Quantifying β-catenin subcellular dynamics and cyclin D1 mRNA transcription during Wnt signaling in single living cells
Figures
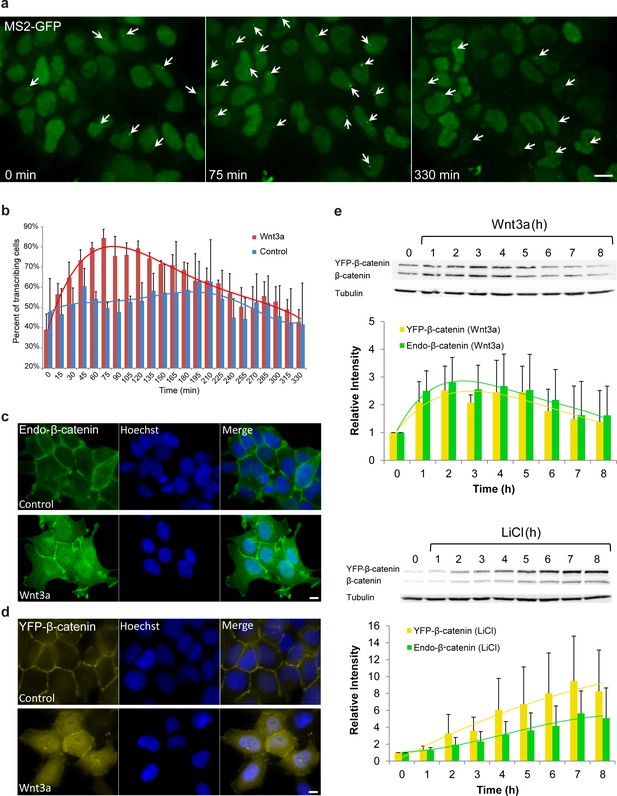
Cell system for following β-catenin intra-cellular dynamics and CCND1 transcription in single living cells.
(a) CCND1-MS2 HEK293 cells stably expressing MS2-GFP-CP were treated with Wnt3a and followed for 6 hr (every 15 min). Several frames from Video 1 are presented. The number of cells exhibiting transcriptionally active CCND1-MS2 genes (green dot in the nucleus, white arrow) was counted over time. Scale bar, 10 µm. (b) Plots showing the percentage of cells in the population with actively transcribing CCND1-MS2 genes in Wnt3a-treated (red, n = 98) and mock treated (blue, n = 128) cells. Mean ± sd from three fields imaged on different days—see Supplementary file 1a for statistics. (c) Immunofluorescence showing that endogenous β-catenin (green) is prominent at the cell membrane in untreated HEK293 cells (top) and accumulates in the cytoplasm and nucleus following activation by Wnt3a for 2 hr (bottom). Hoechst DNA stain is in blue. (d) Similar changes in the subcellular distribution following activation are seen in the YFP-β-catenin low-expressing clone. Bar = 10 μm. (e) Western blot time course of endogenous β-catenin and YFP-β-catenin protein accumulation following either Wnt3a (top) or LiCl (bottom) stimulation. Anti-β-catenin antibody was used for the detection of both β-catenin proteins. Tubulin was used as a loading control. Time 0 is the time point of activator addition. Blots are representative of 3 repeated experiments. The average quantification of 3 repeated experiments is presented in the plots below (mean ± sd). There is no statistical difference between the endogenous and exogenous levels of β-catenin in the two plots.
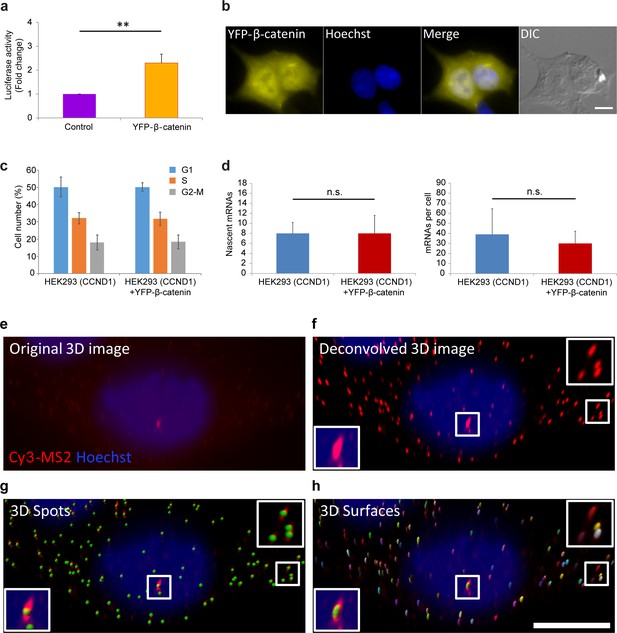
Measuring the effect of YFP-β-catenin expression in HEK293 cells.
(a) Luciferase assay showing the levels of cyclin D1 promoter activation following the transient transfection of YFP-β-catenin into HEK293 cells. p=0.003. (b) Overexpression of YFP-β-catenin shows that overexpressed protein localization does not resemble endogenous β-catenin under non-activated conditions, since it is highly present in the nucleus prior to Wnt activation, and does not appear at the membrane. Bar = 10 µm. (c) Cell cycle analysis of HEK293 CCND1-MS2 cells with and without YFP-β-catenin. (d) Quantification of CCND1-MS2 nascent mRNAs (left) (p=0.8) and cellular mRNAs (right) (p=0.16) levels by RNA FISH in HEK293 CCND1-MS2 cell clones with or without YFP-β-catenin (n = 18, n = 26 respectively). **p<0.01, n.s. = p>0.05. (e–h) Example of single molecule mRNA FISH quantification procedure with a probe that hybridizes to the MS2 region in the 3’UTR of the CCND1-MS2 mRNA. (e) Raw 3D image (76 planes in z stack) showing the active transcription site (red) and single mRNA molecules. Hoechst DNA stain is in blue. (f) Deconvolved 3D image. Boxes show the transcription site (bottom) and single cellular mRNAs (top). (g) Identification of ‘spots’ of single mRNAs and transcription site (green dots) by Imaris. (h) Generation of a 3D shell for each spot to be taken for intensity measurements. Bar = 10 µm. Then the sum of intensity at the transcription site was divided by the frequent intensity value of a single mRNA. This ratio provided the number of mRNAs associated with the transcription unit, as explained in the Materials and methods section.
