Fetal and neonatal hematopoietic progenitors are functionally and transcriptionally resistant to Flt3-ITD mutations
Figures
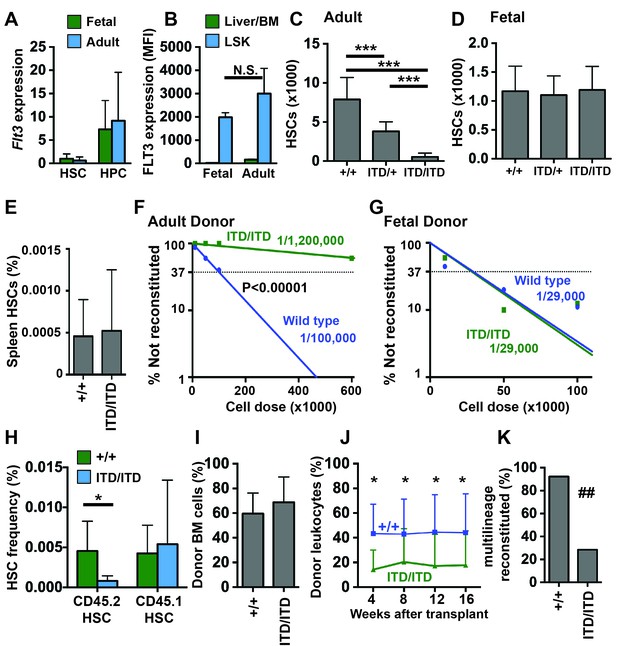
Flt3ITD causes HSC depletion in adult but not fetal mice.
(A) Flt3 transcript expression in fetal and adult HSCs and HPCs relative to fetal HSCs; n = 4–9. (B) FLT3 expression in fetal and adult HSC/HPCs (Lineage-Sca1+c-kit+) and unfractionated fetal liver or bone marrow cells, as determined by flow cytometry (N = 3). (C) HSC numbers in two tibias and femurs from adult wild type and Flt3ITD mice; n = 12–16. (D) HSC numbers in fetal livers from E14.5 wild type and Flt3ITD mice; n = 9–20. (E) Spleen HSC frequency in adult wild type and Flt3ITD/ITD mice; n = 4–5. (F,G) Limiting dilution analyses using adult bone marrow (F) or E14.5 fetal liver cells (G); n = 9–10 recipients per cell dose. Wild type and Flt3ITD/ITD HSC frequencies were calculated by extreme limiting dilution analysis. (H,I) Frequencies of donor (CD45.2) and competitor (CD45.1) HSCs (H) and donor bone marrow cells (I) in primary recipients of 100,000 fetal liver cells; n = 15 per genotype. (J) Frequencies of CD45.2+ peripheral blood cells in secondary recipients of donor cells that originated from wild type or Flt3ITD/ITD fetal livers; n = 12–14. (K) Percentage of secondary recipient mice with multilineage donor reconstitution. In all panels, error bars indicate standard deviations and n reflects biological replicates. *p<0.05, ***p<0.001 by two-tailed Student’s t-test. ## p<0.01 by Fisher exact probability test.
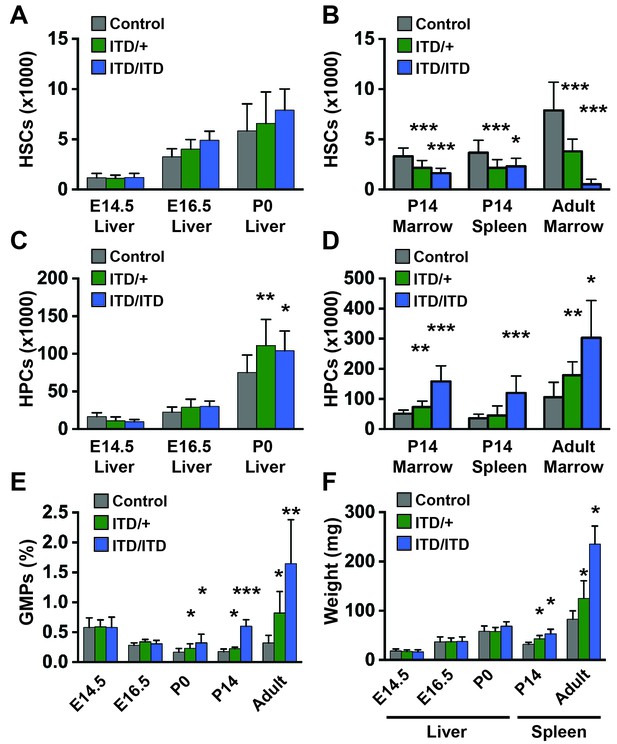
Flt3ITD causes HSC depletion, HPC expansion and GMP expansion at, or shortly after, birth.
