Ring-like oligomers of Synaptotagmins and related C2 domain proteins
Figures
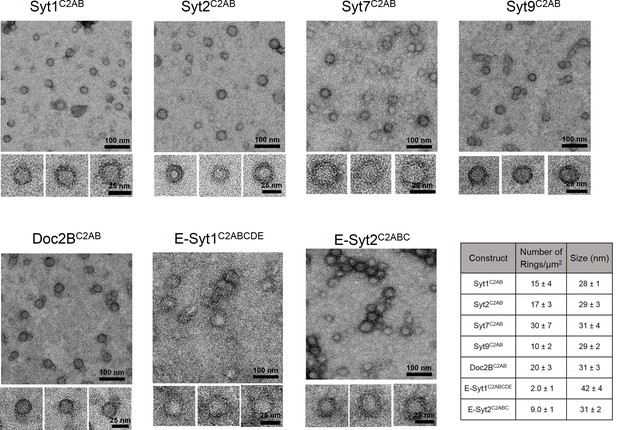
Ring-like oligomers are a common structural feature of C2 domain proteins.
EM analysis showing the C2AB domains of neuronal isoforms of Syt, namely Syt1, Syt2, Syt7, and Syt9 form ring like oligomers on monolayers under Ca2+-free conditions. Similar ring-like structures were observed for other related C2 domain proteins, like Doc2B and E-Syt 1 & 2. The number of ring-oligomers observed on the monolayers varied, but the dimensions of the rings were remarkably consistent (~30 nm). All EM analyses were carried out using 5 µM protein on monolayer containing 40% PS and buffer containing 15 mM KCl and 1 mM free Mg2+. Representative micrographs and average values, along with standard error of the means (SEM) from a minimum of three independent trials are included.
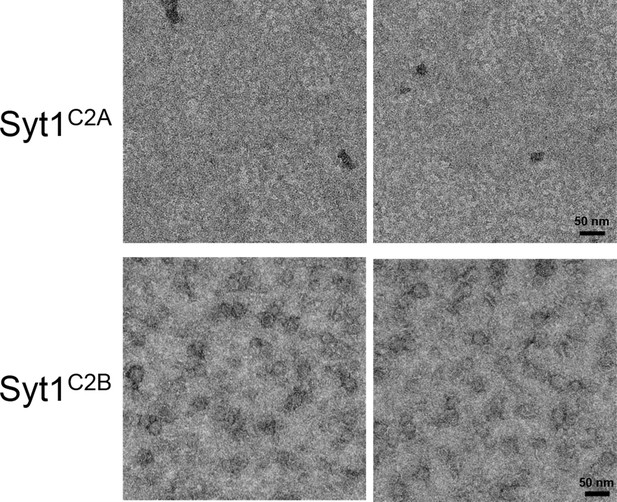
Ring assembly is not a conserved property of all C2 domains.
Negative stain EM analysis showing the Syt1C2B domain alone can form ring-like oligomers, but the Syt1C2A domain cannot. Syt1C2B rings are relatively un-stable and are smaller (~22 nm) compared to the Syt1C2AB ring oligomers (~30 nm). EM analysis were carried out using 5 µM protein on monolayer containing 40% PS and buffer containing 15 mM KCl and 1 mM free Mg2+. Representative micrographs from 3 independent trials are shown.
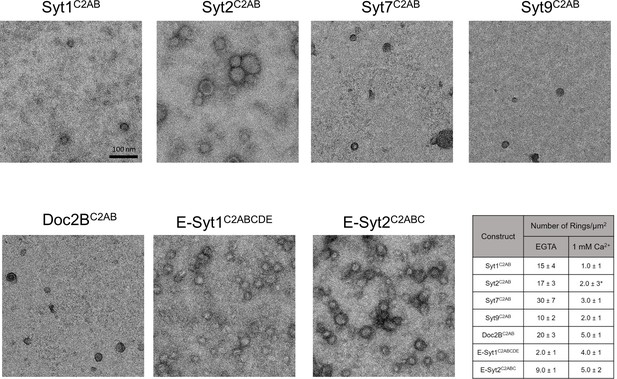
Effect of Ca2+ addition on pre-formed ring-like oligomers of Syt isoforms and other C2 domain proteins.