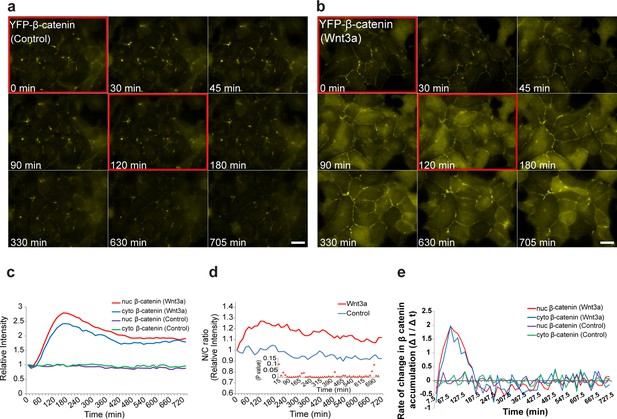
The dynamics of β-catenin accumulation following Wnt3a activation in cell populations.
Frames from live-cell movies (Video 2) showing YFP-β-catenin dynamics in cells treated with (a) mock conditioned medium or (b) Wnt3a for 12 hr. Red bordered frames compare between the 0 min and 120 min time points. Bar = 20 μm. (c) The relative average intensity of β-catenin measured in the cytoplasm (n = 24) and nucleus (n = 31) of cells treated with Wnt3a for 12 hr, compared to mock-treated control cells (n = 13). (d) Nucleus to cytoplasm ratio (N/C) of fluorescence intensities over 12 hr from c. The initial ratio was designated as 1. Inset plot shows the statistical significance p values (t test) at each time point between the two treatments over the experiment time course. (e) The rate of change in β-catenin levels (ΔI/Δt), during accumulation or degradation, in the cytoplasm and nucleus over time in cells from c.
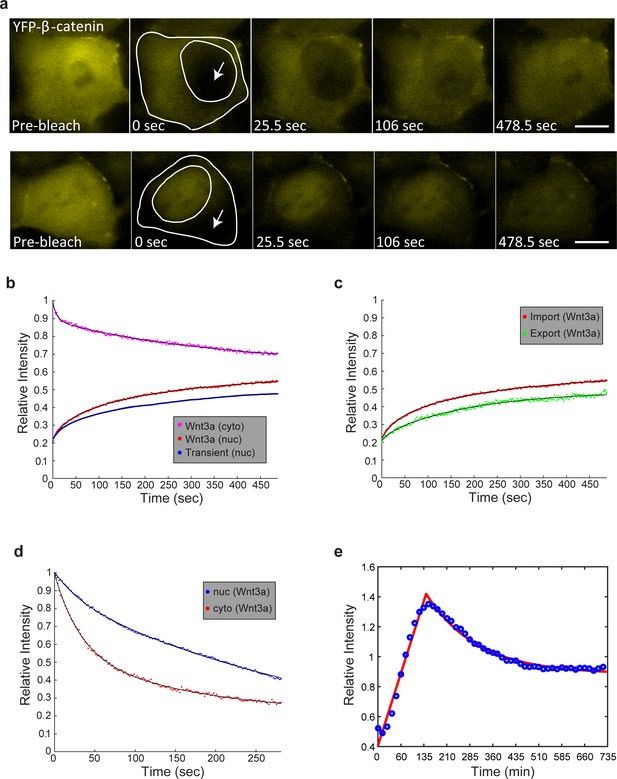
FRAP and FLIP measurements of YFP-β-catenin import and export dynamics.
(a) Frames showing one pre-bleach frame, the bleach of the YFP-β-catenin in the nucleus (top) or cytoplasm (bottom) of a Wnt3a-treated cell (2 hr, arrows point to bleached region), and frames following the recovery of signal over time (frame every 1.5 s for 8 min). Bar = 10 μm. (b) Averaged data plot of FRAP recovery import curves from Wnt3a-treated cells (n = 27, red curve), and transiently overexpressing YFP-β-catenin cells (n = 33, blue curve). Pink curve shows the decline in YFP-β-catenin in the cytoplasm of Wnt3-treated cells concurrent with nuclear import (red curve). (c) Averaged data plot of FRAP recovery export curves (green) from Wnt3a-treated cells (n = 14), compared to the import curve (red). (d) FLIP curves for Wnt3a-treated cells photobleached continuously in the nucleus to show import rates from the cytoplasm (n = 15, red curve) compared with cells photobleached continuously in the cytoplasm to show export rates from the nucleus (n = 16, blue curve). Statistics can be found in Supplementary file 1. (e) Data from Figure 2c (blue dots) of nuclear YFP-β-catenin accumulation were fit with a two-phase exponential (red curve).
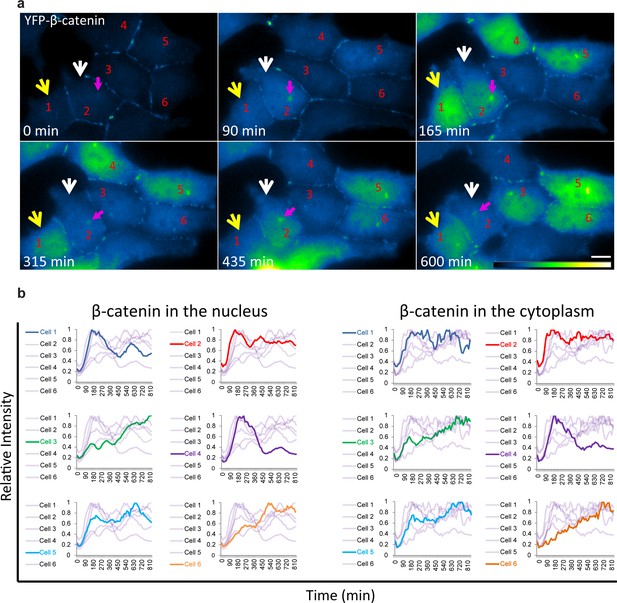
Variability of β-catenin accumulation dynamics following Wnt3a activation in individual cells.
(a) Frames from time-lapse Video 3 showing YFP-β-catenin accumulation in a population of cells. The YFP signal is pseudo-coloured using ImageJ ‘Green Fire Blue’ look-up table. White and yellow arrows point to cells in which β-catenin levels increase and decrease twice during the movie. The pink arrow points to centrosomal accumulation. Bar = 10 μm. (b) β-catenin levels in the nucleus (left) and cytoplasm (right) in individual cells (as numbered in a) are plotted in different colors. The grey background plots show the complete set of plots from all the cells. Maximum β-catenin intensity in each cell was normalized to 1.
