A host basal transcription factor is a key component for infection of rice by TALE-carrying bacteria
Figures
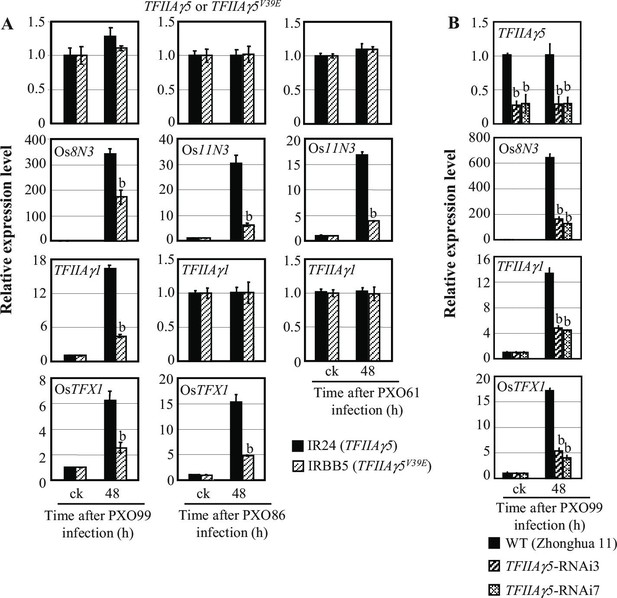
Effects of TFIIAγ5 on the expression of disease susceptibility genes Os8N3, Os11N3, TFIIAγ1, or OsTFX1, after Xoo infection.
Plants were inoculated with Xoo strain PXO99 (harbouring TALEs PthXo1, PthXo7, and PthXo6), PXO86 (harbouring TALE PthXo3) or PXO61 (harbouring TALE AvrXa7) at the booting (panicle development) stage. It is known that PthXo1, PthXo7, and PthXo6 induce Os8N3, TFIIAγ1, and OsTFX1, respectively, and PthXo3 and AvrXa7 all induce Os11N3. Each bar represents mean (three replicates) ± standard deviation. (A) Mutation of TFIIAγ5 (rice line IRBB5). b, significant difference between IR24 and IRBB5 at p<0.01. (B) TFIIAγ5-RNAi lines. b, significant difference between wild-type (WT) and transgenic plants at p<0.01.

Effects of TFIIAγ5 on rice resistance to Xoo strains known to carry TALEs.
Rice plants at the booting (panicle development) stage were inoculated with Xoo. (A) The near-isogenic lines IR24 and IRBB5 showed different responses to the infection of Xoo. IRBB5 in IR24 background carries a natural mutated TFIIAγ5, TFIIAγ5V39E. Each bar represents mean (total 17 to 29 leaves from 5 plants) ± standard deviation. b, significant difference between IR24 and IRBB5 at p<0.01. (B) The enhanced resistance of TFIIAγ5-RNAi plants to strain PXO99 was associated with reduced transcription of TFIIAγ5 but not TFIIAγ1. WT, wild-type Zhonghua 11. Each bar represents mean (3 replicates for gene expression and total 5 to 10 leaves from one plant for lesion length) ± standard deviation. b, significant difference between wild-type (WT) and transgenic plants at p<0.01. (C) The enhanced resistance of TFIIAγ5-RNAi plants co-segregated with reduced TFIIAγ5 transcription in T1 families. Each bar represents mean (3 replicates for gene expression and 5 to 10 leaves from one plant for lesion length) ± standard deviation. b, significant difference between WT and transgenic plants at p<0.01. (D) TFIIAγ5-RNAi plants showed enhanced resistance to all the Xoo strains. Each bar represents mean (total 35 to 40 leaves from 5 plants) ± standard deviation. b, significant difference between WT and transgenic plants at p<0.01.