(A) Absolute HSC numbers in fetal or P0 livers for the indicated genotypes. (B) Absolute HSC numbers in P14 and adult bone marrow (two hind limbs) or P14 spleen. (C,D) Fetal and adult HPC numbers (two hind limbs). (E) GMP frequencies in fetal liver or adult bone marrow. (F) Liver or spleen weights. In all panels, error bars indicate standard deviations; n = 6–20 biological replicates for each age and genotype. *p<0.05; **p<0.01; ***p<0.001 by two-tailed Student’s t-test relative to the wild type control at the same time point.
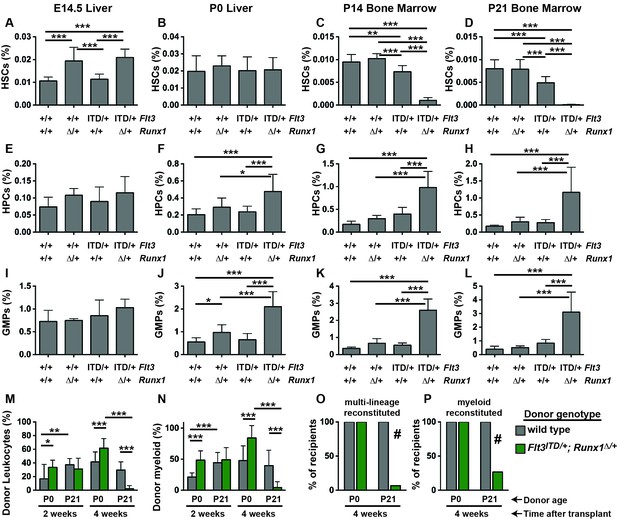
Flt3ITD and Runx1 mutations cooperate to deplete HSCs and expand committed progenitor populations after birth.
(A–D) HSC frequencies in E14.5 fetal liver, P0 liver, P14 bone marrow and P21 bone marrow for the indicated genotypes. (E–H) HPC frequencies in E14.5 fetal liver, P0 liver, P14 bone marrow and P21 bone marrow for the indicated genotypes. (I–L) GMP frequencies in E14.5 fetal liver, P0 liver, P14 bone marrow and P21 bone marrow for the indicated genotypes. (M,N) Percentages of CD45.2+ donor leukocytes (M) or CD11b+Gr1+ myeloid cells (N) in the peripheral blood of recipients of P0 liver or P21 bone marrow cells from control or Flt3ITD/+; Runx1Δ/+ mice. Measurements are shown at 2 and 4 weeks after transplantation. (O,P) Percentage of recipients with multi-lineage (O) or myeloid (P) donor reconstitution at four weeks after transplantation. In all panels, error bars indicate standard deviations. For A-L, n = 8–18 biological replicates per genotype and age. For (M–P), n = 14–15 recipients from three independent donors. Statistical significance was determined with a one-way ANOVA followed by Holm-Sidak’s post-hoc test for multiple comparisons (*p<0.05; **p<0.01; ***p<0.001), or # p<0.0001 by the Fisher exact probability test.
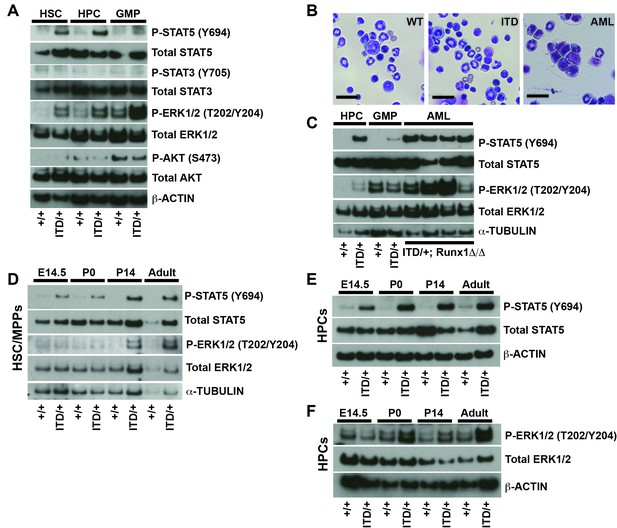
FLT3ITD activates STAT5 in both fetal and adult progenitors, but it activates the MAPK pathway after birth.