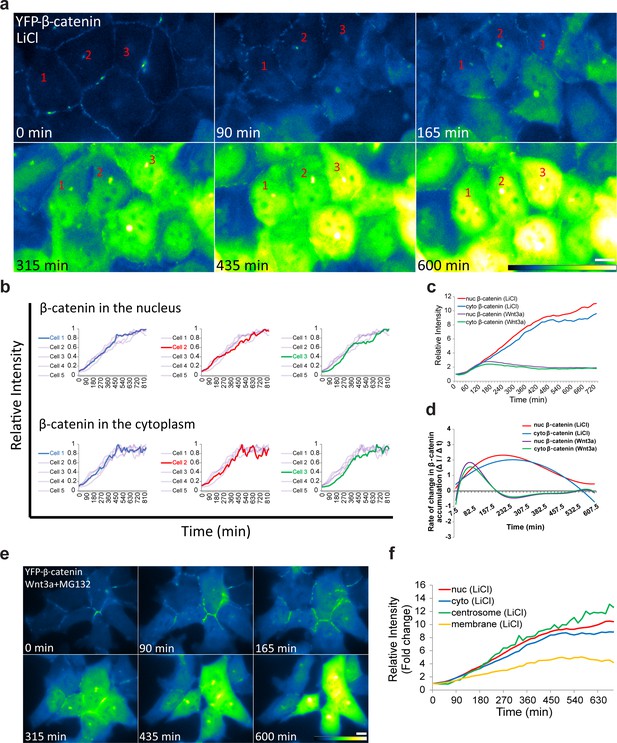
β-catenin accumulation dynamics in response to LiCl activation in individual cells.
(a) Frames from time-lapse Video 4 showing YFP-β-catenin accumulation in a population of cells. The YFP signal is pseudo-colored using the ImageJ ‘Green Fire Blue’ look-up table. Bar = 10 μm. (b) β-catenin levels in the nucleus (top) and cytoplasm (bottom) of individual cells (as numbered in a) are plotted in different colors. The grey background plots show the complete set of plots from all the cells. The maximum intensity of β-catenin in each cell was normalized to 1. (c) The relative average intensity of β-catenin measured in the cytoplasm (n = 17) and nucleus (n = 18) of individual cells treated with LiCl for 12 hr, compared to Wnt3a-treated cells (from Figure 2). (d) The rate of change in β-catenin levels (ΔI/Δt) accumulation or degradation in the cytoplasm and nucleus over time in cells from c. (e) Frames from a time-lapse movie showing YFP-β-catenin accumulation in a population of Wnt3a+MG132-treated cells. The YFP signal is pseudo-colored using the ImageJ ‘Green Fire Blue’ look-up table. Bar = 10 μm. (f) The relative average intensity of β-catenin measured in the centrosome, membrane, cytoplasm and nucleus of LiCl-treated cells.
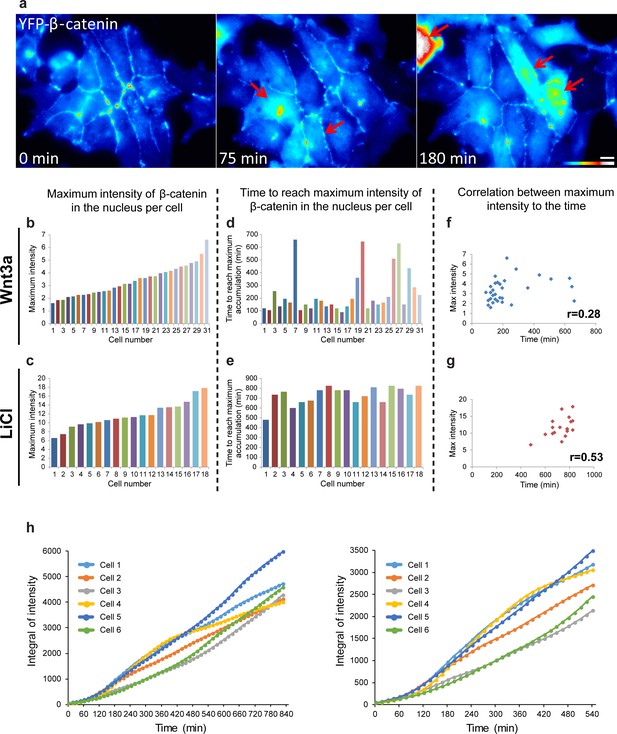
The relationship between YFP-β-catenin levels of accumulation and time of Wnt3a activation.
(a) Frames from a time-lapse movie showing YFP-β-catenin accumulation in a population of cells in the field. The YFP signal is pseudo-colored using the ImageJ ‘Royal’ look-up table. Red arrows point to cells with very high β-catenin levels. Bar = 10 μm. Plots showing the relative maximal levels of β-catenin measured in the nuclei of (b) Wnt3a-treated (n = 31) or (c) LiCl-treated cells (n = 18). Order of cells is according to increasing relative intensities. Plots showing the time from the addition of the activator until reaching the maximal levels of β-catenin in the same set of (d) Wnt3a-treated or (e) LiCl-treated cells. (f,g) The respective correlation plots and scores for a Pearson correlation analysis between the maximum intensity in each cell and the time to reach the highest accumulation. (h) The integral of the fluorescence values in the six cells (from Figure 3) showing the total accumulation levels over time during the whole observation period (left). The right-hand plot shows the differences between accumulation in the cells at earlier times.
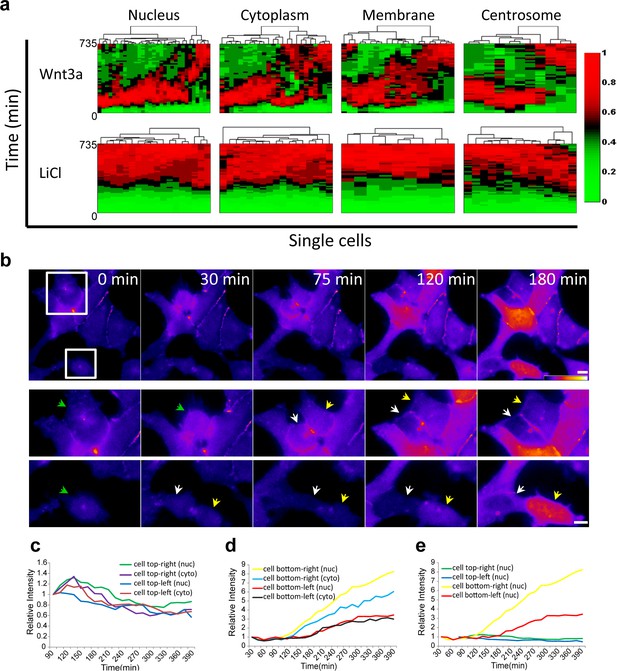
Variability of β-catenin dynamics in the cell population and during the cell cycle.
