A network of epigenetic modifiers and DNA repair genes controls tissue-specific copy number alteration preference
Figures
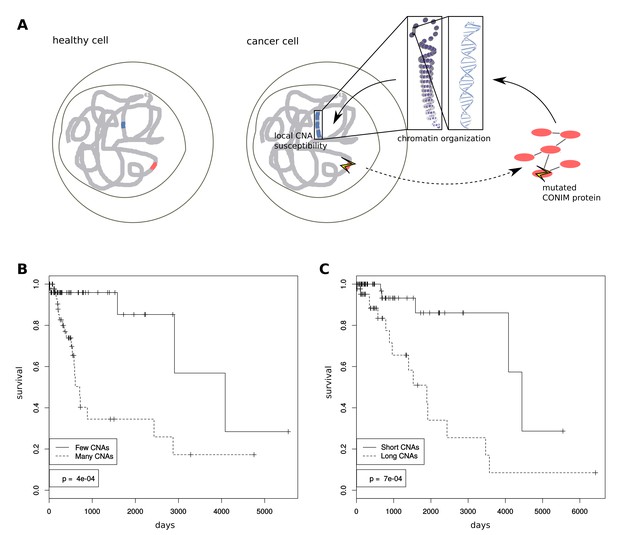
Mechanisms of CNA number modulation and clinical importance.
(A) Schematic showing how CONIM gene mutations can result in a higher or lower CNA number. (B) We performed Kaplan-Meier statistics on data from lower grade glioma (LGG) patients with deviating CNA numbers and lengths. LGG patients with fewer CNAs have a significantly better survival prognosis as compared to patients with many CNAs. (C) LGG patients with shorter CNAs have a significantly better survival prognosis when compared to patients with longer CNAs.
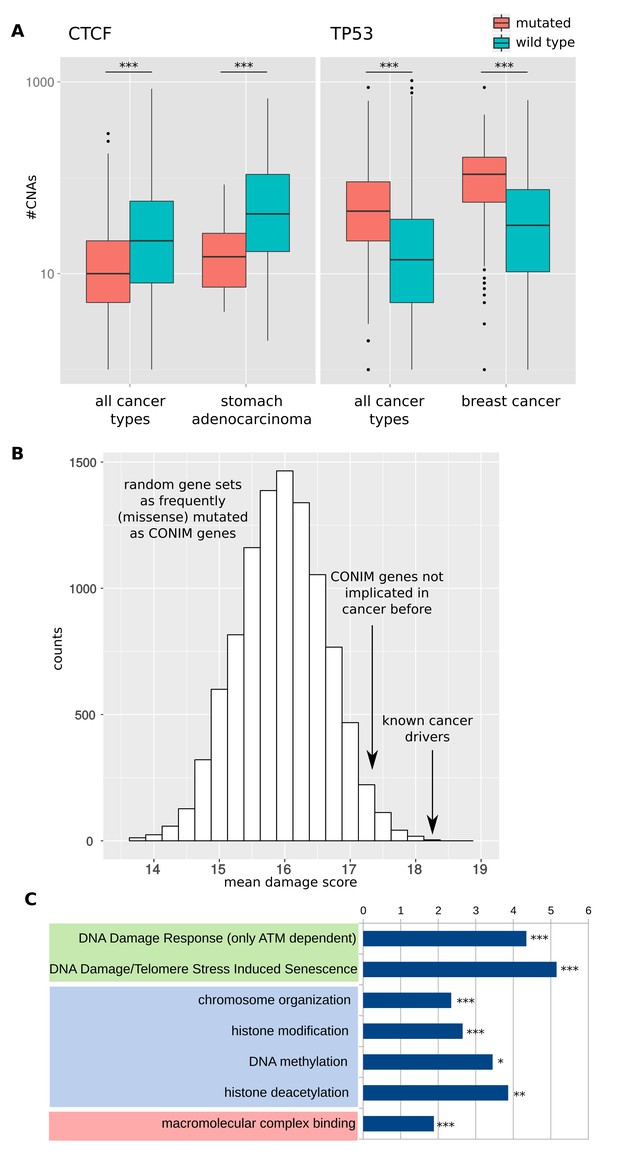
Detection and functional properties of CONIM genes.