(A) Western blot showing phosphorylation of STAT5, ERK1/2, STAT3 and AKT in adult wild type and Flt3ITD/+ HSC/MPPs, HPCs and GMPs. (B) Flt3ITD/+; Runx1Δ/Δ progenitors give rise to AML in adult mice (right panel) that is not observed in wild type or Flt3ITD/+ bone marrow. Scale bars indicate 100 microns. (C) STAT5 and MAPK are hyper-phosphorylated in Flt3ITD/+; Runx1Δ/Δ AML that develops in adult mice. (D) STAT5 and ERK1/2 phosphorylation in wild type and Flt3ITD/+ HSC/MPPs at E14.5, P0, P14 and adulthood. (E) STAT5 phosphorylation in wild type and Flt3ITD/+ HPCs at E14.5, P0, P14 and adulthood. (F) ERK1/2 phosphorylation in wild type and Flt3ITD/+ HPCs at E14.5, P0, P14 and adulthood. Each blot is representative of two (panels A and C) or at least three (panels D–F) independent experiments.
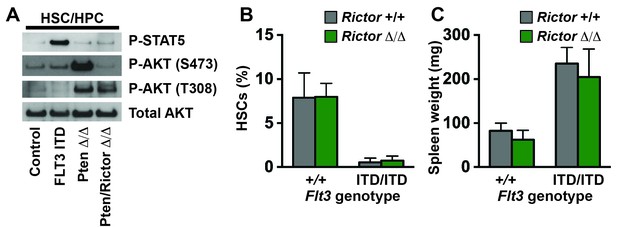
The PI3K/mTORC2 pathway does not mediate HSC depletion or MPN in Flt3ITD mutant mice.
(A) Western blot showing STAT5 and AKT phosphorylation in wild type and Flt3ITD/ITD HPCs. Pten-deficient HPCs were included as a positive control for AKT phosphorylation. Rictor deletion prevented AKT hyper-phosphorylation by mTORC2. (B,C) HSC numbers in two tibias and femurs (B) and spleen weights (C) in Flt3ITD/ITD; Rictorf/f; Vav1-Cre compound mutant mice (and the indicated controls) after conditional Rictor deletion; n = 3–12 per genotype. Error bars indicate standard deviations.
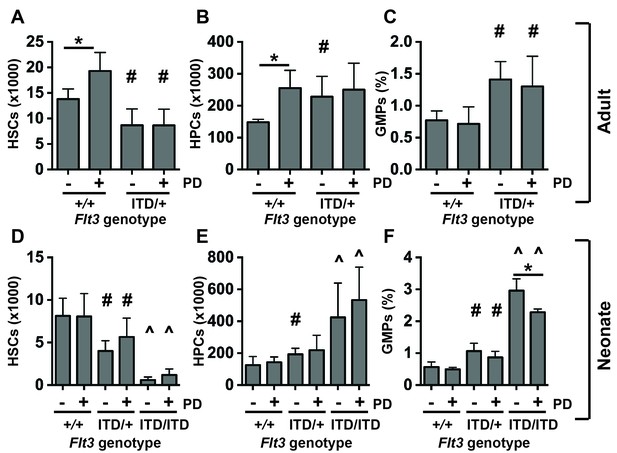
MAPK pathway inhibition does not prevent HSC depletion or committed progenitor expansion in Flt3ITD/+ mice.
(A–D) HSC numbers (A), HPC numbers (B) and GMP frequencies (C) in wild type and Flt3ITD/+ mice that were treated with vehicle or PD0325901 for 10 days beginning at six weeks after birth; n = 4–5 biological replicates per genotype and treatment. (D–F) HSC numbers (D), HPC numbers (E) and GMP frequencies (F) in P19 wild type, Flt3ITD/+ and Flt3ITD/ITD mice whose mothers were given PD0325901 beginning at P1; n = 4–15 biological replicates for each genotype and treatment. In all panels, error bars indicate standard deviation. Statistical comparisons were made with a two-tailed Student’s t-test. *p<0.05 relative to vehicle treated cells with equivalent genotypes; # p<0.05 relative to similarly treated wild type controls; ^ p<0.05 relative to similarly treated wild type and Flt3ITD/+ groups.
