Sparse activity of identified dentate granule cells during spatial exploration
Figures
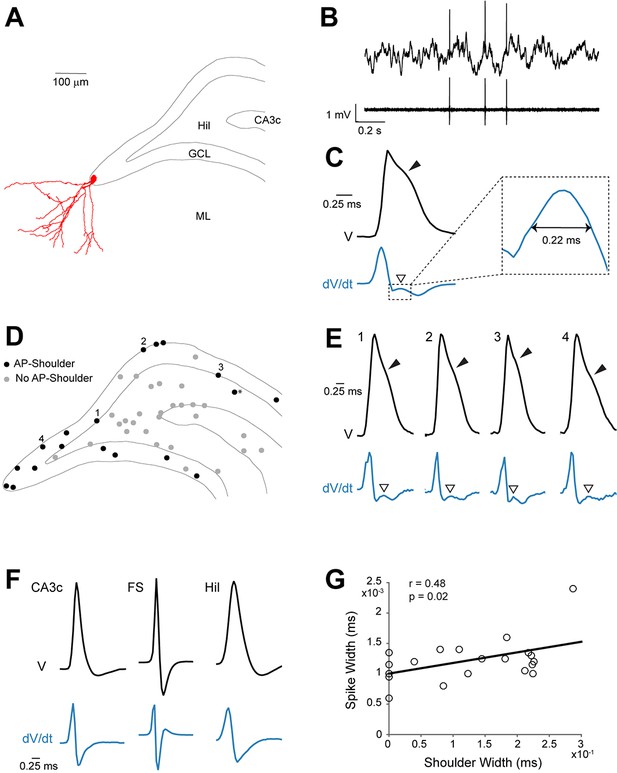
Spike waveform features of identified granule cells.
(A) Reconstruction of the somatodendritic morphology of a morphologically mature GC recorded in an awake, head-fixed animal. (B) Representative raw (top) and high-pass filtered (bottom) spike trace recorded under head-fixation from the cell shown in (A). (C) Average spike waveform (black) of the cell shown in (A). Note the presence of the shoulder (black arrowhead) on the spike waveform and the corresponding peak (white arrowhead) in the first derivative below (blue, dV/dt). The right panel shows a magnification of the inset on the left. The ‘shoulder width’ measured at half-maximum of the average spike waveform’s first derivative is illustrated by the double-headed arrow. The first derivative is scaled up for illustration purposes. (D) Schematic outline of the dentate gyrus showing the somatic location of all active neurons recorded and identified juxtacellularly (n=47 neurons: GCs, n=21; hilar cells, n=17; CA3c cells, n=7; and fast-spiking cells, n=2; see details in Materials and methods). Recordings where spike-shoulders (‘AP-shoulders’) were or were not detected are indicated as black or grey dots, respectively. The asterisk indicates the neuron located below the GC layer, where shoulders were also identified (see Materials and methods for details). (E) Representative normalized average spike waveforms (black) of four identified GCs with their corresponding first derivatives (blue). Black arrowheads indicate the spike-shoulder and white arrowheads the local peak in the first derivatives. First derivatives are scaled up for illustration purposes. The location of these cells within the GC layer is indicated by the corresponding numbers in (D). (F) Representative normalized average spike waveforms (black) of an identified CA3c cell, an identified fast-spiking cell (FS) and an identified hilar cell (Hil) with their corresponding first derivatives (blue). Note the absence of shoulders in the spike waveforms and first derivatives. First derivatives are scaled up for illustration purposes. (G) Correlation between spike width and ‘shoulder width’ for all identified GCs (n=21, see panel D). Regression line (black), Pearson’s correlation coefficient (r) and p-value are indicated. AP=action potential; Hil=hilus; GCL=granule cell layer; ML=molecular layer.
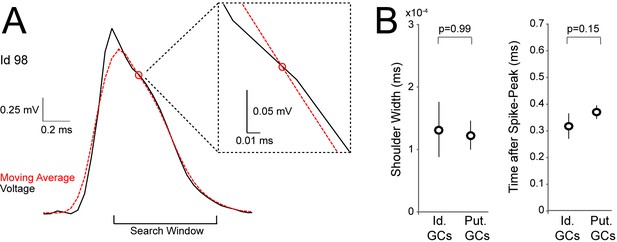
Detection of spike-shoulders.
