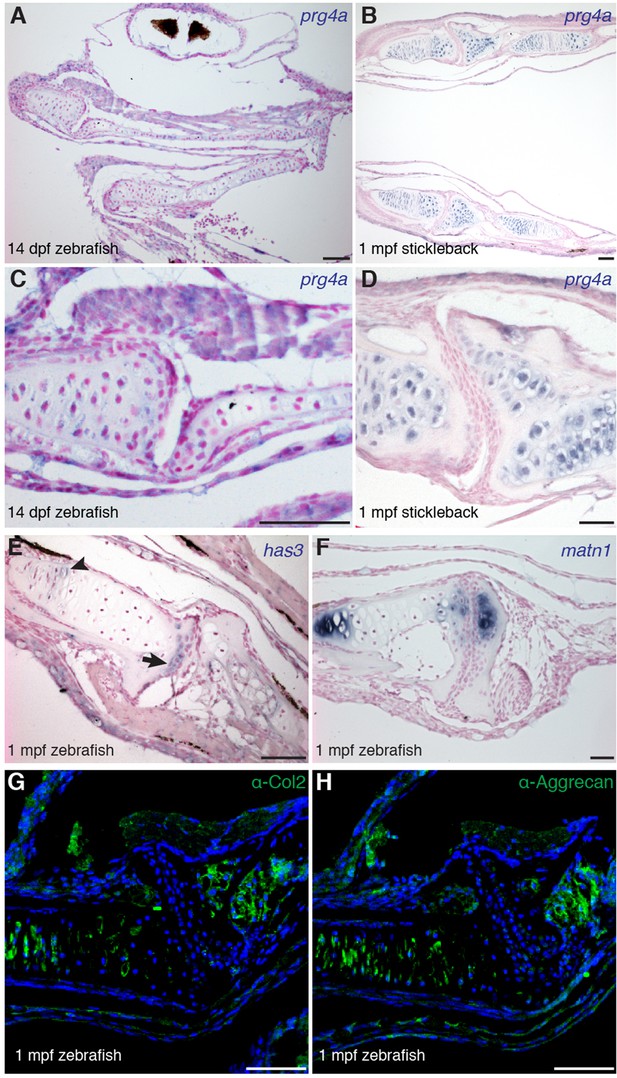
Gene expression within the zebrafish and stickleback jaw joints.
(A–D) In situ hybridization reveals broad chondrocyte expression of prg4a but no enrichment within jaw joint articular chondrocytes (C–D, magnified views). (E) Zebrafish has3 is expressed in chondrocytes just underneath the articular surface (arrow) and in a small number of cells within the growth plate (arrowhead). (F) Expression of matn1 is excluded from superficial chondrocytes of the zebrafish jaw joint at 1 mpf. (G, H) Immunofluorescence staining for Col2a1 and Aggrecan protein reveals broad cartilage expression yet exclusion from jaw articular chondrocytes. n = 3 each panel. Scale bars, 50 μm.
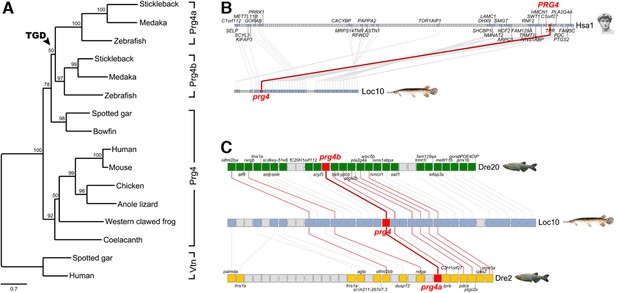
Evolution of vertebrate Proteoglycan 4 (Lubricin).
(A) Maximum likelihood phylogeny of vertebrate Prg4 proteins generated using the LG model in PhyML (Guindon et al., 2009). The tree was rooted with spotted gar and human Vitronectin (Vtn) proteins. In the ray-finned lineage, after divergence of the holostean lineage (spotted gar and bowfin), the teleost genome duplication (TGD) generated Prg4a and Prg4b found in extant teleosts. GenBank/Ensembl accession numbers for sequences in the MAFFT (Katoh and Standley, 2014) alignment: zebrafish: Prg4a, NP_997918, Prg4b, XP_005160745; stickleback: Prg4a, ENSGACP00000019616, Prg4b, concatenate of ENSGACP00000009961/ENSGACP00000009951/ENSGACP00000009946; medaka: Prg4a, ENSORLP00000017289, Prg4b, ENSORLP00000011688; Spotted gar: Prg4, XP_015210531, Vtn, XP_006641129; bowfin: Prg4, AAC_TPR.1.1 [http://phylofish.sigenae.org/]; human: PRG4, NP_005798, VTN, NP_000629; mouse: Prg4, NP_067375; chicken: Prg4, ENSGALP00000008204; Anole lizard: Prg4, XP_008112908; Western clawed frog: Prg4, XP_012817383; coelacanth: Prg4, ENSLACP00000017445. (B) Orthologous pairwise synteny cluster from the Synteny Database (Catchen et al., 2009) (window size: 50 genes) shows extensive conserved synteny of Prg4 gene regions on human chromosome Hsa1 and spotted gar Loc10, supporting their orthology. (C) Composite cluster from the Synteny Database (window size: 50 genes) compares the prg4 gene regions in zebrafish to the TGD outgroup spotted gar. Double conserved synteny of prg4 on gar Loc10 and zebrafish prg4a on Dre2 and prg4b on Dre20, supports the prg4 duplication as result of the TGD.
