Metabolic network rewiring of propionate flux compensates vitamin B12 deficiency in C. elegans
Figures
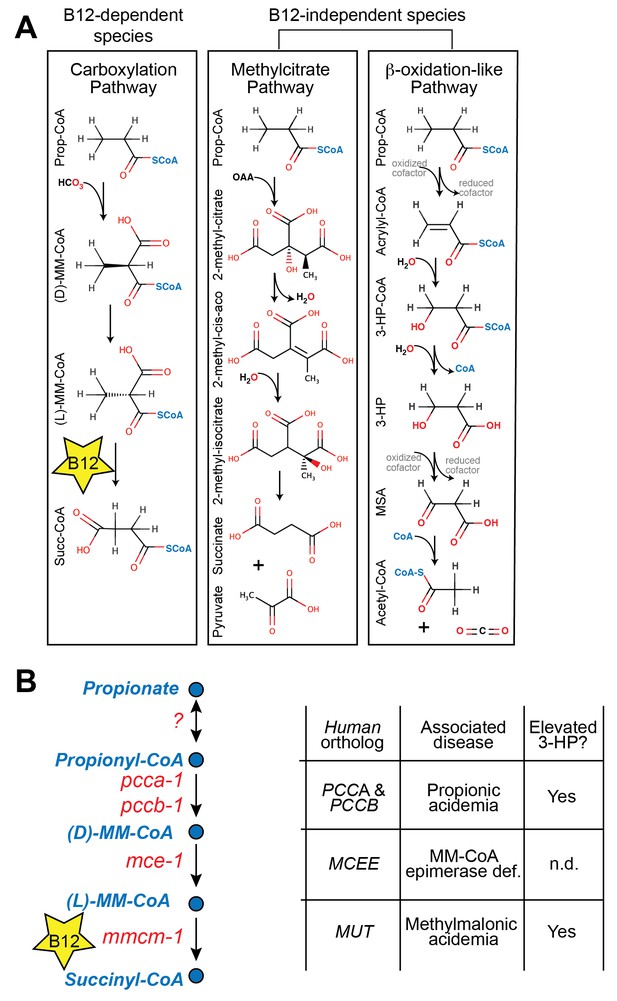
Propionate breakdown pathways in different organisms.
(A) Vitamin B12-dependent species use a propionate carboxylation pathway to breakdown propionate. Other species use either the methylcitrate pathway or a β-oxidation-like pathway. (B) Diagram of canonical vitamin B12-dependent propionyl-CoA breakdown pathway indicating C. elegans and human enzymes and associated genetic diseases. MM – methylmalonyl, 3-HP – 3-hydroxypropionate, MSA – malonic semialdehyde, n.d. – not determined.
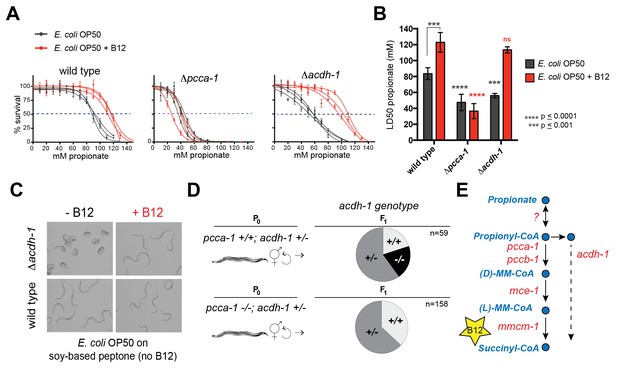
acdh-1 mutants are sensitive to propionate and synthetic lethal with pcca-1 mutants.
(A) Dose-response curves showing that Δpcca-1 and Δacdh-1 mutants exhibit increased sensitivity to propionate compared to wild type animals. Three biological replicate experiments are shown, each with three technical replicates per data point with average and SEM plotted. (B) Average LD50 and standard deviation of data shown in (A). Unpaired student’s T tests were used to calculate p-values. Black asterisks indicate significant difference compared to wild type, red asterisks indicate significant difference compared to wild type plus B12. (C) Δacdh-1 mutants cannot survive on E. coli grown in vitamin B12 deficient media. (D) Δpcca-1 and Δacdh-1 are synthetically lethal because a cross between Δpcca-1 and Δacdh-1 mutants yielded no viable double homozygous mutants. pcca-1 +/+;acdh-1 +/- animals and pcca-1 -/-;acdh-1 +/- animals were grown on E. coli OP50 seeded plates containing 64nM vitamin B12, and individual F1s were picked onto new plates, also containing 64nM vitamin B12. The distribution of acdh-1 genotypes among the viable F1s picked from each P0 genotype is shown. (E) These genetic data support a role for acdh-1 parallel to the canonical propionate breakdown pathway.