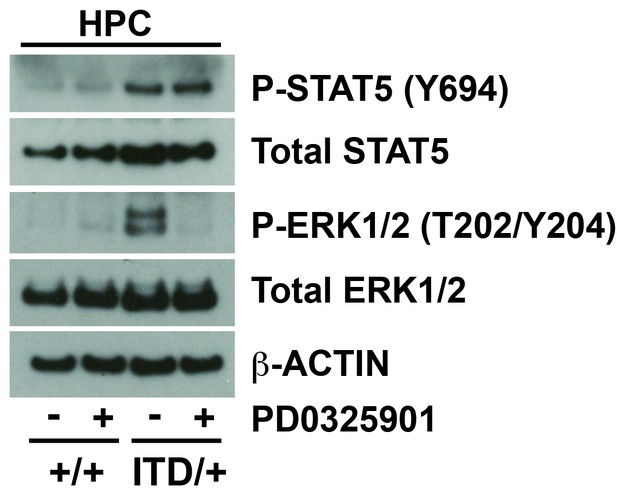
Inhibition of the MAPK pathway fails to rescue FLT3ITD-mediated HSC depletion and myeloid progenitor expansion, but Stat5a/b deletion enhances these phenotypes.
A Western blot was performed with 25,000 HPCs from wild type and Flt3ITD/+, vehicle and PD0325901 treated mice. PD0325901 inhibited ERK1/2 phosphorylation without affecting STAT5 phosphorylation.
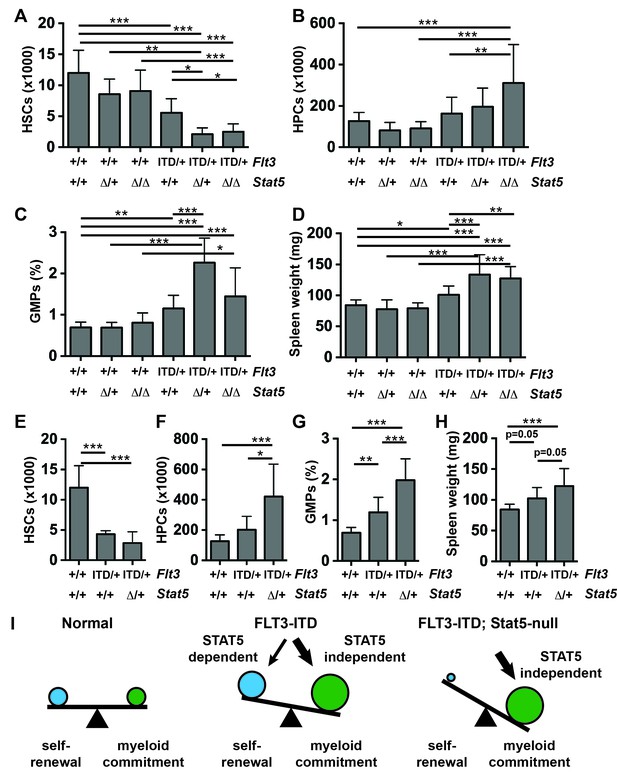
Stat5a/b deletion exacerbates rather than rescues HSC depletion, HPC expansion, GMP expansion and MPN in Flt3ITD/+ mice.
(A–D) HSC numbers (A), HPC numbers (B), GMP frequencies (C) and spleen weights (D) in Flt3ITD/+; Stat5a/bf/f; Mx1-Cre compound mutant mice and littermate controls; n = 6–20 biological replicates per genotype. Stat5a/b was conditionally deleted six weeks after birth, and this caused a complete loss of protein expression (figure supplement 1). (E–H) HSC numbers (E), HPC numbers (F), GMP frequencies (G) and spleen weights (H) in Flt3ITD/+; Stat5a/bf/+; Vav1-Cre compound mutant mice and littermate controls; n = 8–20 biological replicates per genotype. (I) The data suggest that STAT5-dependent pathways promote HSC self-renewal downstream of FLT3ITD, but these effects are outweighed by STAT5-independent myeloid commitment pathways. In all panels, error bars indicate standard deviation. Statistical significance was determined with a one-way ANOVA followed by Holm-Sidak’s post-hoc test for multiple comparisons. *p<0.05; **p<0.01; ***p<0.001.