(a) Heat map and cluster analysis of normalized β-catenin accumulation dynamics in sub-cellular compartments following Wnt3a (top, n(nucleus) = 31, n(cytoplasm) = 24, n(membrane) = 21, n(centrosome) = 11) or LiCl (bottom, n(nucleus) = 18, n(cytoplasm) = 17, n(membrane) = 9, n(centrosome) = 14) treatments. Data were taken from live-cell movies with each column representing one cell, and rows representing time from Wnt addition. Relative levels of β-catenin are depicted from low (green) to high (red). Hierarchical cluster analysis depicted above the plots shows the homogenous behavior in LiCl-treated cells and heterogenous behavior in Wnt3a-treated cells. Most cells reach maximal levels of β-catenin within 2–3 hr. (b) (Top) Frames from time-lapse Video 5 showing YFP-β-catenin accumulation in a population of cells. The YFP signal is pseudo-colored using the ImageJ ‘Fire’ look-up table. Boxes denote cells that go through mitosis, and enlargements are shown below. Green arrows point to mother cells, and yellow and white arrows point to the daughter cells. Bar = 10 μm. Plots showing the relative intensity levels of YFP-β-catenin in the cytoplasm and nucleus of the (c) top and (d) bottom daughter cells of each cell division. (e) Plot comparing the relative intensity levels in the nuclei of the four daughter cells.
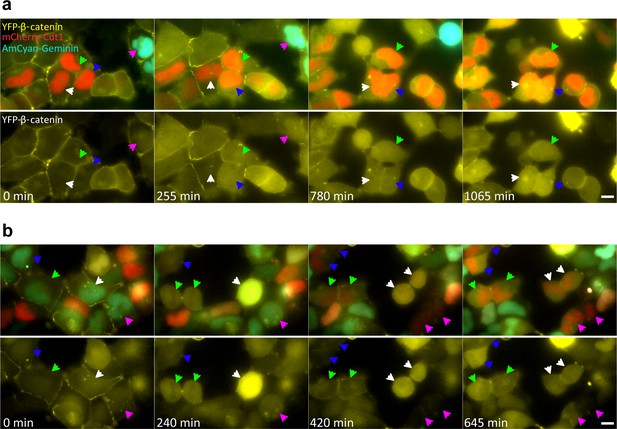
YFP-β-catenin dynamics during the cell cycle in Wnt3a induced cells.
HEK293 CCND1-MS2 YFP-β-catenin cells were stably infected with the Fucci system (mCherry-Cdt1 and AmCyan1-Geminin). Cdt1 levels peak during G1 (red cells), and as cells transition into S, Cdt1 levels decline and Geminin levels rise (cyan cells), remaining high from G2 onwards. (a) Frames from Video 6. Before Wnt3a treatment (time 0 min), β-catenin levels are low in the cytoplasm and the nucleus of all cells marked by arrowheads (cells in G1 or G2). At time 255 min after Wnt3a, the cells marked with blue, green and pink arrowheads show an increase in the β-catenin levels in response to Wnt signaling. The cell marked with a white arrowhead has not responded yet. At time 780 min, the cell marked with a pink arrowhead has gone through mitosis and the three cells marked by blue, green and white arrowheads have similar β-catenin levels. At time 1065 min, the cells marked by white and blue arrowheads are increasing further, while the cell marked with the green arrowhead is not changing. (b) Frames from Video 7. Before Wnt3a treatment (time 0 min), all the cells marked with arrowheads (green, blue, white and pink) are in G2. At time 240 min after Wnt3a, the cell marked by a green arrowhead has gone through mitosis and the daughter cells have similar β-catenin levels. At time 420 min, all four cells have gone through mitosis. β-catenin levels in each of the two daughter cells in all four cases are similar to each other. At time 705 min, the daughter cells marked by blue, green and pink arrowheads are different from each other. The cells marked with white arrowheads have similar levels. Bottom rows are the same frames without Fucci labels. Bar = 10 μm.
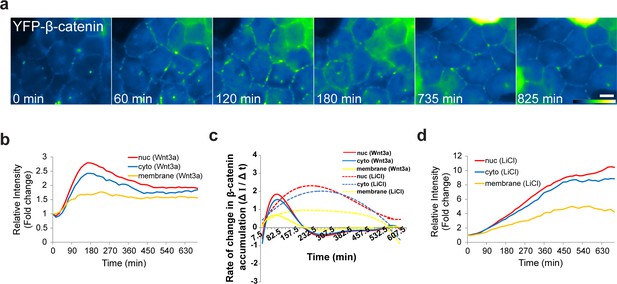
The dynamics of β-catenin accumulation at the membrane following Wnt3a activation.
(a) Frames from time-lapse Video 8 showing YFP-β-catenin accumulation at the cell membrane. The YFP signal is pseudo-colored using the ImageJ ‘Green Fire Blue’ look-up table. Bar = 10 μm. (b) The relative average intensity of β-catenin measured in the membrane (n = 21), cytoplasm and nucleus (from Figure 2) of Wnt3a-treated cells. (c) The rate of change in β-catenin levels (ΔI/Δt) accumulation or degradation in the membrane, cytoplasm and nucleus over time in Wnt3a- and LiCl-treated cells. (d) The relative average intensity of β-catenin measured in the membrane, cytoplasm and nucleus of LiCl-treated cells.
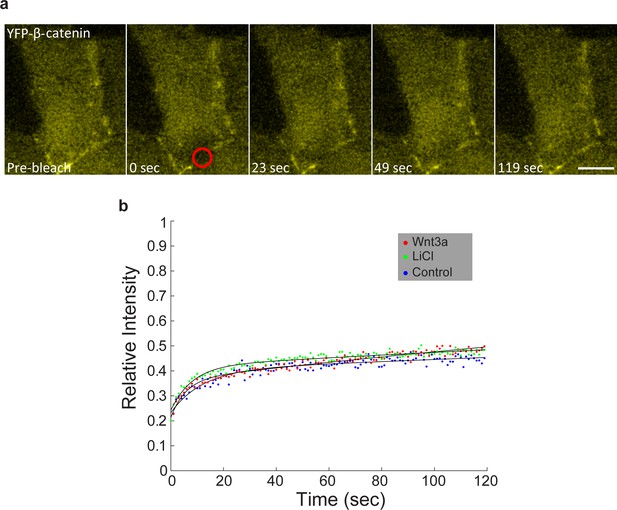
FRAP measurements of YFP-β-catenin dynamics at adherens junctions.
(a) Frames showing one pre-bleach frame, the bleach of the YFP-β-catenin in the membrane region of a Wnt3a-treated cell, and frames following the recovery of the signal over time. Circle denotes the bleached region. Bar = 10 μm. (b) Averaged data plot of FRAP recovery curves from mock-treated (control, n = 21), Wnt3a-treated (n = 32) and LiCl-treated cells (n = 18). Statistics can be found in Supplementary file 1e,f.
