Klf5 regulates muscle differentiation by directly targeting muscle-specific genes in cooperation with MyoD in mice
Figures
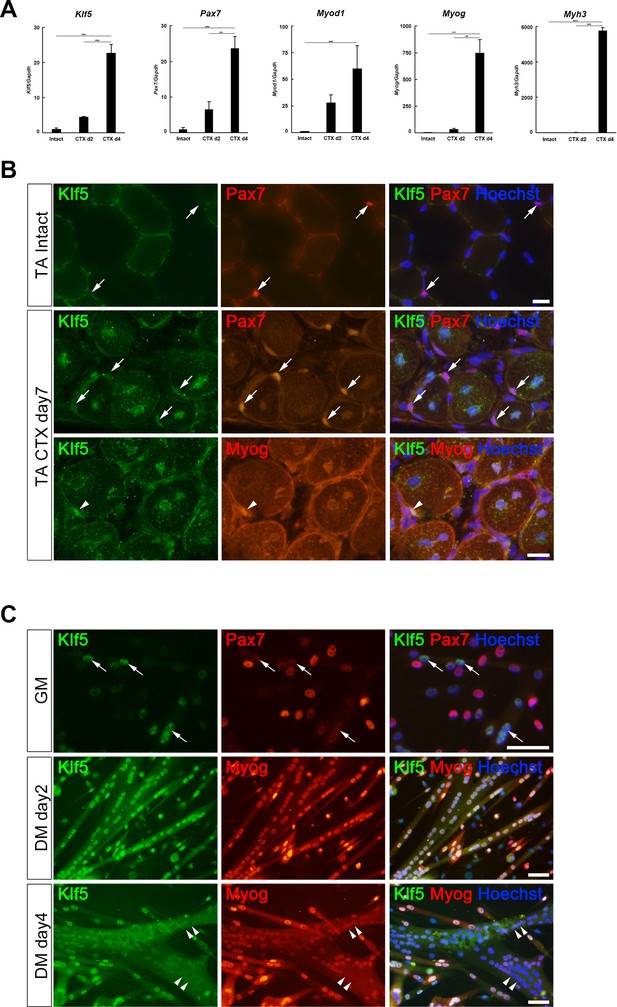
Klf5 is upregulated during myogenesis.
(A) Relative mRNA expression of Klf5, Pax7, Myod1, Myog and Myh3 (embryonic myosin heavy chain) in intact and regenerating TA muscles. The animals were sacrificed on day 4 after CTX injection. Data are means ± SEM. ***p<0.001. **p<0.01. Representative data from three individual mice are shown. (B) Klf5 expression during muscle regeneration. Klf5 was not detectable in quiescent satellite cells (SCs) in intact muscle (arrows in the top panels). During muscle regeneration, Klf5 was highly expressed in the nuclei of differentiating myocytes expressing Myog (arrowheads) and regenerating myofibers, whereas Klf5 expression was not detected or detected very weakly in Pax7-positive SCs (arrows). Representative data from at least three individual mice are shown. Scale bar represents 20 µm. (C) Plated SCs were cultured in growth medium (GM) or differentiating medium (DM; for 2 days or 4 days) after isolation and were co-immunostained for Klf5 with Pax7 or Myog. Klf5 is expressed in the differentiating myocytes, which were negative or very weakly positive for Pax7 (arrows). Klf5 was upregulated during differentiation and frequently co-localized with Myog. After 4 days of culture, Klf5 levels were decreased in the nuclei of large myotubes (arrowheads). Representative data from at least three individual mice are shown. Scale bar represents 50 µm.
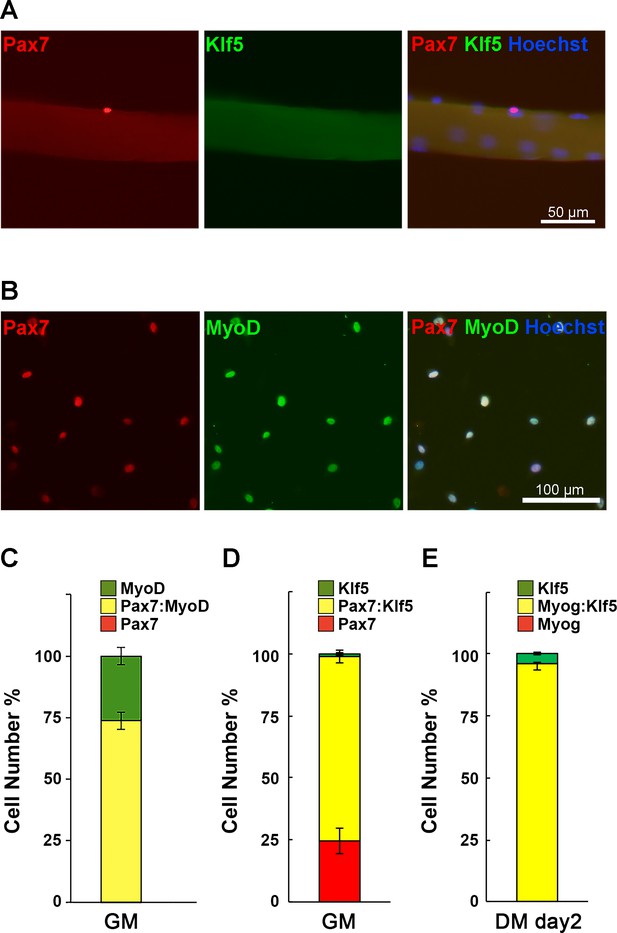
Preparation of SC primary cultures.
(A) Isolated EDL myofibers with their associated SCs were immediately fixed and immunostained for Pax7 and Klf5. (B) SCs were prepared from EDL muscles from wild-type C57/B6 mice and stained for Pax7 and MyoD. (C–D) Proportion of Pax7+, Pax7+: MyoD+ and MyoD+ cells (C) or Pax7+, Pax7+: Klf5+ and Klf5+ cells (D) in the primary SCs cultured in the growth media (GM). Expression of MyoD in every analyzed cells, indicating these cells are composed of pure, myoblast-lineage population. Pax7+: MyoD- cells was not detected. (E) Two days post differentiation, propotion of Myog+, Myog+: Klf5+ and Klf5+ cells were analyzed. Note that all the Myog positive cells were also positive for Klf5. Data represent means ± SEM. Representative data from at least three individual mice are shown. DM: differentiation medium.
