The Ndc80 complex bridges two Dam1 complex rings
Figures
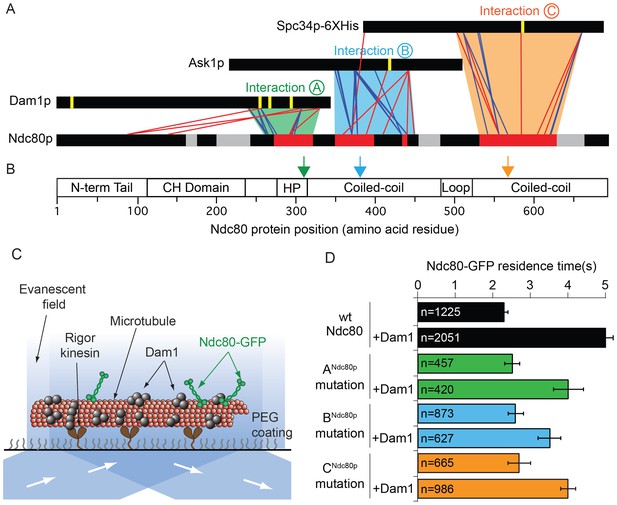
Dam1p, Ask1p, and Spc34p form cross-links to three distinct Ndc80p regions.
(A) Cross-links between the Dam1p, Ask1p, and Spc34p of the Dam1 complex and Ndc80p of the Ndc80 complex. Dam1 and Ndc80 complexes were cross-linked in the presence of microtubules. Horizontal black bars represent proteins and the six vertical yellow lines indicate Aurora B kinase phosphorylation sites on the Dam1 complex. Red and blue lines show cross-links formed with DSS and EDC cross-linkers, respectively. For clarity, only the cross-links between the Dam1 complex proteins and Ndc80p are displayed. Data are shown for peptides with Percolator (Käll et al., 2007) assigned q-values ≤ 0.05. Red bars on Ndc80p indicate regions where clusters of lethal mutations mapped (from Tien et al., 2013) to cross-linked regions. Grey bars on Ndc80p indicate clusters of lethal insertions outside of cross-linked regions. Green, blue, and orange trapezoids represent putative interactions (A, B, and C) between the Dam1 and Ndc80 complexes. (B) Bar diagram of Ndc80p with structural features. Green, blue and orange arrows indicate the positions of lethal mutations in interaction regions ANdc80p, BNdc80p, and CNdc80p used in this study. (CH: calponin homology; HP: hairpin) (C) Diagram showing the setup of TIRF microscopy experiments. Single molecule Ndc80-GFP complex binding on microtubules was visualized in the presence or absence of the Dam1 complex. (D) Lethal mutation in region ANdc80, BNdc80, or CNdc80 partially disrupts the Ndc80 complex’s interaction with Dam1 complex. Average residence time of Ndc80-GFP mutant and wild-type complexes on microtubules in the presence or absence of Dam1 complex. Bars represent average residence time ± error of the mean (estimated by bootstrapping analysis; see Materials and methods for additional details). Ndc80-GFP complex microtubule residence time raw data are included in Figure 1—source data 1. Refer to Supplementary file 1A for statistical analysis of data in part (D).
-
Figure 1—source data 1
Table of Ndc80-GFP microtubule residence times for Figure 1D.
- https://doi.org/10.7554/eLife.21069.004
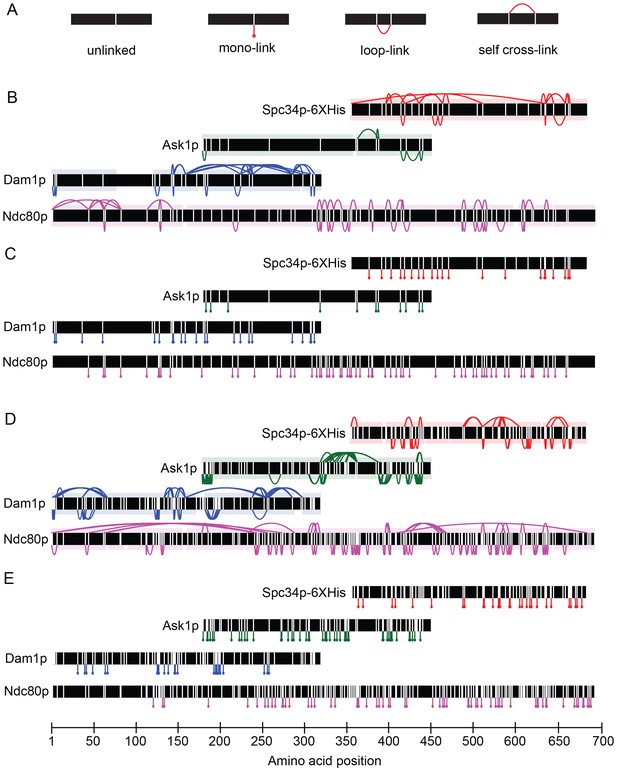

Dam1 and Ndc80 complexes robustly react with DSS and EDC cross-linking agents.
