Dual leucine zipper kinase-dependent PERK activation contributes to neuronal degeneration following insult
Figures
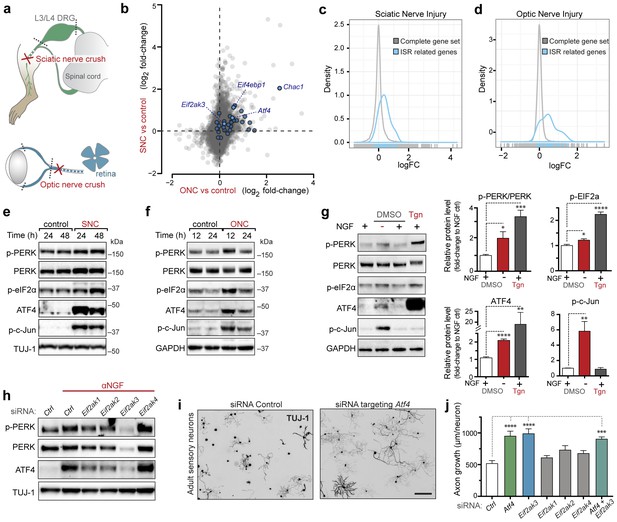
Acute neuronal insults activate the integrated stress response (ISR).
(a) Schematic of sciatic nerve crush (SNC) and optic nerve crush (ONC). Mouse lumbar level 3 and 4 dorsal root ganglia (DRG) and retina were isolated to assess the neuronal stress response after SNC and ONC, respectively. (b) Microarray cross-comparison of injury-regulated mRNAs following SNC or ONC (n = 5 per condition) identifies multiple ISR-associated genes (blue), including Eif4ebp1, Atf4, Chac1 and Eif2ak3 (PERK), upregulated by both insults. (c–d) mRNAs within the ‘ISR-related’ gene set (see Materials and Methods) are observed more frequently amongst upregulated mRNAs than expected by the overall distribution of mRNA expression changes assessed in each microarray study (‘complete gene set’) following SNC (p=2.4 × 10−5, (c)) or ONC (p=9.9 × 10−7, (d)), suggesting the selective activation of the ISR. (e–f) Immunoblots reveal upregulation of the ISR (p-PERK, p-eIF2α, and ATF4), in addition to the JNK-pathway (p-c-Jun) in L3/L4 DRG lysates after SNC (e), and in retina lysates after ONC (f). The time post-injury is indicated in hours. (g) Primary e12.5 mouse DRG cultures deprived of NGF (3 h) or treated with the ER stress inducer thapsigargin (Tgn) in the vehicle DMSO. NGF deprivation engages phosphorylation of PERK (p-PERK/PERK), p-eIF2α and ATF4. Protein levels were normalized to GAPDH and non-NGF deprived vehicle control (n = 6–7/condition, four independent experiments). (h) siRNA targeting each of the four eIF2α kinases differentially impacts ISR activation at 3 h after NGF withdrawal from embryonic DRG neuronal cultures, with only siRNA targeting Eif2ak3 consistently reducing ATF4 protein levels. (i) Representative TUJ-1 immunostainings 42 hr after isolation and siRNA-transfection of adult DRG neurons. (j) OnTarget Plus siRNA-mediated knockdown of Atf4 or Eik2ak3, but not other eIF2α kinases, enhances adult sensory axon regrowth in vitro (n ≥ 8 wells/condition). Molecular weight indicated in kilodaltons (kDa). Data are represented as mean ± SEM. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001, one-way ANOVA with post-hoc Bonferroni test.
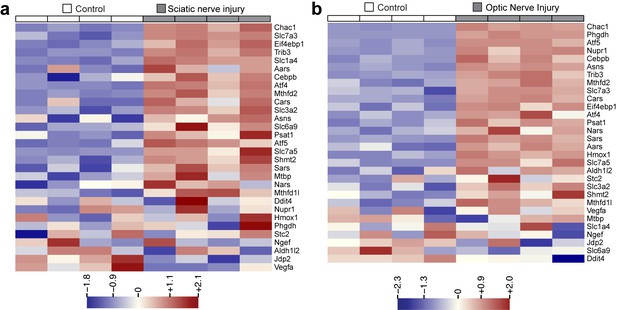
Heat maps showing expression of 30 ISR-related mRNAs that are induced by neuronal oxidative stress in an ATF4-dependent manner (Lange et al., 2008) in L4 DRG after sciatic nerve transection (a) or retina after optic nerve crush (b, data from Watkins et al., 2013) (n = 4 per condition).
