An extended retinotopic map of mouse cortex
Figures
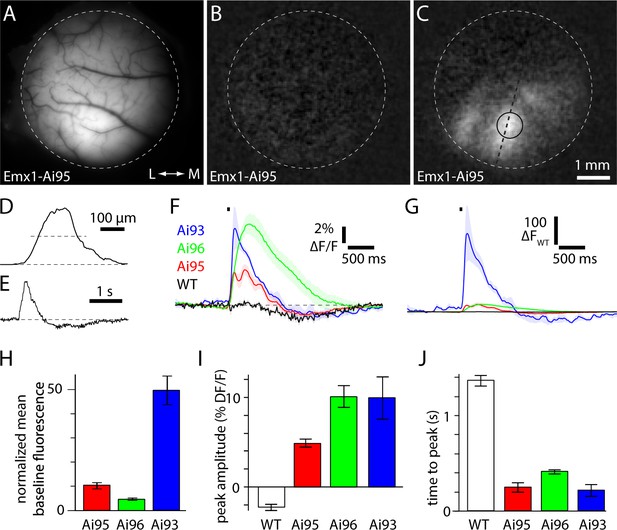
Amplitude and kinetics of fluorescence transients to brief visual stimuli.
(A) Fluorescence image focused on the cortical surface of an Emx1-Ai95 mouse. Approximate edge of 5 mm diameter craniotomy is marked with a dashed line. (B and C) Baseline (B) and peak (C) change in fluorescence for the field of view shown in A. In C, dashed line and circle indicate the regions used to extract values for D and E, respectively. (D) Spatial extent of the fluorescence change along line marked in panel C. Width of response at half height (dashed line) is 190 µm. (E) Fluorescence time course for a single trial, from the region marked in panel C. D and E are in arbitrary fluorescence units. (F) Mean ± SEM fractional fluorescence changes for each mouse line. Four wild-type mice (black), 6 Emx1-Ai95 mice (red), 4 Emx1-Ai96 mice (green), 4 Emx1-Ai93 mice (blue). Stimulus indicated with black bar. (G) Mean change in fluorescence (△F) for each mouse line, normalized to the peak amplitude of the fluorescence change in wild-type mice (△FWT). Shaded areas denote ± SEM. (H) Time-averaged baseline fluorescence, normalized to wild-type mice. Bears denote mean ± SEM from four wild-type mice, 6 Emx1-Ai95 mice, 4 Emx1-Ai96 mice, 4 Emx1-Ai93 mice. (I) Peak fractional fluorescence change. (J) Time to peak fluorescence, measured from the onset of the stimulus.
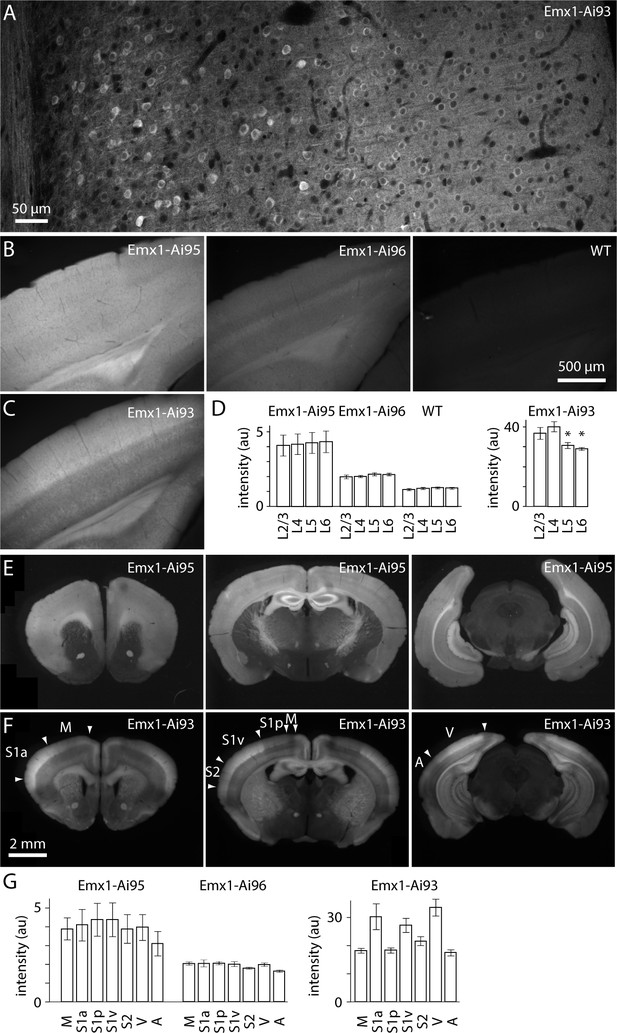
Expression of GCaMP6 in Emx1-Ai95, Emx1-Ai96 and Emx1-Ai93 mice under the Emx1-IRES-Cre driver.
Images of GCaMP6 fluorescence in 100 µm-thick coronal sections. (A) Two-photon image of GCaMP6 fluorescence throughout the depth of cortex in an Emx1-Ai93 mice. White matter is to the left of the image, pia to the right. Three stitched images in a single optical plane. Note the absence of GCaMP6 from the nuclei; we observed nuclear exclusion in all neurons examined (271 neurons in layer 2/3 of visual cortex, 1 Emx1-Ai93 mouse). (B) Widefield images of visual cortices of an Emx1-Ai95, an Emx1-Ai96 and a wild-type mouse. Same spatial and intensity scale for all images. (C) Widefield image from an Emx1-Ai93 mouse. Same spatial scale, but a different intensity scale from the images in B. (D) Summary of laminar variations in fluorescence intensity in visual cortex. Bars represent mean ± SEM fluorescence (arbitrary units). n = 4 (two hemispheres from two mice for each line). Emx1-Ai93: asterisks denote significant difference in intensity compared to layer 4 (p<0.05, two-tailed t-test). (E) Widefield fluorescence images of three 100 µm-thick coronal sections from an Emx1-Ai95 mouse. (F) Widefield fluorescence images of three coronal sections from an Emx1-Ai93 mouse. Approximate of cortical areas (arrow heads) were derived from Franklin and Paxinos (2007). M motor cortex, S1a anterior primary somatosensory cortex (likely jaw and forepaw representations), S1p posterior primary somatosensory cortex (likely trunk and hindpaw representations), S1v vibrissal primary somatosensory cortex, S2 secondary somatosensory cortex, V visual cortex, A auditory cortex. (G) Summary of areal variations in fluorescence intensity. Bars represent mean ± SEM fluorescence (arbitrary units). n = 4 (two hemispheres from two mice for each line). In Emx1-Ai95 mice, GCaMP6 was evenly distributed across the layers of neocortex (Figure 1B,D) and across neocortical areas (Figure 1E,G). Fluorescence displayed laminar and areal variations in Emx1-Ai93 mice (Figure 1C,D,F,G), likely due to use of the CaMK2a-tTA line to enhance expression of GCaMP6 (Mayford et al., 1996; Krestel et al., 2001); Allen Mouse Brain Connectivity Atlas, Transgenic Characterization http://connectivity.brain-map.org/transgenic).