(A) Four types of peptides identified by mass spectrometry after cross-linking and trypsin digestion. Horizontal black bars represent protein sequence and vertical white lines represent residues able to react with the cross-linking agent. Unlinked: peptide that did not react with cross-linking agent; mono-link: only one end of the cross-linking reagent reacted with a residue; loop-link: intra-protein cross-link without a tryptic site in between the two cross-linked residues; self cross-link: intra-protein cross-link with a tryptic site in between the two-cross-linked residues. Dam1 and Ndc80 complexes were cross-linked in the presence of microtubules using DSS (panels B and C) or EDC (panels D and E). (B, D) Cross-links above and below the black bars represent self-crosslinks and loop-links, respectively. Data are shown for peptides with Percolator (Käll et al., 2007) assigned q-values ≤ 0.05. Highlighted regions above and below black bars represent sequence coverage from mass spectrometry. (C, E) Lines below the black bars represent mono-links.
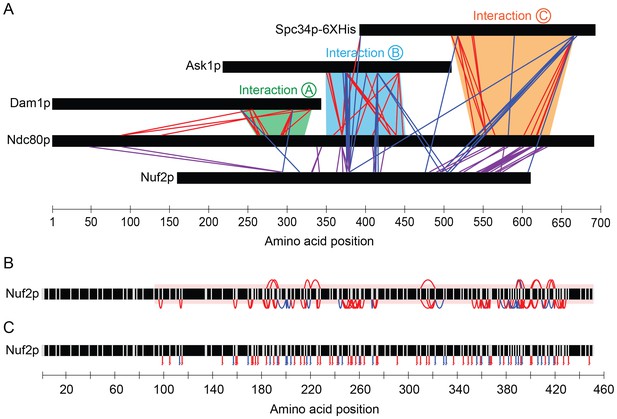
Dam1p, Ask1p, and Spc34p cross-link to Nuf2p, in agreement to the coiled-coil structure of Ndc80p and Nuf2p.
(A) Cross-links between the Dam1p, Ask1p, and Spc34p of the Dam1 complex and Ndc80p and Nuf2p of the Ndc80 complex. For clarity, cross-links between the Dam1 complex proteins are not shown. Red lines represent cross-links between Ndc80p and Dam1p, Ask1p, or Spc34p. Blue lines represent cross-links between Nuf2p and Dam1p, Ask1p, or Spc34p. Purple lines represent cross-links between Ndc80p and Nuf2p. (B) Cross-links above and below the black bar represent self-crosslinks and loop-links, respectively, for Nuf2p. (C) Lines below the black bar represent mono-links for Nuf2p. Data from cross-linking experiments using DSS and EDC cross-linking agents are both shown. Data are shown for peptides with Percolator (Käll et al., 2007) assigned q-values ≤ 0.05.
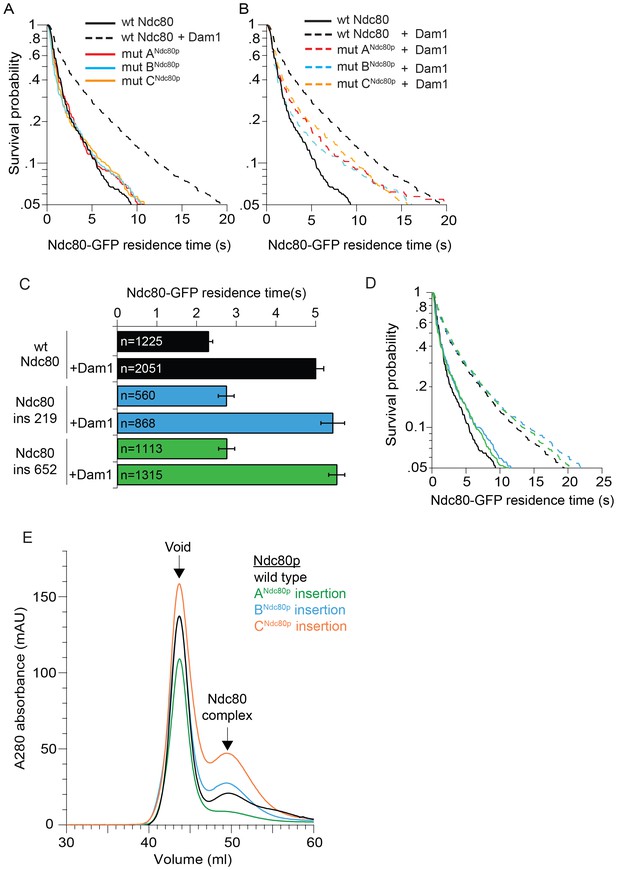
Ndc80 mutations outside of regions A, B, and C do not disrupt the interaction between Dam1 and Ndc80 complexes.