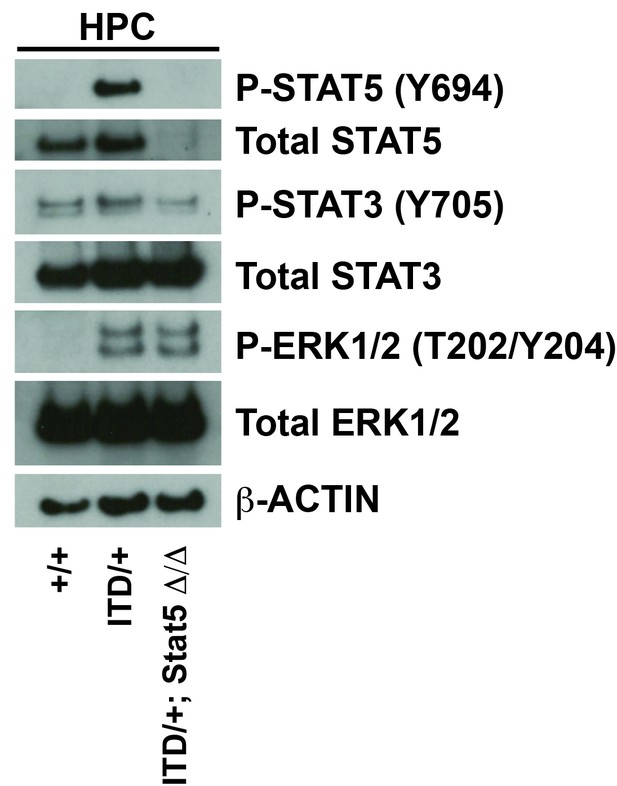
Stat5a/b deletion causes a complete loss of phosphorylated and total STAT5 protein.
A Western blot was performed with 25,000 HPCs from wild type, Flt3ITD/+ and Flt3ITD/+; Stat5Δ/Δ adult mice (representative of two independent experiments). ERK1/2 phosphorylation was not affected by STAT5 deletion. STAT3 phosphorylation was not affected, and STAT1 and AKT phosphorylation were not detectable (not shown).
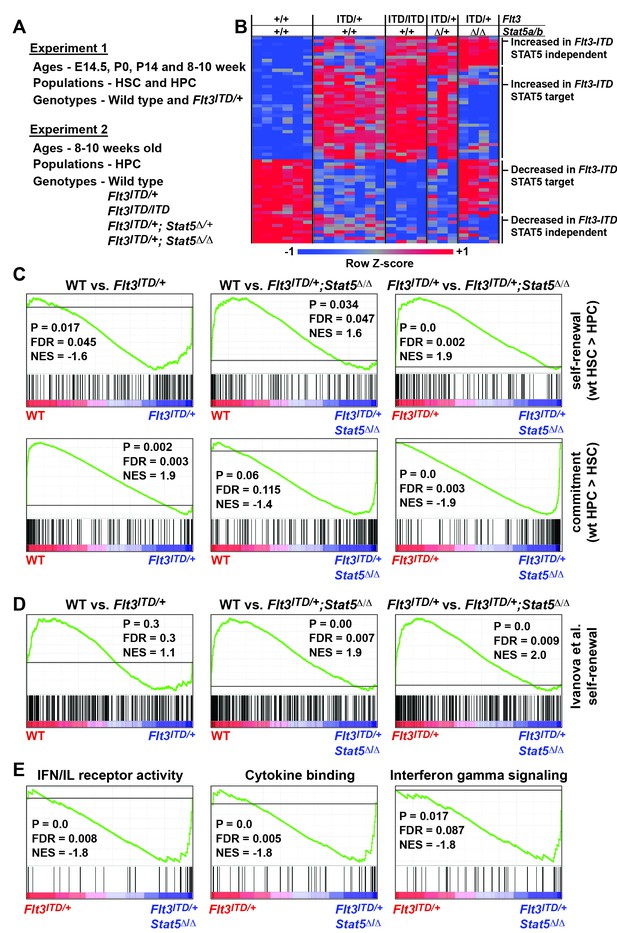
FLT3ITD activates STAT5-dependent self-renewal programs and STAT5-independent commitment programs.
(A) Overview of experimental design. (B) Heatmap representing genes that were differentially expressed in Flt3ITD mutant HPCs relative to wild type HPCs in both experiments 1 and 2. Each column represents an independent sample. The gene names and dendrogram are shown in Figure 7—figure supplement 1 attached to this figure. (C) Self-renewal and commitment-related gene sets were generated by identifying genes that were more highly expressed (>5 fold, adj. p<0.05) in HSCs relative to HPCs (self-renewal), or HPCs relative to HSCs (commitment). GSEA plots show ectopic activation of self-renewal-related genes in HPCs that express FLT3ITD, but these effects are reversed in Stat5a/b-deficient HPCs. (D) An independently curated self-renewal gene set (Ivanova et al., 2002) was similarly enriched in wild type and Flt3ITD/+ HPCs relative to Flt3ITD/+; Stat5Δ/Δ HPCs. (E) GSEA revealed enrichment of gene sets associated with increased inflammatory cytokine signaling.
