Allosteric activation of SENP1 by SUMO1 β-grasp domain involves a dock-and-coalesce mechanism
Figures
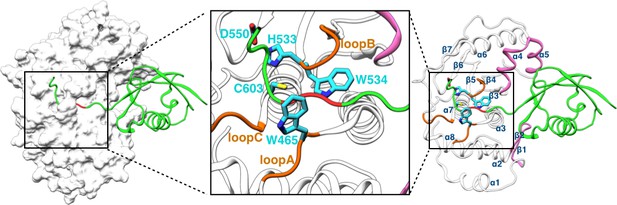
Structure of the SENP1-SUMO1 complex (PDB entry 2IY1).
SENP1 is shown as gray surface and cartoon representations in the left and right panels, respectively, and the closed catalytic channel is boxed and enlarged in the middle panel. Sidechains of the catalytic triad and of two tryptophans shaping the catalytic channel are shown as sticks. Three channel-lining loops (residues 464–466, 530–533, and 599–602) are shown in orange; two exosite interface regions (residues 443–453 and 496–514) are shown in mauve. SUMO1 is shown in green but with the conserved C-terminal Gly-Gly motif in red.
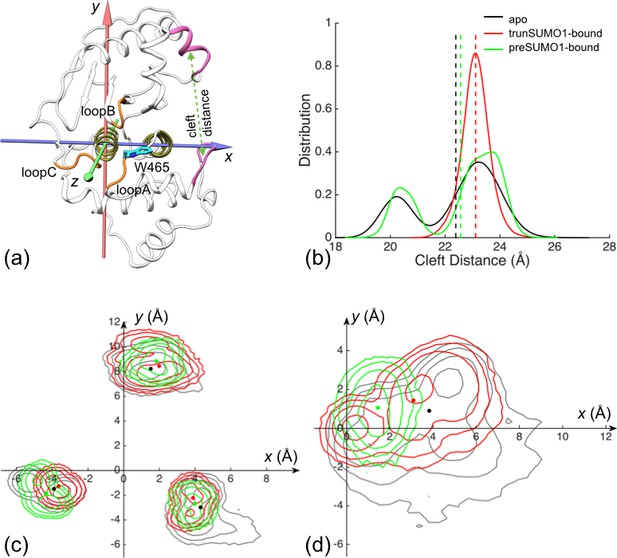
The displacements of the exosite interface regions and channel-lining loops upon SUMO1 binding.
(a) The regions (displayed in mauve) used for defining cleft distance (indicated by double-headed arrow) and the coordinate system used for defining movements of the three channel-lining loops (displayed in orange) and the W465 sidechain (displayed with carbon atoms in cyan). (b) The distributions of the cleft distance in the simulations of the apo and trunSUMO1- and preSUMO1-bound forms of SENP1. The average value of each system is shown as dash with matching color. (c) Distributions of the Cα centers of the three loops in the x-y plane. The average positions of the loops in each system are shown as dots. (d) Corresponding results for the center of W465 sidechain heavy atoms.
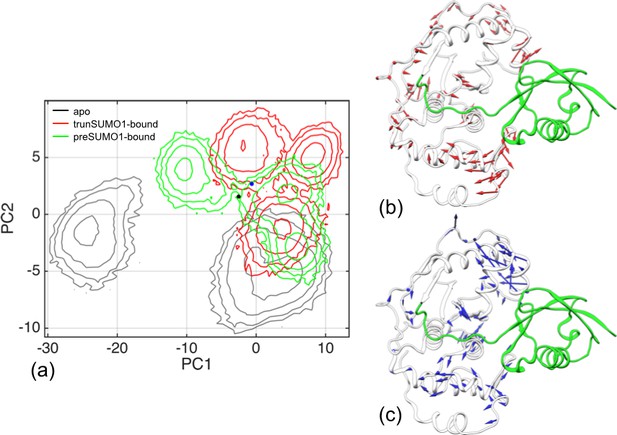
Difference in conformational sampling among apo SENP1 and two SUMO1-bound forms.
(a) Conformational probability densities in the plane of the first two principal components (PC1 and PC2), transformed into free energy surfaces according to the Boltzmann relation. (b–c) Movements represented by PC1 and PC2, respectively.