(A), (B) Survival probability of the data shown in Figure 1D: microtubule residence times of Ndc80-GFP complex with a lethal mutation in region ANdc80p, BNdc80p, or CNdc80p in the absence (A) and presence (B) of wild-type Dam1 complex. Data here have a range of n = 420 to 2051. (C) Average microtubule residence times of Ndc80-GFP complexes with wild-type Ndc80p, or lethal five amino acid insertional mutation at position 219 or 652. Bars represent average residence time ± error of the mean (estimated by bootstrapping analysis; see Materials and methods for additional details). Ndc80-GFP complex microtubule residence time raw data are included in Figure 1—figure supplement 3—source data 1. Refer to Supplementary file 1B for statistical analysis. (D) Survival probability curves of the data shown in (C). (E) Superdex 200 size exclusion chromatography of the Ndc80 complex with wild-type Ndc80p or insertional mutation in ANdc80p, BNdc80p, or CNdc80p.
-
Figure 1—figure supplement 3—source data 1
Table of Ndc80-GFP microtubule residence times for Figure 1—figure supplement 3C.
- https://doi.org/10.7554/eLife.21069.008
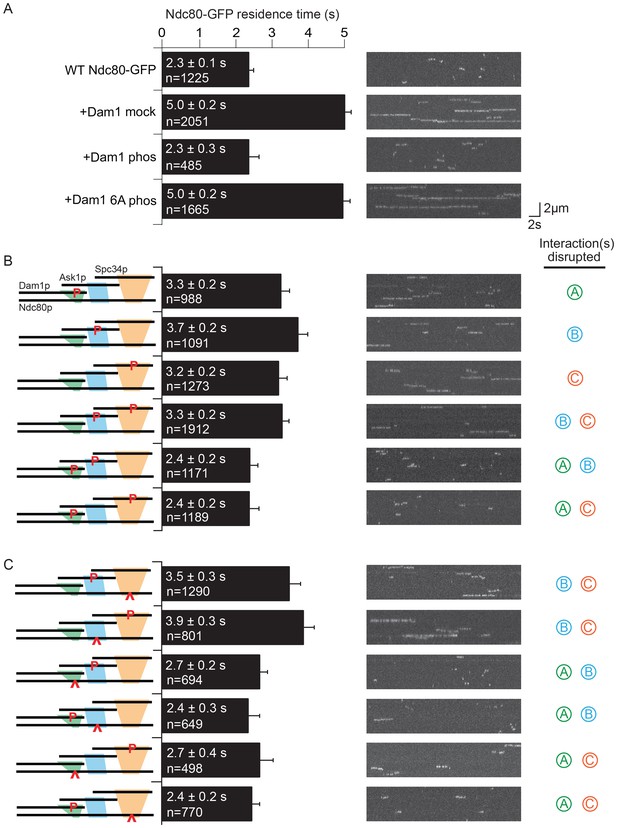
Dam1 and Ndc80 complexes interact through three distinct sites.
(A) Average microtubule residence time of the wild-type Ndc80-GFP complex alone, in the presence of mock treated, phosphorylated Dam1 complex, or phosphorylated Dam1 6A mutant complex. Dam1 6A: all six Dam1 complex phosphorylation sites mutated to Ala. (B) Average microtubule residence time of wild-type Ndc80-GFP complex in the presence of selectively phosphorylated Dam1 complex. Diagram on the left indicates phosphorylated proteins. (C) Average microtubule residence time of Ndc80-GFP complex with lethal mutation in region ANdc80p, BNdc80p, or CNdc80p in the presence of selectively phosphorylated Dam1 complex. Diagrams on the left indicate phosphorylated protein (P) and Ndc80p region with a lethal mutation (∧). On the right are representative kymographs for each experiment. Bars represent average residence time ± error of the mean (estimated by bootstrapping analysis; see Materials and methods or additional details). Ndc80-GFP complex microtubule residence time raw data are included in Figure 2—source data 1. Refer to Supplementary file 1C for statistical analysis.
-
Figure 2—source data 1
Table of Ndc80-GFP microtubule residence times for Figure 2.
- https://doi.org/10.7554/eLife.21069.011
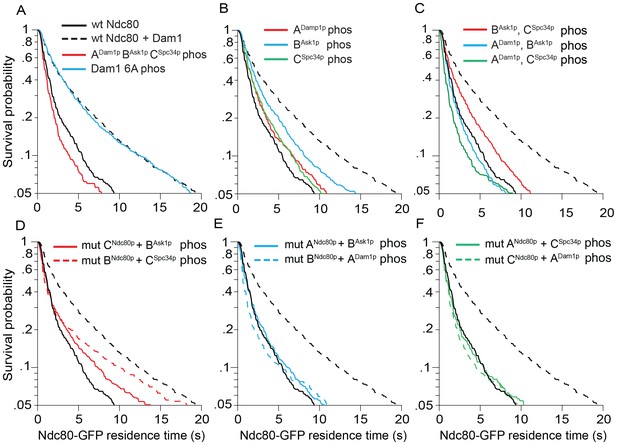
Microtubule residence time survival probability plots of experiments in Figure 2.
For all graphs, solid and dashed black lines represent curves for wild-type Ndc80 complex in the absence and presence of wild-type Dam1 complex, respectively. (A–C) Wild-type Ndc80-GFP complex microtubule residence time in the presence of variably phosphorylated Dam1 complex. Dam1 6A phos: all Dam1 complex phosphorylation sites mutated to Ala and treated with Aurora B kinase. (D–F) Microtubule residence time of Ndc80-GFP complex with a lethal mutation in region ANdc80p, BNdc80p, or CNdc80p in the presence of variably phosphorylated Dam1 complex. Data here have a range of n = 485 to 2051.