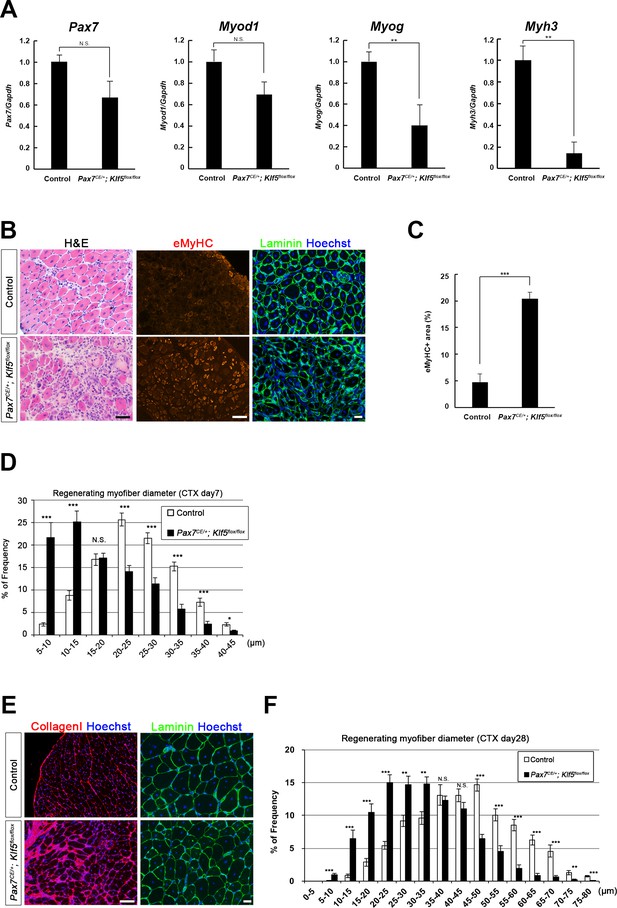
Klf5 is required for muscle regeneration in vivo.
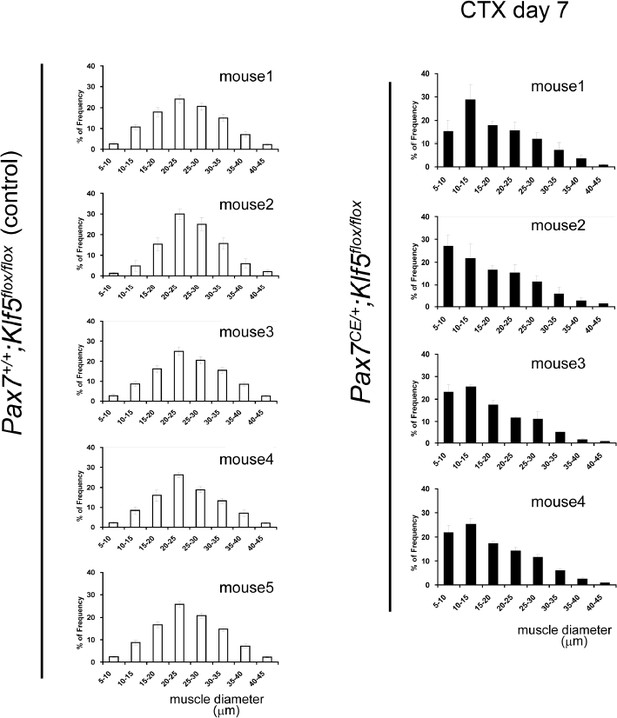
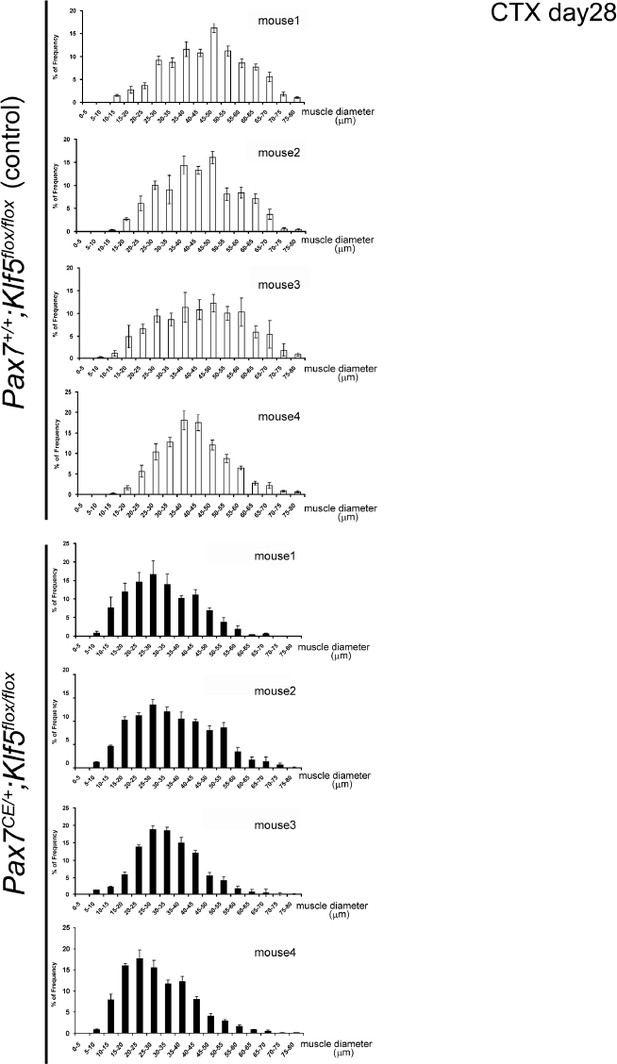
CTX was injected into the TA muscles of SC-specific Klf5 knockout mice (Pax7CE/+;Klf5flox/flox) and control Pax7+/+;Klf5flox/flox mice. The animals were then sacrificed on day 4 (A), 7 (B–D) or 28 (E–F) after CTX injection. (A) Pax7, Myod1, Myog and Myh3 expression in regenerating TA muscle (on day 4 after CTX injection) was analyzed using qRT-PCR (A). Data represent means ± SEM. n = 3 for each group. **p<0.01. N.S., not significant. (B–F) Sections were stained with H&E or immunostained for eMyHC, laminin and collagen I. Representative images of muscle sections on day 7 (B) or 28 (E) after CTX injection are shown. The eMyHC-positive area 7 days after CTX injection was quantified using Olympus cellSense Digital Imaging software (C). Distributions of myofiber diameters on days 7 (D) and 28 (F) after CTX injection are shown. Representative data from four Klf5 knockout and five control (for day 7) or four mice for each genotype (for day 28) are shown. Scale bars represent 100 µm. ***p<0.001. **p<0.01. N.S., not significant.