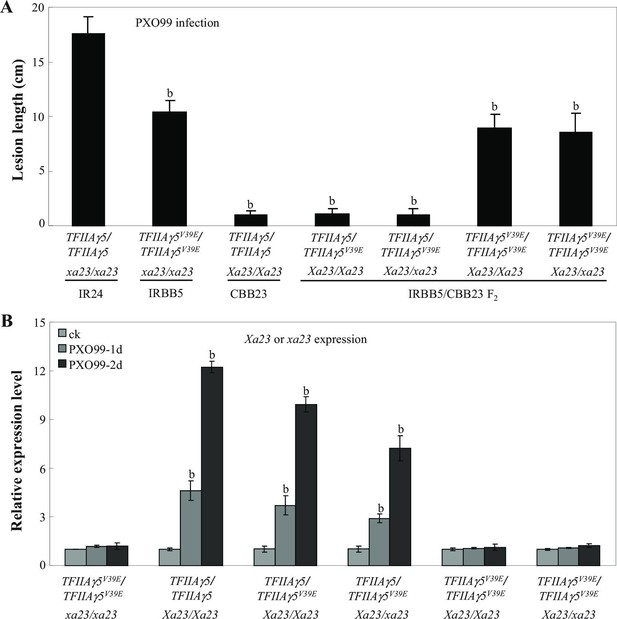
Effect of TFIIAγ5 on Xa23-mediated resistance to Xoo strain PXO99.
Each bar represents mean (total 40 to 45 leaves from 5 plants for lesion length and 3 replicates for gene expression) ± standard deviation. (A) Xa23 Xa23- mediated resistance required the presence of TFIIAγ5/TFIIAγ5 or TFIIAγ5/TFIIAγ5V39E. Rice plants at the booting stage were inoculated with Xoo. b, significant difference between IR24 and other plants at p<0.01. (B) PXO99 infection-induced Xa23 expression required the presence of TFIIAγ5/TFIIA5 or TFIIAγ5/TFIIA5V39E. b, significant difference between non-inoculated plants (ck) and PXO99-inoculated plants within each rice line at p<0.01. 1d and 2d, 1 day or 2 days after inoculation of PXO99.
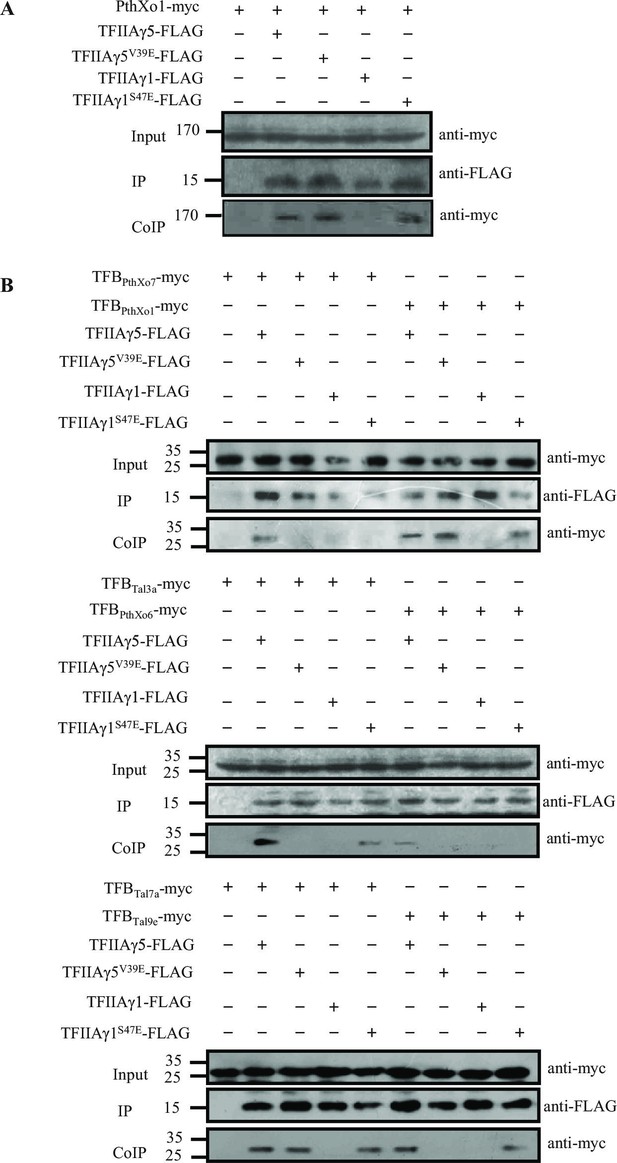
Detection of interactions between rice TFIIAγs and TALEs from Xoo in planta by co-immunoprecipitation.