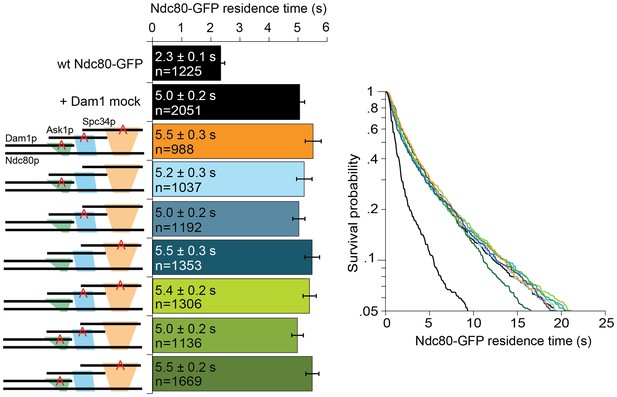
Mutating Dam1 complex Aurora B kinase phosphorylation sites to Ala does not affect the interaction between Dam1 and Ndc80 complexes.
Average microtubule residence time of wild-type Ndc80-GFP complex in the presence of mock treated phosphorylation-blocking Dam1 complex mutant constructs. A red ‘A’ identifies the phosphorylation sites that are mutated to alanine. Mock treatment was carried out by substituting ATP with H20. Survival probability plots of the average microtubule residence time shown on the right. Ndc80-GFP complex microtubule residence time raw data are included in Figure 2—figure supplement 2—source data 1. Refer to Supplementary file 1D for statistical analysis.
-
Figure 2—figure supplement 2—source data 1
Table of Ndc80-GFP microtubule residence times for Figure 2—figure supplement 2.
- https://doi.org/10.7554/eLife.21069.014
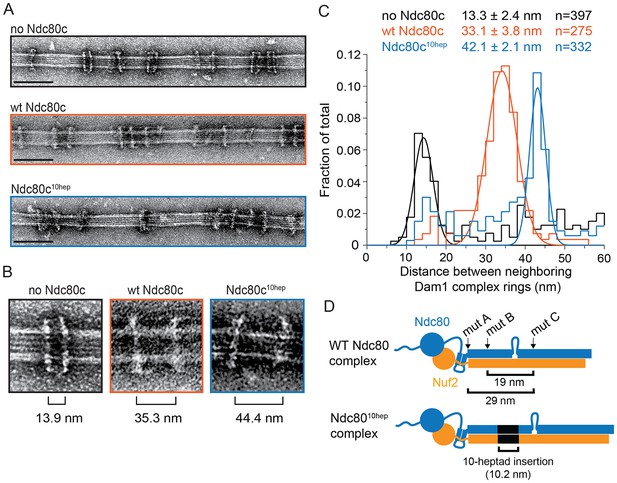
The Ndc80 complex binds two Dam1 complex rings.
(A) Representative EM images of Dam1 complex rings on microtubules in the absence or presence of the Ndc80 complex. Three experiments include no Ndc80 complex (Ndc80c), wild-type Ndc80c, and Ndc80c10hep. Protein concentrations were 20 nM tubulin and 25 nM Dam1 and Ndc80 complexes. Scale bars: 100 nm. (B) Zoomed in images of a Dam1 complex ring doublet for the three experiments. Distances from middle of one ring to that of next ring are indicated. (C) Distribution of closest neighboring inter-ring distances measured for the three different conditions. Measurements made between 0–60 nm are shown. The cluster of distance measurements was fitted with a Gaussian distribution and the vertices ± standard deviations for each fit are listed. The raw data of Dam1 complex inter-ring distance measurements are included in Figure 3—source data 1. (D) Diagram demonstrating the predicted coiled-coil distances between the Ndc80p lethal mutations. Diagram below shows the location of additional 10-heptad insertion between the hypothesized binding sites of two Dam1 complex rings. Refer to Supplementary file 1E for statistical analysis of data in part (C).
-
Figure 3—source data 1
Table of Dam1 complex inter-ring distance measurements for Figure 3C.