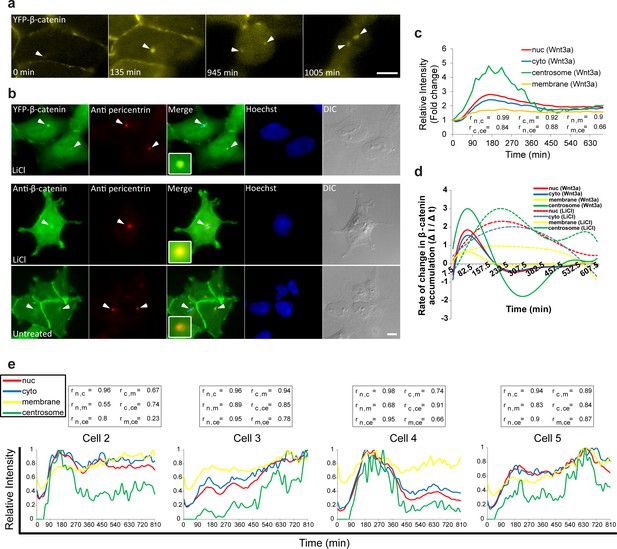
Accumulation of β-catenin at the centrosome after Wnt3a activation.
(a) Frames from time-lapse Video 9 showing YFP-β-catenin accumulation at the centrosome (white arrowheads) and after cell division. Bar = 10 μm. (b) The colocalization (white arrowheads) of YFP-β-catenin (top) or endogenous β-catenin (bottom) with the centrosomal marker pericentrin (red immunofluorescence) in untreated and LiCl-treated cells. Hoechst DNA stain is in blue, and DIC in grey. Boxes show enlarged centrosomal areas. Bar = 10 μm. (c) The relative average intensity of YFP-β-catenin measured in the centrosome (n = 11), membrane, cytoplasm and nucleus (from Figure 2 and 5) of Wnt3a-treated cells. Correlation scores (r) between the nucleus (n), cytoplasm (c), membrane (m) and centrosome (ce) YFP-β-catenin levels are presented at the bottom. (d) The rate of change in YFP-β-catenin levels (ΔI/Δt) accumulation or degradation in the centrosome, membrane, cytoplasm and nucleus over time in Wnt3a- and LiCl-treated cells. (e) Plots of YFP-β-catenin levels in the sub-cellular compartments of individual cells (from Figure 3). Boxes show the correlation scores (r) between the nucleus (n), cytoplasm (c), membrane (m) and centrosome (ce).
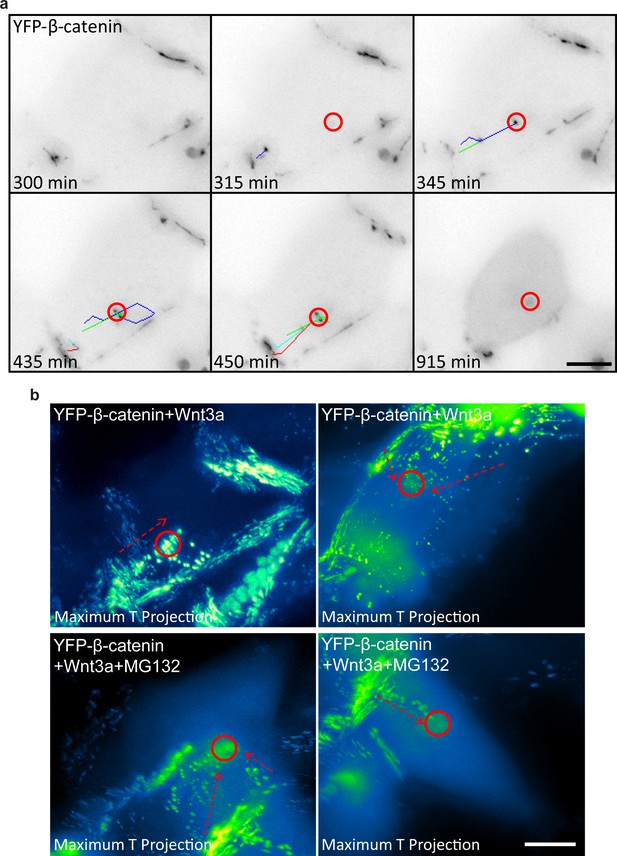

Detachment of membranal YFP-β-catenin puncta and movement towards the centrosome.
(a) Frames from Video 10 showing the tracks of several YFP-β-catenin membranal puncta (colored tracks) moving from the membrane region towards the centrosome area (red circle). Time is minutes after addition of Wnt3a. YFP signal is shown in negative greyscale colors. Bar = 10 μm. (b) Maximum time projections of movements of membranal YFP-β-catenin puncta (arrows) towards the centrosome region (circles) in four different cells. The YFP signal is pseudo-colored using the ImageJ ‘Green Fire Blue’ look-up table. Top row are Wnt3a-treated cells. Bottom row are Wnt3a + MG132-treated cells. Bar = 10 μm.
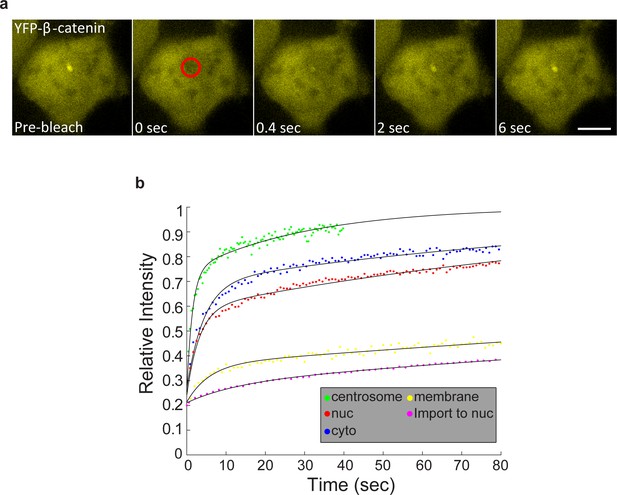
Summary of FRAP measurements of YFP-β-catenin dynamics in subcellular compartments in response to Wnt3a treatment.
(a) Frames showing one pre-bleach frame, the bleach of the YFP-β-catenin in the centrosome of a Wnt3a-treated cell, and frames following the recovery of signal over time. Circle denotes the bleached region. Bar = 10 μm. (b) Averaged data plot of FRAP recovery curves in the cytoplasm (n = 24), nucleus (n = 25), membrane (n = 32) and centrosome (n = 13). Membrane recovery from Figure 5—figure supplement 1 and the import rate to nucleus from Figure 2—figure supplement 1 are also plotted.