Pre-formed ring oligomers were washed briefly (10 s) with buffer containing calcium (final concentration of 1 mM free) and analyzed via negative stain EM . All Syt isoforms and Doc2B were sensitive to the Ca2+ treatment and number of rings observed reduced drastically with Ca2+ treatment. *Note: With Syt2C2AB, we observed irregular patches of protein arrays together with circular structures following Ca2+ treatment. These circular structures were quite different from the Ca2+-free rings as they exhibit a more uniform protein density, indicating these are patches of protein arrays, arranged into round shape probably due to the local buckling of monolayer by insertion of Syt2 calcium loops. Over longer incubation time, the number of 'circular structure' decrease and the size of irregular patches increase and begin to pack orderly, suggesting the circular structure is the intermediate building block of 2D-array formation. Ca2+ addition had divergent effect on the E-Syt isoforms, while the E-Syt2 was largely un-affected, the E-Syt1 rings were stabilized (discussed below). All EM analyses were carried out using 5 µM protein on monolayer containing 40% PS and buffer containing 15 mM KCl and 1 mM free Mg2+. Representative micrographs and averages and SEM from 3–4 independent trials are shown.
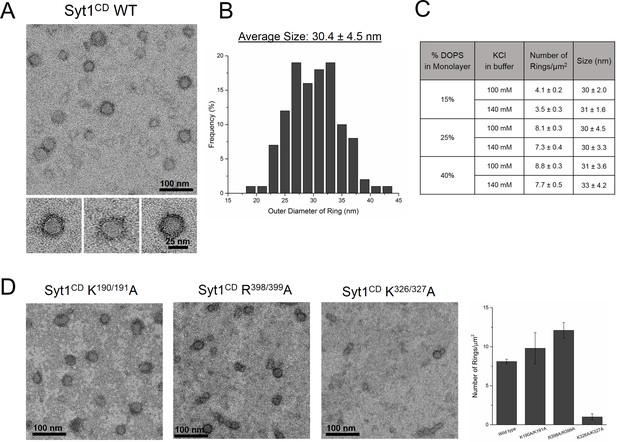
The entire cytoplasmic domain of Syt1 (Syt1CD) forms ring-like oligomers under physiologically relevant conditions.
(A) Negative stain EM analysis shows ring-like oligomers of Syt1CD on PC/PS (3:1 molar ratio) lipid monolayers in buffer containing 100 mM KCl and 1 mM MgCl2. (B) The size distribution of the Syt1CD rings as measured from the outer diameter (n = ~400) under these experimental conditions using ImageJ software. (C) The Syt1CD ring-oligomers were observed under a wide-ranging conditions. Under all conditions tested, the dimension of these ring oligomers were very consistent (~30 nm), but the number of rings observed depended on amount of the anionic lipid in the monolayer and the salt (KCl) concentration of the buffer (D) EM analysis showing that the polylysine (K326/K327) motif of C2B domain is critical to the ring formation, but the other conserved polybasic regions of Syt1, namely K190/K191 on C2A and R398/R399 on C2B are not involved in ring formation. All EM analyses were carried out using 5 µM protein on monolayers containing 25% PS and in buffer containing 100 mM KCl and 1 mM free Mg2+. Representative micrographs and averages/SEM from three independent trials are shown.
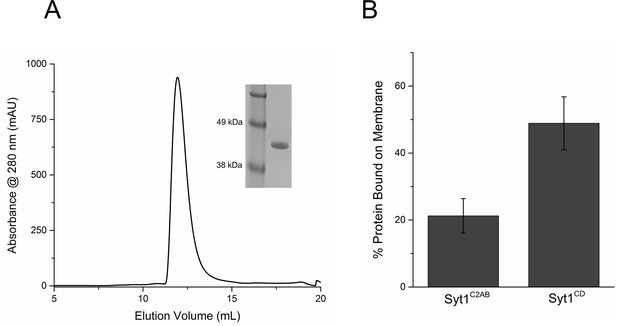
Purification and Characterization of Syt1CD.