(A) CNA numbers in samples in which CTCF (left box) or TP53 (right box) are mutated versus samples in which the respective gene is not mutated. The CNA number distributions are shown for all cancers types (left whiskers within each box) and for a single cancer type (right whiskers within each box). (B) Mutations in CONIM genes tend to have a higher functional impact than mutations found in genes with an equal mutation frequency. Even CONIM genes not previously reported (Lawrence et al., 2014) to be frequently mutated in cancer tend to host mutations with a higher functional impact score (mean 17.23) as compared to random gene sets having matched mutation numbers (p = 0.029; randomisation test). For comparison, the most frequently mutated cancer-driver genes have a mean score of 18.31. (C) The functional categories most significantly overrepresented among the CONIM genes are shown. Among the most highly enriched categories are several terms related to DNA damage (green), chromatin organisation (blue) and complex formation (red). Significance levels are indicated as follows: **q < 0.01, ***q < 0.001.
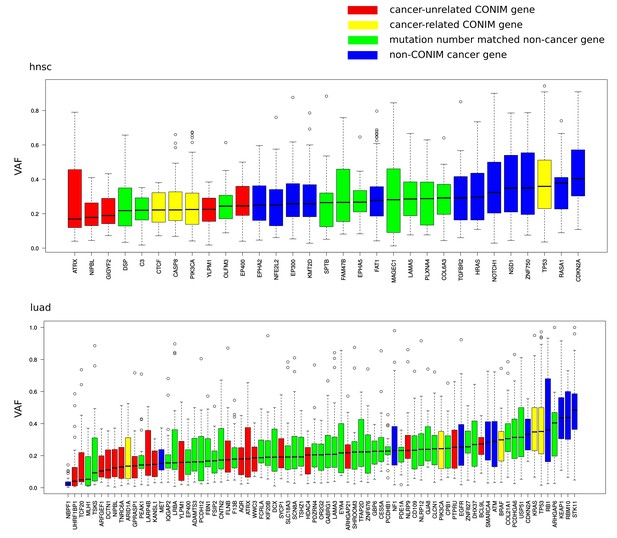
Variant allele fractions (VAFs) of different gene groups.
VAFs of mutations in CONIM genes that have not previously been implicated in cancer (red) are compared to those of a random set of equally often mutated genes (green). Additionally, known cancer driver genes are shown in yellow (CONIM) and blue (non-CONIM). Out of the five cancer types tested, two (hnsc, luad) have non-cancer CONIM genes that are associated with significantly lower VAFs as compared to random genes and cancer drivers. Non-cancer CONIM genes were not associated with significantly higher VAFs in any of the tested cancer types.
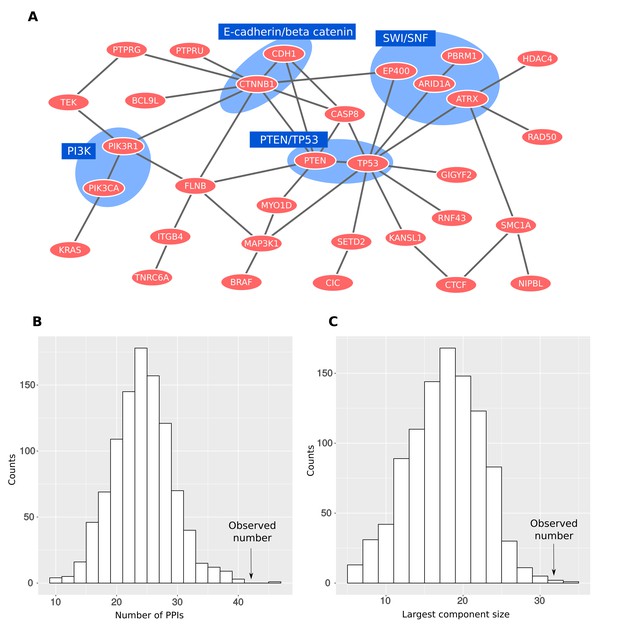
CONIM proteins form a dense network.