-
Figure 7—source data 1
Significantly differentially expressed genes in Flt3ITD/+ HSCs and HPCs.
Fold changes are shown for 270 probes that represent 254 differentially expressed genes in Flt3ITD/+ HPCs relative to controls. HSC fold change data are also shown. P-values and adjusted p-values reflect significance across the entire time course.
- https://doi.org/10.7554/eLife.18882.013
-
Figure 7—source data 2
Self-renewal-related and commitment-related gene sets.
These gene sets were determined by identifying genes that were more highly expressed in HSCs relative to HPCs (self-renewal) or HPCs relative to HSCs (commitment). In each case, a fold change threshold of five and an adjusted p-value threshold of <0.05 were used to define the lists.
- https://doi.org/10.7554/eLife.18882.014
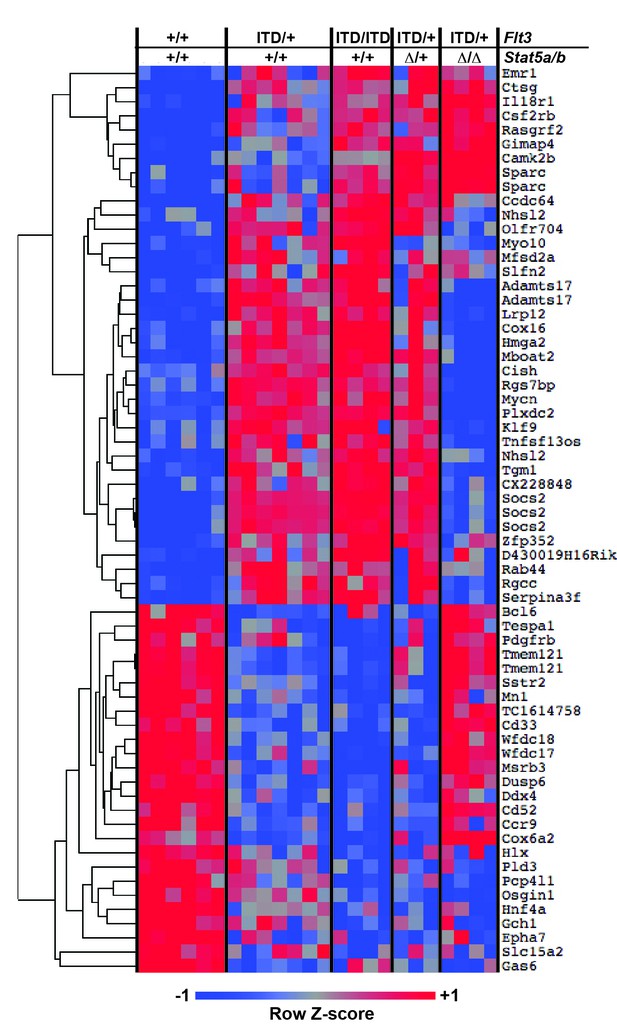
FLT3ITD induces STAT5-dependent and STAT5-independent changes in gene expression.
An expanded version of Figure 7B with the dendrogram and gene names attached to the heatmap.
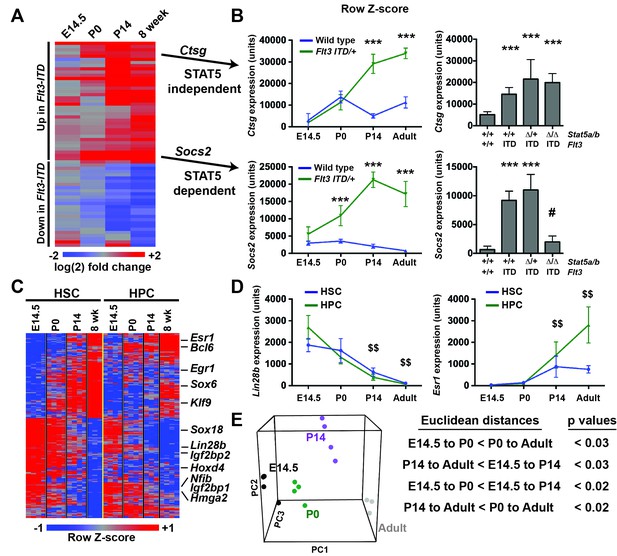
Flt3ITD-mediated changes in gene expression correlate with the normal transition from fetal to adult transcriptional programs.