(A) Size Exclusion Chromatography profile on a Superdex75 10/300 GL column of Syt1CD. Syt1CD was purified using a stringent purification protocol, including benzonase treatment, high salt wash and ion exchange chromatography. The Syt1CD sample shows a single peak confirming that is free of polyacidic impurities, which could trigger aggregation of the protein. Inset: SDS-PAGE Coomaisse analysis of the peak shows a single band consistent with to the size (~41 kDa) of the Syt1CD protein. (B) Lipid binding analysis shows that the inclusion of juxtamembrane domain enhances the Syt1 membrane interaction. To measure binding, 10 µM of Syt1C2AB or Syt1CD were mixed with 1 mM small unilamellar vesicles (SUV) containing 25% PS+3% PIP2 (remainder was PC) and incubated for 1 hr at RT with in buffer containing 100 mM KCl and 1 mM free Mg2+. The SUVs were isolated using discontinuous density gradient and analyzed on SDS-PAGE/ Coomaisse analysis, after adjusting for the amount of lipid recovered. The amount of protein bound was estimated using density measurement using ImageJ software.

Presence of anionic lipid is required to assemble the Syt1CD ring oligomers.
Negative stain EM analysis was carried out using 5 µM protein on monolayer with DOPC alone in buffer containing 100 mM KCl and 1 mM free Mg2+. Representative micrographs from 3–4 independent trials are shown.
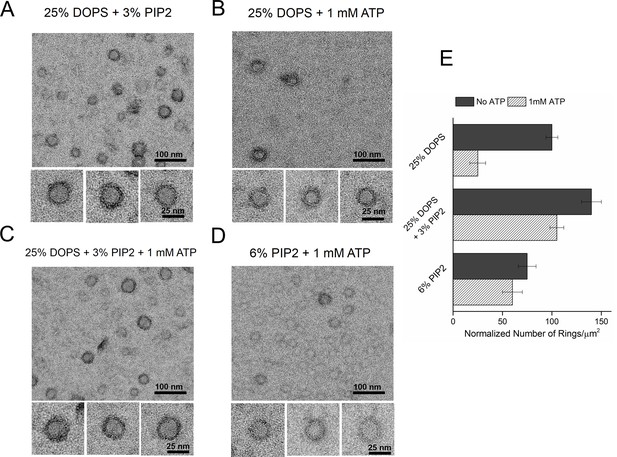
Syt1-PIP2 interaction is key to ring-formation under physiologically relevant conditions.
(A) Inclusion of 3% PIP2 (in addition to 25% PS) in the monolayer stabilized the ring structures and increased the number of rings observed. (B, C) Addition of ATP drastically reduced the number of rings observed in monolayers containing 25% PS only, but not when supplemented with 3% PIP2. (D) PIP2 (6%) as the only anionic lipid on the bilayer was sufficient to assemble ring-like oligomers, even in the presence of 1 mM ATP. All EM analyses were carried out using 5 µM protein in buffer containing 100 mM KCl and 1 mM free Mg2+. Representative micrographs and average values/SEM from a minimum of three independent trials are shown in (E). The rings observed under all conditions shown in (E) were similarly (~30 nm) sized
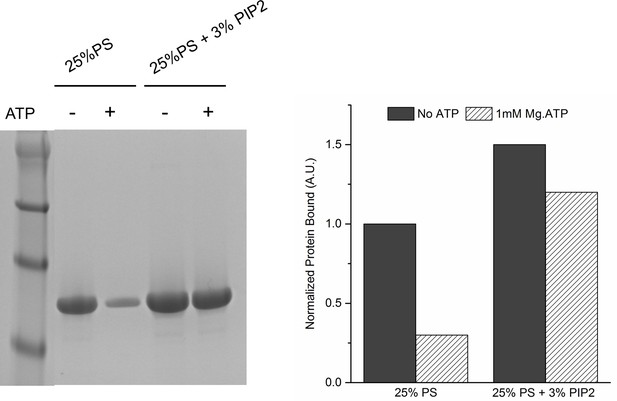
Lipid binding analysis shows that ATP effectively screens the interaction of Syt1CD to PS-only membrane, but not membrane containing 3% PIP2.
To assess Syt1CD-membrane binding, 10 µM of Syt1CD were mixed with 1 mM small unilamellar vesicles (SUV) containing either 25% PS or 25% PS+3% PIP2 (remainder was PC) and incubated for 1 hr at RT with in in buffer containing 100 mM KCl and 1 mM free Mg2+ supplemented with 1 mM Mg2+ or 1 mM Mg-ATP. The SUVs were isolated using discontinuous density gradient and analyzed on SDS-PAGE/ Coomaisse analysis after adjusting for the amount of lipid recovered. The amount of protein bound in each case was estimated using density measurement using ImageJ software.