https://doi.org/10.7554/eLife.20725.003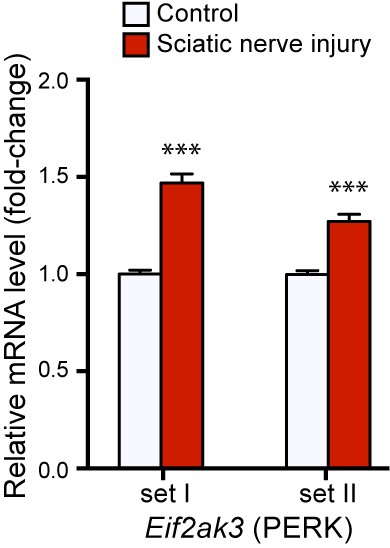
qRT-PCR validation of Perk mRNA upregulation in L4 DRG 24 h after SNC using two different primer sets (n = 4 per condition).
https://doi.org/10.7554/eLife.20725.004
Targeted silencing of ATF4 by OnTarget Plus siRNA pool verified by immunoblotting.
https://doi.org/10.7554/eLife.20725.005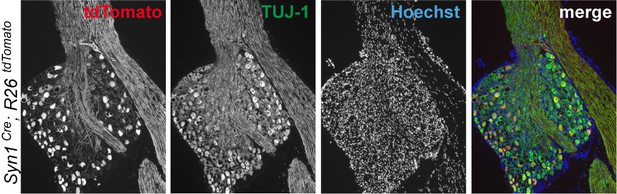
Immunohistochemistry of DRG cryosections from Syn1-Cre;R26LSL.tdTomato mice exhibits labeling of axons from medium- and large-diameter TUJ1-positive sensory neurons.
https://doi.org/10.7554/eLife.20725.006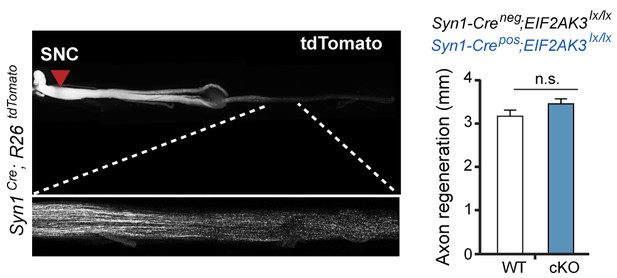
Assessment of PERK influence on axon regeneration in vivo after SNC.
(left) Identification of tdTomato-positive (Syn1-Cre;R26LSL.tdTomato) axon regeneration front by whole-mount sciatic nerve epifluorescence (top), confirmed by two-photon optical sectioning of a sciatic nerve (bottom) seven days after SNC (red arrow indicates injury site). (right) tdTomato-positive axon regeneration front is not significantly improved in Syn1-Cre;R26LSL.tdTomato;Eif2ak3lx/lx (PERK cKO) mice (n ≥ 5 per condition, p=0.12).
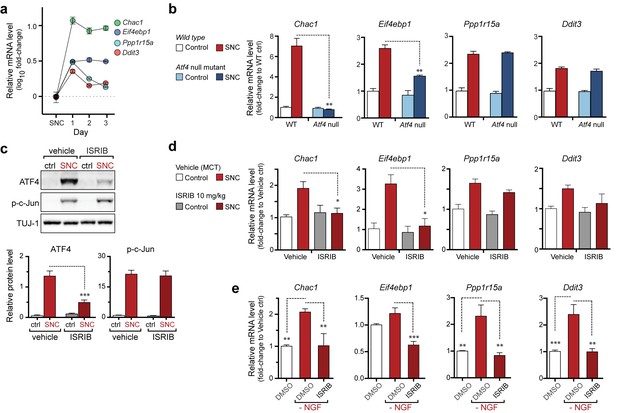
The PERK-mediated ISR regulates mRNA levels of a subset of ATF4 target genes.
(a) qRT-PCR of L4 DRG confirms the upregulation of each of four ISR-related mRNAs (Chac1, Eif4ebp1, Ppp1r15a, and Ddit3) over the first 3 days after SNC (n = 3/condition). (b) Real-time qPCR for four putative ATF4 target genes in ATF4-null and wild-type mouse DRGs 24 h following SNC (n = 3/condition). (c–d) Dosing with ISRIB reduces induction of ATF4 protein and the mRNA of a subset of its putative target genes in associated DRGs following SNC. (c) Immunoblots of L3/L4 DRGs from ISRIB-treated mice 16 h post-SNC (n = 3/condition). (d) Real-time qPCR of mouse DRGs 16 h following SNC (n = 3/condition). Mice were dosed with vehicle or ISRIB (10 mg/kg, 1 h pre-SNC and 12 h post-crush). (e) qPCR of NGF-deprived embryonic DRG cultures reveals upregulation of mRNA of four putative ATF4 target genes, blocked by 400 nM ISRIB (n = 6–8/condition, two independent experiments). Data are represented as mean ± SEM. One-way ANOVA with post-hoc Bonferroni was used for statistical comparisons. *p<0.05, **p<0.01, ***p<0.001.