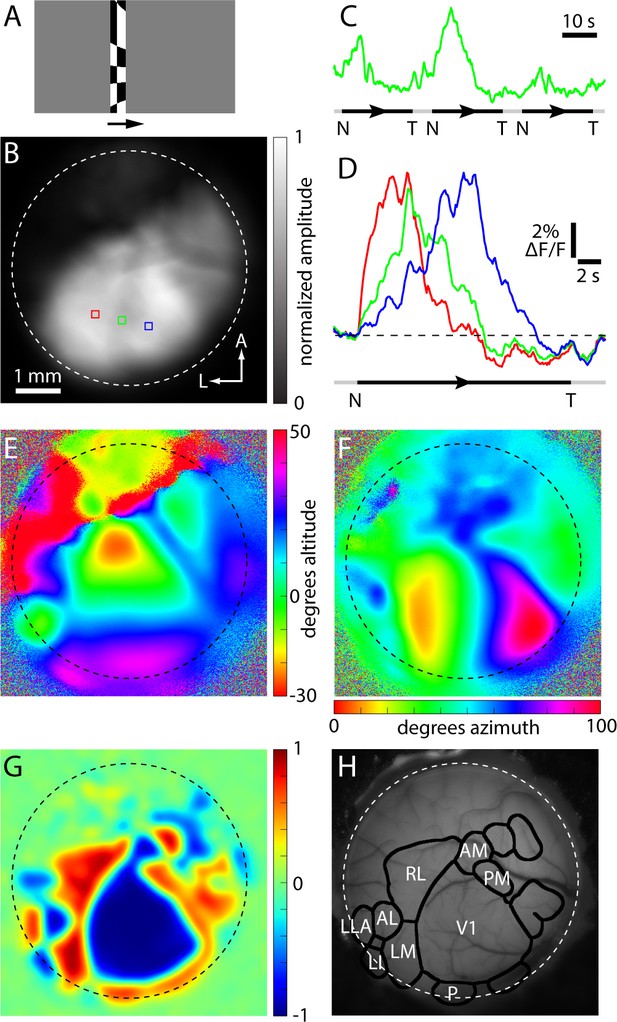
Example of a GCaMP6 fluorescence-based retinotopic map.
Example of a GCaMP6 fluorescence-based retinotopic mapping data set from an awake mouse, generated with the drifting checkerboard stimulus on a grey background. (A) A single image from the visual stimulus movie used to map in the nasal to temporal (azimuth) direction in an Emx1-Ai96 mouse. The checkerboard pattern was swept from left to right (arrow) on a grey background. (B) Greyscale image illustrating the amplitude of the fluorescence change at 0.043 Hz during azimuth mapping, normalized to the maximum amplitude in the image. Dashed white circle approximates the border of the cranial window. (C) Spatial average fluorescence during nasal (N) to temporal (T) mapping from the region of interest outlined in green in panel B. The timing of stimulus presentation is indicated below, where the black segments indicate the center position of the checkerboard bar from −14 to 132 degrees (azimuth) and the grey segments indicate that no stimulus was on the monitor. (D) Fractional fluorescence changes from the three regions marked in panel B. Each trace is the average of 10 presentations of the stimulus. (E and F) Altitude and azimuth maps for the same cranial window. (G) Field sign map derived from the altitude and azimuth maps. (H) The result of automated border identification, drawn on a brightfield image of the brain surface over visual areas. Named visual areas were identified manually, based on published maps of visual areas.
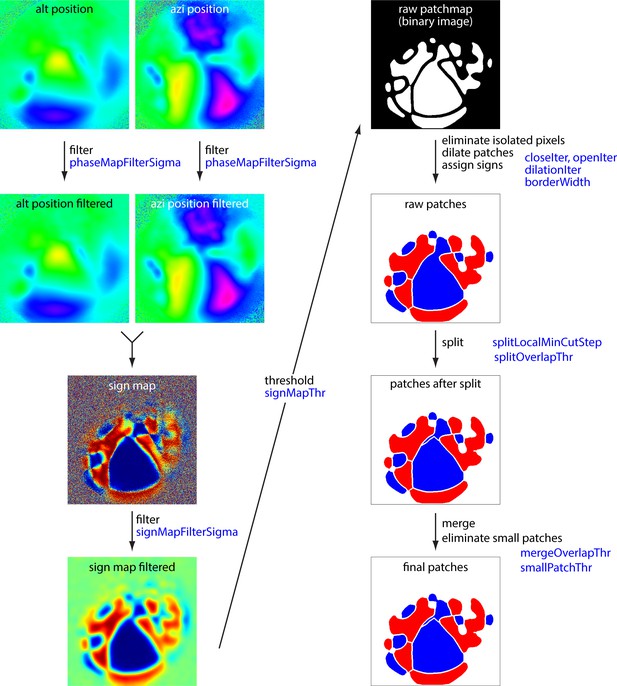
Schematic summary of border identification routine.
Summary of analysis steps for the generation of borders from altitude and azimuth maps. The left column summarizes the steps from altitude and azimuth maps to the sign map. The right column summarizes generation of borders from the sign map. Image names match those in the example Python notebook available at https://github.com/zhuangjun1981/retinotopic_mapping Blue text indicates the main variables employed at each step. Variables are described in the example notebook.
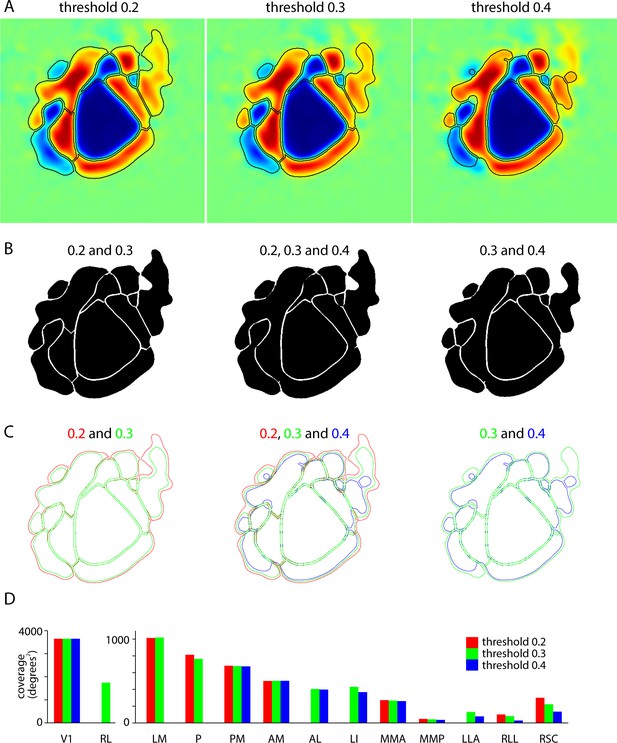
Effects of sign map threshold on patch size, shape and visual coverage.
