Cholesterol activates the G-protein coupled receptor Smoothened to promote Hedgehog signaling
Figures
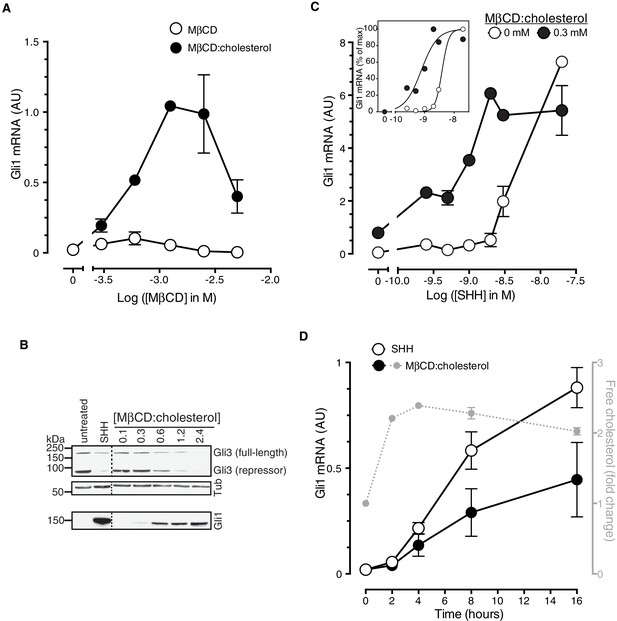
Cholesterol is sufficient to activate Hh target genes in NIH/3T3 cells.
(A) Gli1 mRNA, encoded by a direct Hh target gene, was measured by quantitative real-time reverse-transcription PCR (qRT-PCR) and normalized to mRNA levels of the housekeeping gene GAPDH after treatment (12 hr) with various doses of naked MβCD or a saturated MβCD:cholesterol (8.8:1 molar ratio) complex. In both cases, the concentration of MβCD is plotted on the abscissa. (B) Immunoblotting was used to measure protein levels of GLI1, full-length GLI3 and the repressor fragment of GLI3 after treatment (12 hr) with various concentrations (in mM) of MβCD:cholesterol. Dotted lines demarcate non-contiguous regions of the same immunoblot that were juxtaposed for clarity. (C) Gli1 induction in response to various doses of SHH in the presence or absence of a low dose of MβCD:cholesterol. Inset shows non-linear curve fits to the data after a normalization in which the Gli1 mRNA level in the absence of SHH was set to 0% and at the maximum dose of SHH was set to 100%. (D) Time course of Gli1 induction (left y-axis) after treatment with SHH (265 nM) or the MβCD:cholesterol complex (2.5 mM). The gray circles (right y-axis) show the kinetics of increase in unesterified cholesterol (normalized to total protein) after the addition of MβCD:cholesterol. In all graphs, circles depict mean values from 3 replicates and error bars show the SD.
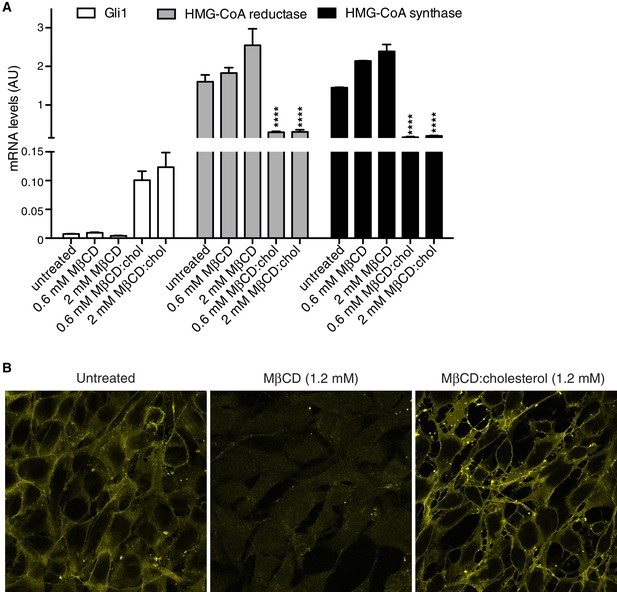
MβCD:cholesterol treatment increases the free cholesterol content of NIH/3T3 cells.