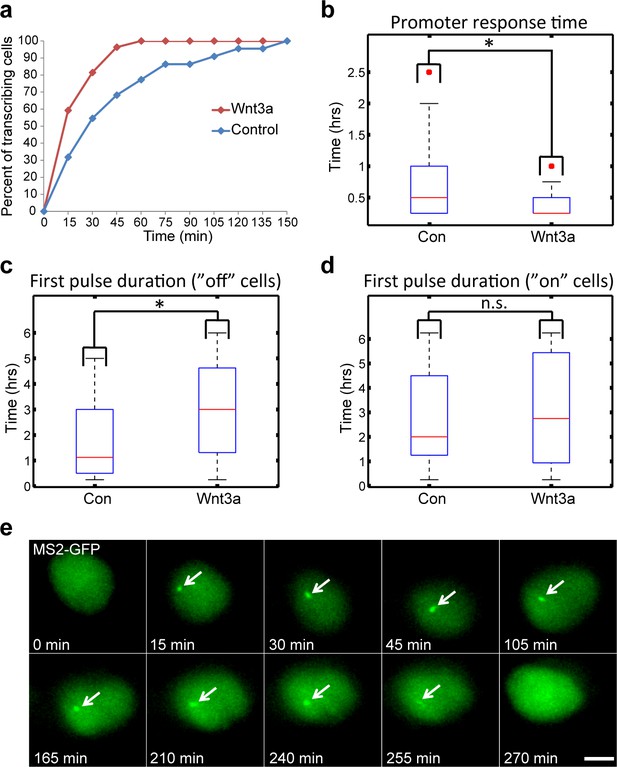
Measuring the transcriptional response of CCND1-MS2 to Wnt3a activation in living cells.
(a) The percentage of cells in a population of either mock-treated (blue) or Wnt3a-activated cells (red) showing an active CCND1-MS2 transcribing gene, over time. (b) The promoter response time of CCND1-MS2 activation from the addition of Wnt3a (n = 27) or in mock-treated conditions (n = 22). In the boxplots, the median is indicated by a red line, the box represents the interquartile range, the whiskers represent the maximum and minimum values, and red dots represent outliers. (p=0.01). (c,d) Periods of gene activity measured in mock-treated and Wnt3a-treated cells. The population was divided into cases where the gene was either not transcribing before the addition of Wnt3a or mock-treatment (‘off’, n(Wnt3a) = 27, n(Con) = 22, p=0.01) or if the gene was already active (‘on’, n(Wnt3a) = 37, n(Con) = 52, p=0.77). *p<0.05, n.s. = p>0.05. (e) Frames from Video 12 showing the activation of the CCND1-MS2 gene detected by MS2-GFP mRNA tagging (arrow) following Wnt3a treatment. Bar = 10 µm.
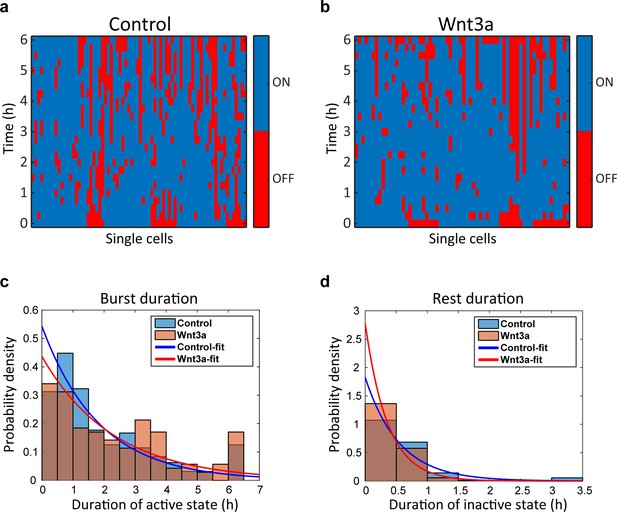
Wnt signaling causes shorter rest duration in addition to an increase in the gene burst duration.
Plots of single cells demonstrate the active (blue) and inactive (red) state of CCND1-MS2 transcribing gene along 6 hr in (a) mock-treated cells (Control, n = 74) and (b) Wnt3a-treated cells (n = 64). Data were taken from live-cell movies with each column representing one cell along 6 hr. Histograms showing the distribution of (c) active and (d) inactive state durations in Wnt3a-treated and mock-treated (Control) cells (p=0.02, p=0.0004 respectively). The curves are a fit to exponential distribution (Golding et al., 2005).
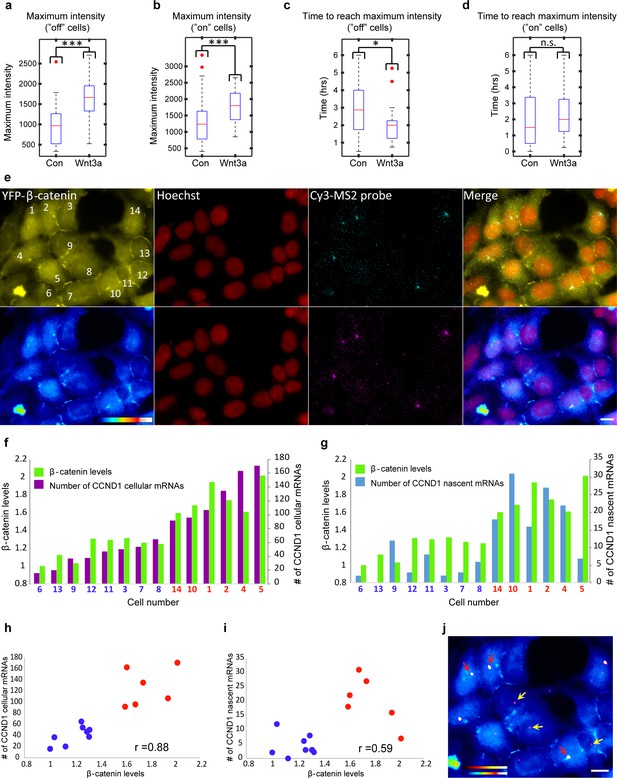
Quantification of CCND1 activity levels following Wnt activation in single fixed and living cells.