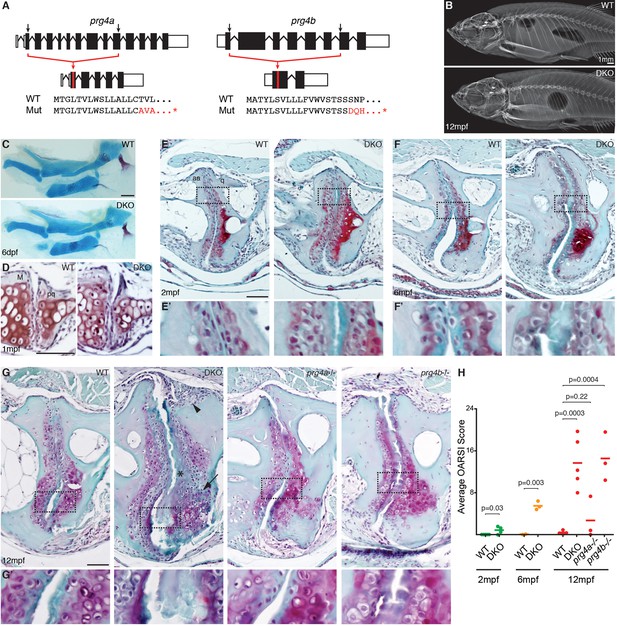
Progressive deterioration of the jaw joint in zebrafish lacking prg4b.
(A) Schematics of prg4a and prg4b TALEN mutants show deleted sequences. (B) X-ray imaging shows no gross morphological bone defects of prg4a-/-; prg4b-/- mutants (DKO) at 12 mpf. (C) Alcian Blue staining shows normal facial cartilages in DKO at 6 dpf. (D) SafraninO staining shows a normal jaw joint between Meckel’s (M) and palatoquadrate (pq) cartilages in DKO at 1 mpf. (E–G) SafraninO staining at 2, 6 and 12 mpf shows increasingly abnormal joints between the jaw anguloarticular (aa) and quadrate (q) bones in DKO. Defects include acellular matrix accumulation in the cavity (asterisks, see magnified regions in E’–G’) at all stages, and synovial hyperplasia (arrowheads) and expanded deep chondrocytes (arrows) at 12 mpf. Single prg4b but not prg4a mutants showed similar jaw joint defects to DKO at 12 mpf. (H) Quantification of jaw joint defects using our modified OARSI system for zebrafish. Scale bar 50 μm, except in (B). See also Figure 4—figure supplements 1,2, and Figure 4—source data 1.
-
Figure 4—source data 1
Quantification of joint defects in zebrafish lacking prg4 genes.
This file includes the average OARSI scores for wild types and mutants related to Figure 4H.
- https://doi.org/10.7554/eLife.16415.010
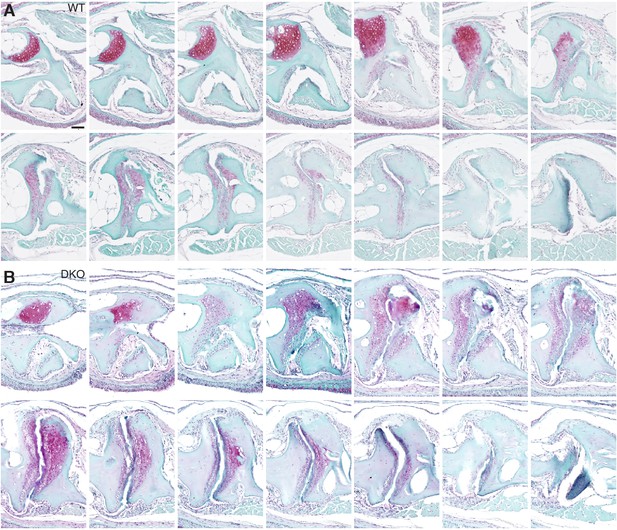
Serial sections through a representative wild-type and prg4a-/-; prg4b-/- mutant jaw joint.
SafraninO staining of a serial series of 5 μm sections were imaged every third section to capture a complete wild-type and prg4a-/-; prg4b-/- (DKO) jaw joint at 12 mpf. See also Figure 4G. Scale bar, 50 μm.