(A) All interactions between CONIM proteins are shown. A total of 32 CONIM proteins are connected to each other via 42 physical interactions. Several complexes are highlighted. (B) The observed number of PPIs between CONIM proteins is greater than that for randomly sampled networks of proteins forming as many interactions as the CONIM proteins (p = 0.001; randomisation test). (C) Using the same network randomisation approach, we establish that the size of the largest connected component exceeds random expectation (p = 0.003).
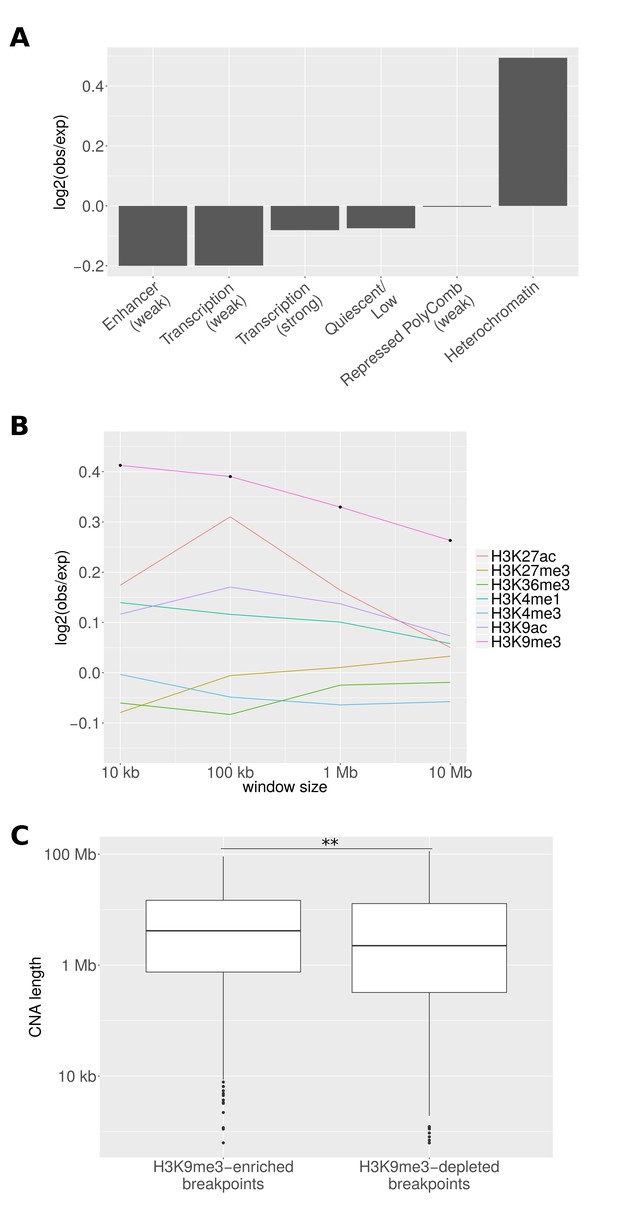
Epigenetic properties of CNA breakpoint regions.
(A) Ratio of the number of breakpoints falling into different chromatin regions in tissues where the CNA event is significantly recurrent to the number in other tissues. States coinciding at least 100 times with breakpoints in non-associated tissues are shown. The number of CNA breakpoints in 'Heterochromatin' is significantly enriched (p = 0.009; chi-square test). (B) The average fraction of genomic windows centering on CNA breakpoints that is associated with different histone marks is compared between tissues where the CNA region drives cancer (observed) and other tissues (expected). Black dots represent bin sizes with significant enrichment (Bonferroni-corrected p < 0.05; Mann-Whitney-Wilcoxon test). (C) CNAs originating from 343 H3K9me3-enriched breakpoints are significantly longer than those originating from 738 H3K9me3-depleted breakpoints (**p < 0.01; Mann-Whitney-Wilcoxon test; 10 kb window).