The protein–protein interaction assays were performed in N. benthamiana leaf cells. Proteins before (input) and after immunoprecipitation (IP) were detected with anti-myc and anti-FLAG antibodies. (A) Interaction of the myc-labelled full-length PthXo1 with FLAG-labelled TFIIAγ5, TFIIAγ5V39E, and mutated rice TFIIAγ1 (TFIIAγ1S47E). (B) Interactions of the myc-labelled TFB regions of six TALEs with FLAG-labelled rice TFIIAγs.
-
Figure 2—source data 1
The defined domains/motifs and sequences of TALE PthXo1 from Xoo strain PXO99.
- https://doi.org/10.7554/eLife.19605.007
-
Figure 2—source data 2
Amino acid sequence alignment of the TFB regions of TALEs from Xanthomonas oryzae strains, composed of either 134 or 145 amino acids.
- https://doi.org/10.7554/eLife.19605.008
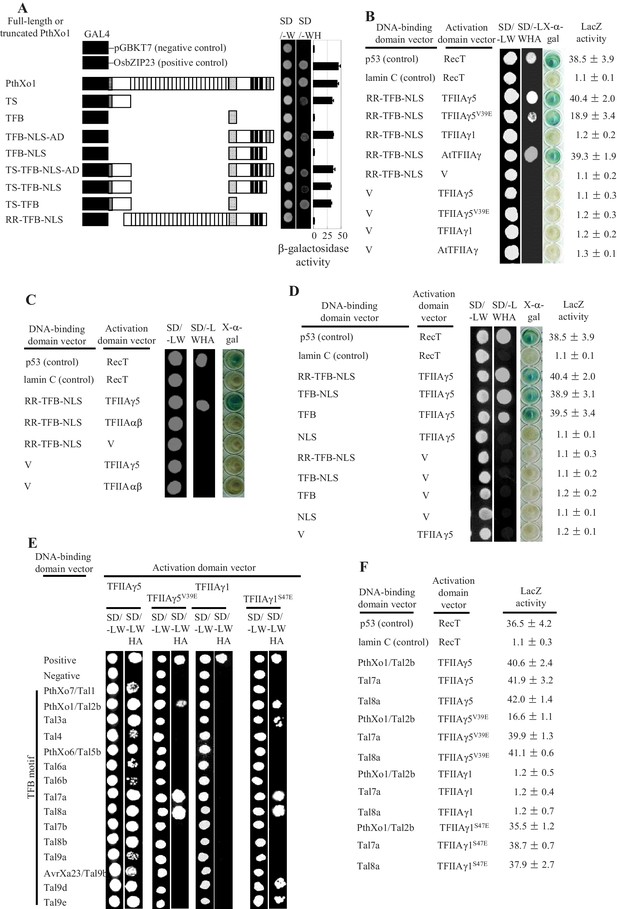
Interactions between Xoo TALEs and plant TFIIAγs in yeast cells.
The interactions were assessed by growth of yeast cells on synthetic defined premixes (SD) medium lacking (-) leucine (L), tryptophan (W), histidine (H), and adenine (A). V, empty vector as control. TS, translocation signal; RR, repeat region; TFB, transcription factor binding region; NLS, nuclear localization signal; AD, transcription activation domain. (A) Examination of transactivation activity of different domains and motifs of TALE PthXo1. The full-length and truncated PthXo1 were separately fused to the DNA-binding domain of GAL4, which is a yeast transcription factor, and transformed into yeast. (B) Truncated PthXo1 (RR-TFB-NLS) interacted with rice TFIIAγ5 and TFIIAγ5V39E (mutated TFIIAγ5) and Arabidopsis AtTFIIAγ analysed by yeast two-hybrid (Y2H) assay. (C) Truncated PthXo1 did not interact with rice basal transcription factor TFIIAαβ analysed by Y2H assay. (D) The TFB region of PthXo1 was required for the interaction with TFIIAγ5 analysed by Y2H assay. (E) The TFB regions of TALEs differentially interacted with rice TFIIAγs. The TFB regions of all the 15 TALEs from Xoo strain PXO99 interacted with rice TFIIAγ5, and the TFB regions of some of the 15 TALEs interacted with the mutated TFIIAγs from rice (TFIIAγ5V39E and TFIIAγ1S47E) analysed by Y2H assay. The Tal6b is a putative non-functional TALE. (F) The TFB regions of TALEs interacted with rice TFIIAγs with different strength based on the analysis of LacZ activity.