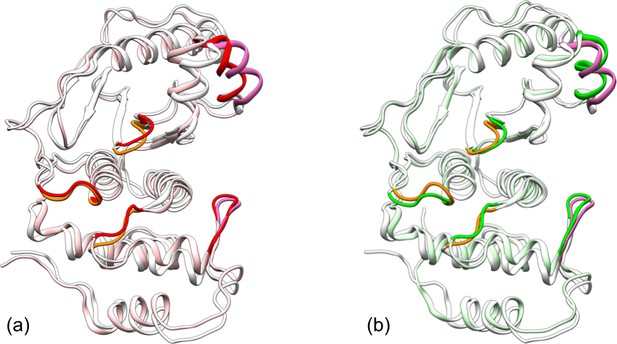
Comparison of representative structures from replicate simulations of different systems.
A structure for apo SENP1 is overlaid with those for the (a) trunSUMO1-bound and (b) preSUMO1-bound forms, by superposition of the α3 and α7 helices. Apo SENP1 is shown in gray except with the three channel-lining loops in orange and the two interface regions in mauve. These portions are in red and green for the trunSUMO1-bound and preSUMO1-bound forms, respectively.
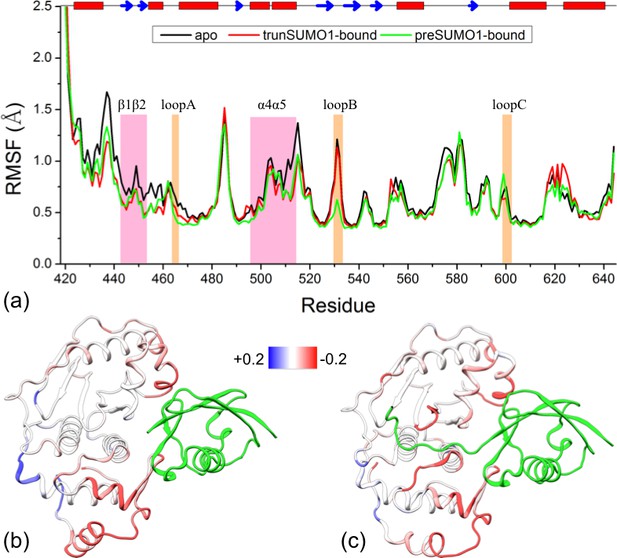
Comparison of flexibility among the three systems, as measured by Cα atom root-mean-square fluctuations (RMSFs).
(a) Variations of RMSF along the amino acid sequence for the apo, trunSUMO1- and preSUMO1-bound forms of SENP1. The two exosite interface regions and three channel-lining loops are highlighted by shading in mauve and orange, respectively. (b–c) Changes in RMSF upon binding trunSUMO1 and preSUMO1, displayed on the bound structures according to a color scale (shown; red and blue corresponding to lower and higher flexibilities, respectively).
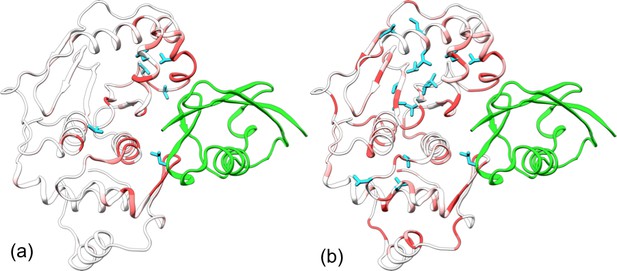
Comparison of experimental and calculated chemical shift perturbations (CSPs) of SENP1 upon trunSUMO1 binding.
(a) NMR results of Chen et al. (2014). (b) Calculated results. Backbone amide CSPs are displayed according to a color scale (red to gray corresponding to high to low CSPs); sidechain methyls with significant CSPs (>0.05 in panel a and >0.08 in panel b) are shown as cyan sticks.