(A) Comparison of borders (black lines), calculated using threshold values (variable signMapThr in analysis code) of 0.2, 0.3 and 0.4. For each panel, borders were calculated (and are overlaid onto) the mean sign map (Figure 3C), with all variables within the analysis code held constant except the sign map threshold. Thresholds of 0.2 and 0.4 are the limits of the range of values employed in the analysis of our data sets and a threshold of 0.3 is close to the mean value (mean = 0.32). (B) Overlaid images of patches. Patches are shown in black, with the result that invariant borders are readily visible in white. Note that borders between two patches are generally stable unless the threshold is raised enough to eliminate the border. For example, all the borders of V1 are virtually invariant across the threshold range from 0.2 to 0.4; borders between RL and LM and between LM and P are unaffected by the change in threshold from 0.2 to 0.3, but are absent at a threshold of 0.4. (C) Borders overlaid in color to emphasize the sensitivity the exterior borders of the map to changes in threshold. Exterior borders typically retract as the threshold is raised. (D) Effects of threshold on the visual coverage of each patch. For most patches, the change in coverage with threshold is mild and is less than the difference in coverage between patches. As expected, the change in coverage is more pronounced for patches on the periphery of the map that display weaker changes in fluorescence to the visual stimulus, such as patches LLA and RLL.
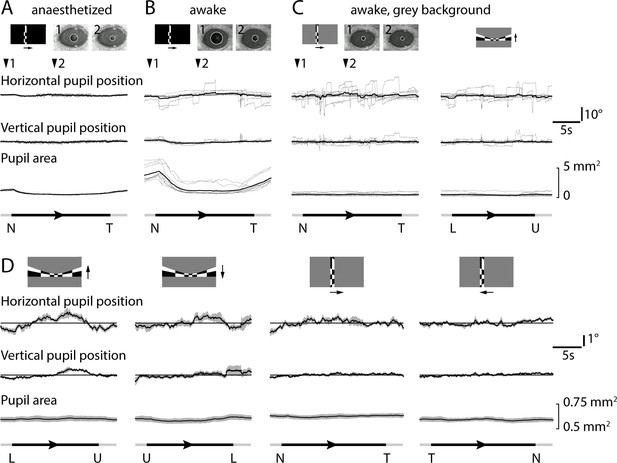
Eye movements and pupil area during retinotopic mapping.
(A) Example of pupil position and area in an anesthetized mouse during azimuth mapping on a black background on which the stimulus moved in the nasal-to-temporal direction. Stimulus is shown schematically (top left). Images of the pupil at time points 1 and 2 are overlaid with circles fit by analysis software that locates the reflection of the infrared LED (inner, black circle) and the perimeter of the pupil (outer, white circle). Pupil position in horizontal and vertical dimensions were measured relative to the mean position, with ten individual traces in grey and the mean in black. Stimulus position is indicated as in Figure 4. (B) Example of pupil position and area in an awake mouse during azimuth mapping on a black background. (C) Example of pupil position and area in an awake mouse during azimuth and altitude mapping on a grey background of 50% of maximum luminance. (D) Mean pupil location and area during grey-background retinotopic mapping with all four stimulus directions. Mean (black line) ± SEM (grey area) results from 11 mice. Horizontal lines indicate mean horizontal or vertical pupil position.
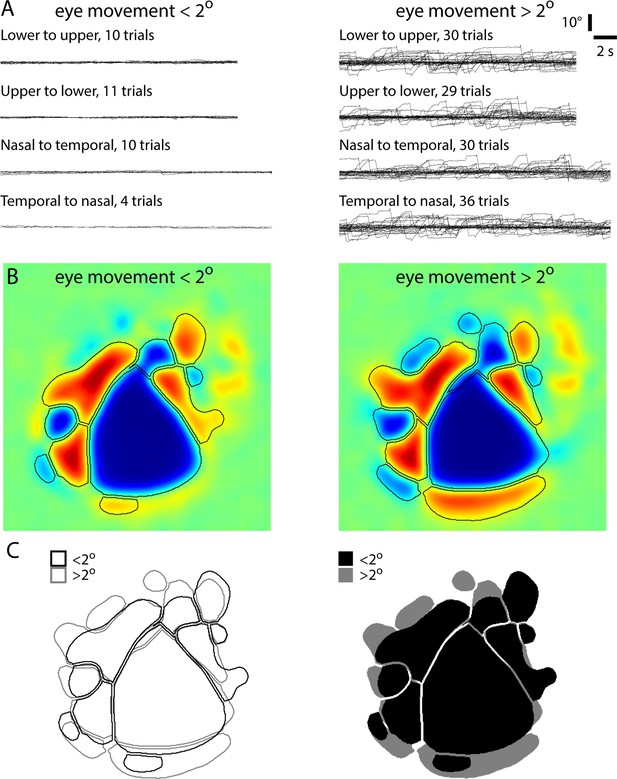
Comparison of maps with and without eye movements.
Maps from an Emx1-Ai96 mouse, generated after sorting trials into those with and those without eye movement of two degrees or greater. (A) Eye position for individual trails, sorted by stimulus direction. (B) Resulting sign maps and border locations. (C) Comparison of patches. Borders between visual areas (e.g. between V1 and LM/RL) were largely unaffected by eye movements. The exterior borders of the sign map display greater differences, either as a result of eye movements or of averaging across the relatively small number of trials without eye movements.
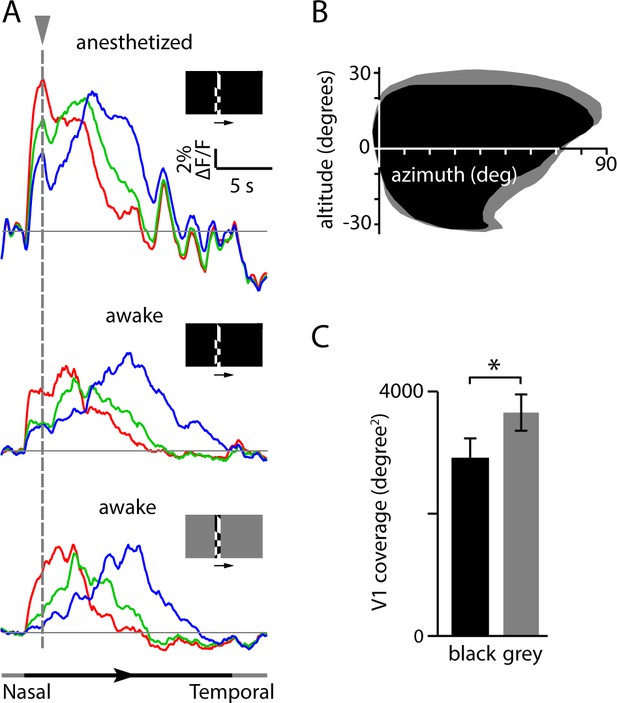
Comparison of fluorescence changes and V1 coverage with stimuli on black and grey backgrounds.