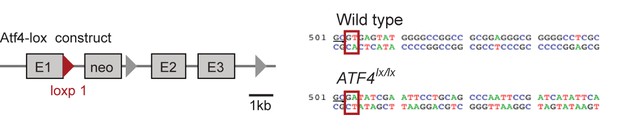
The structure of a targeted ATF4 allele (Yoshizawa et al., 2009), indicating the placement of a loxp-flanked neo cassette at the 3’ end of the exon 1 (E1).
Red triangle represents the first loxp site. Sequences comparing the wildtype allele at the end of exon 1 and the loxp allele (underlined GC bases) show the disruption of the splice site (red boxes) at the beginning of the first intron upon insertion of the neo cassette.
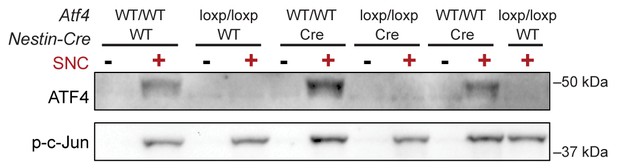
Immunoblots indicate a lack of ATF4 protein induction in L3/L4 DRG 24 hr post-SNC in ATF4loxp/loxp mice, irrespective of the presence of Cre recombinase.
https://doi.org/10.7554/eLife.20725.010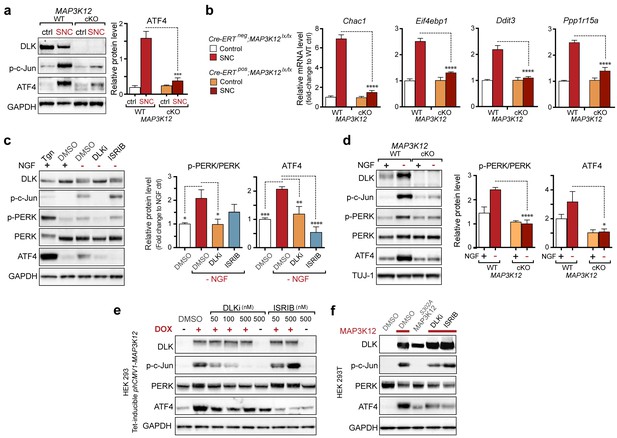
DLK is necessary and sufficient for PERK activation.
(a–b) DLK-deficient (CAG-ERTpos;Map3k12lx/lx, DLK cKO) mice exhibit reduced activation of the ISR and JNK pathways following SNC compared to controls (CAG-ERTneg;Map3k12lx/lx). (a) Activation of ATF4 is reduced in DLK cKO L3/L4 DRGs 24 h after SNC (n = 3/condition). (b) qRT-PCR of ISR-associated genes in L4 DRG following SNC in DLK cKO mice compared to controls (n = 4/condition). (c) Inhibition of DLK (DLKi, 500 nM) reduces p-PERK and ATF4 induction, in addition to p-c-Jun, compared to DMSO vehicle controls following NGF-withdrawal in primary DRG cultures. ISRIB (500 nM) treatment specifically reduces ATF4 upregulation. (n = 6/condition, aggregated data from four independent experiments). Thapsigargin (Tgn, 100 nM) provides a positive control for PERK activation. (d) Primary DRG cultures deficient in DLK (DLK KO) lack activation of PERK and ATF4 following NGF deprivation (n = 4/condition). (e) Doxycycline (DOX)-inducible DLK expression in stably-transfected HEK293 cells activates PERK (assessed by molecular weight shift, as anti-p-PERK does not recognize human p-PERK) and ATF4, which can be blocked by DLKi or ISRIB treatment. (f) Transient transfection of mouse wildtype DLK, but not kinase dead DLK-S302A control, in HEK293T cells activates the ISR. One-way ANOVA with post-hoc Bonferroni was used for statistical comparisons. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001.
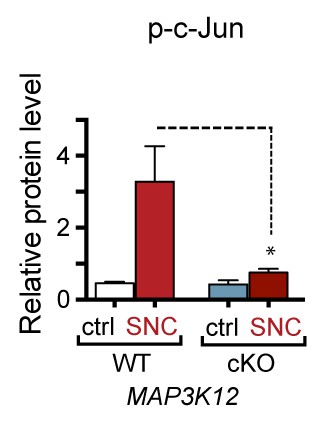
Quantification of p-c-Jun immunoblots presented in Figure 3a (DLK-deficient L3/L4 DRGs 24 hr after SNC, n = 3/condition).