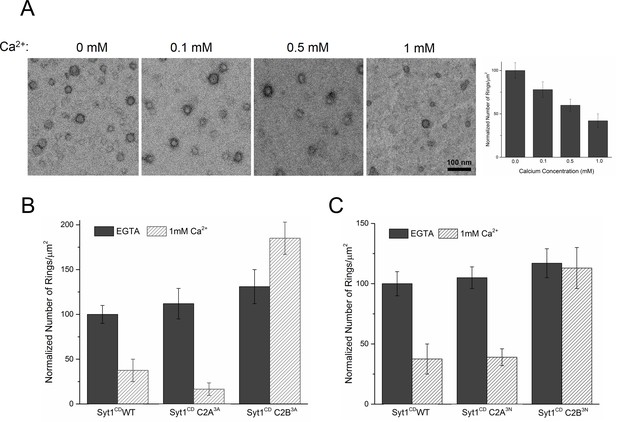
Ca2+ binding and subsequent re-orientation of the C2B domain into the membrane are needed to disassemble the Syt1 ring oligomer.
(A) Syt1CD ring oligomers were sensitive to Ca2+ and brief treatment (10 s) of the pre-formed rings with physiological levels of Ca2+ greatly reduced the number of rings observed. (B) Ca2+sensitivity of the Syt1CD rings maps to the C2B domain as disrupting Ca2+ binding to C2B (Syt1CD D309A, D363A, D365A; C2B3A) but not C2A (Syt1CD D178A, D230A, D232A; C2A3A) rendered the rings Ca2+ insensitive. (C) Ca2+-induced insertion of just the C2B domain is necessary to disrupt the ring oligomers as hydrophilic mutation that blocks its insertion of the C2B loop (Syt1CD V304N, Y364N, I367N; C2B3N) but not the C2A loop (Syt1CD F231N, F234N, S235N; C2A3N) makes the rings insensitive to Ca2+. All EM analyses were carried out using 5 µM protein on monolayers containing 25% PS and in buffer containing 100 mM KCl and 1 mM free Mg2+. Effect of addition of 1mM Ca2+ (final concentration) is shown in (B) & (C). Representative micrographs and average values and deviations (SEM) from 3–4 independent trials are shown.
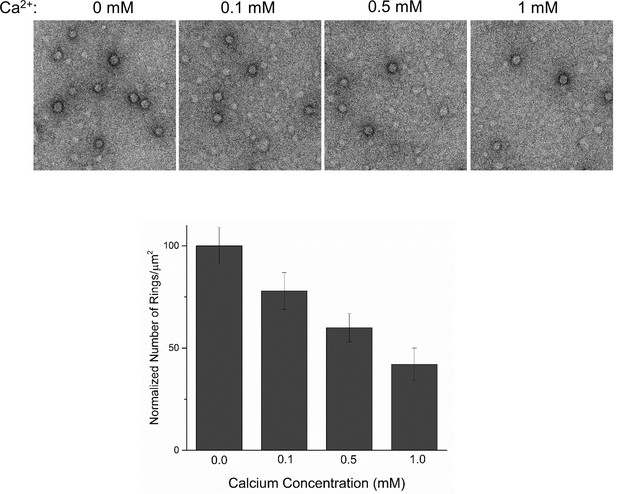
Inclusion of PIP2 does not change the Ca2+ sensitivity of the Syt1CD ring oligomers.
The pre-formed rings were sensitive to Ca2+ and brief treatment (10 s) of the pre-formed rings with physiological levels of Ca2+ drastically reduced the number of rings observed, even when 3% PIP2 (in addition to 25% PS) was included in the monlayer.The reduction in the number of ring oligomers observed was comparable to no PIP2 condition. (Figure 4A). Representative micrographs and averages and SEM from 3–4 independent trials are shown.