(A) Fractional fluorescence changes from three regions in V1 (see Figure 2B) from mice mapped under three conditions: mouse anesthetized, stimulus on black background; mouse awake, stimulus on black background; and mouse awake, stimulus on grey background. Each trace is the average of 10 presentations of the stimulus. Dashed vertical line marks the initial fluorescence transient following appearance of the stimulus at the nasal edge of the monitor. The timing of stimulus presentation is indicated below. When presented on a black background, the initial appearance of the checkerboard stimulus evoked an increase in fluorescence across much of cortex (and a decrease in pupil size; see Figure 2—figure supplement 3A,B). This global fluorescence transient could shift the calculated phases of the peak responses, resulting in miscalculation of retinotopic position. (B) Comparison of coverage for V1 in one mouse, mapped with black and grey backgrounds. The apparent coverage of V1 was reduced when the stimulus was presented on a black background. The coverage was 3481 degrees2 with black background and 4032 degrees2 with grey background. (C) Population average of V1 coverage for 11 mice (mean ± SEM, black and grey bars indicate black and grey backgrounds, respectively, p<0.05, two-tailed t-test). Mapping using a black background resulted in a 21% reduction in coverage of V1 relative to mapping with a grey background (coverage 2902 ± 349 degrees2 with a black background and 3655 ± 313 degrees2 with a grey background, 11 mice, p<0.05, paired t-test).
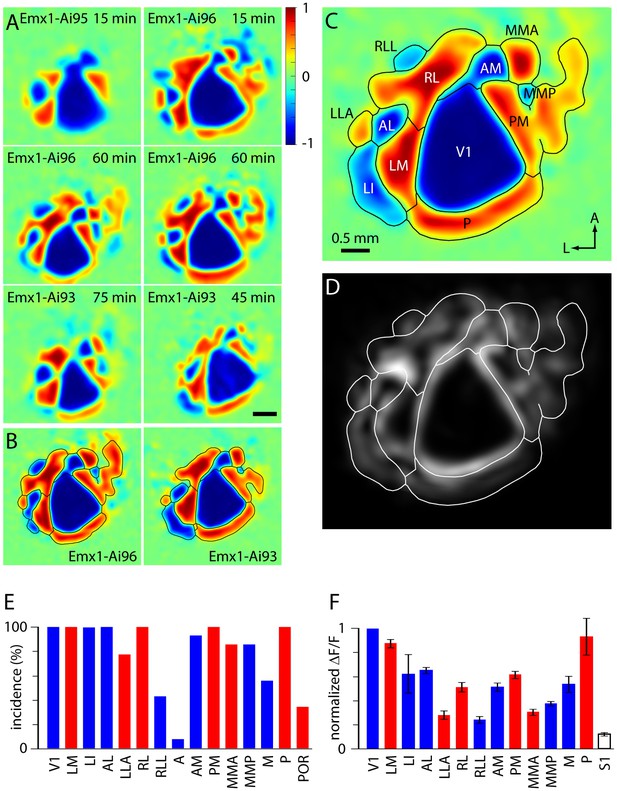
Retinotopic organization of mouse visual cortex.
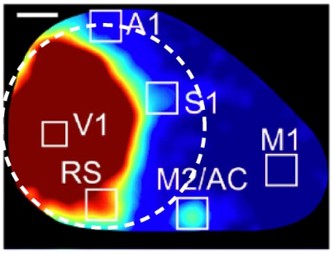
(A) Field sign maps from six mice, illustrating differences between mouse lines and individual mice. The mouse line and duration of imaging are indicated on each map. Scale bar 0.5 mm. (B) Mean field sign maps for 4 Emx1-Ai96 and 10 Emx1-Ai93 mice, from 30–75 min of imaging. (C) Mean of the Emx1-Ai96 and Emx1-Ai93 field sign maps in panel B, with borders and area labels. (D) Map of variance of the visual field sign. Variance was calculated from visual field sign maps from 14 mice, after alignment as described for calculation of the mean field sign map. Whiter areas denote higher variance. Area borders are overlaid in white. (E) The probability of mapping different visual areas with GCaMP6 fluorescence in awake mice. Blue and red bars denote areas with negative and positive field signs, respectively. Results were derived from 14 mice (4 Emx1-Ai96, 10 Emx1-Ai93). Mouse numbers: V1 14/14, LM 14/14, LI 10/10, AL 14/14, LLA 10/13, RL 14/14, RLL 6/14, AM 13/14, PM 14/14, MMA 14/14, MMP 14/14, M 5/9, P 14/14, POR 1/3, where, for each area, the denominator indicates the number of mice in which the area was visible within the cranial window, determined manually. (F) Mean ± SEM fluorescence change for each visual area, derived from the △F/F spectral power. For each map power was normalized to that in V1. S1 region was drawn manually towards the anterior extent of the cranial window. Mouse numbers: LM 14, LI 10, AL 14, LLA 10, RL 14, RLL 5, AM 12, PM 14, MMA 12, MMP 12, M 5, P 14, S1 14; from 14 mice (4 Emx1-Ai96, 10 Emx1-Ai93).
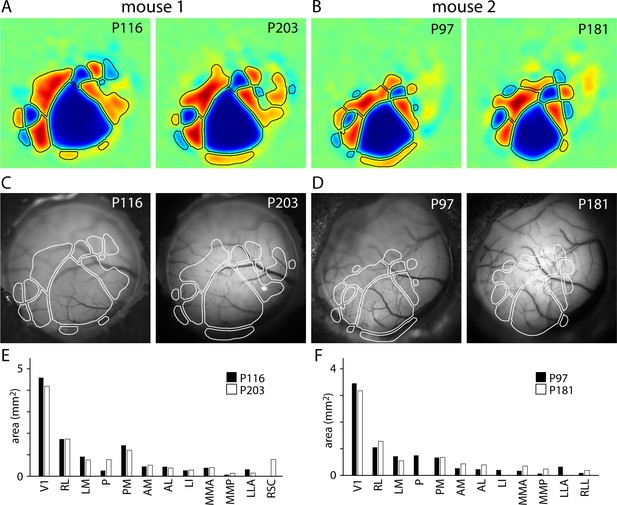
Map stability.