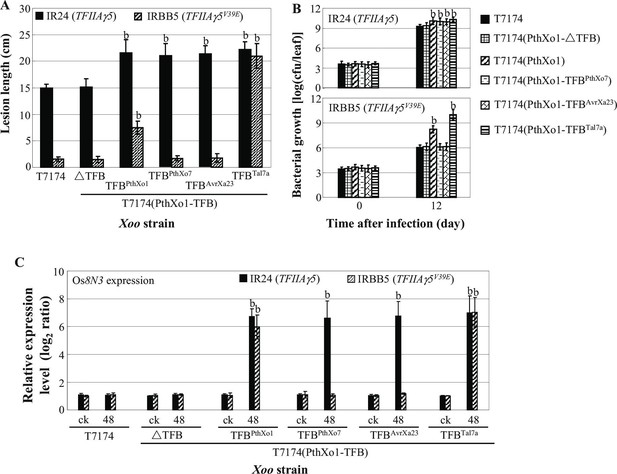
Effects of the TFB region of TALE PthXo1 on the virulence of Xoo strains and on the expression of rice susceptibility gene in rice−Xoo interaction.
Each bar represents mean (total 30 to 35 leaves from five plants for lesion length; three replicates for gene expression and bacterial growth rate) ± standard deviation. (A) Virulence of wild-type strain T7174 and recombinant strains carrying PthXo1 and its derivatives in IR24 and IRBB5. b, significant difference between T7174 and recombinant strains in each rice line at p<0.01. (B) Growth of different Xoo strains in rice leaves. b, significant difference between 0 day (30 min after infection) and 12 days after infection of each strain at p<0.01. (C) Expression of susceptibility gene Os8N3 after infection of different strains. b, significant difference between non-inoculated (ck) and inoculated (at 48 hr after infection of a strain) plants in each rice line at p<0.01.
-
Figure 3—source data 1
Effects of leucine residues of PthXo1 TFB region on TALE-mediated infection.
- https://doi.org/10.7554/eLife.19605.011

Effects of the TFB region of TALE PthXo1 on the virulence of Xoo strains and on the expression of rice susceptibility gene in rice−Xoo interaction.
Each bar represents mean (total 30 to 35 leaves from five plants for lesion length; three replicates for gene expression and bacterial growth rate) ± standard deviation. (A) Virulence of wild-type strain KACC10331 and recombinant strains carrying PthXo1 and its derivatives in near-isogenic lines IR24 and IRBB5. b, significant difference between T7174 and recombinant strains in each rice line at p<0.01. (B) Growth of different strains in rice leaves. b, significant difference between 0 day (30 minutes after infection) and 12 days after infection of each strain at p<0.01. (C) Expression of susceptibility gene Os8N3 after infection of different strains. b, significant difference between non-inoculated (ck) and inoculated (at 48 hr after infection of a strain) plants in each rice line at p<0.01.