(A) Heatmap showing expression of FLT3ITD target genes at E14.5, P0, P14 and adult stages. Each column shows average fold change in Flt3ITD/+ HPCs relative wild type HPCs at the indicated time point; n = 3–4 independent arrays per genotype. The gene names and dendrogram are shown in figure supplement one attached to this figure. (B) Representative examples of expression of STAT5-independent (Ctsg) and STAT5-dependent (Socs2) FLT3ITD targets. Error bars reflect standard deviation. ***adj. p<0.05 relative to wild type at the same time point, # adj. p<0.05 relative to Flt3ITD/+ at the same time point. (C) Heterochronic genes began transitioning from fetal to adult expression patterns between P0 and P14, concordant with sensitivity to FLT3ITD. Genes that encode transcription factors and RNA binding proteins are noted to the right of the heatmap. A complete gene list is provided in Figure 8—source data 1 attached to this figure. (D) Representative examples of heterochronic genes that show decreased (Lin28b) or increased (Esr1; Estrogen Receptor α) expression in adult relative to fetal HSCs and HPCs. Error bars reflect standard deviation. $$ adj. p<0.05 relative to E14.5 for both HSCs and HPCs. (E) Principal component analysis and Euclidean distance measurements show that gene expression in P0 HSCs more closely resembles fetal HSCs than adult HSCs, and gene expression in P14 HSCs more closely resembles that of adult HSCs. Similar calculations for HPCs are shown in figure supplement 2.
-
Figure 8—source data 1
Heterochronic gene expression in wild type HSCs and HPCs.
Source data (log2) are shown for the 250 most differentially expressed probes in developing HSCs (228 unique genes).
- https://doi.org/10.7554/eLife.18882.017
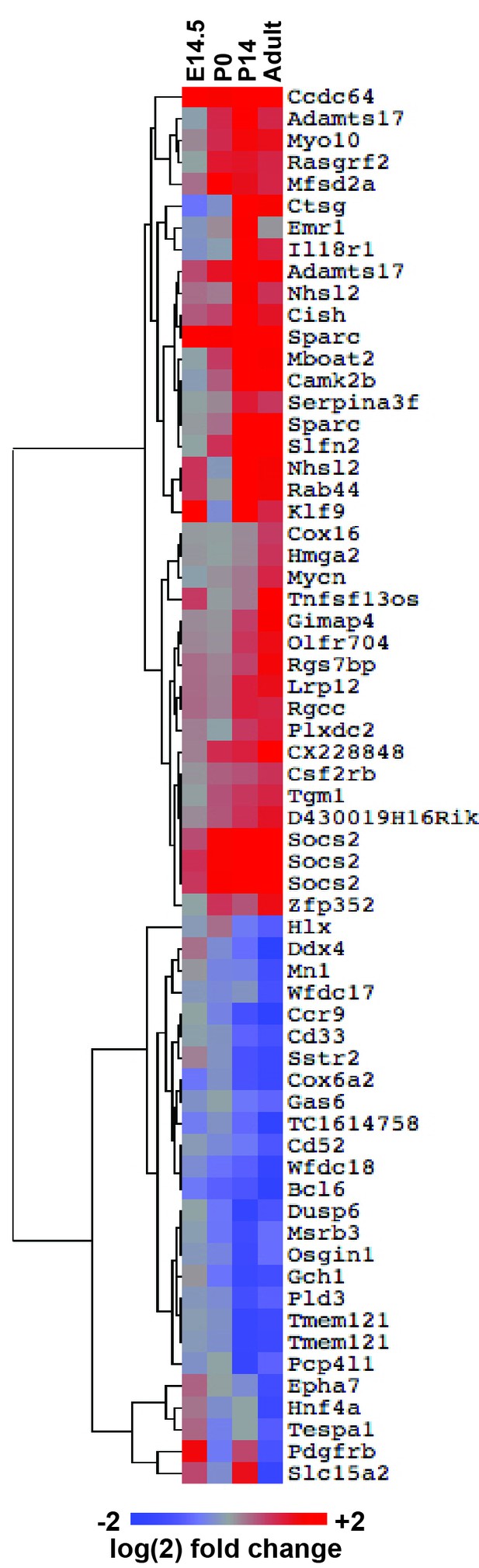
FLT3ITD-mediated changes in gene expression correlate temporally with a transition from fetal to adult transcriptional states.