(A and B) Field sign maps generated from two Emx1-Ai96 mice, one at postnatal days 116 and 203 and a second at postnatal days 97 and 181. (C and D) Images of surface vasculature, with border positions overlaid in white. (E and F) Comparison of the area of each patch across imaging sessions.
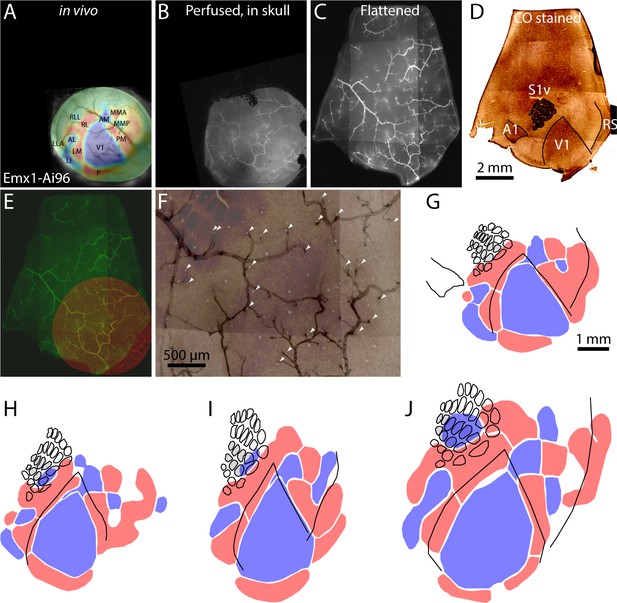
Registration of functional retinotopic maps to chemoarchitectonic borders.
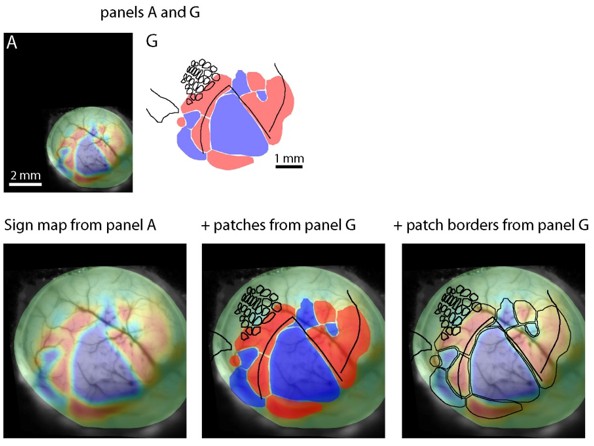
(A–D) Images from key stages in the processing of tissue from an Emx1-Ai96 mouse, each aligned to the cytochrome C oxidase (CO) image. (A) Brightfield image of surface vasculature with overlaid field sign map. (B) Fluorescence image of whole-mount brain, after perfusion, in which a subset of the surface vasculature is labeled with DyLight 649-lectin conjugate. (C) fluorescence image of the flattened cortex. D: brightfield image of a section through layer four after CO staining. (E) Overlaid fluorescence images of surface vasculature in whole-mount (red, panel B) and after flattening (green, panel C). (F) Overlaid images of the surface vasculature and CO staining in posterior barrel cortex and anterior V1. The contrast of the vasculature image is inverted for clarity. Arrowheads indicate small, circular regions that do not stain for CO and likely result from transverse cuts through ascending/descending vessels. Note the alignment of these putative vessels with likely locations of ascending/descending vessels in the fluorescence image of surface vasculature. (G) Field sign map (panel A) aligned to chemoarchitectonic borders from the CO image (panel D). Borders of primary visual cortex, auditory cortex, and of barrels in primary somatosensory cortex) were drawn manually. Barrels in putative columns B and C are shaded grey. (H–J) Alignment of functional retinotopic maps and chemoarchitectonic borders for three additional Emx1-Ai96 mice.
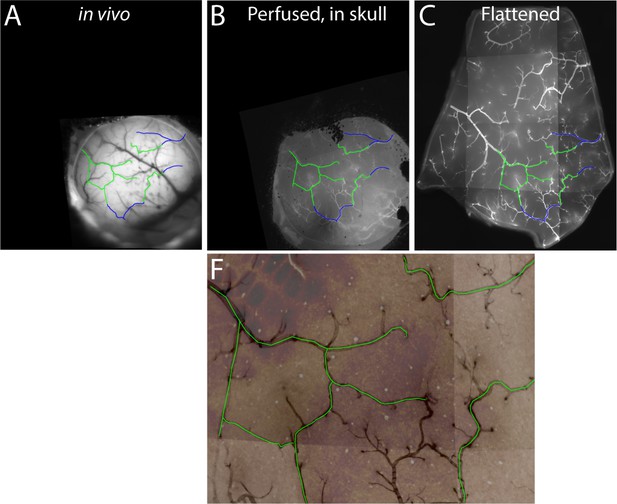
Vessels common to images from live and fixed tissue.
Example of vessel tracing through the series of images used for registration of sign maps to fixed tissue. Major vessels common to multiple images are traced on the images of Figure 4. Green lines mark vessels that are present in all four images. Blue lines mark vessels that extend beyond the borders of the cytochrome oxidase image, but are visible in the other images. Note that not all vessels are labeled after fixation, with the result that some large vessels are visible only in the in vivo image.
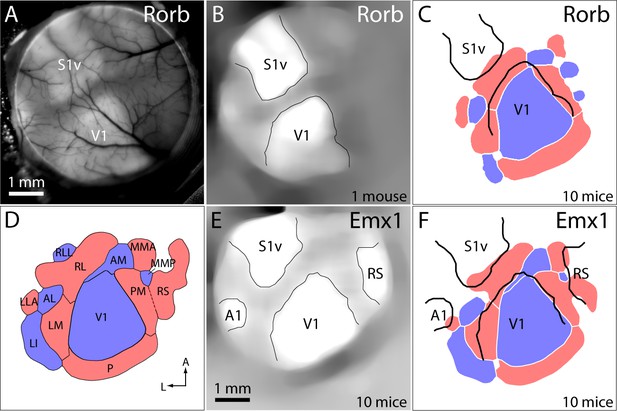
Registration of functional retinotopic maps to cytoarchitectonic borders.
(A) GCaMP6 fluorescence image from a Rorb-Ai93 mouse. Primary sensory areas are marked: S1v barrel cortex, V1 primary visual cortex. (B) Image in panel A after filtering and semi-automated identification of major cytoarchitectonic borders. (C) Mean retinotopic map/cytoarchitectonic border registration for 10 Rorb-Ai93 mice. Retinotopic maps and cytoarchitectonic borders were pooled across mice as described in the Materials and methods. Cytoarchitectonic borders are shown in black. (D) Mean sign map with patch notation, from Figure 3C. (E) Mean fluorescence image from 10 Emx1-Ai93 mice, after filtering, alignment and semi-automated identification of the borders of primary sensory areas and retrosplenial cortex (RS). (F) Mean map/border registration for 10 Emx1-Ai93 mice.