(A) Mean (±SD, n = 4) mRNA levels of Gli1 or of two genes, encoding HMG-CoA reductase and synthase, in the cholesterol biosynthetic pathway that are negatively regulated by cellular cholesterol levels are shown after treatment with the indicated concentrations of MβCD or the MβCD:cholesterol complex. Asterisks denote statistical significance for difference from the untreated sample using two-way ANOVA with a Holm-Sidak post-test. (B) Levels of free cholesterol in the plasma membrane were assessed by staining with fluorescently labeled Perfringolysin O (PFO), a toxin that preferentially binds to the accessible (or chemically active) pool of cholesterol in membranes.
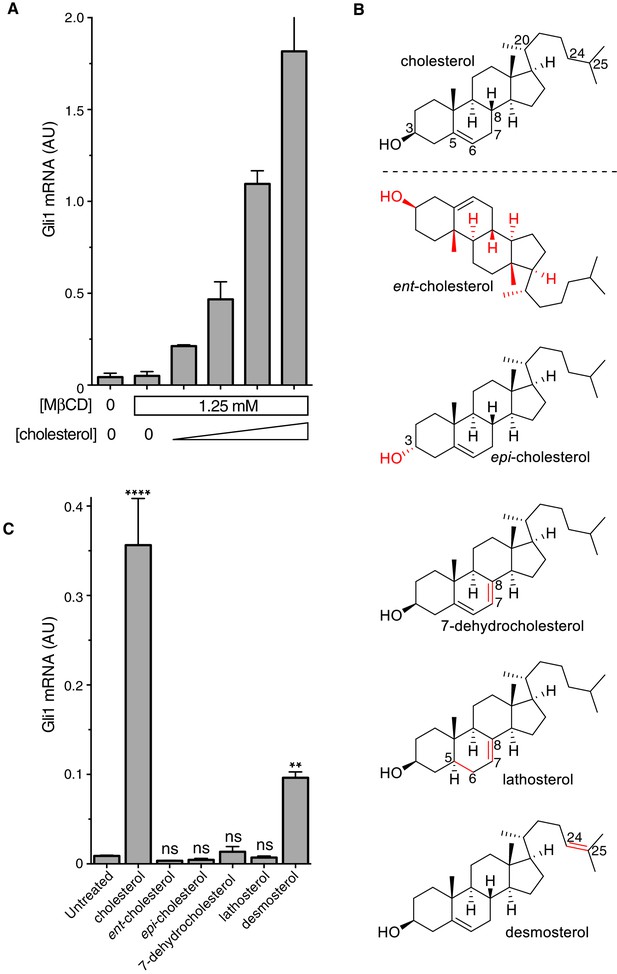
The cholesterol in MβCD:cholesterol complexes activates Hedgehog signaling.
(A) Mean (±SD, n = 3) Gli1 mRNA levels after 12 hr of treatment of NIH/3T3 cells with a series of inclusion complexes in which the MβCD concentration was clamped at 1.25 mM while the cholesterol concentration was varied to yield MβCD:cholesterol molar ratios of 12:1, 9:1, 7:1 and 6:1. (B) Structures of cholesterol analogs tested for Hh signaling activity as inclusion complexes with MβCD. Structural differences from cholesterol are highlighted in red: ent-cholesterol is the mirror-image of cholesterol with inverted stereochemistry at all 8 stereocenters; epi-cholesterol is a diastereomer with inverted stereochemistry only at the 3 carbon postion; 7-dehydrocholesterol, lathosterol and desmosterol are naturally occurring cholesterol precursors. (C) Mean (±SD, n = 4) Gli1 mRNA levels after treatment (12 hr) with inclusion complexes of MβCD (1.25 mM) with the indicated sterols (see B for structures). Asterisks denote statistical significance for difference from the untreated sample using one-way ANOVA with a Holm-Sidak post-test.