(A) Representative average spike waveform of an identified GC (Id 98), showing how the moving-average approach was used to detect the presence of spike-shoulders. The moving average (red dotted trace) was used to smooth high-frequency fluctuations in the spike trace (black) and to capture the underlying slow trends. The spike trace in the repolarization phase was compared to its time-lagged average and slower changes (shoulders) were detected as intersections between them. The right panel shows a close-up magnification of the intersection (indicated by a red circle) between spike voltage trace and its moving average. (B) Variability of spike-shoulder prominence (quantified as the half-width of the local maximum on the first derivative of the spike trace; see Materials and methods) and location relative to the spike peak (quantified as the time difference between the spike-peak and the intersection with the moving average, as in A) for identified (Id.; n=6) and classified GCs (Put.; n=18) recorded in freely moving rats. Error bars indicate SEM. The variability of spike shoulders was not significantly different between identified and classified GCs (p-values are indicated; Mann-Whitney test).
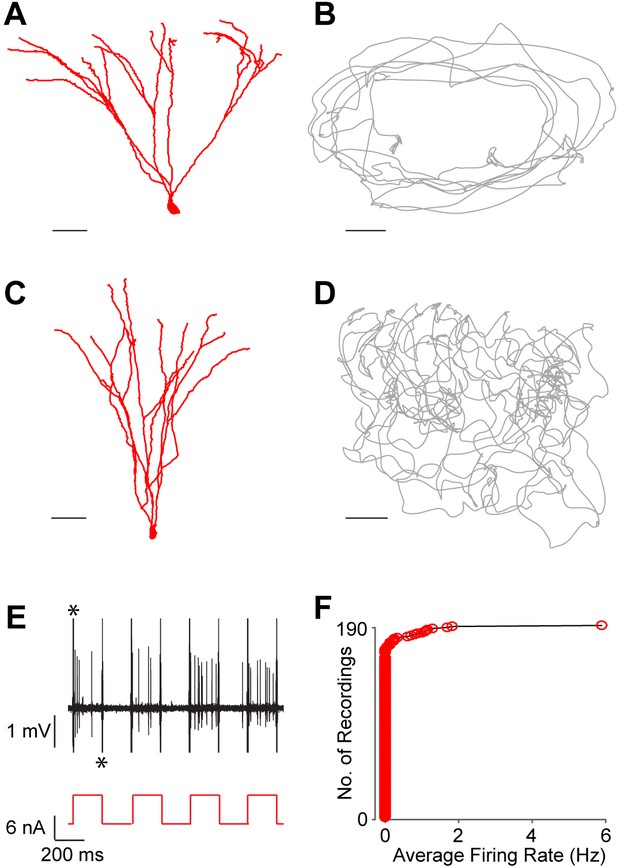
Silent granule cells during spatial exploration.
(A and C) Reconstruction of the dendritic morphology of two silent, morphologically mature GCs recorded during freely moving behavior (cell ids 924 and 991, respectively). Scale bars=50 µm. (B and D) Trajectory of a rat (grey) running on an O-shaped arena (B) or on an open-fieldarena (D). The two recordings correspond to the neurons in (A and C), respectively. Scale bars=10 cm. (E) Unlike extracellular recordings, juxtacellular sampling is not biased towards active cells, as silent neurons can also be recorded and their presence confirmed by current injection. This is shown here for the silent cell in (C): action potential firing (black, top) induced by squared pulses of injected current (red, bottom). These pulses were delivered at the end of the freely moving recording session to confirm the presence of the silent celland to label the cell (C). Asterisks indicate stimulation artifacts (truncated for display purposes). (F) Cumulative plot showing the firing-rate distribution within the GC layer. Each red circle represents one neuron, sampled juxtacellularly within the GC layer (see Materials and methods for details). Note the large proportion of silent neurons (163 out of 190) compared to that of active cells; this is likely to be an underestimate of the true silent proportion (see Results and Materials and methods for details).