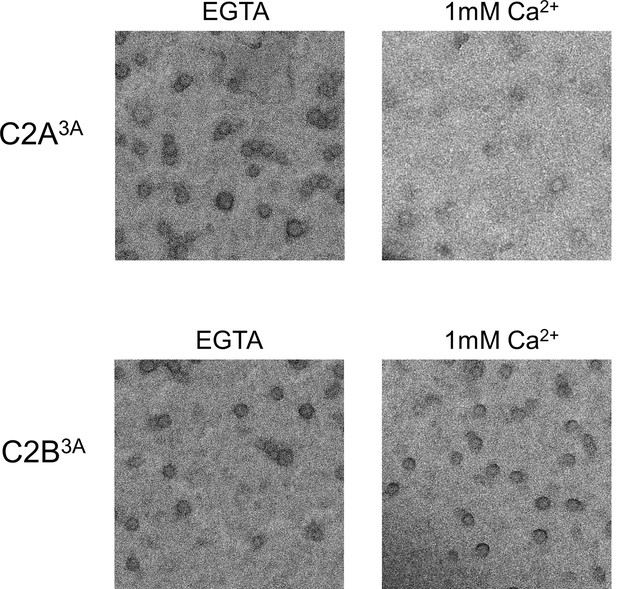
Disrupting the calcium binding to C2B (Syt1CD D309A, D363A, D365A; C2B3A), but not C2A (Syt1CD D178A, D230A, D232A; C2A3A) renders the Syt1CD rings insensitive to calcium.
All EM analyses were carried out using 5 µM protein on monolayers containing 25% PS and in buffer containing 100 mM KCl and 1 mM free Mg2+. Pre-assembled ring oligomers were briefly (10 s) washed with buffer containing 1 mM free Ca2+ to assess the calcium sensitivity. Representative micrographs from 3–4 independent trials are shown.
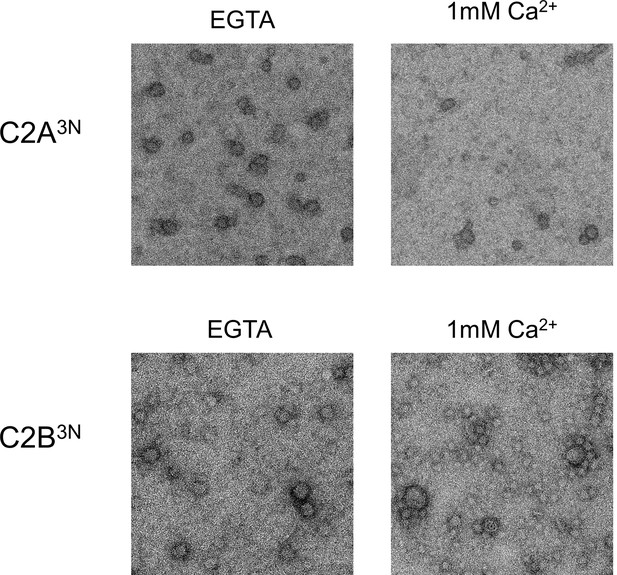
Disrupting the Ca2+-induced membrane insertion of C2B loop (Syt1CD V304N, Y364N, I367N; C2B3N) but not the C2A loop (Syt1CD F231N, F234N, S235N; C2A3N) makes the rings insensitive to calcium.
All EM analyses were carried out using 5 µM protein on monolayers containing 25% PS and in buffer containing 100 mM KCl and 1 mM free Mg2+. Pre-assembled ring oligomers were briefly (10 s) washed with buffer containing 1 mM free Ca2+ to assess the calcium sensitivity. Representative micrographs from 3–4 independent trials are shown.
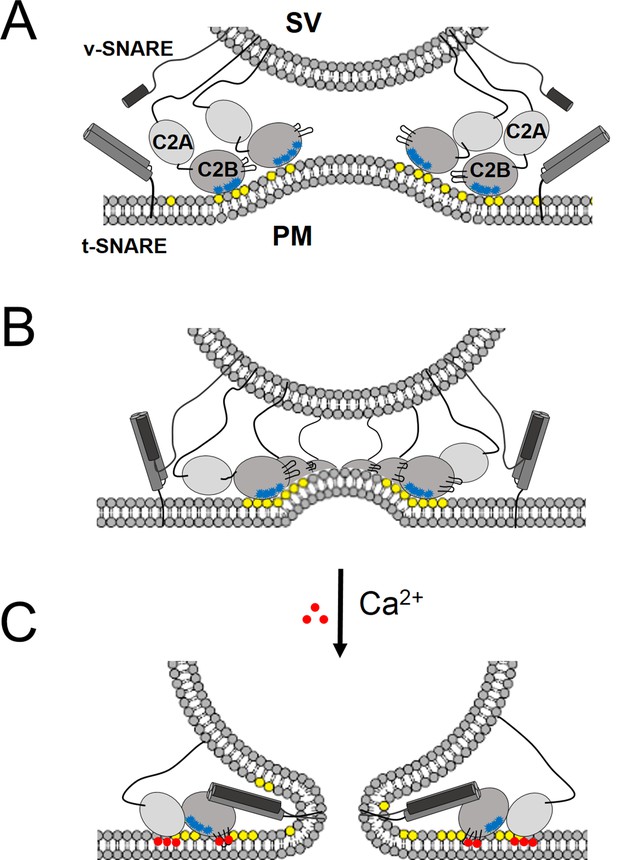
‘Washer’ model for the regulation of neurotransmitter release by Syt1.