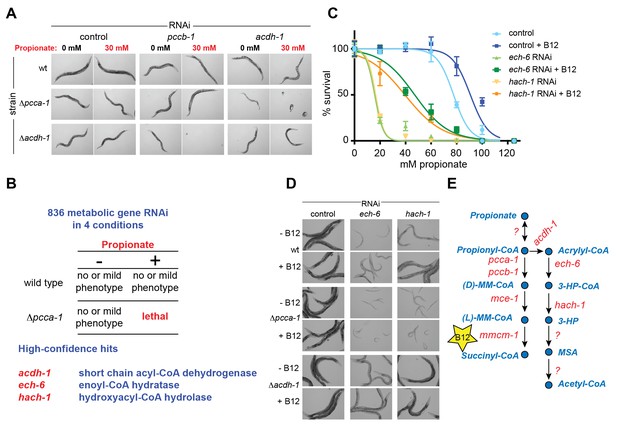
A synthetic genetic interaction screen identifies candidate genes involved in the propionate shunt.
(A) RNAi of acdh-1 is lethal in Δpcca-1 mutants supplemented with 30 mM propionate. (B) Synthetic genetic interaction screen of 836 metabolic genes in presence or absence of 30 mM added propionate, in wild type and Δpcca-1 mutant animals identifies three candidate genes, including acdh-1. (C) Propionate toxicity dose response curve showing that the two candidate genes identified in the screen, ech-6 and hach-1, phenocopy acdh-1 loss-of-function. (D) Genetic buffering of ech-6 and hach-1 RNAi phenotypes by loss of acdh-1. Representative images of animals subjected to two generations of RNAi knockdown are shown. (E) Our data indicate that ech-6 and hach-1 function downstream of acdh-1 in the propionate breakdown shunt.
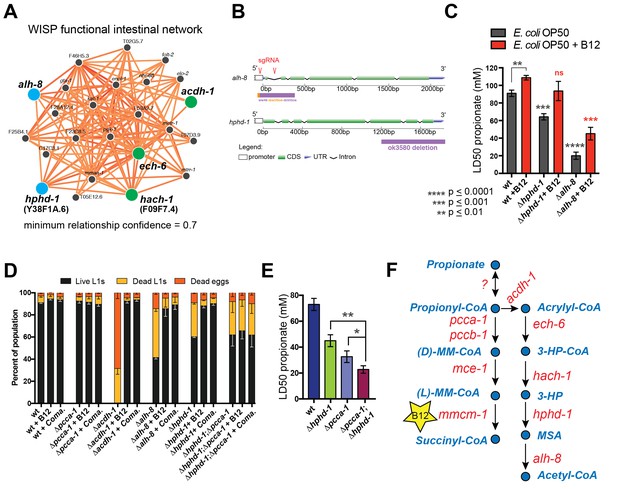
Identifying additional putative propionate shunt genes.
(A) hphd-1 and alh-8 (blue) are tightly connected to acdh-1, ech-6 and hach-1 (green) in a C. elegans intestinal functional network and are candidates to catalyze the fourth and fifth reactions of the propionate shunt, respectively. (B) Structure of CRISPR/Cas9-generated alh-8 mutant. Diagram of the mutation generated by CRISPR/Cas9-mediated genome editing using an sgRNA (red sequence) targeting alh-8. The alh-8(ww48) mutation consists of a 23 bp insertion and 399 bp deletion, and removes a part of the 5’UTR, the start codon, the first and second exons, and part of the third exon. Also shown is the Δhphd-1(ok3590) mutation. (C) Propionate toxicity dose response showing that Δhphd-1 and Δalh-8 mutants phenocopy acdh-1, ech-6 and hach-1 perturbation. (D) Δhphd-1 and Δalh-8 mutants exhibit partial lethality on low-B12 conditions. Like the Δacdh-1 mutant phenotype, Δhphd-1 and Δalh-8 mutant phenotypes were rescued by 64nM B12 supplementation or by Comamonas aquatica DA1877 (Coma.). The partial lethal phenotype of the Δhphd-1;Δpcca-1 double mutant was not rescued by B12. (E) Combined deletion of hphd-1 and pcca-1 renders the animals more sensitive to propionate than mutation in either gene alone. Note that Δhphd-1 may not be a null allele. (F) The C. elegans propionate breakdown shunt pathway comprises five genes: acdh-1, ech-6, hach-1, hphd-1 and alh-8.