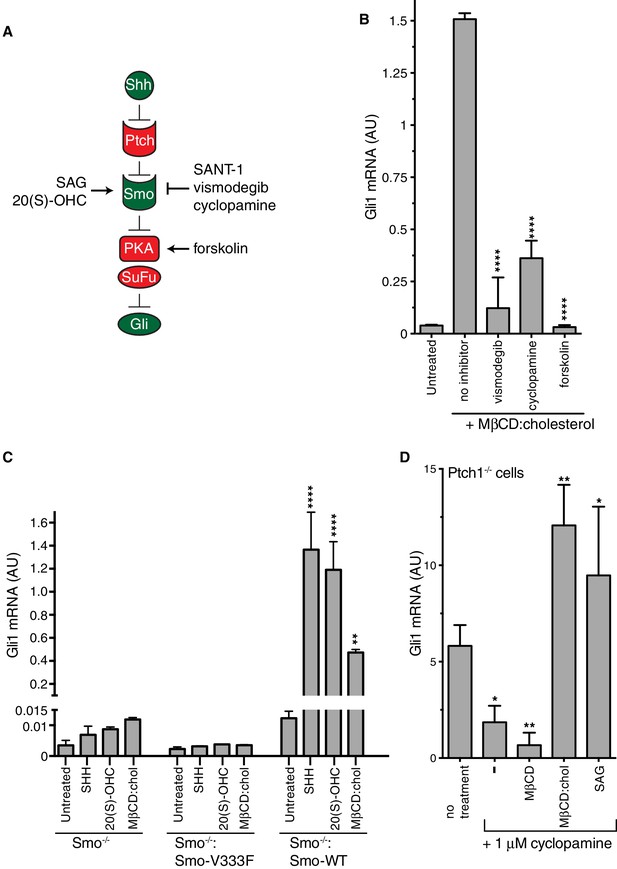
Smoothened activity is necessary for cholesterol to activate Hh signaling.
(A) Schematic of the Hh signaling pathway showing the sequence in which core components function to transmit the signal from the cell surface to the nucleus. SAG and 20(S)-OHC are agonists and SANT-1, vismodegib, and cyclopamine are antagonists that bind and modulate the activity of SMO. Forskolin blocks signaling by elevating cAMP levels, which increases the activity of Protein Kinase A. (B) Mean (±SD, n = 3) Gli1 mRNA levels after treatment with MβCD:cholesterol (1.25 mM, 12 hr) in the presence of vismodegib (1 μM), cyclopamine (10 μM) or forskolin (10 μM). (C) Mean (±SD, n = 3) Gli1 mRNA levels after addition of agonists (12 hr) to Smo-/- cells, in which both Smo alleles have been genetically inactivated, or Smo-/- cells stably expressing a wild-type (WT) SMO protein or a variant SMO protein carrying an inactivating mutation (V333F) in its 7TMD (Byrne et al., 2016). SHH was used at 265 nM, 20(S)-OHC at 5 μM, and MβCD:cholesterol at 1.25 mM. (D) Mean (±SD, n = 4) Gli1 mRNA levels in Ptch1-/- cells after treatment with cyclopamine alone or cyclopamine in the presence of SAG (100 nM), MβCD (1.25 mM) or MβCD:cholesterol (1.25 mM). Asterisks denote statistical significance for differences from the “no inhibitor” sample in B, the V333F-expressing cell line in C, and the “no treatment” sample in D using one-way (B, D) or two-way (C) ANOVA with a Holm-Sidak post-test.