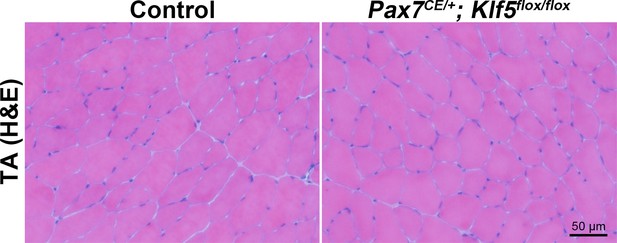
Tamoxifen treatment does not affect TA muscle morphology.
TA muscles were obtained from Klf5 knockout mice (Pax7CE/+; Klf5flox/flox) and control Pax7+/+; Klf5flox/flox mice on day 28 after tamoxifen injection. Muscle morphology was assessed by H–E staining.
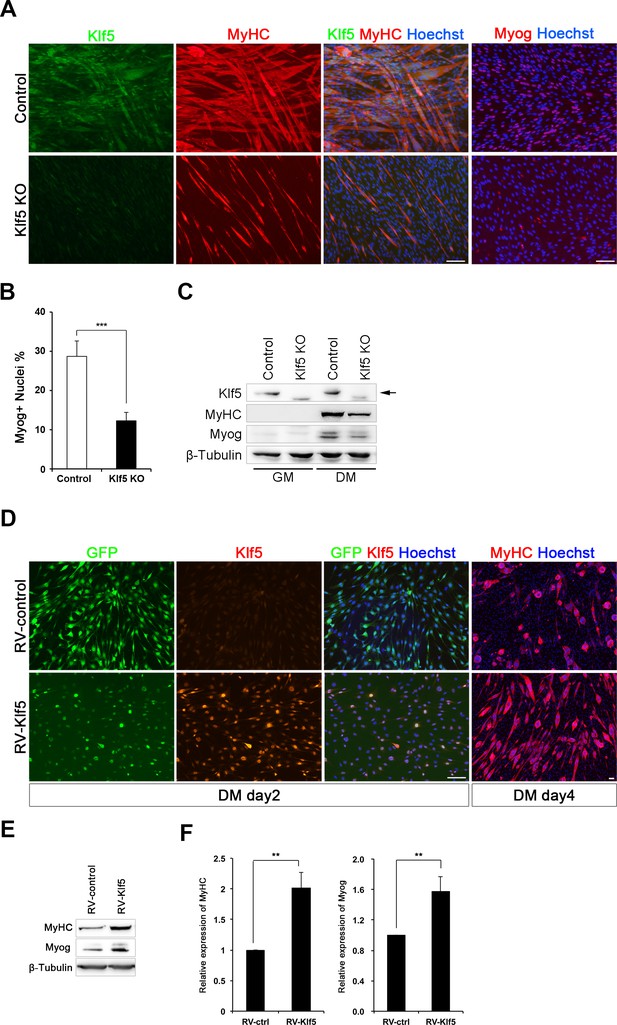
Klf5 is essential for muscle differentiation.
(A) Establishment of Klf5 knockout (KO) myoblasts using a CRISPR-Cas9 system. Klf5 KO C2C12 cells or Control cells were immunostained for MyHC and Klf5 or myogenin. Klf5 KO cells do not express Klf5 during muscle differentiation and exhibit severely reduced myotube formation. MyHC: myosin heavy chain, Myog: myogenin. Scale bar represents 100 µm. (B) Percentage of Myog-positive cells among total cells. Klf5 KO cells exhibited less Myog expression than Control. Data are means ± SEM. (***p<0.001) Representative data from at least three individual experiments are shown. (C) Western blots showing reduced MyHC and Myog expression in Klf5 KO C2C12 cells during differentiation. Representative data from at least three individual experiments are shown. (D) Klf5 KO cells were infected with a Retro-viral vector (RV-Klf5) harboring Klf5 or empty vector (RV-control), after which differentiation was induced for 2 or 4 days. The cells were then fixed and immunostained for Klf5 or MyHC. Impairment of myotube formation, as evidence from the loss of MyHC expression in Klf5 KO cells was rescued by exogenous Klf5 expression. Representative data from at least three individual experiments are shown. (E–F) Western blots revealing the reduction of MyHC and Myog expression in Klf5 KO C2C12 cells and its rescue by RV-Klf5. The expression levels were normalized to β-Tubulin (F). Data represent means ± SEM. (**p<0.01) Representative data from at least three individual experiments are shown.
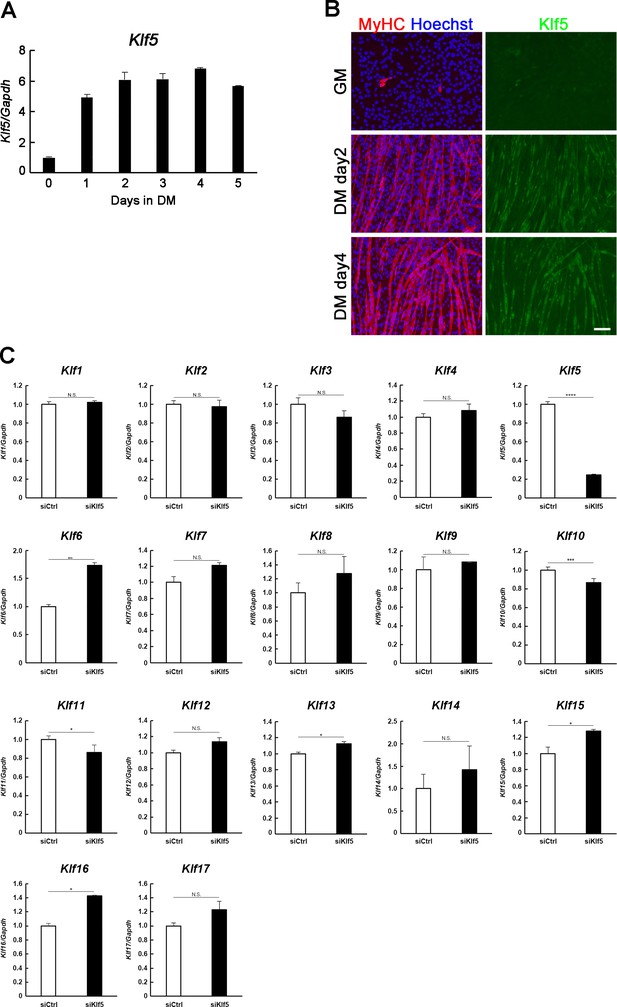
Klf5 knockdown impairs myoblast differentiation.