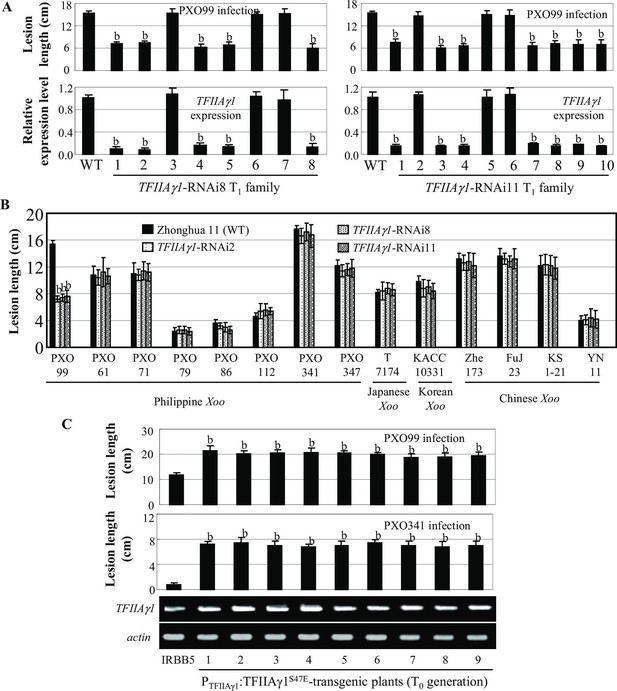
Effects of TFIIAγ1 on response to infections by different Xoo strains.
Plants were inoculated with Xoo at the booting stage. Each bar represents mean (three replicates for gene expression and total 35 to 40 leaves from five plants for lesion length) ± standard deviation. (A) Suppressing TFIIAγ1 enhanced rice resistance to strain PXO99. b, significant difference between wild-type (WT) Zhonghua 11 and transgenic plants at p<0.01. (B) Suppressing TFIIAγ1 did not change rice response to other strains. b, significant difference between WT and transgenic plants at p<0.01. (C) PTFIIAγ1:TFIIAγ1S47E-transgenic plants showed susceptibility to PXO99 and PXO341 compared to IRBB5. b, significant difference between IRBB5 and transgenic plants at p<0.01.
-
Figure 4—source data 1
Amino acid sequence alignment of basal transcription factor IIA gamma subunit (TFIIAγ) from different species.
- https://doi.org/10.7554/eLife.19605.014
-
Figure 4—source data 2
Single nucleotide polymorphisms in the TFIIAγ1 coding region of 1419 rice accessions from RiceVarMap (http://ricevarmap.ncpgr.cn).
- https://doi.org/10.7554/eLife.19605.015
-
Figure 4—source data 3
Single nucleotide polymorphisms in the TFIIAγ5 coding region of 1419 rice accessions from RiceVarMap (http://ricevarmap.ncpgr.cn).
- https://doi.org/10.7554/eLife.19605.016

Expression profiles of TFIIAγ5 and TFIIAγ1 in 28 tissues covering the entire life cycle of rice varieties Minghui 63 and Zhenshan 97.
Data were obtained from a microarray database (http://www.ncbi.nlm.nih.gov). E3, endosperm at 21 days after pollination; E2, endosperm at 14 days after pollination; E1, endosperm at 7 days after heading; Spi, spikelet at 3 days after pollination; Sta, stamen at 1 day before flowering; H, hull at 1 day before flowering; P5, panicle at heading stage; P4, panicle at 4- to 5-cm young panicle stage; P3, panicle at pollen–mother cell formation stage; P2, panicle at pistil and stamen primordium differentiation stage; P1, panicle at secondary branch primordium stage; Ste2, stem at heading stage; Ste1, stem at 5 days before heading; FL2, flag leaf at 14 days after heading; FL1, flag leaf at 5 days before heading; L2, leaf at 4- to 5-cm young panicle stage; L1, leaf at secondary branch primordium stage; She2, sheath at 4- to 5-cm young panicle stage; She1, sheath at secondary branch primordium stage; Sh, shoot ofseedling with two tillers; R, root of seedling with two tillers; L & R, leaf and root at three-leaf stage; Em & Ra, embryo and radicle at 3 days after germination; Pl1, plumule at 48 hr after emergence under light; Pl2, plumule at 48 hr after emergence under dark; Ra1, radicle at 48 hr after emergence under light; Ra2, radicle at 48 hr after emergence under dark; S, germinating seed at 72 hr of imbibitions. Expression levels (log2 transformations of average signal values) are color-coded: yellow and blue denote high and low expression, respectively.