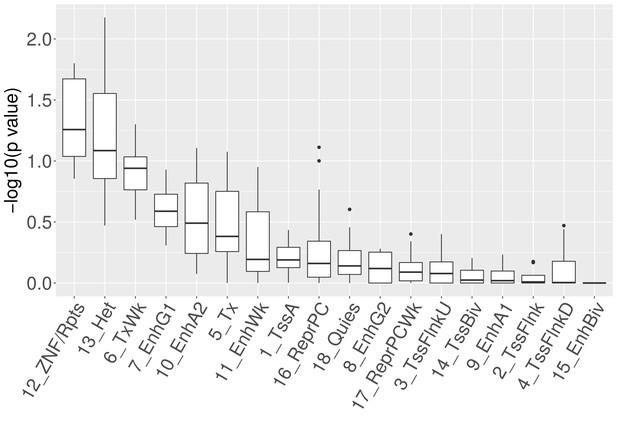
Enrichment of chromatin states at breakpoints for different cell-of-origin associations.
For the original and alternative selections of reference epigenomes, the states 'ZNF genes and repeats' (12_ZNF/Rpts) and ‘Heterochromatin’ (13_Het) show the most significant enrichments [chi-square test; chromatin state labels according to 18-state model (Kundaje et al., 2015).
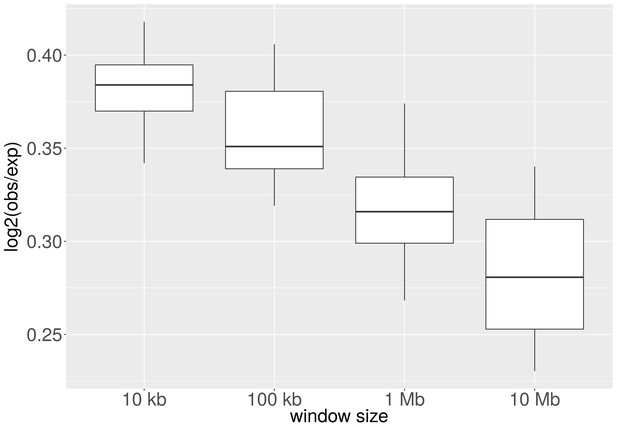
Enrichment of H3K9me3 for different cell-of-origin associations.
Independently of the reference epigenome selection, H3K9me3 at CNA breakpoints is enriched in tissues where the CNA event is recurrent as compared to other tissues (p < 0.003; Mann-Whitney-Wilcoxon test).
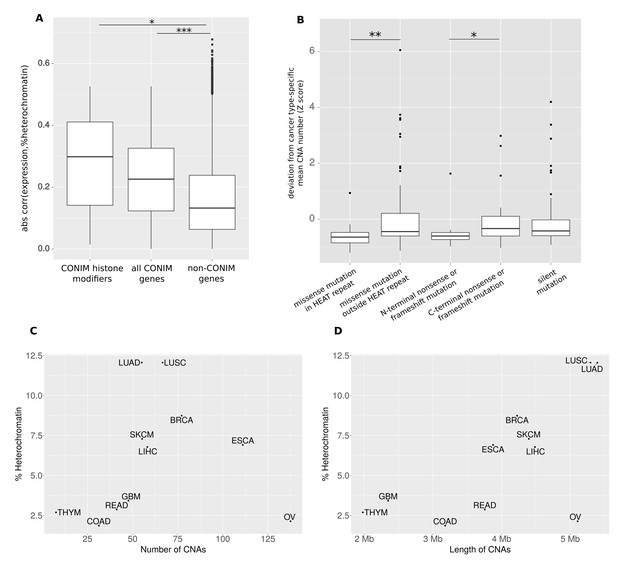
CONIM genes modify the CNA amount via the epigenome.
(A) The absolute correlation between heterochromatin amount and expression of either CONIM histone modifiers or all CONIM genes is significantly larger than that of non-CONIM genes. (B) In the NIPBL gene, nonsense or frameshift mutations in the N-terminal third of the protein, and missense mutations in the HEAT repeat, have a stronger effect on the CNA number in the respective samples than do those mutations that have a smaller effect on protein structure and function. The average (C) CNA number and (D) CNA length per cancer type is correlated with the percentage of heterochromatin in the associated healthy tissue. Significance levels are indicated as follows: *: q < 0.05, **: q < 0.01, ***: q < 0.001.