(A) Klf5 expression during C2C12 differentiation. Bars depict means ± SEM. n = 3, biological replicates. GM: growth medium, DM: differentiation medium. (B) Induction of Klf5 expression during C2C12 differentiation was assessed immunostaining for MyHC and Klf5. Klf5 is highly expressed in the nuclei of differentiating myoblasts expressing MyHC and in myotubes. However, little Klf5 expression was seen in proliferating C2C12 cells in GM. Representative data from at least three individual experiments are shown. MyHC: Myosin heavy chain. Scale bar represents 100 µm. (C) Relative mRNA expression of Klf family members in C2C12 myotube on day 3 by QPCR. Bars depict means ± SEM. ****p<0.0001, ***p<0.001, **p<0.01, *p<0.05, N.S., not significant. n = 3, biological replicates.
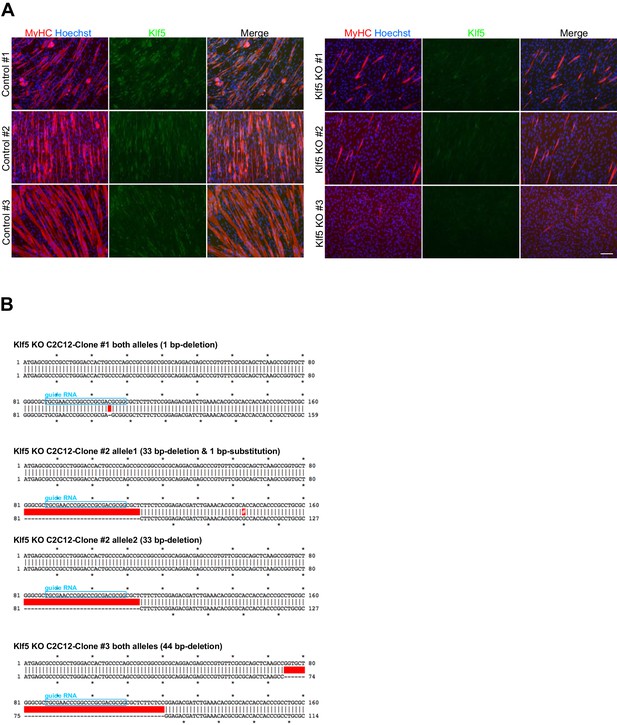
Establishment of Klf5 knockout C2C12 cells.
(A) C2C12 cells were transfected with pSpCas9-Klf5 or GFP-targeted control vector. After puromycin selection, the cells were cloned using the limiting dilution method. Three Klf5 knockout (KO) clones and three control C2C12 clones were obtained. Klf5 KO was confirmed by the absence of immunofluorescence. Compared to the control cells, all Klf5 KO cells exhibited defective myotube formation, as indicated by MyHC staining. Scale bar represents 100 µm. (B) Confirmation of mutation in the Klf5 gene locus. The position of guide RNA is indicated by the blue boxes. The red boxes indicate deleted sequences in the mutant allele, and the white # in the red box indicates the substituted nucleotide in the mutant allele. Clones #1 and #3 have a 1-bp deletion mutation in both alleles. The one allele of the clone #2 contains a 33-bp deletion, and the other allele contains a 33-bp deletion plus a 1-bp substitution mutation.
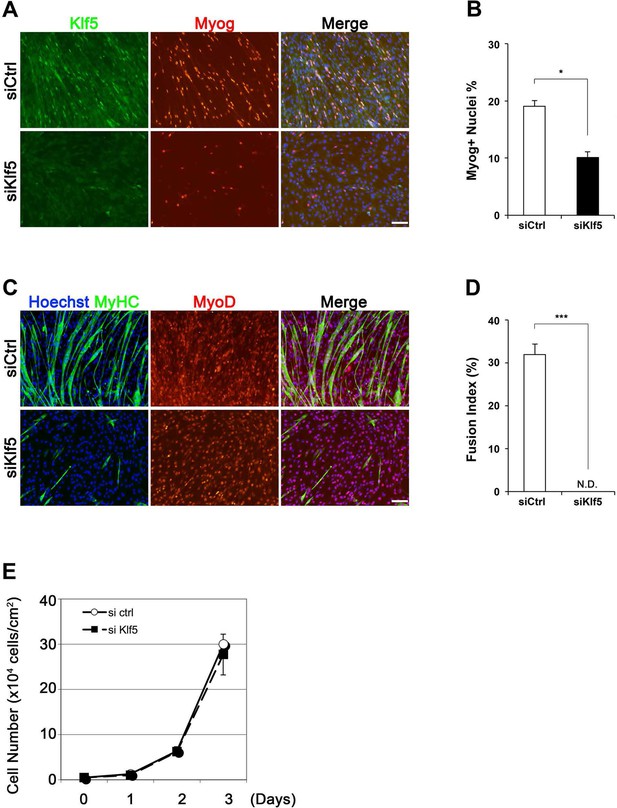
Phenotype of Klf5 knockdown cells.
(A–C) Klf5, myogenin (Myog), and MyHC expression in C2C12 cells after siRNA treatment. C2C12 cells were treated with either siKlf5 or siControl (siCtrl). After siRNA treatment in GM for 24hr, the medium was replaced with DM. After 2 days in DM, the cells were fixed and co-immunostained. Note the ratio of Myog-positive cell was significantly decreased by siKlf5 treatment (B). Data represent means ± SEM. (*p<0.05) Representative data from at least three individual experiments are shown. (D) Fusion index indicating fibers containing more than 3 nuclei. The fusion index was significantly reduced in cells transfected with siKlf5, though similar levels of MyoD positivity was detected. Data represent means ± SEM. (***p<0.001) Representative data from at least three individual experiments are shown. Scale bar represents 100 µm. N.D.: not detected. (E) Growth curve of siCtrl- and siKlf5-treated C2C12 cells in GM. Data represent means ± SEM. Representative data from four individual experiments are shown.