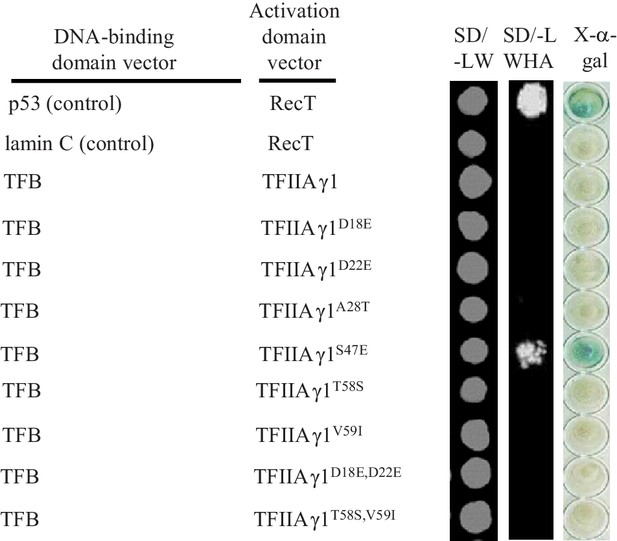
Interactions between the TFB region of TALE PthXo1 and mutated TFIIAγ1s in yeast cells.
The interactions were assessed by growth of yeast cells on synthetic defined premixes (SD) medium lacking leucine (L), tryptophan (W), histidine (H), and adenine (A).
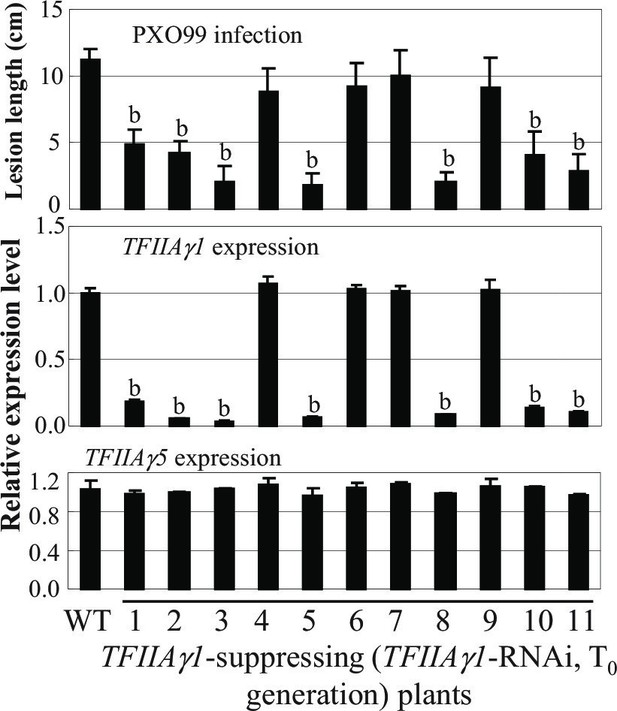
Effect of suppressing TFIIAγ1 on rice resistance to Xoo strain PXO99.
Plants were inoculated with PXO99 at the booting stage. RNA was isolated from the flag leaves. WT, wild-type Zhonghua 11. Each bar represents mean (3 replicates for gene expression and 5 to10 leaves from one plant for lesion length) ± standard deviation. b, significant difference between WT and transgenic plants at p<0.01.

Effect of mutation of TFIIAγ1 on the expression of disease susceptibility gene Os8N3 after Xoo infection.
Plants were inoculated with Xoo PXO99 at the booting stage. Each bar represents mean (3 replicates) ± standard deviation. b, significant difference between IRBB5 and transgenic plants at p<0.01. ck, before Xoo inoculation.
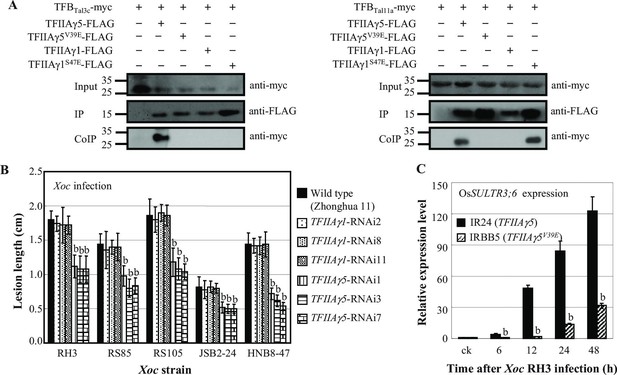
Effect of TFIIAγ on rice-Xoc interaction.