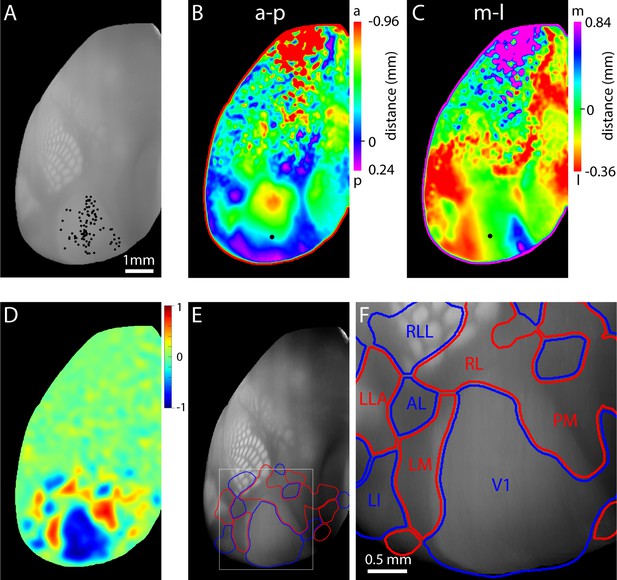
Retinotopy of projections from V1.
(A) Locations of injections into V1 in 99 mice, selected from the Allen Brain Connectivity Atlas. Each point indicates an injection. Injection locations were registered to a 3D model of the mouse brain generated from 1675 brains and injections are illustrated on a top projection of the mean autofluorescence from the 3D model. Variations in autofluorescence clearly delineate major architectonic boundaries, including barrels in primary somatosensory cortex, primary auditory cortex, primary visual cortex and retrosplenial cortex. (B and C) Projection-based maps of connectivity with ipsilateral V1. Colors indicate the distance from the geometric center of V1 (black circle) from which the strongest projection arises, along the anterior (a) – posterior (p) and medial (m) – lateral (l) axes of V1. These maps are projection-based homologues of the azimuth and altitude maps generated from functional mapping (Figure 4E,F). (D) Projection sign map generated from the maps in panels B and C. (E) Automated borders generated from the projection sign maps of panel D, overlaid onto the autofluorescence top projection from panel A. (F) Subregion corresponding to the box in panel E. A lower threshold was used to generate the visual area borders from the sign map, eliminating the gap between areas V1, AL, RL and LM.
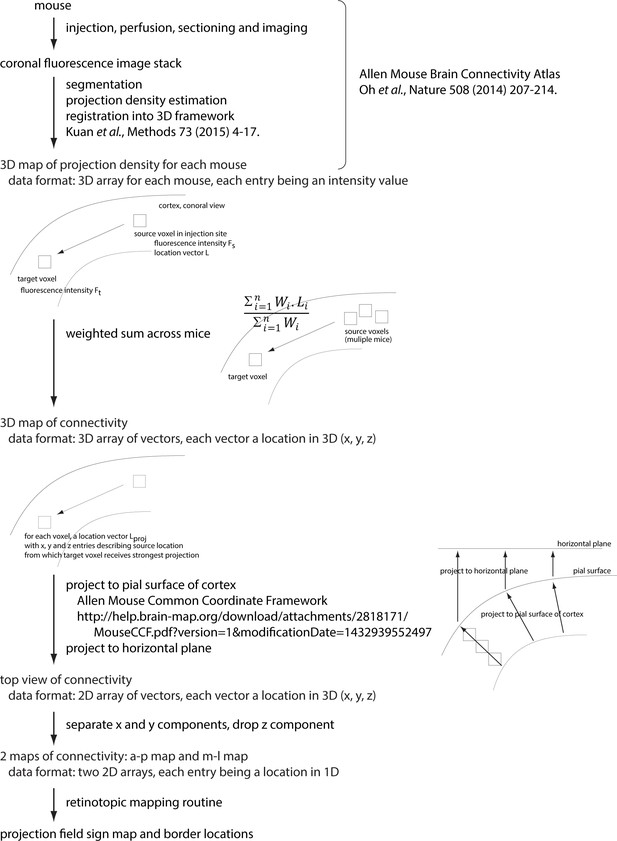
Generation of projection-based retinotopy maps.
Schematic indicating the steps in generating the projection-based retinotopy map. References are provided where methods are published (including the collection of the source data set). Information on data formats are provided and key steps are illustrated graphically.
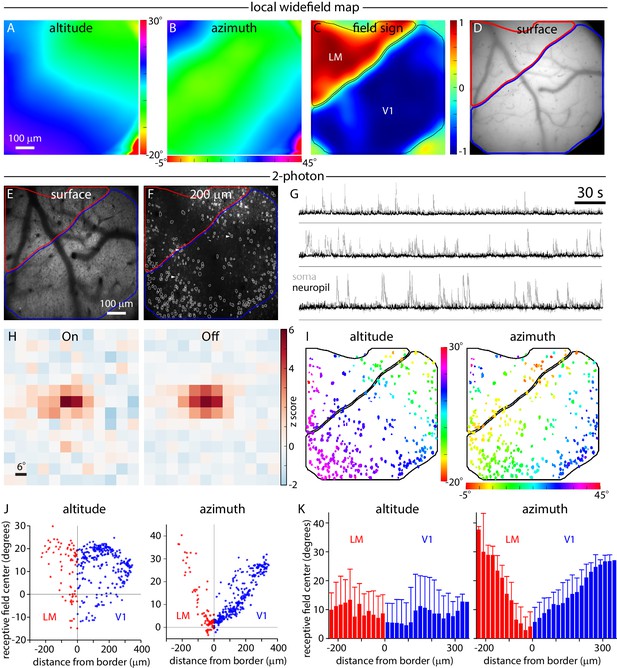
Widefield borders match single-cell retinotopy at the V1-LM border.