https://doi.org/10.7554/eLife.20725.012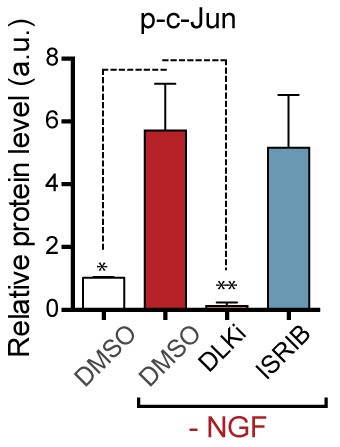
Quantification of p-c-Jun immunoblots presented in Figure 3c (NGF-withdrawal in primary DRG cultures, n = 6/condition, aggregated data from four independent experiments).
https://doi.org/10.7554/eLife.20725.013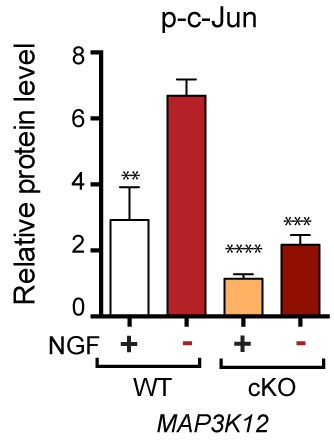
Quantification of p-c-Jun immunoblots presented in Figure 3d (NGF-withdrawal in primary DRG cultures, n = 4/condition).
https://doi.org/10.7554/eLife.20725.014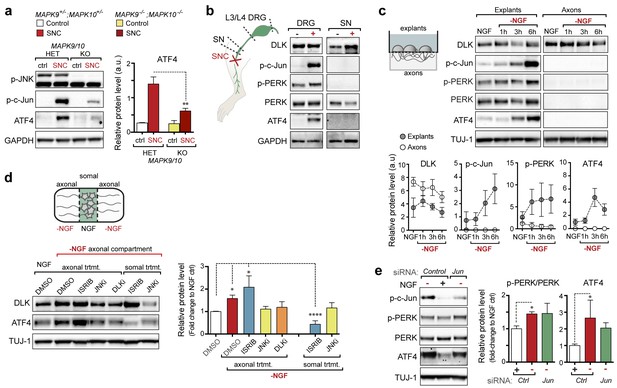
PERK activation is dependent on JNK-mediated retrograde signaling but not c-Jun.
(a) Immunoblot of JNK2/3-deficient (Mapk9−/−Mapk10−/−, Mapk9/10 KO) L3/L4 DRGs 24 h after SNC demonstrates reduced ATF4 induction compared to double heterozygous controls (Mapk9+/− Mapk10+/−, Mapk9/10 HET) (n = 3/condition). (b) Immunoblots of L3/L4 DRGs and 5-mm sciatic nerve (SN) segments 2–3 mm proximal to the injury site at 20 h post-SNC (+) illustrate elevation of DLK within the nerve and associated DRGs compared to uninjured DRGs and SN. PERK signaling is not detected within the SN after injury, in contrast to the associated L3/L4 DRGs. (c) Illustration of experimental setup for the isolation of DRG axons from explants (left, see methods). Immunoblots of DRG explants and enriched axons following NGF deprivation at the indicated time points. Phosphorylation of PERK and induction of ATF4 and p-c-Jun is observed in explant (grey), but not axonal (white), lysates quantified by immunoblotting (n = 5/condition, two experiments). (d) Schematic of compartmentalized Campenot chamber setup (top, see methods) displaying isolated axons in the outer compartment and cell bodies/proximal axons in the central compartment. Immunoblots of central compartment lysates after axonal NGF deprivation display elevated ATF4 levels. JNKi and DLKi prevent ATF4 induction when applied to either the central somal compartment or outer axonal compartments, whereas ISRIB only reduces ATF4 upregulation when applied to the central compartment (n = 6–9/condition, five independent experiments). (e) Immunoblots of siRNA-transfected embryonic DRG neurons suggests that c-Jun does not contribute to PERK activation following NGF withdrawal (n = 4–5/condition, three experiments). One-way ANOVA with Bonferroni’s post-hoc test was used for statistical analysis. Data are mean ± SEM. *p<0.05, **p<0.01, ****p<0.0001.
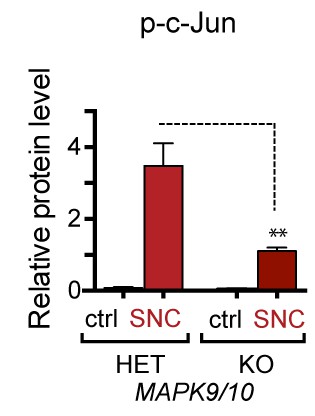
Quantification of p-c-Jun immunoblots presented in Figure 4a (JNK2/3-deficient L3/L4 DRGs 24 hr after SNC compared to HET controls, n = 3/condition).
https://doi.org/10.7554/eLife.20725.016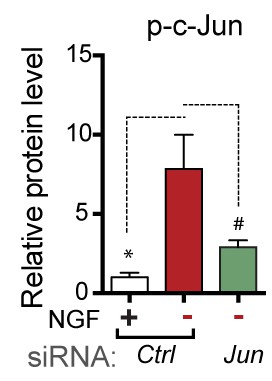
Quantification of p-c-Jun immunoblots presented in Figure 4e (NGF-withdrawal in siRNA-transfected primary DRG cultures, n = 4–5/condition, three experiments).