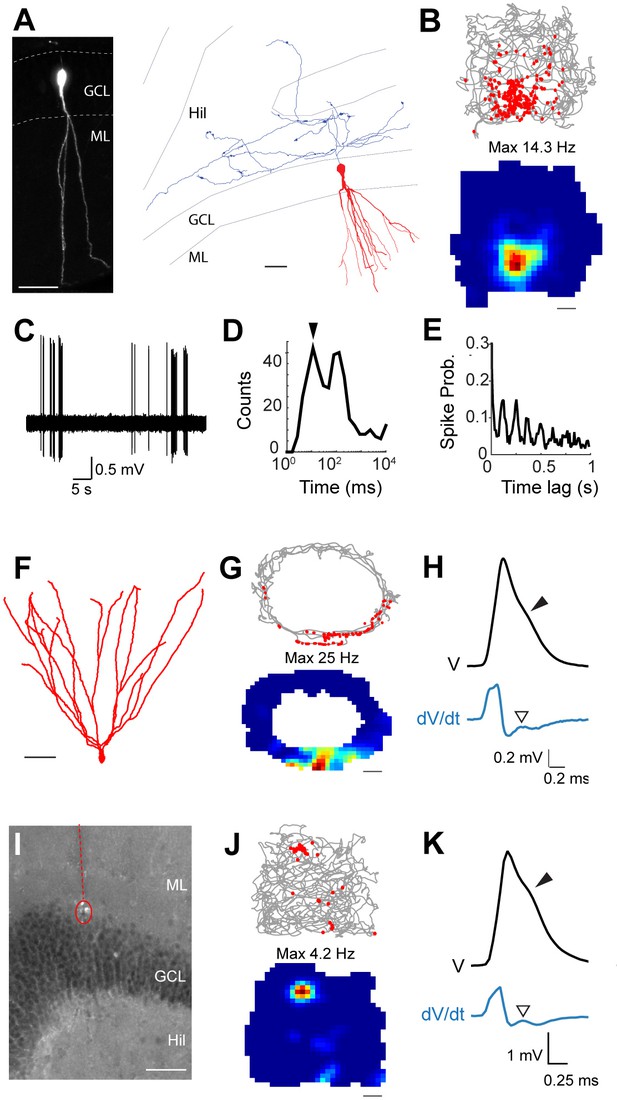
Spatial and temporal firing properties of identified active granule cells.
(A) Left, single-plane confocal image showing a morphologically mature GC (cell id 103) located in the infrapyramidal blade, recorded and labeled in a freely moving animal. Right, reconstruction of dendritic (red) and axonal (blue) morphologies; only the axon extending within the hilus (Hil) is shown. Close anatomical proximity (< 1 µm; compatible with putative synaptic contacts) was observed between mossy boutons and large hilar neurons (4 out of 13 analyzed boutons; not shown), suggesting that hilar neurons might integrate spatial signals from upstream GCs. Scale bars=50 μm. GCL=granule cell layer; ML=molecular layer. (B) Spike-trajectory plot (top) and rate map (bottom) for the neuron shown in (A), recorded in a square arena. Scale bar=10 cm. (C) Representative high-pass filtered spike trace, recorded during free behavior from the neuron in (A). (D) Interspike interval (ISI) distribution for the neuron in (A). Note the early peak (arrowhead) representing short ISI within bursts. (E) Spike-autocorrelogram for the neuron in (A). Note the prominent rhythmicity in the theta-frequency range (~8 Hz during awake behavior). (F) Reconstruction of the dendritic morphology of an active morphologically mature GC (cell id 104) recorded during freely moving behavior. Scale bar=50 µm. (G) Spike-trajectory plot (top) and rate map (bottom) for the neuron shown in (F) recorded on an O-shaped linear arena. Scale bar=10 cm. (H) Normalized average spike waveform (black) of the neuron shown in (F), with corresponding first derivative (blue). Black arrowhead indicates the spike-shoulder and white arrowhead the local peak in the first derivative. The first derivative is scaled up for illustration purposes. (I) Fluorescence micrograph showing a histologically verified electrode track (red dotted line) and recordings site (dotted circle), which was localized to the superficial portion of the GC layer (end of the track). Note the presence of small cell debris at the labeling site (dotted circle). (J) Spike-trajectory plot (top) and rate map (bottom) for the recording location shown in (I) (putative GC recording, cell id 80) during exploration of a square open arena. Scale bar= 10 cm. (K) Same as in (H), but for the putative GC recording in (I).