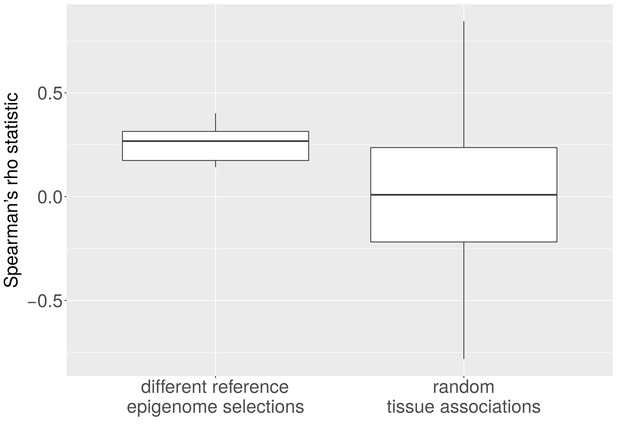
Average CNA number and heterochromatin percentage for alternative reference epigenomes.
The Spearman correlation between the average number of CNAs per cancer type and the heterochromatin percentage in the tissue-of-origin is significantly larger when all possible combinations of reference epigenomes are considered than for 1,000 randomly sampled associations between cancer types and healthy tissues (p < e-10; Mann-Whitney-Wilcoxon test).
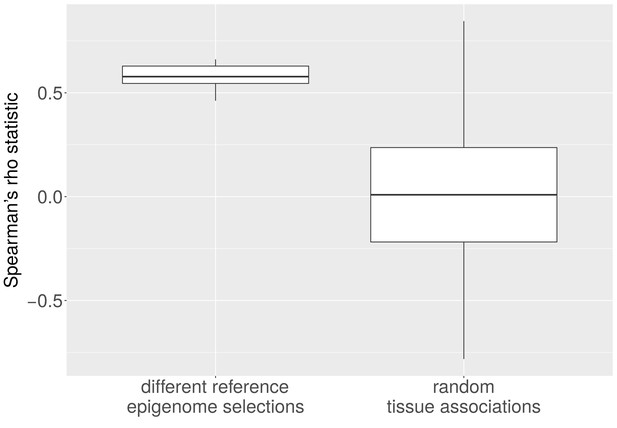
Average CNA length and heterochromatin percentage for alternative reference epigenomes.
The Spearman correlation between average length of CNAs per cancer type and heterochromatin percentage in the tissue-of-origin is significantly larger when all possible combinations of reference epigenomes are considered than for 1,000 randomly sampled associations between cancer types and healthy tissues (p < e-16; Mann-Whitney-Wilcoxon test).
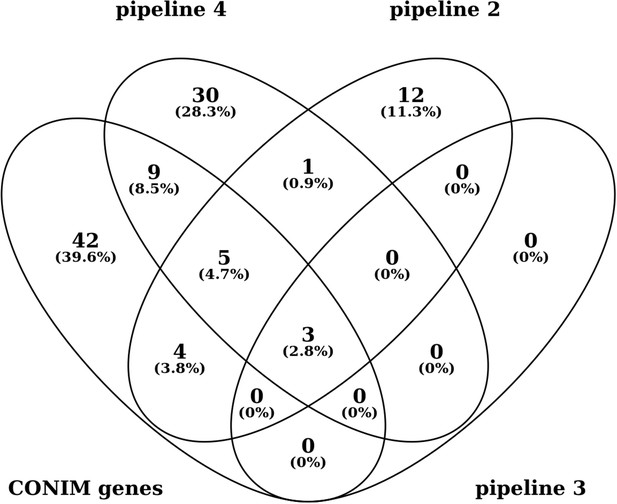
Additional files
-
Supplementary file 1
All CONIM proteins.
- https://doi.org/10.7554/eLife.16519.013
-
Supplementary file 2
Association of cancer types to tissues of origin.
- https://doi.org/10.7554/eLife.16519.014