(A, B and C) Local altitude and azimuth and field sign maps generated under the 2-photon microscope by widefield imaging through the x16 objective. Images were acquired with the objective focused 200 µm below the pial surface of cortex. (D and E) Images of surface vasculature acquired with the objective focused on the pial surface. Borders of V1 and LM derived from the field sign map are marked in blue and red, respectively. (F) 2-photon fluorescence image acquired with the microscope objective focused 200 µm below the pial surface of cortex. Borders of V1 and LM derived from the field sign map are marked in blue and red, respectively. White outlines indicate 366 somatic ROIs. (G) Example fluorescence traces extracted from the three somatic regions (before neuropil subtraction) marked with arrowheads in panel F (grey traces) and the corresponding neuropil regions (black traces). Fluorescence scale is in arbitrary units. A black horizontal line indicates zero fluorescence for each pair of traces. (H) On and Off receptive fields from an example cell, extending from −6 to 60 degrees in azimuth and −24 to 54 degrees in altitude. (I) Altitude and azimuth maps (from summed receptive fields) for the experiment illustrated in panel F. The color of each soma represents its receptive field center location. Of the 366 somata identified in this field of view, 336 displayed significant receptive fields (maximum z-score ≥ 2) and are illustrated in panel I. Black lines mark the borders of V1 and LM. (J) Plots illustrating the distribution of single-cell altitude and azimuth as a function of minimum distance to the V1-LM border. Each point represents a single soma from the field of view illustrated in panel F. (K) Single-cell altitude and azimuth as a function of distance to the V1-LM border. Results from three experiments were pooled, yielding 964 somata with receptive fields. Each bar represents the mean and standard deviation of cells binned by distance from the border, in 20 µm bins.
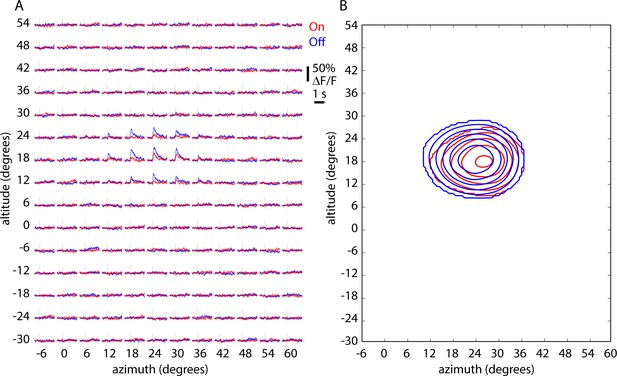
On and Off receptive fields for an example cell.
(A) ΔF/F traces displaying On (red) and Off (blue) responses for each pixel in the nasal receptive field. Each pixel of the stimulus subtended six degrees in altitude and six in azimuth. Each trace represents the mean (line) and standard error (shaded area) of 60 trials. Horizontal lines indicate zero ΔF/F and vertical line the onset of the stimulus. (B) Receptive field maps derived from the example in panel A. Lines represent 40%, 50%, 60%, 70%, 80% and 90% of the maximum z-score.
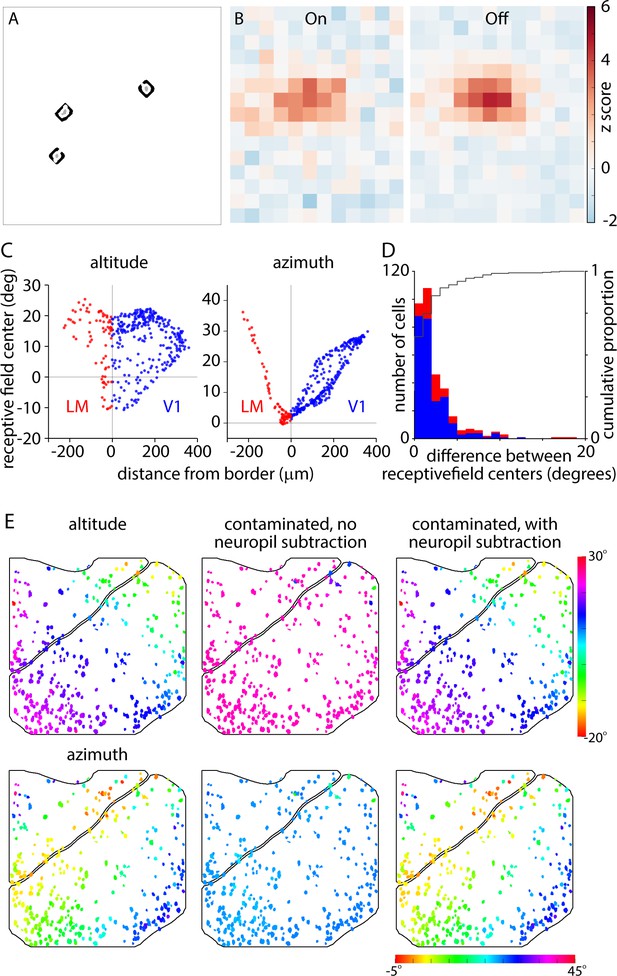
Neuropil tuning and subtraction.
(A) Somatic ROIs (grey) and corresponding neuropil ROIs (black) for the three cells highlighted in Figure 7F and G. (B) Neuropil On and Off receptive fields for the cell illustrated in Figure 7H. (C) Plots illustrating the distribution of neuropil altitude and azimuth as a function of distance to the V1-LM border. Each point represents a single neuropil ROI from the field of view illustrated in Figure 7F and can be directly compared to the equivalent plot for somatic tuning in Figure 7J. (D) Distribution of the difference in somatic and neuropil tuning for V1 (blue bars) and LM (red bars). Black line is the cumulative distribution (right axis). These results indicate that most somata display similar tuning to the local neuropil, with a difference of <5 degrees for >90% of neurons. (E) Results of a numerical simulation to illustrate the effectiveness of neuropil subtraction. We examined the effects of adding an additional contaminating signal to each somatic trace. The additional signal was a neuropil trace tuned to 24.15 and 24.28 degrees in altitude and azimuth. To increase the effect of this additional contamination, we first multiplied the amplitude of the contaminating trace by 10. From the modified traces, we recalculated the somatic altitude and azimuth maps with and without neuropil subtraction. As expected, in the absence of neuropil subtraction, the added signal shifted all somatic tuning towards the coordinates of the added signal, whereas with neuropil subtraction enabled, the somatic altitude and azimuth maps were almost identical to the unmodified maps). Left column: somatic altitude and azimuth maps. Center column: somatic altitude and azimuth maps after addition of enlarged neuropil signal, with no neuropil subtraction. Right column: somatic altitude and azimuth maps after addition of enlarged neuropil trace, with neuropil subtraction. The results of this test indicate that our neuropil subtraction routine corrects for neuropil contamination several times larger than that observed in our results.
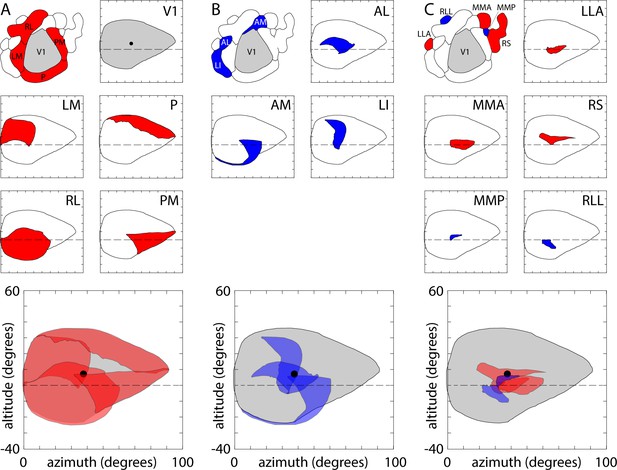
Visual coverage across areas.