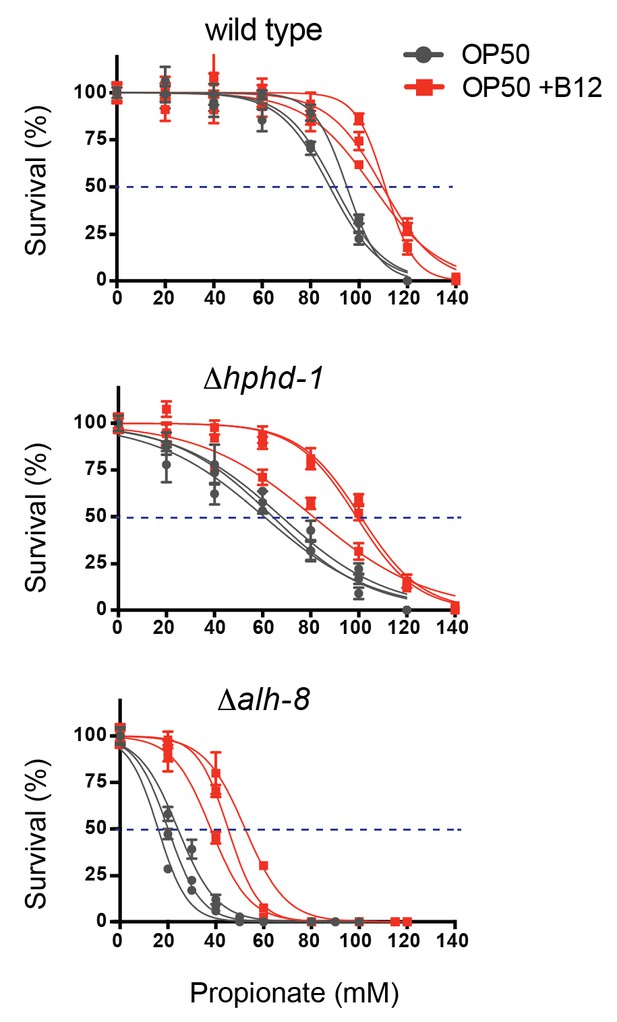
Δhphd-1 and Δalh-8 mutants exhibit increased sensitivity to propionate compared to wild type animals.
Three biological replicate corves are shown.
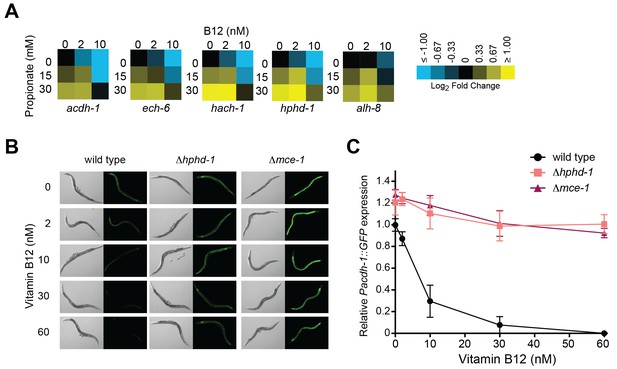
Transcriptional activation of the propionate shunt.
(A) The expression of all five propionate shunt genes is repressed by vitamin B12 and activated by propionate. Condition matrices are shown for each shunt gene. Expression is normalized to levels in the control condition (no vitamin B12, no propionate). (B) Vitamin B12 reduces GFP levels in Pacdh-1::GFP transgenic animals, but not in those carrying a deletion in the canonical propionate pathway gene mce-1 or in the propionate shunt gene hphd-1. (C) Quantification of GFP levels from part (B)
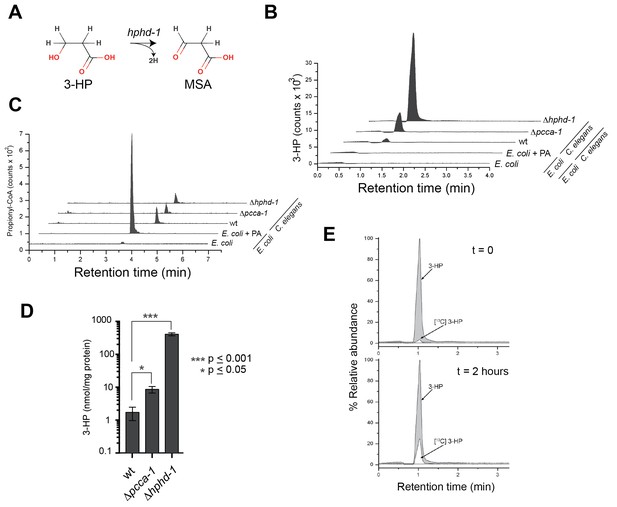
3-Hydroxypropionate is a substrate for HPHD-1.