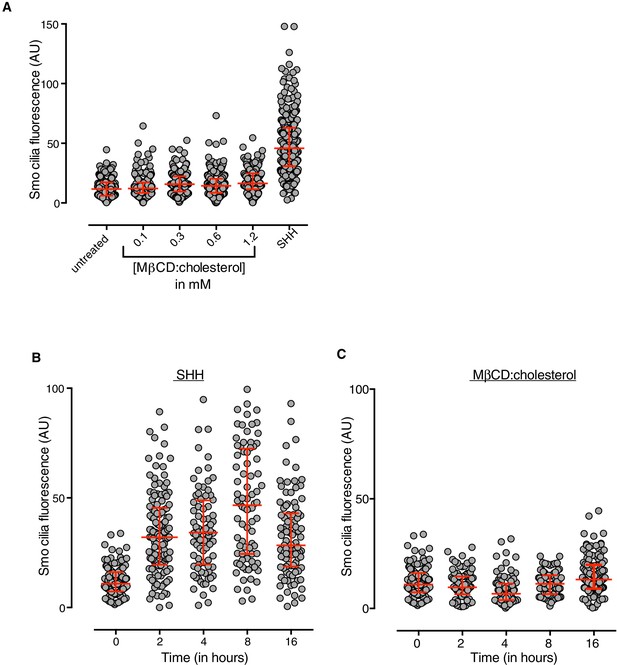
MβCD:cholesterol fails to drive SMO accumulation in the ciliary membrane.
(A) SMO protein levels in primary cilia were determined by immunostaining NIH/3T3 cells after treatment (12 hr) with SHH (265 nM) or the indicated concentrations of MβCD:cholesterol. The kinetics of SMO accumulation in cilia were measured after treatment with SHH (B, 265 nM) or MβCD:cholesterol (C, 1.2 mM). Each point depicts SMO fluorescence at a single cilium and the red bars show the median and interquartile range of measurements from ~250 cilia per condition for A and ~100 cilia per condition for B and C.
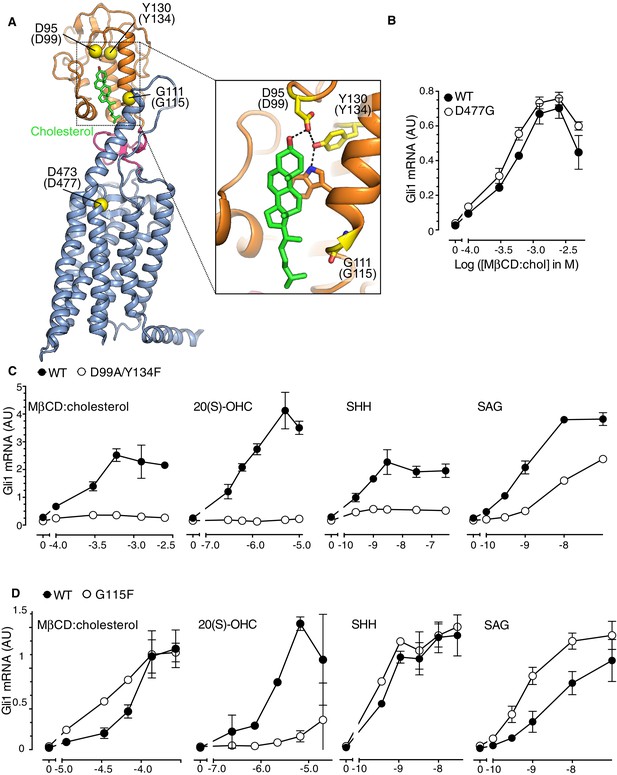
The Smoothened cysteine-rich domain is required for cholesterol-mediated activation of Hh signaling.
(A) Structure of human SMO (PDB 5L7D), with the CRD in orange, the 7TMD in blue, the linker domain in pink, and the cholesterol ligand bound to the CRD in green. The Cα positions of the gatekeeper residues in the two ligand binding sites are highlighted as yellow spheres and numbered, with the mouse numbering shown in parenthesis. The inset shows a close-up of the cholesterol-binding site. D95 and Y130 form part of a hydrogen-bonding network (dotted lines) with the 3-hydroxyl of cholesterol, G111 abuts the iso-octyl chain of cholesterol, and D473 is a critical residue in the 7TMD binding-site. (B, C and D) Dose-response curves for the indicated agonists in Smo-/- cells stably expressing WT SMO (always solid black circles) or the indicated SMO variants (open circles) carrying mutations in the 7TMD ligand-binding site (B) or at two opposite ends of the CRD binding groove (C and D). All agonists were applied to cells for 12 hr and mean (±SD) values for Gli1 mRNA are plotted based on 3 replicates. In C and D, values on the abscissa represent Log ([Agonist] in M) and the ordinate for all four graphs is only shown once at the left.