(A) Coverage map of visual space by V1 and surrounding positive field sign patches. Top left, overview map of V1 (grey) and the four surrounding positive field sign patches (red; LM, RL, PM, P). Top right, V1 coverage map. Locations represented in V1 are indicated in grey. Circle indicates the center of coverage of V1 at 7.4° altitude, 37.8° azimuth. Dashed line indicates the horizontal meridian. Center panels, coverage maps of positive field sign patches that border V1 (LM, RL, PM, P). Each coverage map illustrates the one positive field sign patch (in red), overlaid on the coverage of V1 (black outline). Note that coverage of PM excludes retrosplenial cortex. Lowest panel, overlapping coverage of 5 areas (V1 in grey; LM, RL, PM, P in red). (B) Coverage of negative field sign patches LI, AL and AM. Coverage of each patch (blue) is overlaid on the coverage of V1 (black outline). (C) Coverage of remaining positive and negative field sign patches.
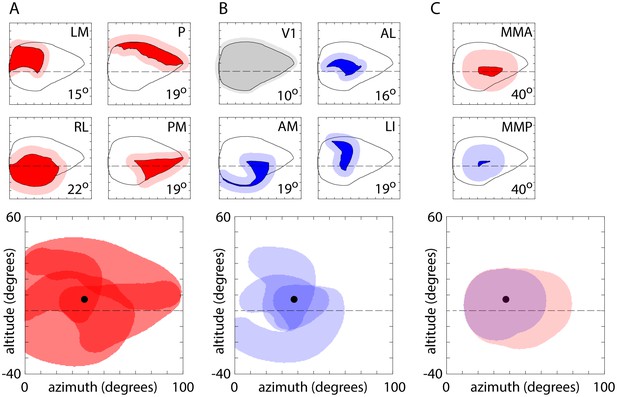
Expanded coverage.
Plots illustrating the expansion of visual field coverage, based on published receptive field sizes. (A) Upper panels: visual field coverage for areas LM, P, RL and PM (from Figure 8A) and expanded coverage (red shading). For each area, coverage expanded by the mean radius of single-cell receptive fields, taken from Wang and Burkhalter (2007). The receptive field radius employed is in the lower right corner of each plot. Lower panel: overlapping coverage of LM, P, RL and PM. Circle indicates the center of coverage of V1 at 7.4° altitude, 37.8° azimuth. (B) Coverage and expanded coverage of V1 (grey) and of AL, AM and LI. (C) Coverage and expanded coverage of MMA and MMP. For MMA and MMP, the receptive field radius employed (40°) was that of MM in Wang and Burkhalter (2007). No expanded coverage estimates are provided for LLA, RS and RLL as their receptive field sizes were not provided by Wang and Burkhalter (2007).
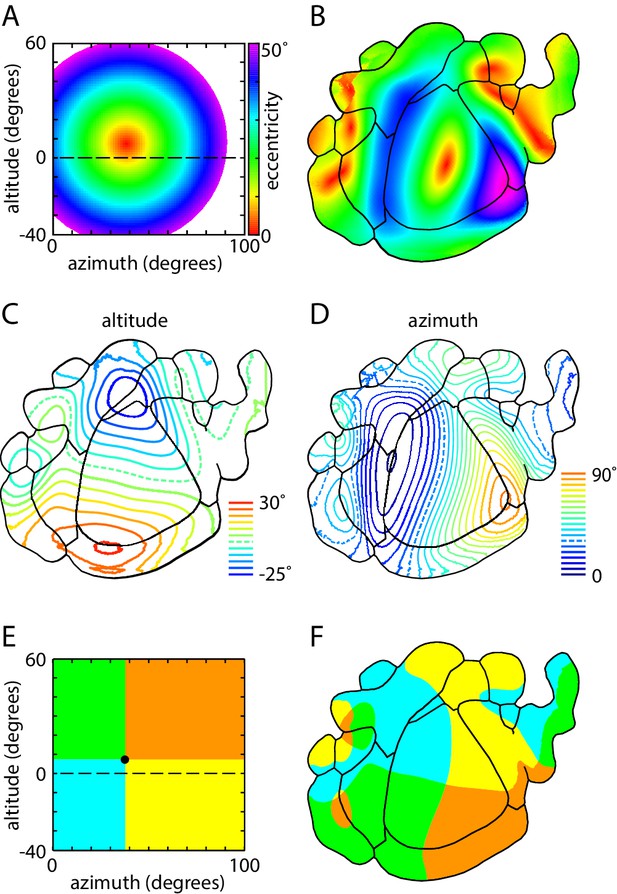
Magnification and representation of visual space across visual cortex.
(A) Color map of visual space in eccentricity coordinates. Color (bar) indicates the distance (in degrees) from the center of coverage of V1 (7.4 ± 2.1 degrees altitude, 37.8 ± 1.4 degrees azimuth). (B) Eccentricity map of mouse visual areas, using the color scheme indicated in panel A. (C,D) Altitude and azimuth contour plots of mouse visual cortex, overlaid on mean field sign borders. Dashed line represents horizontal meridian. Contours are at 5° intervals from −25 to 30° in altitude and 0 to 90° in azimuth. (E) Colored sector map of visual space, with the division between upper and lower visual fields at 7.4° altitude, 37.8° azimuth, which corresponds to the center of coverage of V1. (F) Colored sector map of mouse visual cortex, with colors corresponding to those in panel B and denoting representation of the four quadrants of the visual field.

Videos
Example fluorescence movies from retinotopic mapping experiment.
Example of fluorescence changes during retinotopic mapping. Left: response to checkerboard travelling from the lower to the upper visual field. Right response to checkerboard travelling from the nasal to temporal visual field. Each movie is the mean change in fluorescence (△F) of 40 trials.
Lower-to-upper visual field visual stimulus.
Lower to upper visual field moving checkerboard stimulus used for retinotopic mapping.
Nasal-to-temporal visual stimulus.
Nasal-to-temporal moving checkerboard stimulus used for retinotopic mapping.
Additional files
-
Supplementary file 1
Retinotopic mapping example analysis.
Example of derivation of a field sign map from altitude and azimuth maps, in HTML format. The same example is available as a Jupyter notebook at https://github.com/zhuangjun1981/retinotopic_mapping.
- https://doi.org/10.7554/eLife.18372.027