# corresponds to p=0.19.
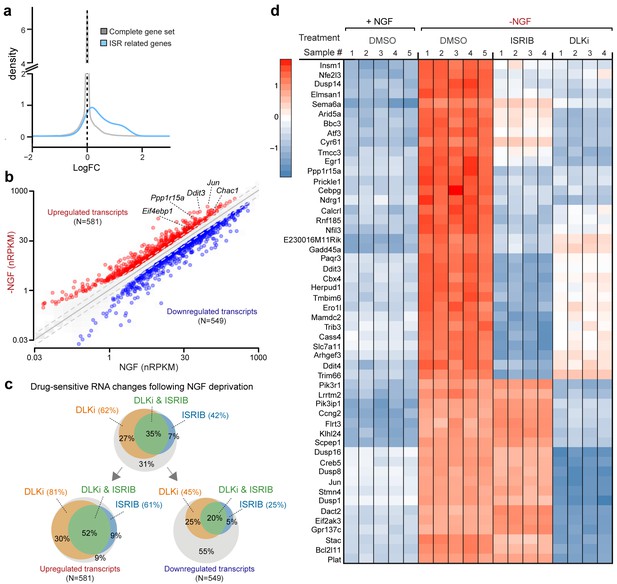
RNA-seq reveals ISRIB- and DLK inhibitor-sensitive expression changes following NGF deprivation.
(a–b) Global expression analysis indicates an enrichment in ISR-related genes upregulated 4.5 h after NGF withdrawal from embryonic DRG cultures in the presence of DMSO vehicle (n = 5/condition). (a) Density plot showing ‘ISR-related’ genes (blue, see Materials and methods) are more frequently upregulated compared to the distribution of all mRNAs expression changes (‘complete gene set’) (p<1 × 10−5, one-tailed Student-t test). (b) Scatterplot of gene expression levels (nRPKM) in NGF-containing and NGF-deprived samples. Transcripts of ATF4 target genes Chac1, Eif4ebp1, Ddit3, and Ppp1r15a are among 581 robustly upregulated (red) RNAs following NGF deprivation (n = 5 per condition, >1.5 fold, adjusted-p<0.001, nRPKM = Reads Per Kilobase per Million mapped reads) (c) Venn diagrams reveal the portions of these NGF-responsive genes that exhibit DLKi (GNE-3511, 400 nM) and/or ISRIB (400 nM) sensitivity (see Materials and methods). (d) ISRIB-sensitive and ISRIB-insensitive RNAs are among those represented in a heat map of top 50 transcripts strongly upregulated in a DLKi-sensitive manner following NGF withdrawal.
-
Figure 5—source data 1
RNA-seq analysis of primary sensory neurons following NGF deprivation, in the presence of ISRIB or DLK inhibitor GNE-3511.
- https://doi.org/10.7554/eLife.20725.019
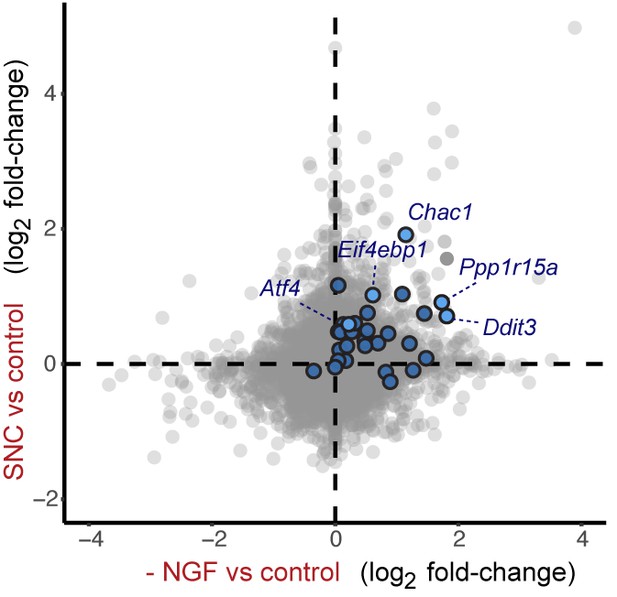
Cross-platform analysis of stress-regulated mRNAs following SNC or NGF withdrawal identifies multiple ISR-associated genes (blue), including Eif4ebp1, Ppp1r15a, Ddit3, and Chac1, upregulated by both insults.