(A) Interactions of myc-labelled TFB regions of TALEs from Xoc RH3 and FLAG-labelled rice TFIIAγs in N. benthamiana leaf cells analysed by CoIP assays. Proteins before (input) and after immunoprecipitation (IP) were detected with anti-myc and anti-FLAG antibodies. (B) TFIIAγ5-RNAi but not TFIIAγ1-RNAi plants showed enhanced resistance to Xoc strains. Each bar represents mean (total 30 to 35 leaves from five plants) ± standard deviation. b, significant difference between wild-type and transgenic plants after infection of a strain at p<0.01. (C) Mutation of TFIIAγ5 (rice line IRBB5) reduced expression of disease susceptibility gene OsSULTR3;6 after infection. Each bar represents mean (three replicates) ± standard deviation. b, significant difference between IR24 and IRBB5 at p<0.01.
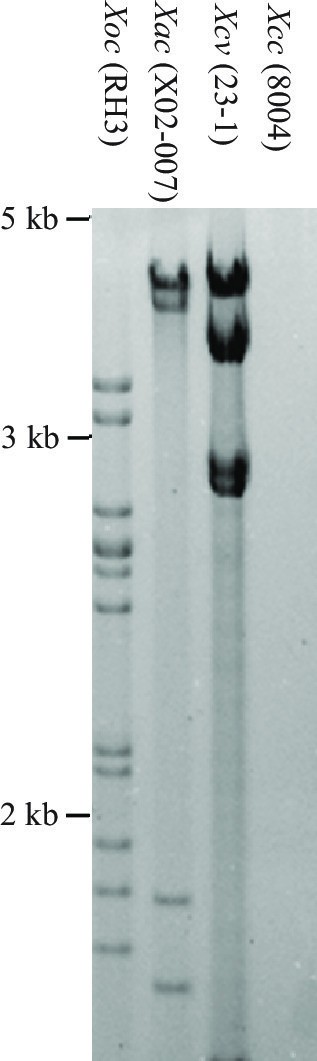
Southern hybridization analysis of TALE genes in different Xanthomonas species.
DNA was digested with SphI and probed with the 2.9-kb SphI fragment of TALE gene pthXo1 from Xoo strain PXO99. The size positions of DNA markers are indicated at left. Xac, Xanthomonas axonopodis pv. citri; Xcv, Xanthomonas euvesicatoria; Xoc, Xanthomonas oryzae pv. oryzicola; Xcc, Xanthomonas campestris pv. camperstris. The Xcc strain 8004 is TALE-free (Qian et al., 2005, Genome Res. 15:757-767).

Analysis of interactions between Xoc TALEs and rice TFIIAγs.
The physical interactions between TFB regions of TALEs from Xoc RH3 and TFIIAγs were assessed by growth of yeast cells on synthetic defined premixes (SD) medium lacking leucine (L), tryptophan (W), histidine (H), and adenine (A). Among the 18 TFB regions, the sequences of TFB11, TFB15, TFB23, TFB24, and TFB27 from RH3 were the same as the TFB regions of Tal9b, Tal11a, Tal5b, Tal3c, and Tal12 from sequenced Xoc strain BLS256, respectively. (A) The TFB regions from RH3 interacted with rice TFIIAγ5 analysed using yeast two-hybrid assay. (B) A randomly chosen 7 of the 18 TFB regions did not interact with TFIIAγ5V39E (the mutated TFIIAγ5), but three interacted with mutated rice TFIIAγ1S47E analysed by yeast two-hybrid assay. (C) Mutation of TFIIA5 enhanced resistance to Xoc. Plants were inoculated with Xoc at the booting stage. Each bar represents mean (total 40 to 50 leaves from 5 plants) ± standard deviation. b, significant difference between IR24 and IRBB5 plants at p<0.01.