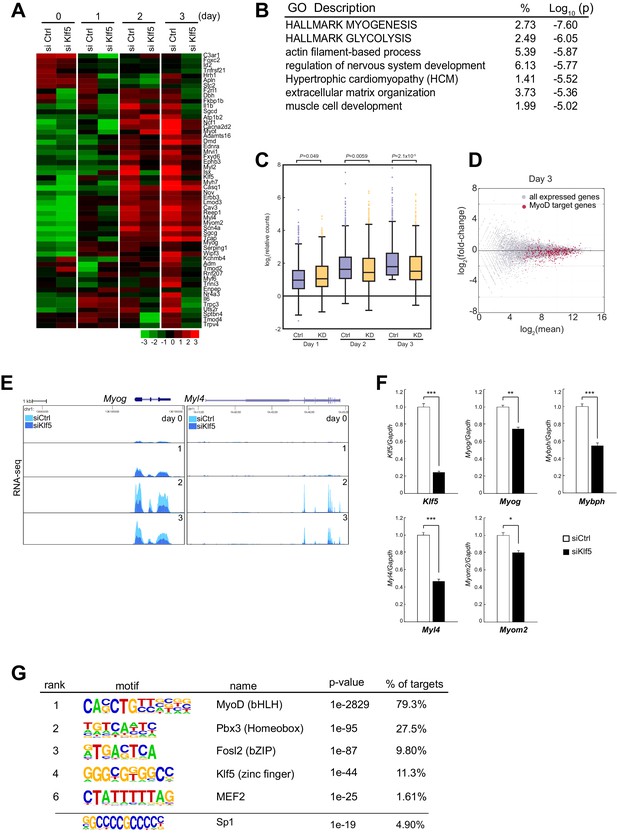
Transient Klf5 knockdown results in a repression of the myogenic program.
(A) Hierarchical clustering and heatmap of the expression levels (log2 normalized counts) of genes involved in the Hallmark Myogenesis (hallmark gene sets M5909) in C2C12 myoblasts transfected with control or Klf5-specific siRNA after induction of differentiation for the indicated times. (B) Functional annotations associated with genes that were inhibited in Klf5 knockdown cells. (C) Relative distribution of RNA-seq tags of genes with MyoD binding during the course of myoblast differentiation. Box-and-whisker plot showing log2 counts of the MyoD-regulated genes and the remaining RefSeq genes normalized to the counts in control siRNA-transfected cells on day 0. P values are between control siRNA (Ctrl) and siKlf5 (KD) cells analyzed using the Mann-Whitney U test. (D) MA plot depicting the relationship between fold changes in myotube differentiation-related genes, comparing RNA-seq from differentiating myoblasts transfected with control siRNA or siRNA targeting Klf5. Red dots represent genes with MyoD binding, and the Gray dots represents genes without MyoD binding. (E) UCSC genome browser images illustrating normalized RNA-seq reads for the representative myogenesis-related genes Myog and Myl4 in control or Klf5 siRNA-transfected C2C12 cells at the indicated days after induction of differentiation. (F) Relative mRNA expression of Klf5, Myog, Mybph, Myl4 and Myom2 in C2C12 cells transfected with control siRNA or siRNA targeting Klf5. Data represent means ± SEM. (*p<0.05, **p<0.01, ***p<0.001, n = 3, biological replicates). (G) de novo motifs identified in regions bound by MyoD in myotubes. The enrichment for a known Sp1 motif was also analyzed.
-
Figure 4—source data 1
Gene ontology analysis on the genes upregulated by Klf5 knockdown.
Functional annotations associated with genes that were induced >1.5-fold with Klf5 knockdown at day 3 post differentiation.
- https://doi.org/10.7554/eLife.17462.011
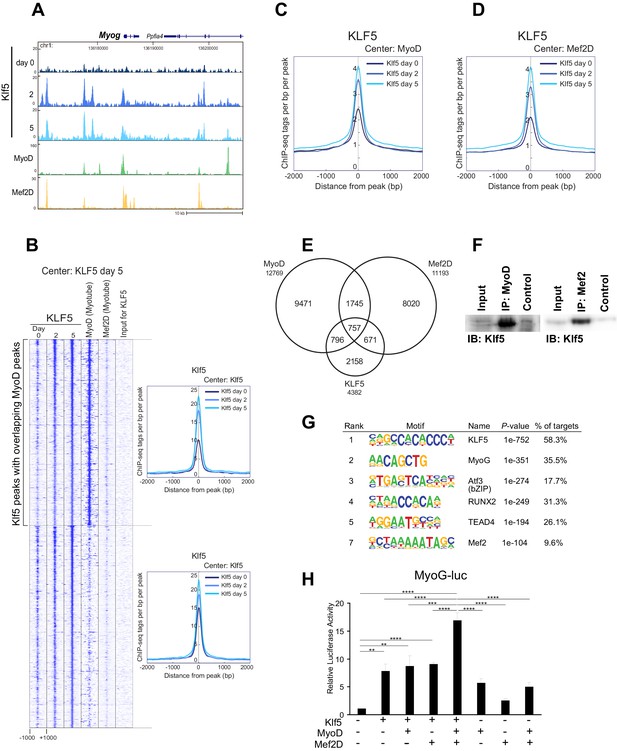
Klf5 regulates myogenesis-related target genes in concert with MyoD.
(A) UCSC genome browser images illustrating normalized tag counts for Klf5 (post-differentiation day 0, 2, 5), MyoD (myotube, differentiated for 2 days) and Mef2D (myotube, differentiated for 5 days) at Myog loci in differentiating C2C12 myotubes. (B) Heatmap for binding of Klf5 (post-differentiation day 0, 2, and 5), MyoD (myotubes, differentiated for 2 days) and Mef2D (myotubes, differentiated for 5 days) within 2 kb around the center of the top 1000 Klf5-binding sites on day 5. The Klf5 binding sites were subdivided into 2 groups based on the presence of significant MyoD binding peaks in myotubes within 500 bp of the Klf5 binding regions. The distribution of Klf5 tag densities in each subgroup were shown as histogram. (C) Distribution of Klf5 tag densities in the vicinity of MyoD-bound loci in myoblasts differentiated for 0, 2 and 5 days. (D) Distribution of Klf5 tag densities in the vicinity of Mef2D-bound loci in myoblasts differentiated for 0, 2 and 5 days. (E) Venn diagram showing the overlap between the 12,769 MyoD peaks, 11,193 Mef2D peaks and 4382 Klf5 peaks identified by ChIP-seq. Note over half the Klf5 peaks (2224 out of 4382 peaks) are also bound by MyoD and/or Mef2. (F) Direct interaction of Klf5 and MyoD or Mef2 shown by co-immunoprecipitation. Whole cell extract from C2C12 cells differentiated for 5 days were immunoprecipitated using anti-MyoD or anti-Mef2 antibody. The interaction between Klf5 and MyoD or Mef2 is shown compared to the 1% Input or control samples without primary antibodies. (G) de novo motifs identified in regions bound by Klf5 in myotubes after differentiation for 5 days. (H). HEK-293T cells were co-transfected with Myog-luciferase; Klf5, MyoD, and Mef2D expression plasmids, as indicated; and a control CMV-renilla luciferase plasmid. Luciferase activities relative to the basal level of Myog-luciferase cotransfected with empty expression vectors are shown. Data represent means ± SEM. (**p<0.01, ***p<0.001, ****p<0.0001). Representative data from three individual experiments are shown.