(a,b) Boxplots showing the maximal MS2-GFP intensity levels reached on actively transcribing CCND1-MS2 genes during 6 hr in Wnt3a-treated and mock-treated (Con) cells, when (a) the gene was either not transcribing before the addition of Wnt3a or mock-treatment (‘off’, n(Wnt3a) = 27, n(Con) = 22, p=0.0001) or (b) if the gene was already active (‘on’, n(Wnt3a) = 37, n(Con) = 52, p=0.0006). The median is indicated by a red line, the box represents the interquartile range, the whiskers represent the maximum and minimum values, and red dots represent outliers. (c,d) Boxplots showing the time required to reach the maximal intensity levels when (c) the gene was either not transcribing before the addition of Wnt3a (‘off’, p=0.03) or (d) if the gene was already active (‘on’, p=0.42). (e) YFP-β-catenin (yellow) together with RNA FISH images obtained with a probe hybridizing to the MS2 region in the 3’UTR of the CCND1-MS2 mRNA (cyan), showing CCND1 nascent mRNAs on active genes (large dots) and cellular mRNAs (small dots) in Wnt3a-treated cells (2 hr), in comparison to YFP-β-catenin levels. Nuclei are stained with Hoechst (pseudo-colored red). Bottom row is the pseudo-colored YFP signal using the ImageJ ‘Royal’ look-up table. Cells are numbered. Bar = 10 μm. (f) Quantification of the number of cellular CCND1-MS2 mRNAs (ordered from low to high) compared to YFP-β-catenin levels. (g) Quantification of the number of nascent CCND1-MS2 mRNAs compared to YFP-β-catenin levels. (h,i) Correlation analysis between (h) the number of cellular CCND1-MS2 mRNAs and YFP-β-catenin levels and (i) between the number of nascent CCND1-MS2 mRNAs and YFP-β-catenin levels. Blue dots – subpopulation with low nuclear YFP-β-catenin levels and low numbers of cellular/nascent CCND1-MS2 mRNAs. Red dots – subpopulation with high nuclear YFP-β-catenin levels and high numbers of cellular/nascent CCND1-MS2 mRNAs. Total correlation score between the number of cellular/nascent CCND1-MS2 mRNAs and YFP-β-catenin levels is 0.88 and 0.59, respectively. (j) The field from panel e demonstrating higher intensity of active CCND1-MS2 genes in cells with high nuclear YFP-β-catenin levels (red arrows) compared to cells with low nuclear YFP-β-catenin levels (yellow arrows). Active genes are pseudo-colored using the ImageJ ‘Red Hot’ look-up table. The fluorescent signal of the active genes was enhanced using ImageJ 'Spot Enhancing Filter 2D'. This enhancement led to the reduced detectability of single mRNAs in this presentation of the image, in order to emphasize the difference in transcriptional activity between low and high levels of nuclear YFP-β-catenin. Bar = 10 μm.
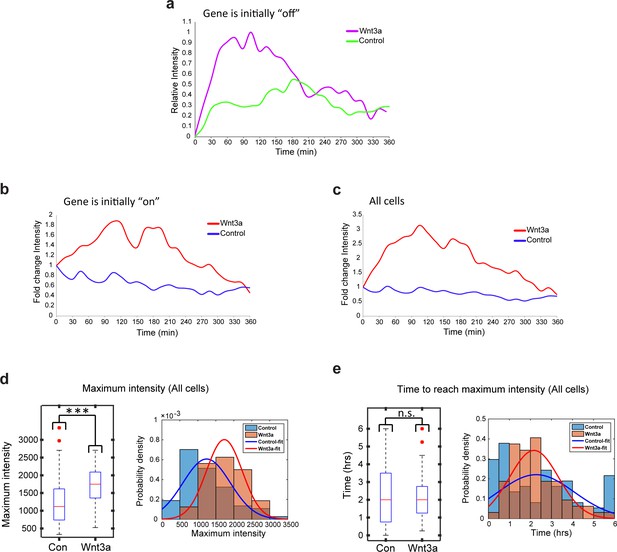
Transcription site intensity levels in living cells following Wnt3a activation.
Plots showing the MS2-GFP average intensity levels measured on active CCND1-MS2 transcription sites during 6 hr in Wnt3a-treated and in mock-treated cells, when (a) the gene was either not transcribing before addition of Wnt3a or mock-treatment (‘off’, n(Wnt3a) = 27, n(Con) = 22) (y axis is ‘relative intensity’ going from an ‘off’ state to an ‘on’ state') or (b) if the gene was already active to begin with (‘on’, n(Wnt3a) = 37, n(Con) = 52) (y axis is the fold change compared to the beginning of the movie). Results were normalized to the intensity at time 0. (c) Combined data from a and b. (d) Boxplot (left) showing the MS2-GFP maximal intensity levels reached on active CCND1-MS2 transcription sites during 6 hr in Wnt3a-treated and in mock-treated cells. In the boxplots, the median is indicated by a red line, the box represents the interquartile range, the whiskers represent the maximum and minimum values, and red dots represent outliers. The histograms (right) show the distribution of maximal intensity levels in these cells (combined data from Figure 8a and Figure 8b [p=2.13e-06]). The histograms show normalized data such that the area of each bar is relative to the number of observations (i.e. graph height is the probability density of the bar value, and graph area is equal to the probability of obtaining the bar value). The sum area of all bars is 1. The data were fitted with a Gaussian curve. (e) Boxplot (left) showing the time required to reach the maximal intensity in Wnt3a-treated and in mock-treated cells. The histograms showing the distribution of this time in these cells (combined data from Figure 8c and Figure 8d p=0.69). ***p<0.001, ns = p>0.05.
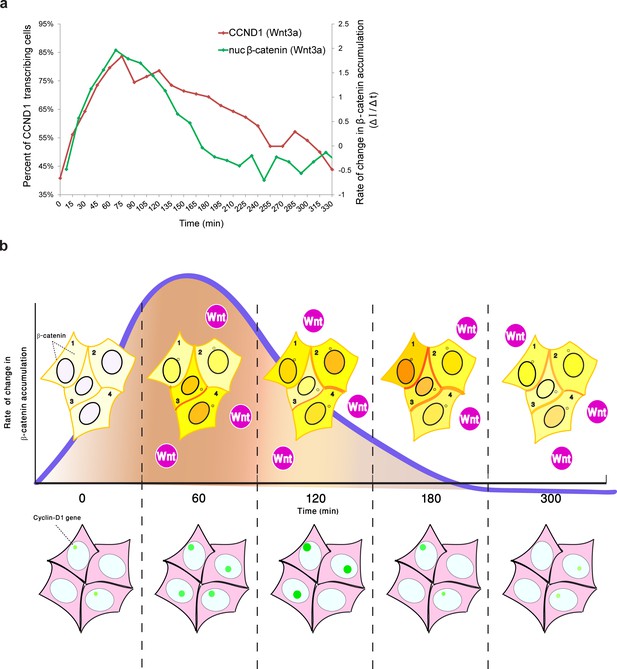
Comparing the kinetics of CCND1 transcriptional activation to the dynamics of β-catenin nuclear accumulation rate of change following Wnt signaling in living cells.