An expanded version of Figure 8A with the dendrogram and gene names attached to the heatmap.
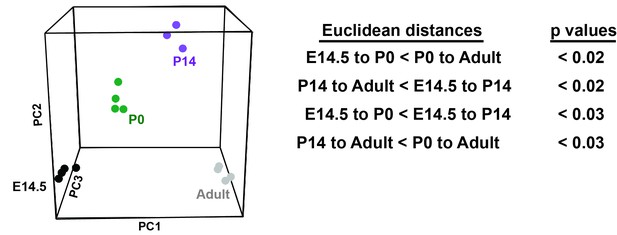
HPCs express heterochronic genes and begin to transition from fetal to adult transcriptional programs by P14.
Principal component analysis shows temporal changes in HPC gene expression before and after birth. Euclidean distance measurements and permutation testing show that P0 HPCs more closely resemble fetal HPCs than adult HPCs. P14 HPCs more closely resemble adult HPCs.
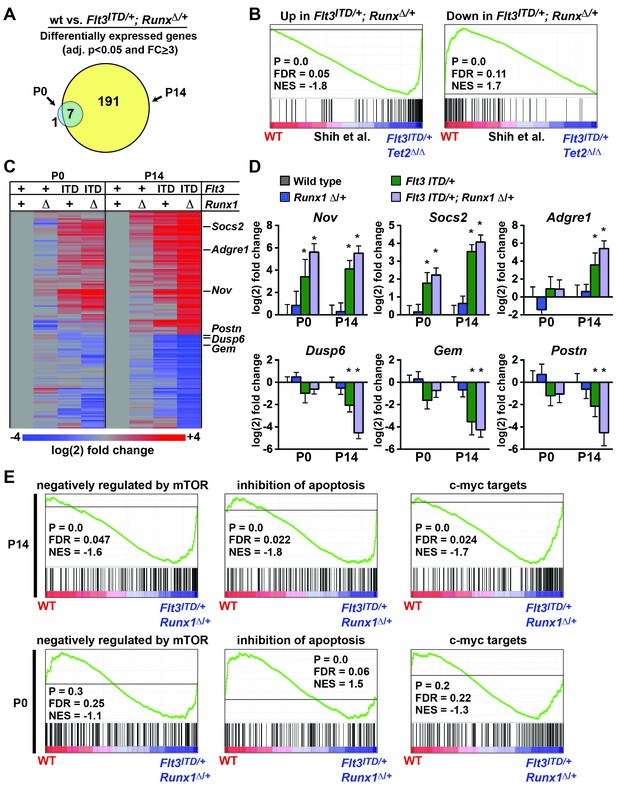
Flt3ITD and Runx1 mutations cooperatively induce changes in gene expression at P14, yet they have a much smaller effect at P0.
(A) Venn diagram showing overlap between genes that were significantly differentially expressed (adj. p<0.05, fold change ≥ 3) in Flt3ITD/+; Runx1Δ/+ HPCs relative to wild type at P14 or P0; n = 4–5 arrays per genotype and age. (B) GSEA shows that differentially expressed genes in Flt3ITD/+; Runx1Δ/+ HPCs overlap significantly with genes that are differentially expressed in Flt3ITD/+; Tet2Δ/Δ HPCs (Shih et al., 2015). (C) Heatmap showing expression of genes that were differentially expressed in Flt3ITD/+; Runx1Δ/+ HPCs relative to wild type HPCs. Each column indicates the average fold change relative to the wild type samples from the same time point. The gene list is shown in Figure 9—source data1 attached to this figure. (D) Representative examples of genes that are among the most differentially expressed in Flt3ITD/+; Runx1Δ/+ HPCs relative to wild type HPCs at P14. Most show much smaller changes in expression at P0. Error bars reflect standard deviations, * adj. p<0.05. (E) GSEA identified several gene sets that were enriched in Flt3ITD/+; Runx1Δ/+ HPCs relative to wild type HPCs at P14. Three of the most significantly enriched gene sets are shown for P14 and P0.
-
Figure 9—source data 1
Flt3ITD and Runx1 mutations cooperatively induce changes in gene expression in post-natal HPCs.
Genes that are differentially expressed (Adjusted p<0.05, fold change ≥ 3) in Flt3ITD/+; Runx1Δ/+ HPCs at P0 and P14 are shown in the indicated tables.
- https://doi.org/10.7554/eLife.18882.021