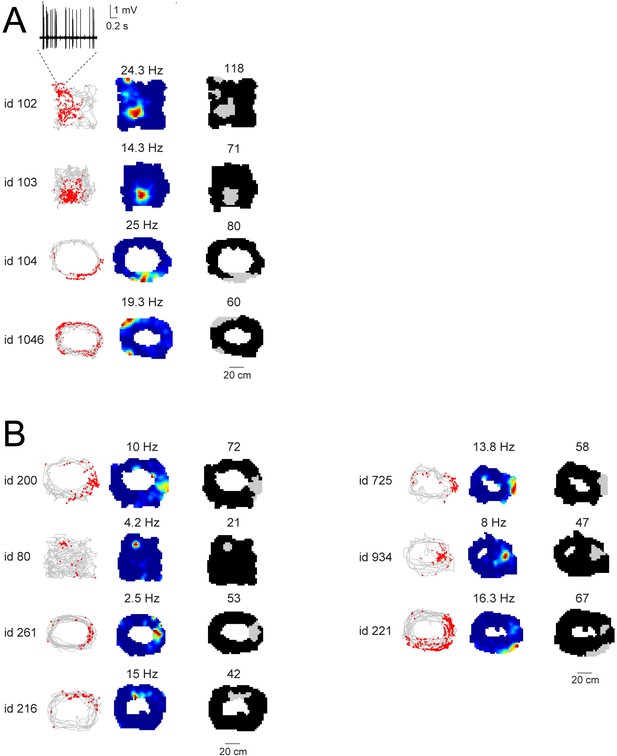

Spatial firing activity of identified and putative GCs.
(A) Spike-trajectory plots (left) and rate maps (middle) for the identified GCs which were classified as spatially modulated (see Materials and methods for details). Cell id and maximal firing rates are indicated. In the right panels, firing fields are outlined as grey areas (see Materials and methods). The total size of the firing fields (expressed in number of pixels) is indicated. For cell id 102, the multiple firing fields might result from suboptimal spatial coverage, as indicated by the close-up magnification on a single-pass at the upper-left corner of the maze. The corresponding high-pass filtered juxtacellular spike trace is indicated. (B) Same as (A), but for ‘putative’ GCs.
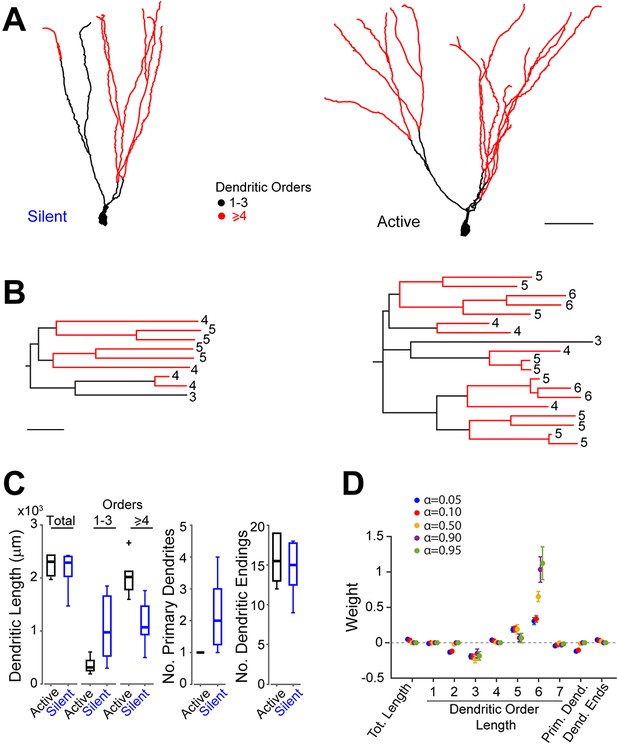
Morphological analysis of identified active and silent granule cells.
(A) Somatodendritic reconstructions of silent (left, cell id 993) and active (right, cell id 102) GCs, recorded in freely moving rats. Dendritic branch orders 1–3 are indicated in black, while high-order branches (≥ 4) are indicated in red. Scale bar=50 µm. (B) Dendrograms for the cells in (A). The branch orders of the terminal dendritic tips are indicated. Color codes same as in (A). Scale bar=50 µm. (C) Box-plots comparing primary dendritic parameters (active GCs, n=6; silent GCs, n=7). Whiskers represent 1.5 IQR. Outliers are shown as crosses. (D) Weights of logistic regression classifier (mean ± SEM over cross-validation folds) used for classifying GCs as active or silent for different levels of sparseness (α=0.05 corresponds to dense weights, α=0.95 corresponds to sparse weights).