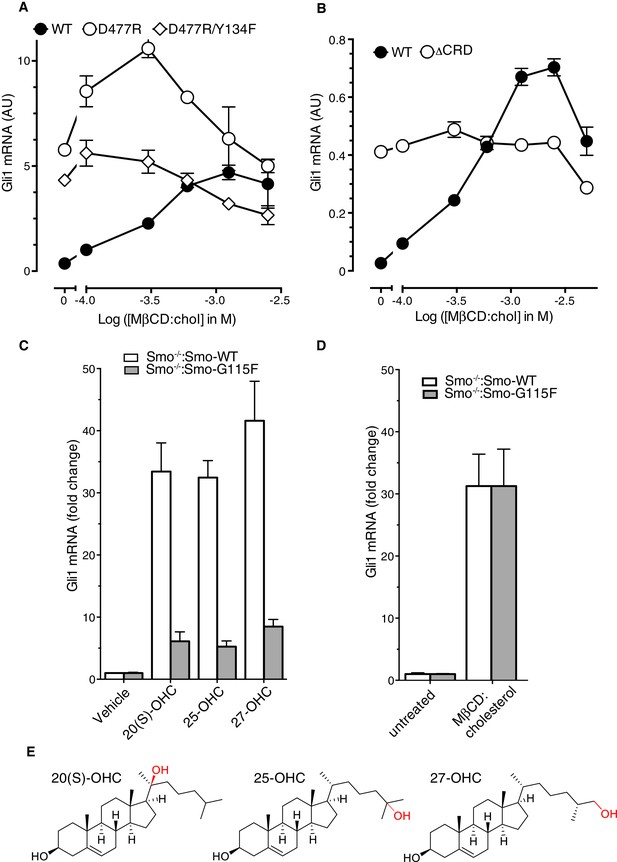
Role of the cysteine-rich domain of Smoothened in responses to cholesterol and side-chain oxysterols.
(A and B) Dose-response curves for MβCD:cholesterol in Smo-/- cells stably expressing WT SMO (solid black circles) or the indicated SMO mutants. D477R (A) is an activating mutation in the 7TMD, Y134F (A) is a mutation in the CRD (see Figure 5) that abrogates cholesterol and oxysterol responses, and ΔCRD (B) is an activating N-terminal truncation mutant that lacks the entire CRD. (C) Gli1 induction in Smo-/- cells expressing SMO-WT or SMO-G115F (see Figure 4 and associated discussion) treated with the indicated side-chain oxysterols, each applied at 5 μM as an inclusion complex with 44 μM MβCD. The activity of MβCD:cholesterol (1.2 mM) in both cell lines is shown in (D) for comparison. (E) Structures of the various side-chain oxysterols used in C, with differences from cholesterol highlighted in red.
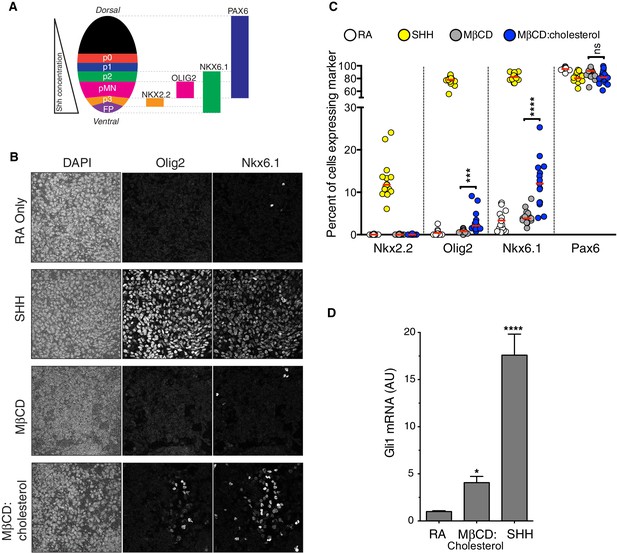
Cholesterol induces the differentiation of neural progenitors.