https://doi.org/10.7554/eLife.20725.020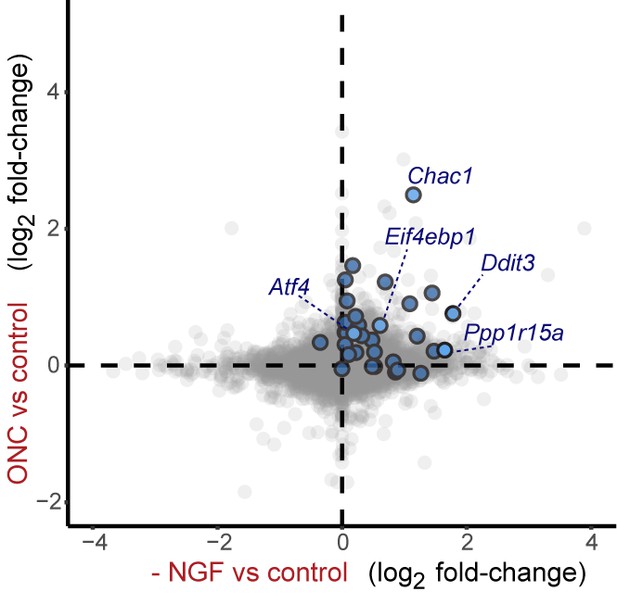
Cross-platform analysis of stress-regulated mRNAs following ONC or NGF withdrawal identifies multiple ISR-associated genes (blue), including Eif4ebp1, Ppp1r15a, Ddit3 , and Chac1, upregulated by both insults.
https://doi.org/10.7554/eLife.20725.021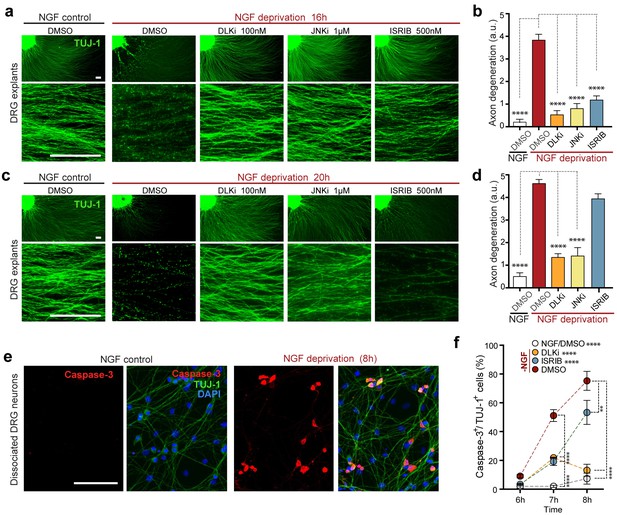
Inhibition of the ISR delays neurodegeneration following NGF withdrawal.
(a–b) NGF-deprived e12.5 mouse DRG explants exhibit reduced axon degeneration (TUJ-1, green) after 16 h NGF withdrawal in the presence of ISR or JNK pathway inhibitors [ISRIB (500 nM), DLKi (GNE-3511, 100 nM) and JNKi (AS601245, 1 µM)] compared to vehicle (DMSO). (n = 8–18 explants/condition, three independent experiments). (c–d) Following 20 h of NGF deprivation, the protective effect with ISRIB is lost, in contrast to JNK pathway inhibitors. (n = 9–13 explants, two independent experiments). Blinded scoring of axon degeneration from 0–5. (e) Representative images of cleaved Caspase-3 (red) immunolabeling following NGF deprivation (8 h) in dissociated DRG cultures (TUJ-1, green), indicative of apoptosis. (f) ISRIB treatment (blue dots) reduces the proportion of DRG neurons exhibiting activated Caspase-3 immunolabeling 6–8 h following NGF withdrawal compared to vehicle control (DMSO, red dots), though not to the same extent as DLKi (orange dots) (n = 4–8 samples/condition). Scale bars = 25 µm. Data are mean ± SEM. One-way ANOVA with Bonferroni’s post-hoc test was used for statistical analysis. In f, statistical significance over time for each treatment compared to –NGF/DMSO, is presented next to the condition legend and statistical significance for individual time points presented in the graph. **p<0.01, ****p<0.0001.