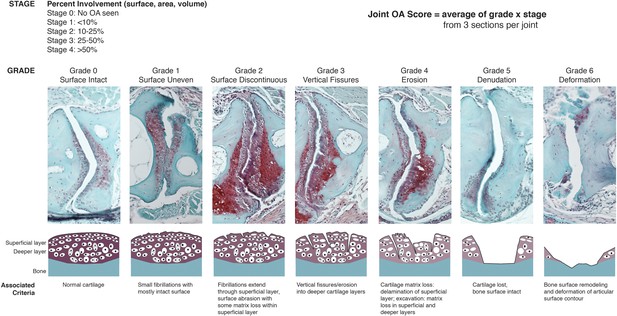
OARSI scoring system for zebrafish.
SafraninO-stained histological sections demonstrate the defining features for each grade of joint damage at the zebrafish jaw. Grade 0 – a smooth intact surface with normal superficial and deeper cartilage layers (12 mpf, wild-type). Grade 1 – an uneven surface cartilage with small fibrillations limited to the superficial layer (12 mpf, wild-type). Grade 2 – the superficial layer is disrupted with focal fibrillations and some matrix loss (12 mpf, DKO). Grade 3 – vertical clefts or fissures extend beyond the superficial layer, disrupting the deeper cartilage (12 mpf, DKO). Grade 4 – erosion of the superficial layer of cartilage with matrix loss extending into deeper cartilage layers (12 mpf, DKO). Grade 5 – cartilage is completely lost, exposing the bone surface (12 mpf, prg4b-/-). Grade 6 – the exposed bone surface is deformed, showing altered contour of the joint (12 mpf, DKO). Individual grade and stage values were determined for both the anterior and posterior sides of the left and right jaw joints from three histological sections per joint. Overall joint OA scores were generated as an average of (grade) x (stage) for each animal.
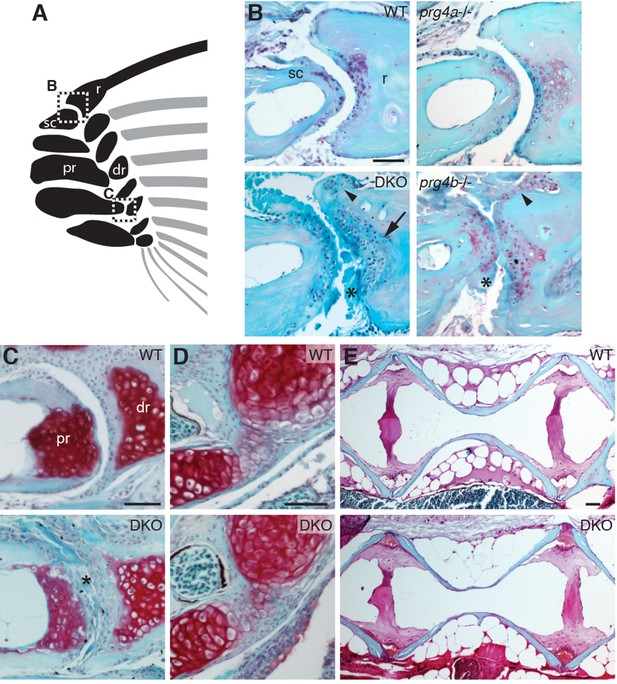
Requirement of Prg4 gene function for fin joints of zebrafish.
(A) Schematic of pectoral fin joints. (B–E) SafraninO staining at 12 mpf shows abnormal joints between the ray (r) and scapula (sc) joint of the pectoral fin (B) in prg4b but not prg4a single mutants and prg4a; prg4b double mutants (DKO), and defects in the proximal radial (pr) and distal radial (dr) joint of the pectoral fin (C) in DKO. Defects include acellular matrix accumulation in the cavity (asterisks), synovial hyperplasia (arrowheads), and expanded deep chondrocytes (arrow). In contrast, the hyoid joint (D) and intervertebral discs (E) were normal in DKO. Phenotypes were consistently observed in three animals of each genotype. Scale bar 50 μm.
Videos
Bone μCT of the adult zebrafish head.
3D reconstruction of bone shows the structure and relative position of the jaw joint within the adult zebrafish head (12 mpf). Color-coding at the end of the movie illustrates the anguloarticular (pink) and quadrate (yellow) bones that articulate at the jaw joint.
Additional files
-
Supplementary file 1
DNA sequences for in situ hybridization probes and genotyping.
(1A) Forward and reverse primers used to amplify zebrafish cDNA for in situ hybridization probe generation. (1B) Sequences used to generate in situ hybridization probes for stickleback and spotted gar prg4 genes. (1C) Forward and reverse primers used to genotype prg4a and prg4b mutant zebrafish.
- https://doi.org/10.7554/eLife.16415.014