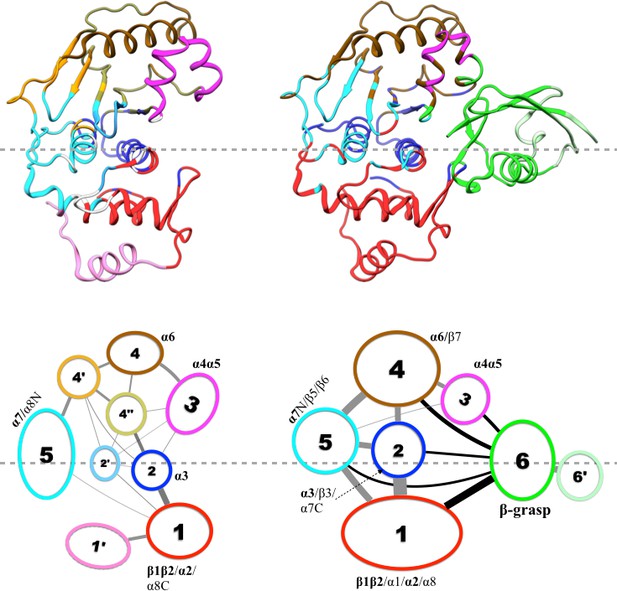
Results of community analysis for apo and trunSUMO1-bound SENP1 sim1, displayed on the left and right panels, respectively.
Communities are displayed either by different colors on the structures (upper row) or as numbered ovals with matching colors (lower row). In the lower row, inter-community cumulative betweennesses are displayed by the thickness of the lines connecting communities. The community analysis was performed using the NetworkView plugin in VMD (Sethi et al., 2009).
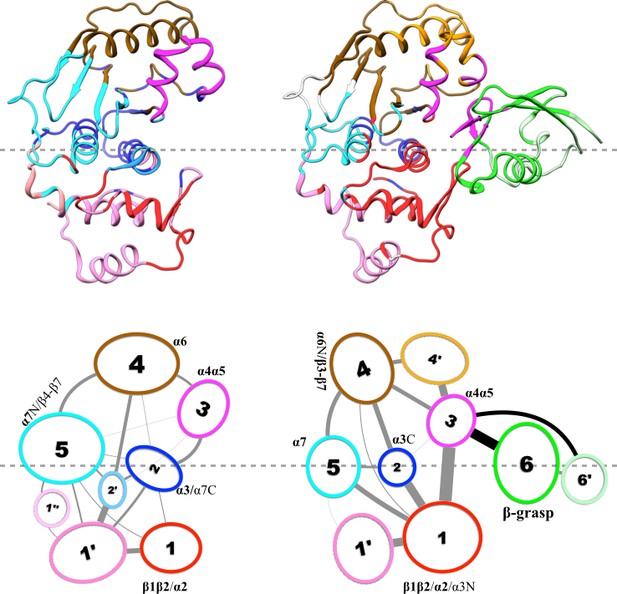
Results of community analysis for apo and trunSUMO1-bound SENP1 sim2.
For caption, see Figure 5. Note that, for trunSUMO1-bound SENP1, β-grasp residues in the lower interface (and upper interface) are collected into community 3; hence the coupling between communities 1 and 6 seen in the other two replicate simulations is now part of the coupling between communities 1 and 3.
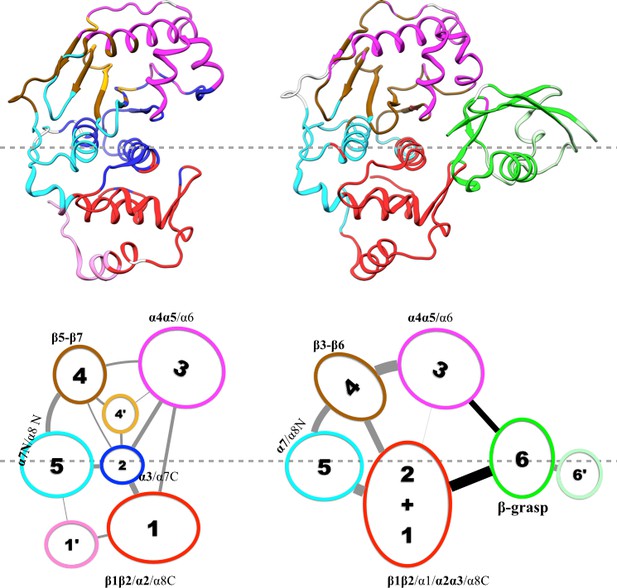
Results of community analysis for apo and trunSUMO1-bound SENP1 sim3.
For caption, see Figure 5.