(A) The SV docking interaction of the Syt1 polylysine motif (blue dots) with the PIP2 (yellow dots) on the plasma membrane positions the Syt1 on the membrane to promote the ring-oligomer formation. The ring assembly might precede the engagement of the SNARE proteins. (B) Syt1 ring-oligomers assembled at the SV-PM interface act as a spacer or ‘washer’ to separate the two membranes. The height of the ring (~4 nm) would allow the partial assembly of the SNARE complex, but prevent complete zippering and thus, block fusion. NOTE: The positioning and occupancy of SNAREs on Syt1 ring is not known and are shown for illustrative purposes only. (C) Upon binding calcium ions (red dots), the Ca2+ loops that locates to the oligomeric interface, re-orients and inserts into the membrane, thus disrupting the ring oligomer to trigger fusion and release neurotransmitters. Thereby, the Syt1 ring oligomers will synchronize the release neurotransmitters to the influx of calcium ions.
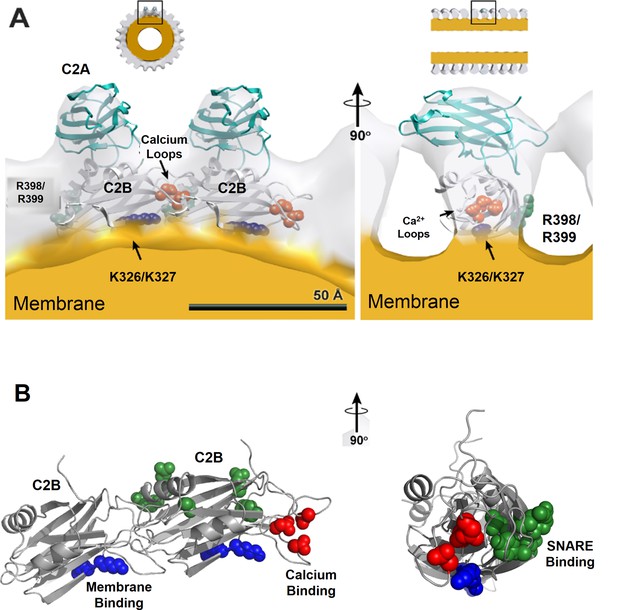
Organization of Syt1 C2 domains in the ring oligomers.
(A) X-ray structure of Syt1C2AB fitted into the EM density map of the Syt1C2AB coated monolayer tubes [Adapted from (Wang et al., 2014)] shows that C2B domain (gray) mediates both the membrane (yellow) association and the ring assembly and C2A domain (cyan) projects away from the ring. Surface representation of the tube is sectioned to the thickness of a single strand of the 4-start helix to reveal the shape of individual asymmetric unit. (B) Organization of the C2B domain in the Syt1 ring oligomer shows that the PIP2 binding polylysine motif (K326, K327, blue) are packed against the membrane surface, which holds back the Ca2+ binding site/loop (red) of C2B from the membrane and it localizes to the protein-protein interface involved in ring formation. This arrangement explains the Ca2+-sensitivity of the Syt1 rings as Ca2+ binding would include reorientation of the C2B domain from the ring geometry. Further, in this model, the recently identified (Zhou et al., 2015) primary SNARE binding interface on the C2B (R281, E295, Y338, R398 & R399, green) is accessible and free to interact with the SNAREs. However, the occupancy and orientation of the SNAREs on the Syt1 ring is not known.