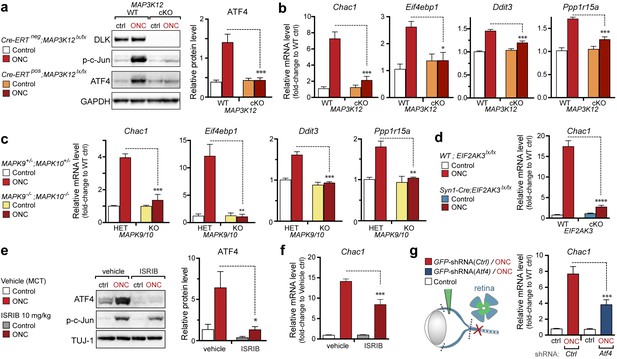
DLK/PERK signaling regulates ATF4 target gene Chac1 mRNA following optic nerve crush.
(a–b) DLK-deficient (CAG-ERTpos;Map3k12lx/lx, DLK cKO) mice exhibit reduced activation of the ISR and JNK pathway following ONC compared to controls (CAG-ERTneg;Map3k12lx/lx). (a) DLK-deficient retina lysates 3 d after ONC display no induction of ATF4 protein by immunoblot (n = 4/condition). (b) qRT-PCR of ISR-associated genes in retina following ONC in DLK cKO mice compared to Cre-negative controls (n = 5–6/ condition). (c) qRT-PCR of ISR-associated genes in retina following ONC (red) in JNK2/3-deficient mice (n = 5–6/condition). (d) Induction of Chac1 mRNA is suppressed in PERK cKO retinas (Syn1-Cre;Eif2ak3lx/lx mice) compared to Cre-negative littermates following ONC (n = 6). (e) Dosing with ISRIB reduces upregulation of ATF4 protein in mouse retinal lysates 16 h post-ONC (immunoblot and quantification, n = 4/condition, 2.5 mg/kg ISRIB b.i.d.). (f) Chac1 mRNA (qPCR, n = 4/condition) in mouse retina 48 h post-ONC (10 mg/kg ISRIB b.i.d.). (g) Intravitreal injection of an AAV2 vector directing expression of GFP and an shRNA targeting mouse Atf4, GFP-shRNA(Atf4), reduces Chac1 mRNA induction following ONC in transduced regions of the retina compared to a similar AAV2 vector with a non-targeting shRNA sequence, GFP-shRNA(Ctrl) (see Materials and methods, qPCR, n = 8/condition). One-way ANOVA with Bonferroni’s post-hoc test was used for statistical analysis. Data are mean ± SEM. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001.
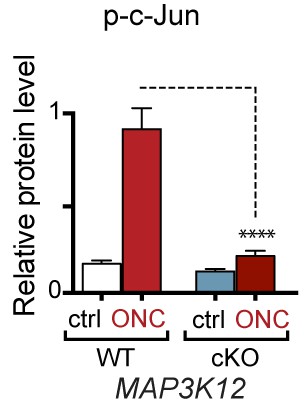
Quantification of p-c-Jun immunoblots presented in Figure 7a (DLK-deficient retina lysates three days after ONC, n = 4/condition).
https://doi.org/10.7554/eLife.20725.024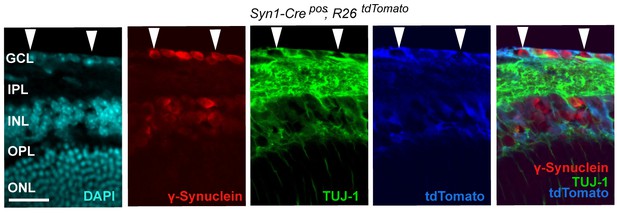
Syn1-Cre activity in the retina, illustrated by Syn1-Cre;R26LSL.tdTomato (blue) expression co-localizing with TUJ-1 (green) and γ-synuclein (red) immunolabeled RGCs in the GCL.
DAPI labeled nuclei (cyan). GCL = ganglion cell layer.
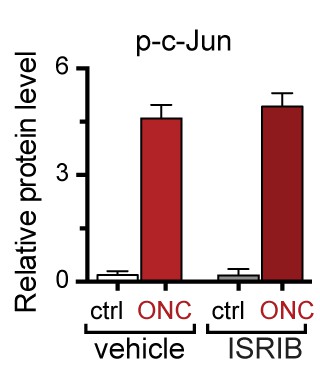
Quantification of p-c-Jun immunoblots presented in Figure 7e (ONC in mice dosed with ISRIB).
https://doi.org/10.7554/eLife.20725.026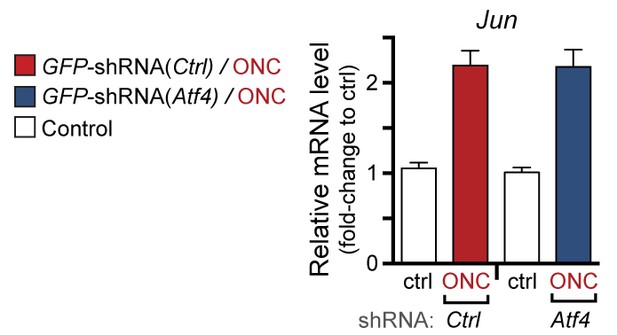
qPCR of Jun mRNA following ONC in AAV2-shRNA-GFP-injected mice (corresponding to Figure 7g).