(A) Conversion of 3-hydroxypropionate (3-HP) into malonicsemialdehyde (MSA). (B) 3-HP mass spectrometry chromatogram for wild type, Δpcca-1 and Δhphd-1 animals. 3-HP was not detected in E. coli OP50 with or without supplemented propionate. (C) Propionyl-CoA chromatograms from E. coli and C. elegans samples. Propionyl-CoA quantifications are as follows: 1.86, 0.20, 0.14, and 0.13 nmol/mg protein for E coli + PA, wild type C. elegans, Δpcca-1 and Δhphd-1 mutants, respectively. For E. coli -PA, propionyl-CoA was detectable but not quantifiable in our assay. (D) Average 3-HP quantities normalized to total protein levels from three biological replicates, +/- SEM. Animals were grown on E. coli OP50. (E) 13C-labeled propionate fed to Δhphd-1 mutant animals for 2 hr yielded 13C-labeled 3-HP, demonstrating that C. elegans oxidizes propionate to 3-HP. Shown are SRM (MS2) chromatograms specific for 3-HP. The peak corresponding to the natural 13C isotope distribution (~ 1.1% of 12C signal) is illustrated for comparison in t = 0.
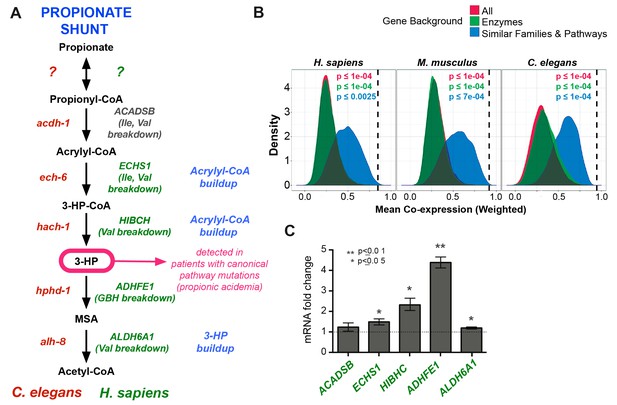
Comparison between putative human and C. elegans propionate shunts.
(A) Comparison between C. elegans propionate shunt genes (red) and candidate human shunt genes (green, gray). Green text indicates higher confidence annotations based on patient mutations and metabolomics, or in the case of ADHFE1 one-to-one orthology of unique enzymes in both genomes. 3-HP is marked in magenta to indicate that it is a biomarker for impaired flux in the canonical, vitamin B12-dependent propionate breakdown pathway, such as occurs in patients with propionic or methylmalonic acidemia. (B) C. elegans propionate shunt genes and orthologs in mouse and humans are strongly co-expressed as a group compared to 10,000 random permutations of five genes from either the whole genome, a subset of only metabolic genes, or a subset of related metabolic genes from connected pathways, including BCAA breakdown and the TCA cycle. The expression data used for this analysis was compiled and weighted using the SEEK and modSEEK databases. Distributions of co-expression scores are shown for each set of randomizations, and vertical dashed lines indicate actual weighted co-expression score for propionate shunt genes and orthologs in human, mouse and C. elegans. (C) The expression of several candidate human genes is activated in response to propionate in HepG2 liver carcinoma cells. qRT-PCR experiment showing the average of four replicate experiments, each containing three technical replicates.
Additional files
-
Supplementary file 1
Summary of acdh-1 genotypes in F1 generation of P0 pcca-1 -/-; acdh-1 +/- animals.
The acdh-1 genotype (homozygous, heterozygous, or wild type) as determined by PCR for each F1 animal chosen from P0 pcca-1 +/+; acdh-1 +/- or pcca-1 -/-; acdh-1 +/- parents is shown, as well as an indication of whether the F1 animal laid non-viable embryos among the F2 offspring.
- https://doi.org/10.7554/eLife.17670.011
-
Supplementary file 2
Metabolic gene lists. A manually curated list of putative metabolic genes in C. elegans.
The 836 genes that were screened as part of the metabolic RNAi library are indicated.
- https://doi.org/10.7554/eLife.17670.012
-
Supplementary file 3
Evidence summary for dual roles of human orthologs in propionate oxidation shunt and other metabolic pathways.
- https://doi.org/10.7554/eLife.17670.013
-
Supplementary file 4
Genotyping and qPCR primers.
- https://doi.org/10.7554/eLife.17670.014