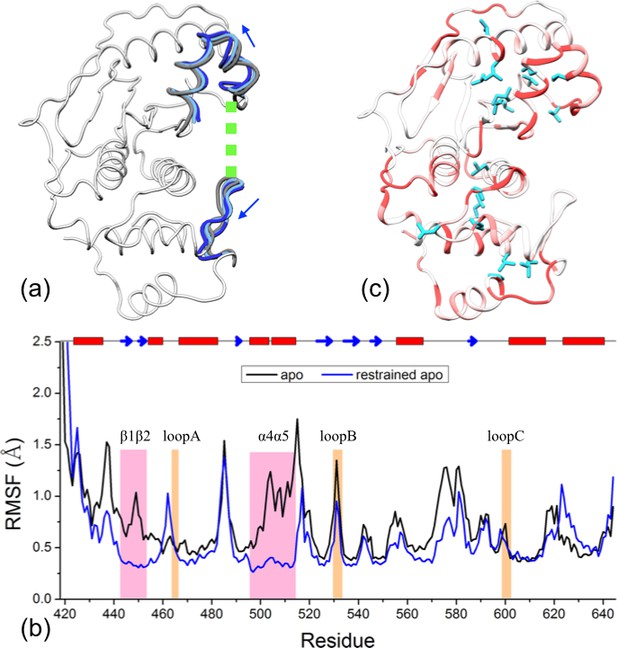
Reproduction of allosteric effects by restraining the exosite interface regions of apo SENP1 at a widened separation.
(a) Illustration of the restraint. Blue arrows indicate the widening of the exosite cleft, and green dash indicates the subsequent restraint. (b) Comparison of RMSFs between apo sim1 and the restrained apo simulation. (c) Backbone amide and sidechain methyl CSPs of the restrained simulation (to be compared with Figure 4b).
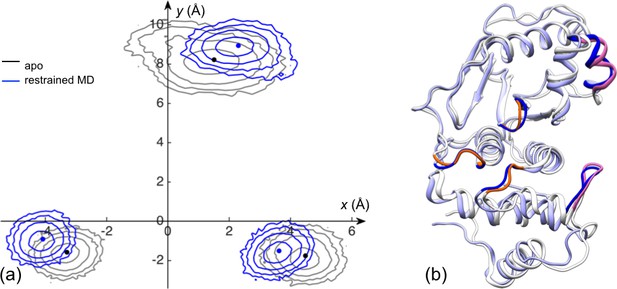
The displacements of the channel-lining loops for a control simulation in which the exosite interface regions of apo SENP1 are restrained at a widened separation.
(a) Distributions of the Cα centers of the three loops in the x-y plane. Results for apo sim1 and the restrained simulation are shown in black and blue, respectively. The average positions of the loops in each system are shown as dots. (b) Comparison of representative structures from apo sim1 and the restrained simulation. For caption, see Figure 2—figure supplement 2.
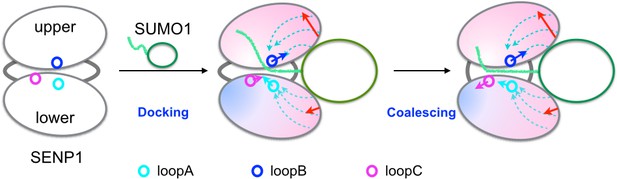
Illustration of the dock-and-coalesce mechanism for SENP-catalyzed SUMO C-terminal cleavage.
SENP is shown as two ovals (representing two subdomains) connected by two strings, with the three channel-lining loops highlighted as small circles; SUMO is shown as an oval (representing the β-grasp domain) with a tail (the C-terminus). In the docking step, as the β-grasp domain wedges into the exosite cleft, the cleft separation widens (indicated by red arrows), loopA and loopB move (cyan and blue arrows) to make stronger inter-subdomain contact and also to create space for docking the proximal portion of the SUMO C-terminus; the two SENP subdomains lose flexibility on the fast timescale except for a distal region (indicated by red to blue shading), and finally the proximal portion of the SUMO C-terminus docks into the catalytic channel. Two hydrophobic pathways (bundles of dashed arrows) propagate the allosteric effects from the exosite interface regions to the catalytic center. In the coalescence step, the wedged β-grasp domain and the docked C-terminus cooperate to reinforce allosteric effects, initiating inter-subdomain correlated slow motions to allow for proper alignment of the substrate around the catalytic center.