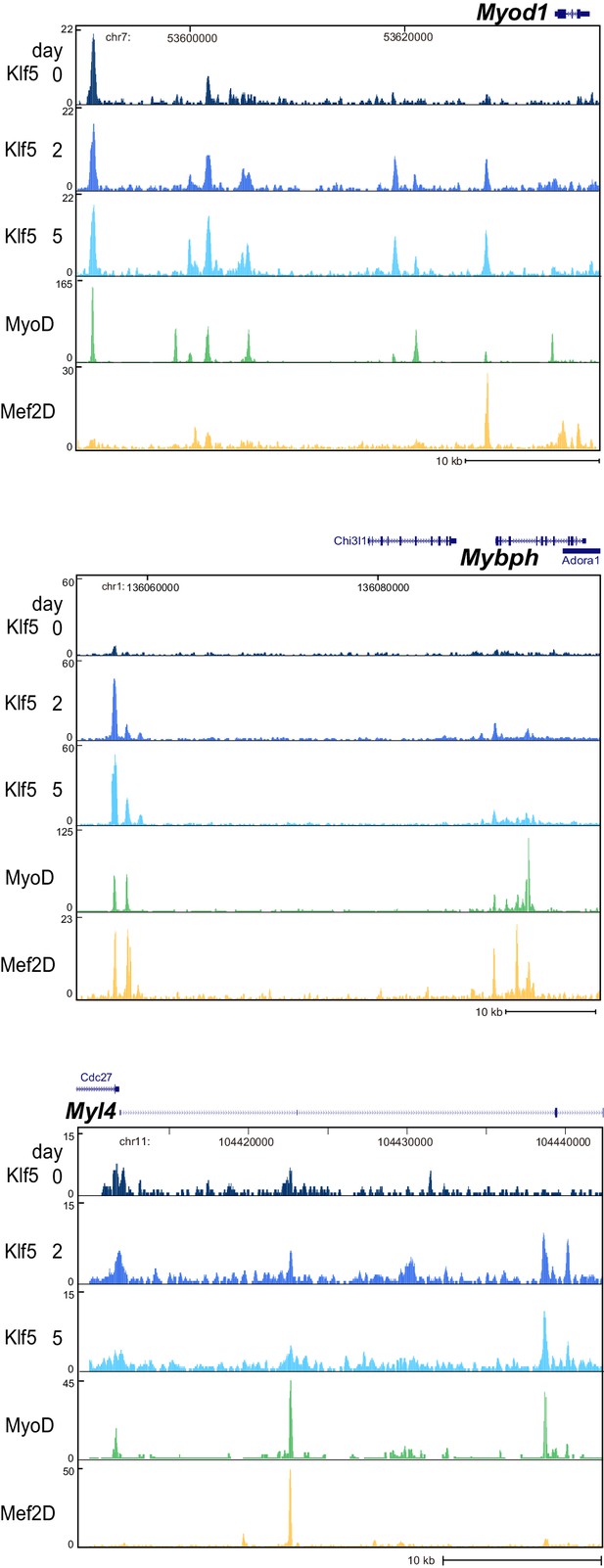
Klf5, MyoD and Mef2D are colocalized in the myogenesis-related gene loci.
UCSC genome browser images illustrating normalized tag counts for Klf5 (post differentiation day 0, 2, 5), MyoD (myotubes, differentiated for 2 days) and Mef2D (myotubes, differentiated for 5 days) at Myod1, Mybph and Myl4 loci in C2C12 myotubes.
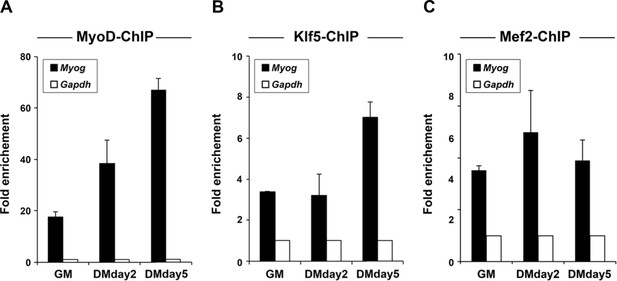
MyoD and Klf5 are recruited to the E-box-containing enhancer of the Myog locus.
Recruitment of MyoD (A), Klf5 (B) and Mef2 (C) to the Myog promoter was monitored using ChIP assays with chromatin prepared from C2C12 myoblasts after differentiation for 0, 2 or 5 days. The Gapdh locus served as a negative control. n = 3, biological replicates.
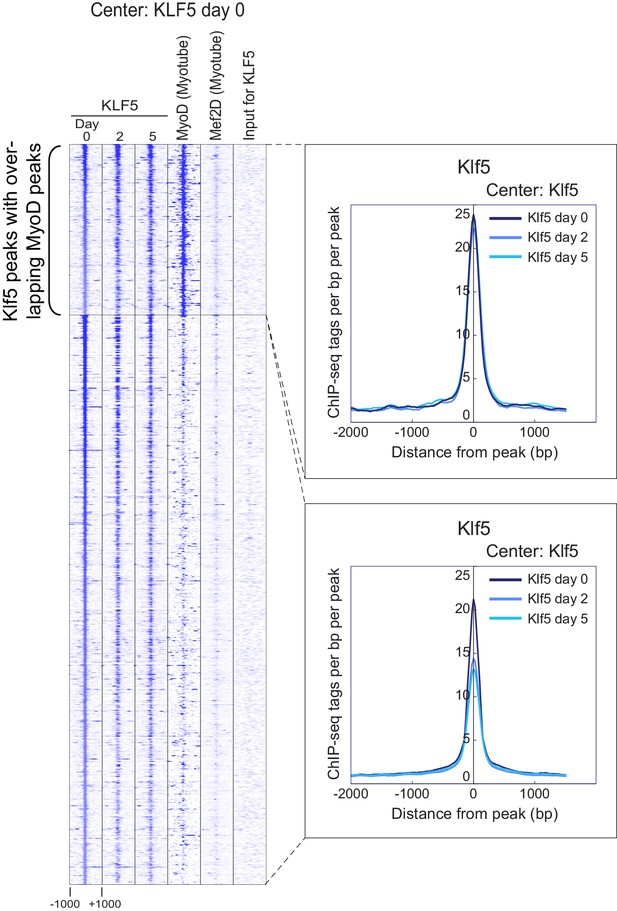
Klf5 binding regions in undifferentiated myoblasts.