- https://doi.org/10.7554/eLife.21069.016
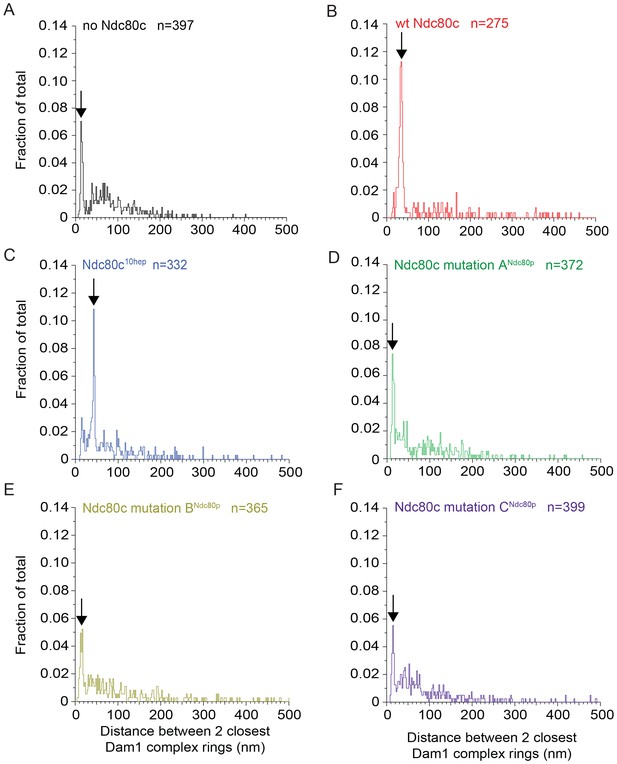
Full distributions of Dam1 complex inter-ring measurements for various experiments shown in Figure 3 and Figure 3—figure supplement 2.
Six conditions include Dam1 complex rings on microtubules in the absence of Ndc80 complex (Ndc80c) (A), presence of wild-type Ndc80c (B), Ndc80c10hep (C), Ndc80c lethal mutation ANdc80p (D), BNdc80p (E), and CNdc80p (F). The arrows identify the sharp peaks that contain at least 20% of the events.
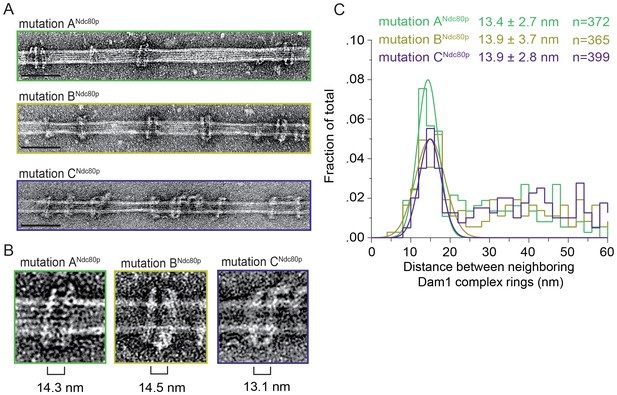
Mutations in regions ANdc80p, BNdc80p, or CNdc80p disrupts the Ndc80 complex’s ability to bind two Dam1 complex rings.
(A) Representative EM images of Dam1 complex rings on microtubules in the presence of the Ndc80 complex with a lethal mutation in region ANdc80p, BNdc80p, or CNdc80p. Scale bars: 100 nm. Protein concentrations were 20 nM tubulin and 25 nM Dam1 and Ndc80 complexes. (B) Zoomed in images of a Dam1 complex ring doublet for the three experiments. Distances from middle of one ring to those of the closest two rings were measured. (C) Distribution of closest inter-ring distances measured for the three experiments. Measurements made between 0–60 nm are shown. The cluster of distance measurements was fitted with a Gaussian distribution and the vertex ± standard deviation for each fit is listed. The raw data of Dam1 complex inter-ring distance measurements are included in Figure 3—figure supplement 2–source data 1. Refer to Supplementary file 1E for statistical analysis of data in part (C).
-
Figure 3—figure supplement 2—source data 1
Table of Dam1 complex inter-ring distance measurements for Figure 3—figure supplement 2C.
- https://doi.org/10.7554/eLife.21069.019
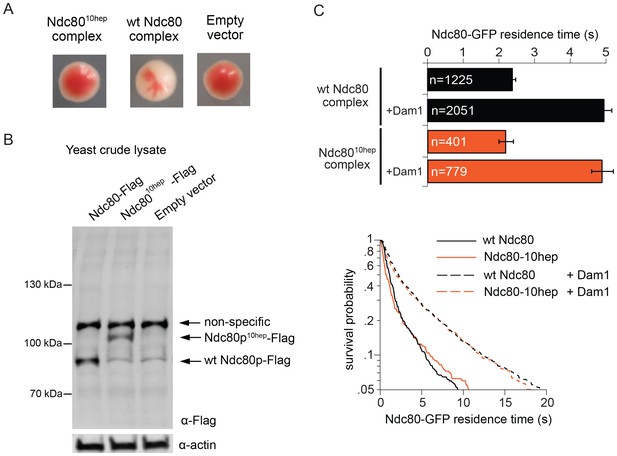
The Ndc8010hep complex does not support growth.
(A) Red/white plasmid shuffle assay to test the viability of the Ndc8010hep complex. Solid red colonies indicate the inability of the Ndc8010hep complex and empty vector to support growth. Sectoring white colony indicates the ability of the wild-type Ndc80 complex to support growth. (B) α-FLAG immunoblot of crude lysate of cells expressing either wild-type Ndc80p, Ndc80p10hep, or empty vector. α-actin immunoblot of the same lysates shows equal loading of the lysates. (C) Wild-type or Ndc8010hep complex average residence time (top) on microtubules in the absence and presence of Dam1 complex as derived from the survival probability curves (bottom). Bars represent average residence time ± error of the mean (estimated by bootstrapping analysis; see Materials and methods or additional details). Ndc80-GFP complex microtubule residence times raw data are included in Figure 4—source data 1. Refer to Supplementary file 1F for statistical analysis of data.