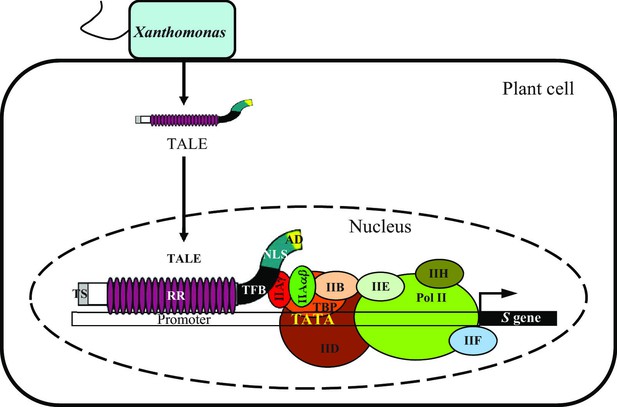
A model showing TFIIAγ5 as a key component of rice infection by Xanthomonas bacteria.
The bacteria hijack rice basal transcription factor TFIIAγ5 (IIAγ) by the transcription factor binding (TFB) region of their TALEs to induce host susceptibility (S) genes for infection. TS, amino-terminal translocation signal; RR, central repeat region; NLS, nuclear localization signal; AD, carboxyl-terminal transcription activation domain. The IIAγ belongs to the transcription pre-initiation complex. This complex consists of transcription factors IIA, which is composed of IIAβα subunit and IIAγ subunit, IIB, IID, IIE, IIF, and IIH, RNA polymerase II (Pol II), and TATA-binding protein (TBP). The binding of transcription pre-initiation complex to the TATA box of promoter was adopted and modified based on Yudkovsky et al. (2000).
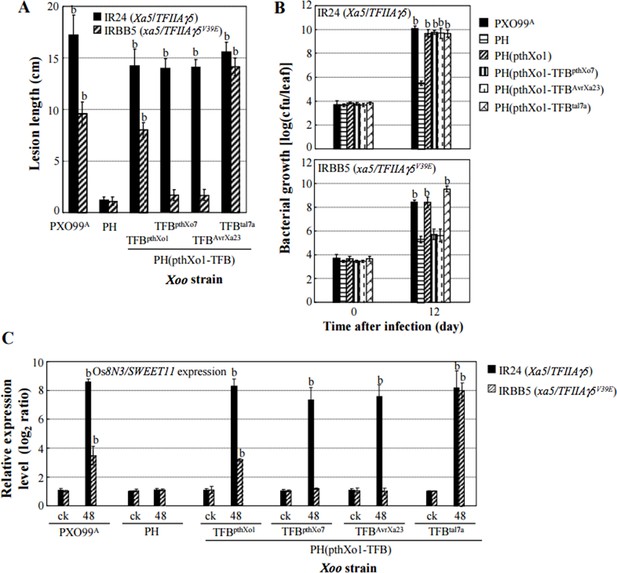
Effects of the TFB region of TALE pthXo1 on the virulence of Xoo strains and on the expression of rice susceptibility gene in rice−Xoo interaction.
Each bar represents mean (total 30 to 35 leaves from five plants for lesion length; three replicates for gene expression and bacterial growth rate) ± standard deviation. (A) Virulence of strain PH and its derivatives carrying pthXol or modified pthXo1 in near-isogenic lines IR24 and IRBB5. PH is an engineered TALE-free strain with the genetic background of strain PXO99A, which carries pthXo1. b, significant difference between PH and its derivatives in each rice line at P < 0.01. (B) Growth of different strains in rice leaves. b, significant difference between 0 day (30 minutes after infection) and 12 days after infection of each strain at P < 0.01. (C) Expression of susceptibility gene Os8N3/SWEET11 after infection of different strains. b, significant difference between non-inoculated (ck) and inoculated (at 48 h after infection of a strain) plants in each rice line at P < 0.01.
Additional files
-
Supplementary file 1
Measurements of agronomic traits of rice lines IR24 and IRBB5 under natural field conditions.
- https://doi.org/10.7554/eLife.19605.025
-
Supplementary file 2
PCR primers used for construction of vectors for protein–protein interactions and transformation, and detection of positive transgenic plants.
- https://doi.org/10.7554/eLife.19605.026
-
Supplementary file 3
PCR primers used for quantitative RT-PCR assays.
- https://doi.org/10.7554/eLife.19605.027