https://doi.org/10.7554/eLife.20725.027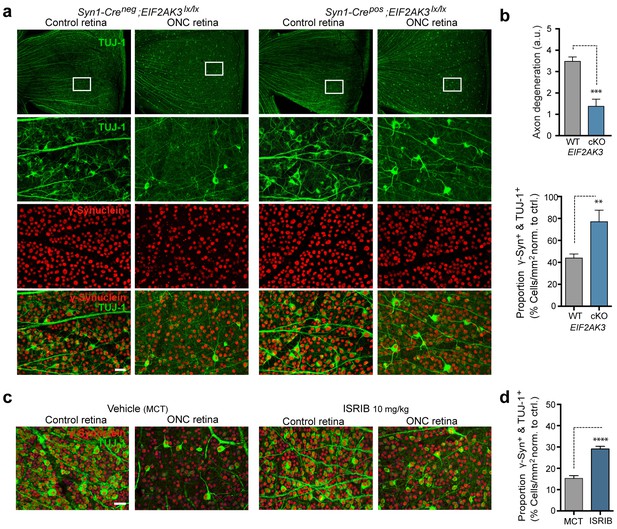
PERK contributes to neurodegeneration following nerve injury.
(a–b) Reduced loss of PERK-deficient RGCs 14 d after ONC. (a) TUJ-1 (green)/ γ-synuclein (red) immunolabeled whole mount retinas from control Syn1-Creneg;Eif2ak3lx/lx (PERK WT) and neuronal PERK-deficient Syn1-Crepos;Eif2ak3lx/lx (PERK cKO) mice 14 d after ONC, high magnification (expanded from boxed region). (b) Blinded scoring of axon degeneration from 0–5 in whole-mounted retinas and automated image quantification of TUJ-1/γ-synuclein double-immunolabeled RGCs normalized to uncrushed retina of each mouse (n = 8 WT, n = 6 PERK cKO). (c–d) Reduced loss of RGCs 10 d post-ONC in ISRIB-treated mice (10 mg/kg ISRIB in MCT vehicle b.i.d) compared to vehicle control. (c) Automated image analysis of TUJ-1 (green, outlined in cyan)/ γ-synuclein (red, outlined in blue) immunolabeled cells. (d) Quantification of TUJ-1/γ-synuclein double-immunolabeled RGCs (outlined in green) in crushed retinas normalized to uncrushed control (n = 8). Scale bars = 25 µm. Student’s t-test was used for statistical analysis. Data are mean ± SEM. **p<0.01, ***p<0.001, ****p<0.0001.
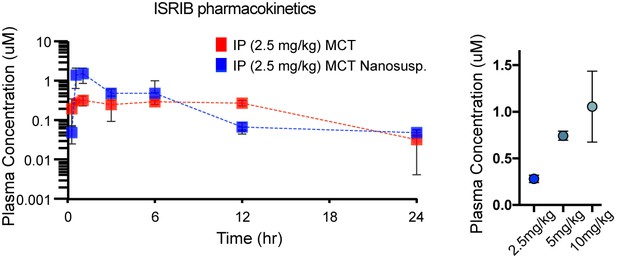
Pharmacokinetic analysis in mice dosed IP with ISRIB formations in MCT (red) and nanosuspenion in MCT (blue) over one day, as determined by plasma concentration (left).
Plasma concentrations measured in a separate cohort of mice 8 h post IP administration of ISRIB in MCT at 2.5, 5 and 10 mg/kg (right), suggesting the use of 10 mg/kg.
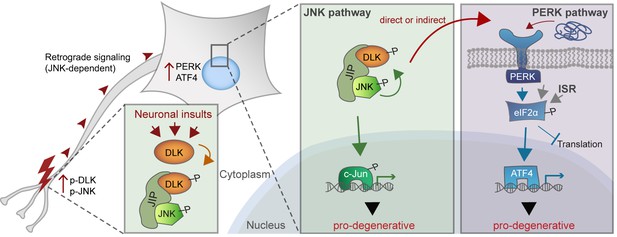
A working model of DLK as a master regulator of multiple neuronal stress response pathways.
DLK (orange) regulates apoptotic transcriptional changes through both JNK signaling (green) and the PERK-driven ISR (blue) (which can also be activated by misfolded proteins in the ER). One potential mechanism of cross-talk may be the activation of PERK by DLK activity (either directly or indirectly) following retrograde signaling mediated by a complex that includes DLK, JNK, and the JNK-interacting protein (JIP).