-
Figure 4—source data 1
Table of Ndc80-GFP microtubule residence times for Figure 4C.
- https://doi.org/10.7554/eLife.21069.021
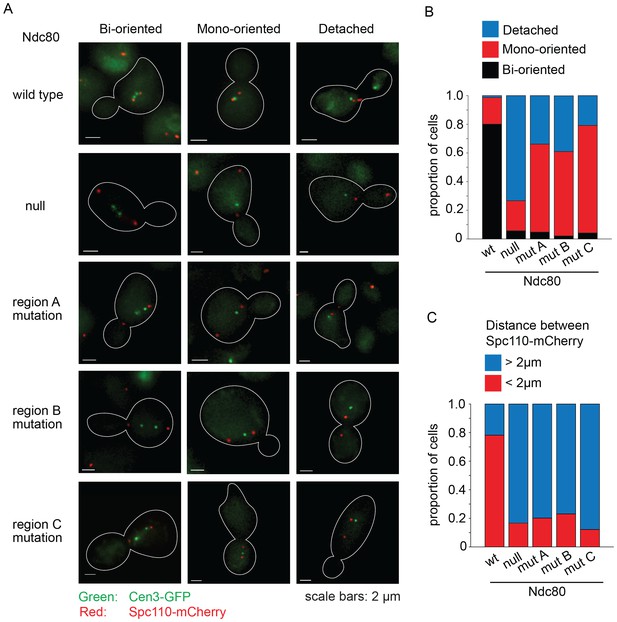
Lethal mutations in ANdc80p, BNdc80p, and CNdc80p have biorientation and microtubule attachment defects in vivo.
(A) Representative images from Ndc80-AID degron experiments with cells carrying an extra copy of NDC80 wild-type, no NDC80 (null), or a mutations in ANdc80p, BNdc80p, or CNdc80p. Cells were treated with 6 μM α-factor for two hours prior to treatment with 6 μM α-factor and 0.5 mM auxin for one additional hour. At time 0, cells were released from α-factor and incubated in YPD medium containing 0.5 mM auxin. Images were taken at 60 min after release from α-factor arrest. Representative images are shown. (B) Stacked bar graphs showing proportion of the cells containing bioriented, monooriented or detached CEN3-GFP. Spindles with two CEN3-GFP puncta positioned between the two SPBs were counted as bioriented. Spindles with one CEN3-GFP puncta positioned between the two SPBs were counted as monooriented and spindles with one (or rarely two) CEN3-GFP puncta positioned off the spindle axis were counted as detached. Only cells with two Spc110-mCherry puncta and at least one CEN3-GFP punctum were included in the analysis. (C) Stacked bar graph showing the distance between SPBs. Wild-type NDC80 n = 236 cells; no NDC80 n = 210 cells; ANdc80pn = 210 cells; BNdc80pn = 190 cells; CNdc80pn = 260 cells.
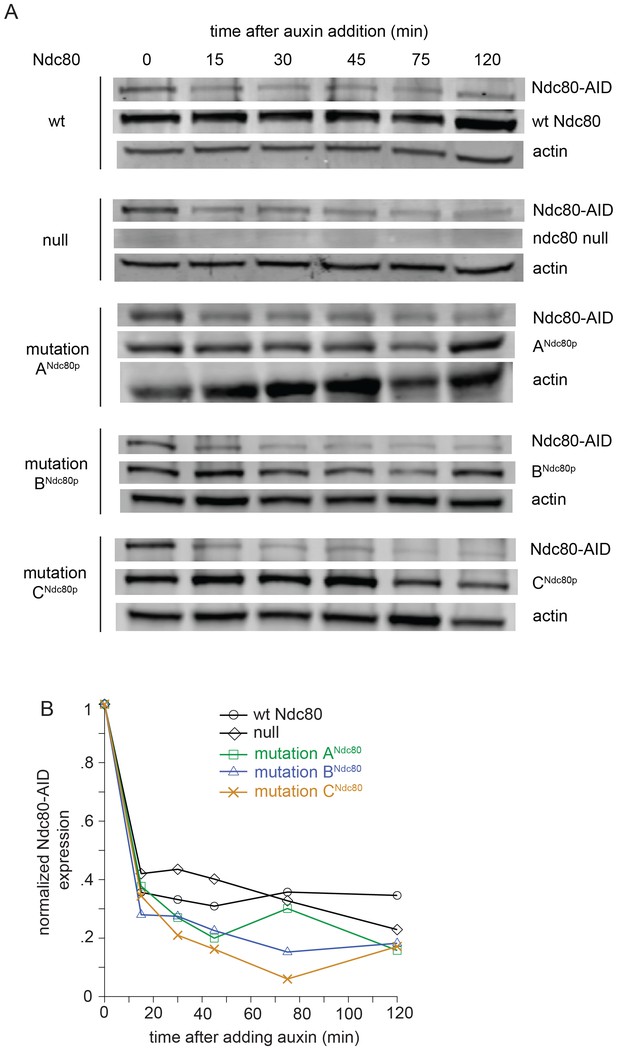
Wild-type Ndc80-AID is degraded upon the addition of auxin.