(A) A schematic illustrating the relationship between marker proteins used to assess differentiation and progenitor cell populations in the embryonic neural tube (taken from (Niewiadomski et al., 2014)). FP – floor plate progenitors, MN – motor neuron progenitors, p0, p1, p2, p3 – ventral interneuron progenitors. (B) Differentiation of neural progenitors was assessed by immunostaining for Nkx6.1+ and Olig2+ expression (see A) after treatment (48 hr) with Retinoic Acid (RA, 100 nM) alone or RA plus SHH (25 nM), MβCD (2 mM) or the saturated MβCD:cholesterol inclusion complex (2 mM). The percentage of nuclei (stained with DAPI) positive for four differentiation markers (see A) in 15 different images is plotted in (C), with each point representing one image of the type shown in (B) and the red line drawn at the median value. Asterisks denote statistical significance (unpaired t-test, Holm-Sidak correction, n = 15) for the comparison between cells treated with RA+MβCD and RA+MβCD:cholesterol. (D) Gli1 mRNA (mean ± SD, n = 3) after 48 hr of the indicated treatments. Asterisks denote statistical significance for difference from the RA-treated sample using one-way ANOVA with a Holm-Sidak post-test.
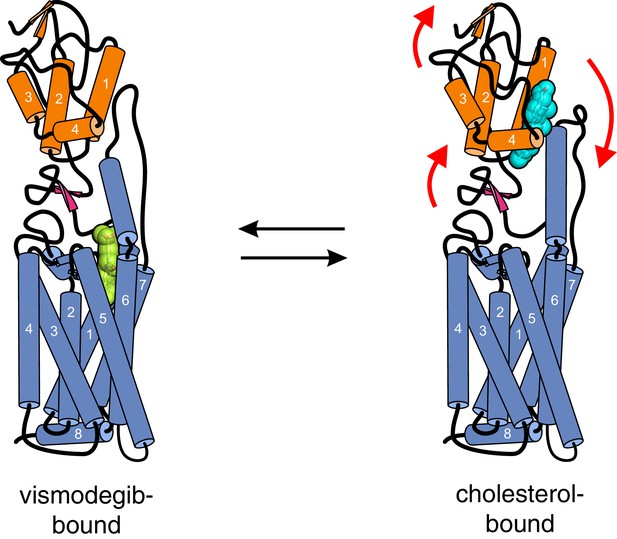
Conformational changes in Smoothened induced by cholesterol binding.
A comparison of the structures of SMO△C bound to vismodegib (PDB ID 5L7I, left), representing an inactive state, and cholesterol (PDB ID 5L7D, right), highlights a rotation of the CRD and the helical extracellular loop 3 relative to the 7TMD. This rotation may communicate ligand binding at the CRD to conformational changes in the 7TMD.
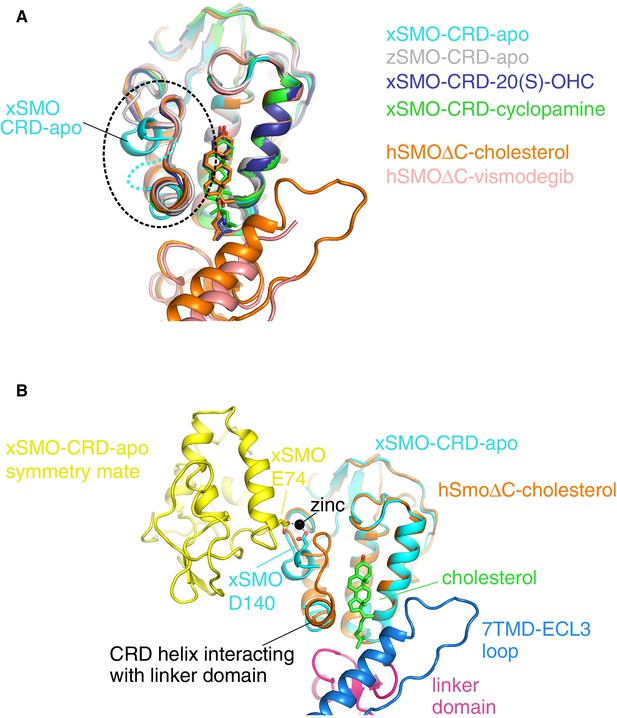
A comparison of all Smoothened CRD structures.