-
Figure 4—source code 1
Classification analysis.
Matlab script to load Figure 4—source data 1, run the logistic regression classification analysis on primary morphological features (Figure 4D), perform the shuffle analysis and run the control analysis with all morphological features.
- https://doi.org/10.7554/eLife.20252.008
-
Figure 4—source data 1
Electrophysiological and morphological parameters of the identified GCs.
The table summarizes the main electrophysiological and morphological properties of the identified and reconstructed silent and active GCs (source data for Figure 4). Morphologies of the neurons are shown in Figure 4—figure supplement 1.
- https://doi.org/10.7554/eLife.20252.009
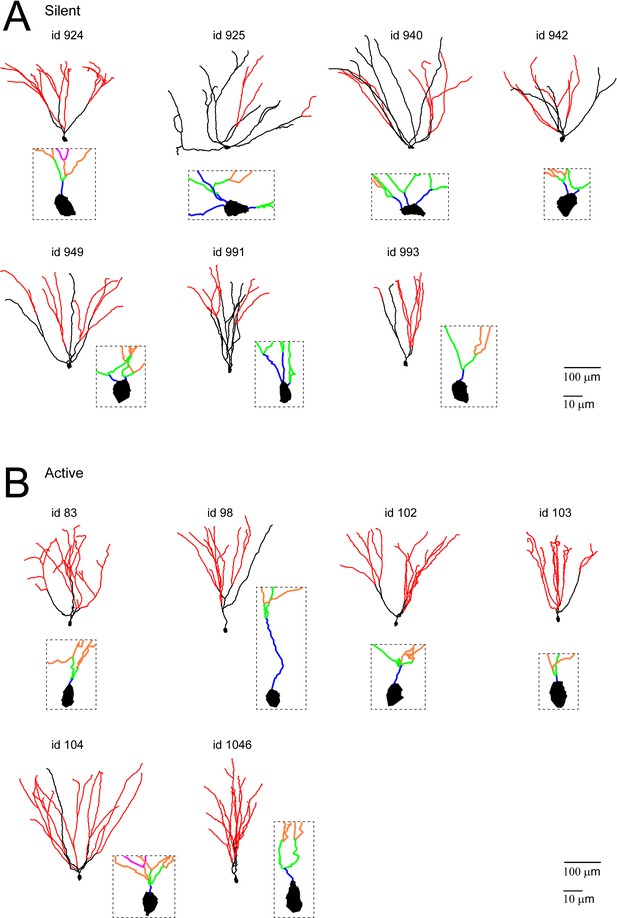
Morphological reconstructions of GCs recorded and labeled in freely moving rats.
(A, B) Morphological reconstructions of silent (A) and active (B) GCs, recorded in freely moving rats. Dendritic branch orders 1–3 are indicated in black, while high-order branches (≥4) are indicated in red (same color-code as in Figure 4A). The insets show close-up magnifications of the soma and the proximal dendrites of each cell (first-order dendrites are in blue, second-order in green, third-order in orange and fourth-order in magenta). Scale bars=100 µm for the low magnification and 10 μm for the insets. Cell ids are indicated.
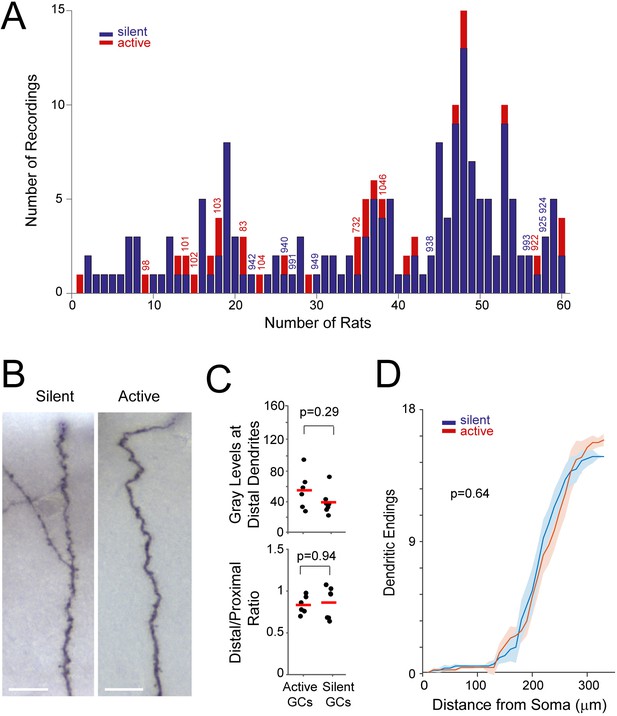
Distribution of recordings across animals and quantification of single-cell labeling quality.