(A) Western blot images from Ndc80-AID degron experiments, blotting for Ndc80-AID and β-actin. Whole cell protein precipitation was carried out for Ndc80-AID cells carrying an extra copy of NDC80 wild-type, no NDC80 (null), lethal mutation in ANdc80p, BNdc80p, or CNdc80p. Samples were collected at various time points after adding auxin (Images were taken 120 min after auxin addition). (B) Normalized Ndc80-AID protein levels analyzed by western blot band intensities.
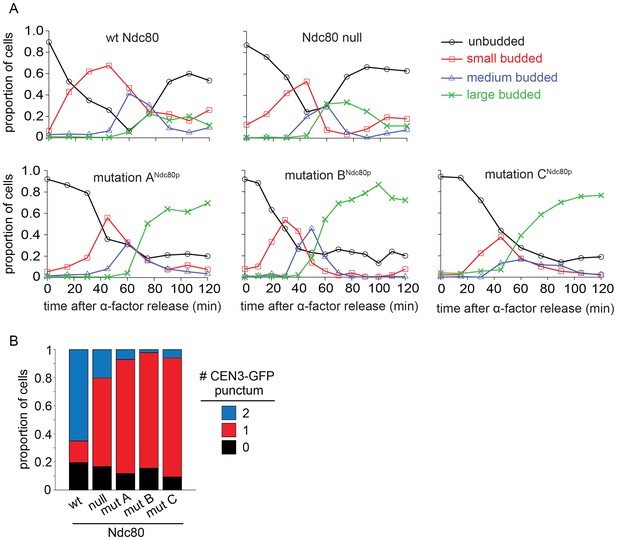
Lethal mutation in region ANdc80p, BNdc80p, or CNdc80p causes mitotic arrest.
(A) Budding index analyses of Ndc80-AID degron experiments. Ndc80-AID cells carrying an addition copy of NDC80 wild-type, no NDC80 (null), lethal mutation in region ANdc80p, BNdc80p, or CNdc80p were fixed at various time points after release from α-factor arrest (time 0). (B) Stacked bar graphs showing the percentage of cells with zero, one, or two CEN3-GFP spots for the various cell types.
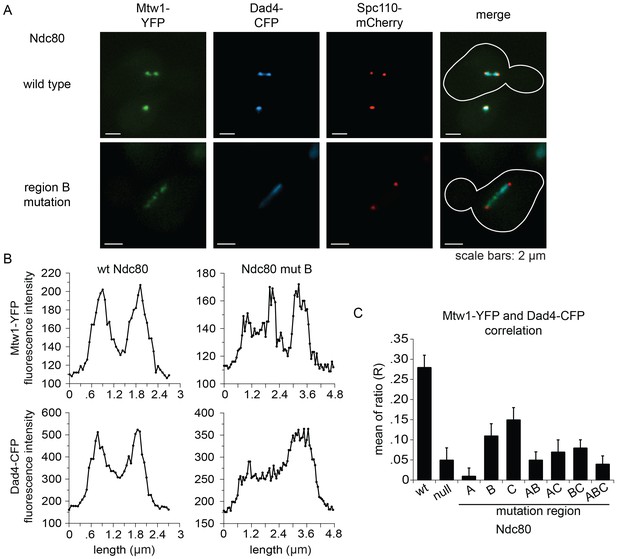
Lethal mutations in ANdc80p, BNdc80p, and CNdc80p confer defects in Dam1 complex recruitment to the kinetochore.
(A) Representative images from NDC80-AID degron experiments with cells also containing wild-type NDC80 or mutation in region BNdc80p. The inner kinetochore was visualized with Mtw1-YFP, the Dam1 complex with Dad4-CFP, and the SPB with Spc110-mCherry. Cells were treated with 6 μM α-factor for two hours prior to the addition of 0.5 mM auxin for one additional hour. At time 0, cells were released from α-factor and incubated in YPD medium containing 0.5 mM auxin. Images were taken at 60 min after release from α-factor arrest. (B) Line scan traces of Mtw1-YFP and Dad4-CFP from the images in (A). Only cells with two SPBs in the same plane of focus were selected for analysis. Each line scan extended from one SPB to the other. Each point represents one pixel. (C) Summary of correlation analysis carried out between Mtw1-YFP and Dad4-CFP. For each line scan, a correlation of the positive or negative changes of pixel intensity along the line between the Mtw1-YFP and Dad4-CFP channels was calculated to examine co-localization along the spindle (see Materials and methods). Between 18 and 33 cells were analyzed for each mutant. Mtw1-YFP and Dad4-CFP fluorescence intensity raw data are included in Figure 6—source data 1. Refer to Supplementary file 1G for statistical analysis of data.
-
Figure 6—source data 1
Tables of Mtw1-YFP and Dad4-CFP fluorescence intensities for cells analyzed in Figure 6C.