Heatmap for binding of Klf5 (post-differentiation day 0, 2, and 5), MyoD (myotubes, differentiated for 2 days) and Mef2D (myotubes, differentiated for 5 days) within 2 kb around the center of the top 1000 Klf5-binding sites on day 0. The Klf5 binding sites were subdivided into 2 groups based on the presence of significant MyoD binding peaks in myotubes within 500 bp of the Klf5 binding regions. The distribution of Klf5 tag densities in each subgroup were shown as histogram.
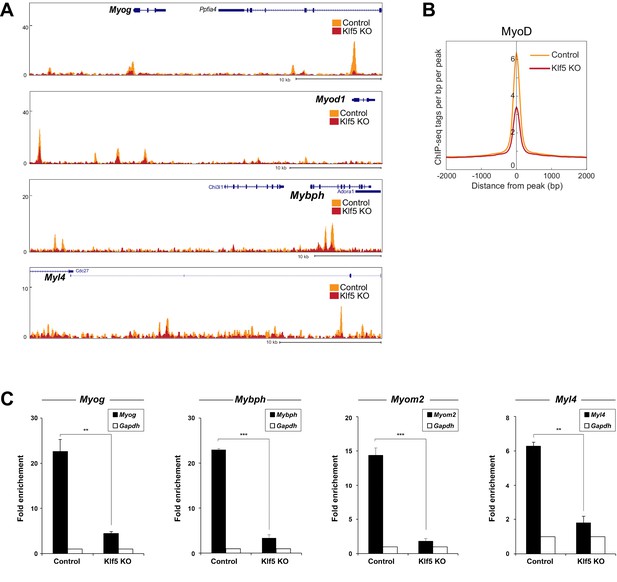
MyoD function is inhibited in Klf5-null C2C12 myoblasts.
(A) UCSC genome browser images illustrating normalized tag counts for MyoD at Myod1, Myog, Myl4 and Mybph gene loci in GFP-targeted control (orange) or Klf5-null (red) C2C12 myotubes differentiated for 3 days. (B) Distribution of MyoD tag densities in the vicinity of MyoD-bound enhancers in the GFP-targeted control or Klf5-null C2C12 myotubes differentiated for 3 days. (C) Comparison of MyoD recruitment at E-box containing enhancers of the Myog, Mybph, Myom2 and Myl4 gene loci in the GFP-targeted control and Klf5 null C2C12 myotubes differentiated for 3 days. Data represent means ± SEM. (**p<0.01, ***p<0.001, n = 3, biological replicates).
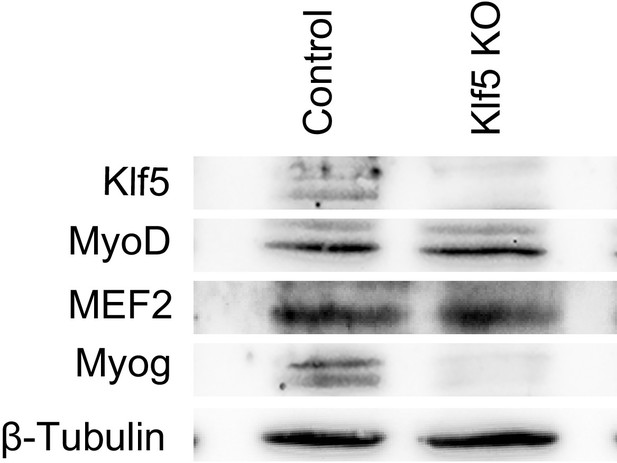
Levels of MyoD and Mef2 protein were not altered by Klf5 deletion.
Levels of MyoD, Mef2, Klf5 and Myog protein were analyzed by Western blotting using GFP-targeted control or Klf5 knockout myoblasts differentiated for 5 days. Representative data from four individual experiments are shown.
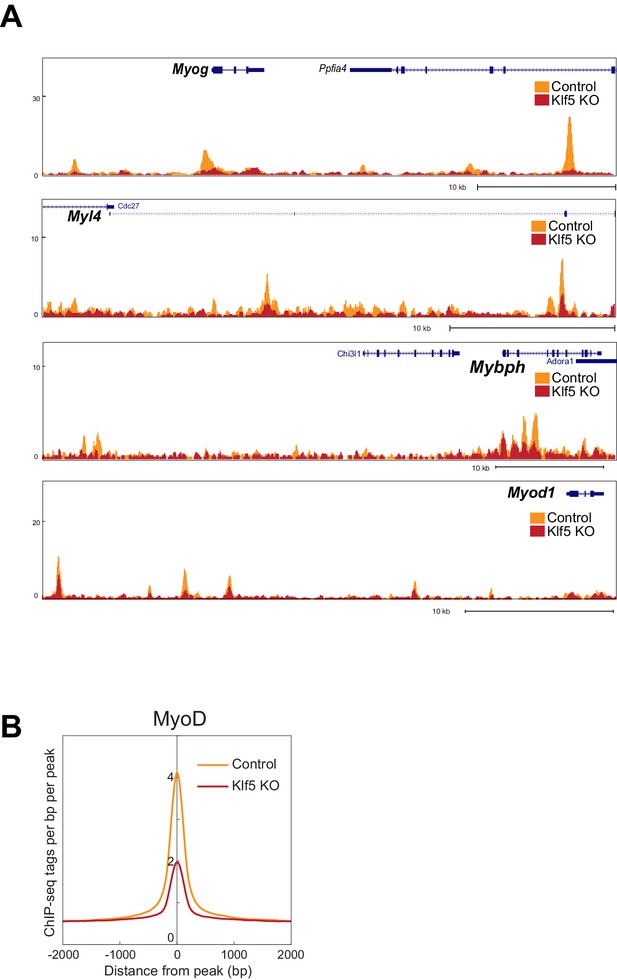
MyoD function is inhibited in Klf5-null C2C12 myoblasts -analysis in the another set of control and Klf5-null cells.
(A) UCSC genome browser images illustrating normalized tag counts for MyoD at Myog, Myl4, Mybph and Myod1 gene loci in GFP-targeted control or Klf5-null C2C12 myotubes differentiated for 3 days. (B) Distribution of MyoD tag densities in the vicinity of MyoD-bound enhancers in GFP-targeted control or Klf5-null C2C12 myotubes differentiated for 3 days.