(A) Superposition of four isolated CRD structures (turquoise, gray, blue, green) and two CRD structures (orange, pink) solved in the context of a complete SMO△C molecule also containing the linker domain and the 7TMD. The dotted circle highlights the location of the unique conformation and associated disorder seen in unliganded, Xenopus apo-CRD (turquoise). ‘x’ denotes Xenopus laevis, ‘z’ Danio rerio, and ‘h’ human SMO. ‘apo’ denotes un-liganded structures; otherwise, the specific ligands are noted in the legend. PDB IDs used to generate this superposition are as follows: 5KZZ (xSMO-CRD-apo), 5KZV (xSMO-CRD-20(S)-OHC), 5KZY (xSMO-CRD-cyclopamine), 4C79 (zSMO-CRD-apo), 5L7D (hSMO△C-cholesterol) and 5L7I (hSMO△C-vismodegib). (B) Superposition of the CRDs in xSMO-CRD-apo (turquoise) and hSMO△C-cholesterol (orange) is shown along with a symmetry partner (yellow) of xSMO-CRD-apo seen in the crystal. The region of altered conformation is partially disordered and forms a zinc-stabilized contact between the two symmetry related xSMO-CRD-apo molecules.
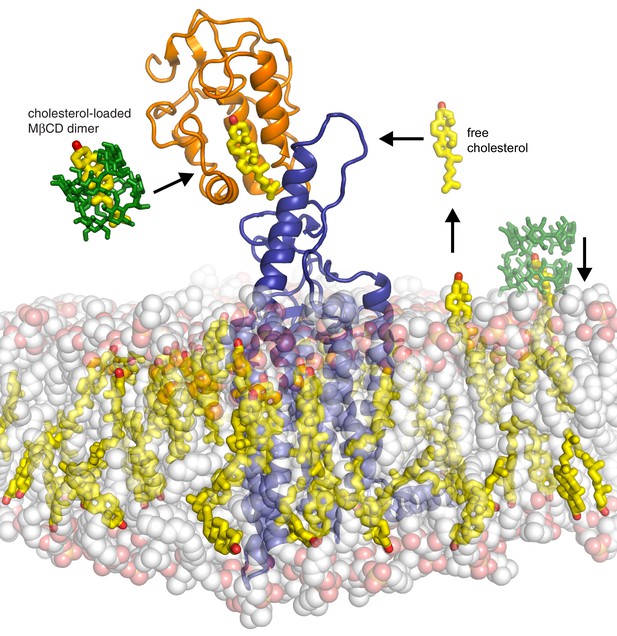
Models for how cholesterol may gain access to its binding-site in the SMO cysteine-rich domain.
The structure of SMO bound to cholesterol (PDB 5L7D) is shown embedded in a lipid bilayer composed of 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) and cholesterol in a ratio of 3:1 (Byrne et al., 2016). The SMO CRD is colored orange; the linker domain and 7TMD are colored blue. Two molecules of MβCD (PDB QKH, shown as green sticks) form an inclusion complex with each molecule of cholesterol (PDB CLR, colored yellow in stick representation with the 3-hydroxyl shown red). MβCD could deliver cholesterol directly to the CRD binding pocket (left) or to the outer leaflet of the plasma membrane (right), which would subsequently require a second transfer step from the membrane to the CRD. The activation energy for the direct delivery mechanism on the left (<10 kcal/mole) is much lower than for the mechanism on the right (~20 kcal/mole), where cholesterol has to desolvate from the membrane without a carrier to access the CRD site (Lopez et al., 2011; Yancey et al., 1996).