(A) Histogram showing the distribution of active (red) and silent recordings (blue) across animals (only recordings lasting > 60 s are included in the analysis; see Materials and methods). Identified recordings are indicated by the corresponding ids. Note that to ensure unequivocal cell identification, a low number of electrode penetrations and recordings were performed for each rat (average number of recordings per rat = 3.2 ± 2.7). In the animals with a relatively large number of recordings (> 5), either labeling was not attempted or cell identification could not be assured with certainty because of multiple consecutive electrode penetrations. We note that, given the relatively small number of observations for each rat, a dependency between rat identity and morphological properties cannot be rigorously tested and thus formally ruled out. (B) Representative micrographs of distal dendritic segments for an active and a silent neuron. Scale bars=10 µm. (C) Maximum gray scale values for distal dendritic compartments (top) and the ratio between distal and proximal dendrites (bottom) for active (n=6) and silent neurons (n=7). p-values are indicated (Mann-Whitney U test). See Materials and methods for details. (D) Distribution of terminal dendritic endings as a function of distance from the soma for active (n=6, red) and silent neurons (n=7, blue). Lines represent means, shadows represent SD. The distances of dendritic endings were not significantly different between active and silent neurons (Mann Whitney U Test, p-value is indicated).
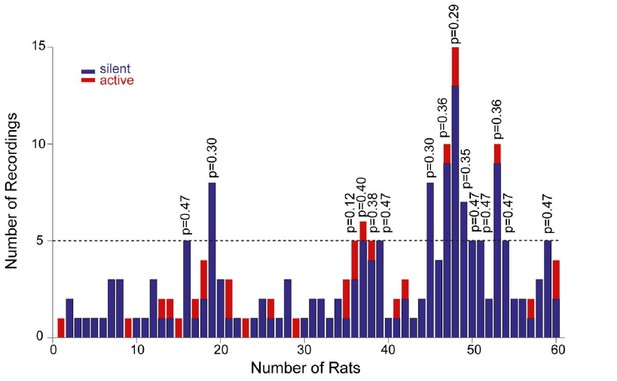
Distribution of recordings across animals.
Histogram showing the distribution of active (red) and silent recordings (blue) across animals. In the animals with a relatively larger number of recordings (≥5, n=15 rats), the occurrence of active neurons was not significantly different from their expected proportion from the population data (~14%). Corresponding p values from binomial test are shown.
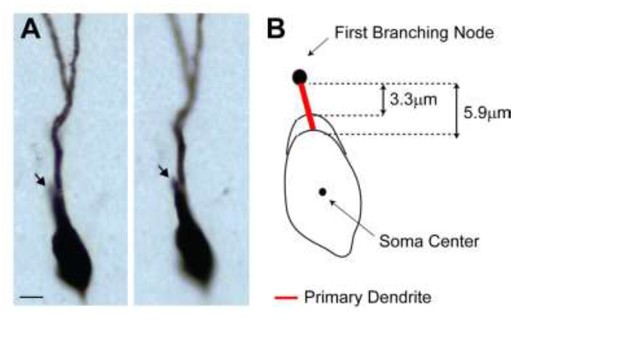
(A) Representative pictures of the cell id 103 at two different focal planes.
Note the presence of a single apical primary dendrite. The arrow points to the initial segment of a secondary dendrite, which continues in the neighboring section. (B) Schematic diagram showing the jitter in estimating the length of short primary dendritic segments, depending on the shape (i.e. more or less elongated) of the somatic contour.