(a) Plots of the average transcriptional activation kinetics of CCND1-MS2 (red) following Wnt3a activation, compared to the plot of rate of change in β-catenin nuclear accumulation (green). (b) Scheme of the dynamic changes occurring in the studied cell system following Wnt signaling. Top - from left to right: Levels of β-catenin (yellow) in the nucleus are normally low but after the addition of Wnt3a to the medium a significant and rapid increase in the nucleus is observed, peaking after 2–3 hr. β-catenin levels later decline in the nucleus and cytoplasm due to degradation. While this is the average behavior in the population (e.g. cells 1 and 2), when examining individual cells, different dynamics such as multiple pulsations (e.g. cell 3) and rapid initial accumulation (e.g. cell 4) are observed. β-catenin levels increase simultaneously at the membrane and at the centrosome. Bottom- β-catenin induces cyclin D1 transcriptional activity (green dot), and modulation of the transcriptional reaction can be observed as the gene reaches higher levels of activity, for longer periods of time. The rate of change in β-catenin accumulation (blue curve, top), rather than the actual levels of β-catenin in the nucleus, correlate with the kinetics of transcriptional activation.
Videos
Transcriptional activation of CCND1 in response to Wnt3a.
HEK293 CCND1-MS2 cells stably expressing MS2-GFP (green) were treated with Wnt3a. The transcribed CCND1 mRNA on the active gene is seen as a bright green dot. The fluorescent signal on the active genes was enhanced using ImageJ 'Spot Enhancing Filter 2D' in order to clearly present the active sites in the movie. Cells were imaged every 15 min for 6 hr.
YFP-β-catenin dynamics at steady state and after Wnt3a activation.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin were treated with Wnt3a (top) and showed nuclear and cytoplasmic accumulation of YFP-β-catenin, followed by slow egress. No change in YFP-β-catenin levels was seen in mock-treated cells (bottom). Right – The YFP signal is pseudo-colored using ImageJ ‘Royal’ look-up table to show YFP-β-catenin levels. Cells were imaged every 15 min for 510 min.
YFP-β-catenin dynamics in individual cells.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin were treated with Wnt3a, and the dynamics of the protein were observed in individual cells. The YFP signal is pseudo-colored using ImageJ ‘Green Fire Blue’ look-up table to show YFP-β-catenin levels. Cells were imaged every 15 min for 825 min.
YFP-β-catenin dynamics in response to LiCl.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin were treated with LiCl and increased accumulation of the protein was observed. The YFP signal is pseudo-colored using ImageJ ‘Green Fire Blue’ look-up table to show YFP-β-catenin levels. Cells were imaged every 15 min for 825 min.
YFP-β-catenin dynamics following Wnt3a activation during cell division.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin were treated with Wnt3a, and the dynamics of the protein in the nucleus were followed over time. Two cells that undergo mitosis were observed in the field. The levels of the protein in the daughter cells formed from the upper cell were low (also note the appearance and division of the centrosome detected via YFP-β-catenin). In comparison, in the bottom mitotic cell, one daughter cell accumulated high YFP-β-catenin levels very rapidly, while the other responded slowly and had very low levels. The YFP signal is pseudo-colored using ImageJ ‘Fire’ look-up table to show YFP-β-catenin levels. Cells were imaged every 15 min for 225 min.
YFP-β-catenin dynamics following Wnt3a activation during the cell cycle.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin (yellow) and the Fucci markers for G1 (red) and G2 (cyan), were treated with Wnt3a, and the dynamics of the protein were followed over time. Cells were imaged every 15 min for 1065 min.
YFP-β-catenin dynamics following Wnt3a activation during cell division.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin (yellow) and the Fucci markers for G1 (red) and G2 (cyan), were treated with Wnt3a, and the dynamics of the protein were followed over time in four cells that undergo mitosis. Cells were imaged every 15 min for 705 min.
YFP-β-catenin dynamics at the cell membrane following Wnt3a activation.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin were treated with Wnt3a, and the dynamics of the protein at the membrane were followed over time, and were similar to the nucleus and cytoplasm accumulation. The YFP signal is pseudo-colored using ImageJ ‘Green Fire Blue’ look-up table to show YFP-β-catenin levels. Cells were imaged every 15 min for 1065 min.
YFP-β-catenin accumulation at the centrosome following Wnt3a activation.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin were treated with Wnt3a, and the dynamics of the protein at the centrosome were observed in parallel to the accumulation in the nucleus and cytoplasm. The separation of the centrosome in a cell during division can be seen after the 960 time point. Cells were imaged every 15 min for 1005 min.
YFP-β-catenin puncta move from the membrane to the centrosome.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin were treated with Wnt3a. At the 300 min time point, a series of YFP-β-catenin puncta can be tracked (track colors) moving from the membrane to the centrosome. An inverted presentation of the movie shows the movie puncta (black dots). Cell was imaged every 15 min for 1005 min.
YFP-β-catenin puncta move from the membrane to the centrosome.
HEK293 CCND1-MS2 cells stably expressing YFP-β-catenin were treated with Wnt3a and MG132. At the 165 min time point, a series of YFP-β-catenin puncta can be tracked (track colors) moving from the membrane to the centrosome. The YFP signal is pseudo-colored using ImageJ ‘Green Fire Blue’ look-up table to show YFP-β-catenin levels. Cells were imaged every 15 min for 1065 min.
Prolonged activation of CCND1 after Wnt activation.
HEK293 CCND1-MS2 cells stably expressing MS2-GFP (green) were treated with Wnt3a. CCND1 mRNA transcription could be detected 15 min after Wnt3a (green dot, transcription site) and continued for 4 hr. Cells were imaged every 15 min for 270 min.
Additional files
-
Supplementary file 1
Statistical analysis performed in this study.
(a) The statistical significance p values (t test) at each time point for the percentage of cells showing an active CCND1-MS2 gene (refers to Figure 1b) between control and Wnt3a-treated cells. (b-f) Mann-Whitney test for comparison between two independent FRAP/FLIP experiments. A statistical comparison between all datasets of two individual FRAP/FLIP experiments are depicted in each plot and are illustrated as a single red circle which marks the p-value (y axis) for all intensity values measured for each time point (x axis). The top and bottom dotted lines indicate where p-value equals 0.05. (b) Statistically significant difference between the FRAP dynamics of YFP-β-catenin in the nucleus under Wnt3a treatment versus overexpression of YFP-β-catenin that enters the nucleus without signal, and (c) between the FRAP and (d) FLIP import and export dynamics (refers to Figure 2—figure supplement 1). (e) No statistically significant difference between YFP-β-catenin at the cell membrane between mock-treated and Wnt3a-treated cells (refers to Figure 5—figure supplement 1). (f) No statistically significant difference between YFP-β-catenin at the cell membrane between mock-treated and LiCl-treated cells (refers to Figure 5—figure supplement 1).
- https://doi.org/10.7554/eLife.16748.034