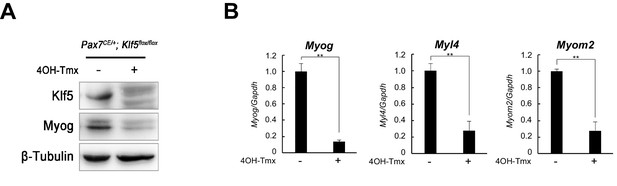
Klf5 is required for satellite cell differentiation.
(A) Isolated SCs from Klf5 knockout mice (Pax7CE/+;Klf5flox/flox) were cultured in the presence or absence of 4OH-Tmx in growth medium for 2 days. Purified protein derived from primary myotubes differentiated for 3 days were analyzed by western blotting. Representative blots from three individual experiments are shown. (B) Relative mRNA expression of Myog Myl4 and Myom2 in SCs differentiated for 3 days. Data represent means ± SEM (**p<0.01). Representative data from three individual experiments are shown.
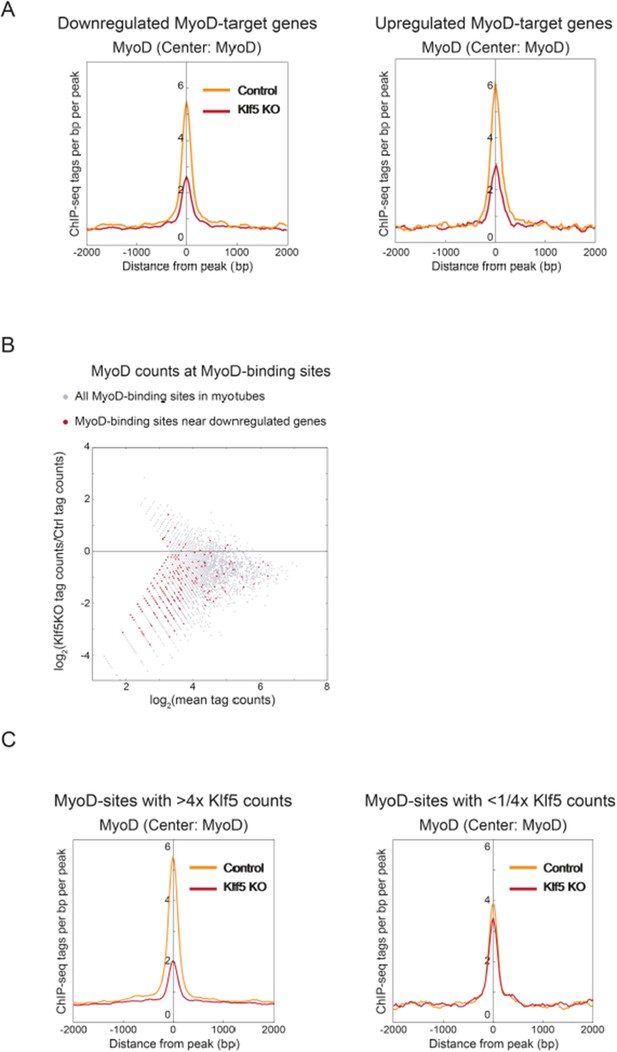
(A) Histograms of MyoD tag densities around MyoD binding peaks proximal to MyoD target genes that were upregulated or downregulated (>1.5-fold) by Klf5 deletion. (B) MA plot of normalized tag counts at MyoD biding regions in Klf5 knockout and control cells. MyoD binding regions proximal to genes whose expression was decreased (>1.5-fold) in Klf5 knockout cells as compared with control cells are shown as red dots. (C) Histograms of MyoD tag densities around MyoD binding peaks that had >4-fold increase or decrease in counts nearby (400 bp) Klf5 between day 0 and 5. MyoD densities in Klf5 knockout and control mice are shown.
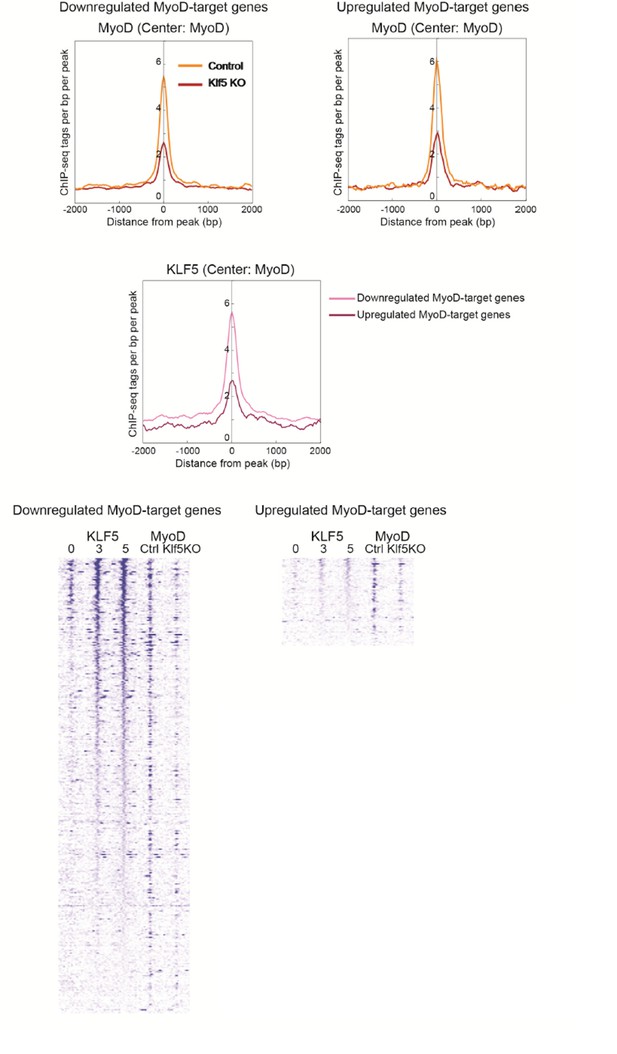
(A) Histograms of MyoD tag densities at MyoD binding regions proximal to MyoD target genes that were upregulated or downregulated (>1.5-fold) by Klf5 deletion. (B) Klf5 binding to MyoD-binding regions proximal to MyoD-target genes whose expression was downregulated or upregulated by Klf5 deletion. (C) Heatmaps for Klf5 and MyoD binding to MyoD-binding regions.
Additional files
-
Supplementary file 1
Table 1: list of oligonucleotides, Table 2: list of antibodies
- https://doi.org/10.7554/eLife.17462.020