- https://doi.org/10.7554/eLife.21069.026
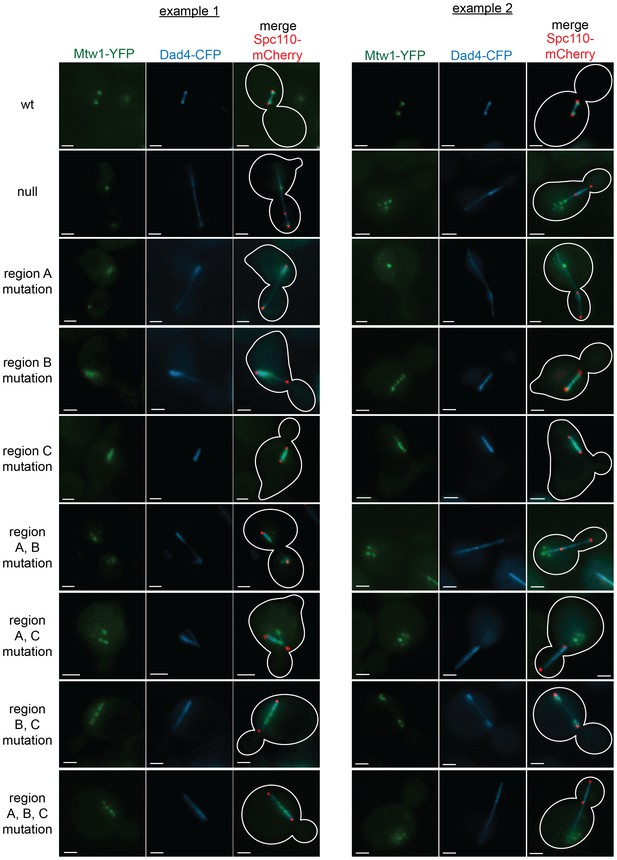
Mutations in regions ANdc80p, BNdc80p, or CNdc80p results in severe defects in kinetochore biorientation and attachment.
Examples of commonly observed phenotypes in experiments with NDC80-AID degron cells carrying an additional copy of ndc80 containing single or multiple mutations in regions ANdc80p, BNdc80p, and CNdc80p, or not carrying a second copy of NDC80 (null). Each example shows Mtw1-YFP, Dad4-CFP, and Spc110-mCherry and the same experiment procedure was carried out as in Figure 6. The majority of mutant cells with two SPBs exhibited detached kinetochores or had kinetochores spread along spindle microtubules. All scale bars: 2 µm.
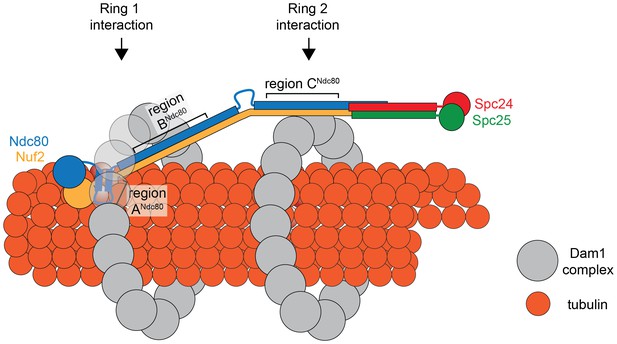
Model of the Ndc80 complex bridging two Dam1 complex rings, separated by 33 nm.
Results in this study support ring 1 interaction being comprised of interaction A and B, and ring 2 interaction being comprised of interaction C (refer to Figure 1). This image depicts only one Ndc80 complex across two Dam1 complex rings, but multiple Ndc80 complexes would be present in vivo.
Tables
The interacting regions in the Dam1 and Ndc80 complexes.
| Interaction | Protein region | Amino acids | Phosphorylated residues | Five amino acid mutation (insertion) position |
|---|---|---|---|---|
| A | ADam1p | 241–330 | S257, S265, S292 | n/a |
| ANdc80p | 262–322 | n/a | 314 | |
| B | BAsk1p | 133–225 | S200 | n/a |
| BNdc80p | 350–448 | n/a | 383 | |
| C | CSpc34p | 118–274 | T199 | n/a |
| CNdc80p | 532–630 | n/a | 563 |
Additional files
-
Supplementary file 1
Tables of pairwise statistical comparisons.
- https://doi.org/10.7554/eLife.21069.029
-
Supplementary file 2
Table of yeast strains and plasmids used in this study.
- https://doi.org/10.7554/eLife.21069.030
-
Source code 1
LabVIEW TIRF analysis step 1.
- https://doi.org/10.7554/eLife.21069.031
-
Source code 2
LabVIEW TIRF analysis step 2.
- https://doi.org/10.7554/eLife.21069.032
-
Source code 3
LabVIEW TIRF analysis step 3.
- https://doi.org/10.7554/eLife.21069.033
-
Source code 4
Igor Pro compile TIRF data.
- https://doi.org/10.7554/eLife.21069.034
-
Source code 5
Igor Pro analyze TIRF data.
- https://doi.org/10.7554/eLife.21069.035
-
Source code 6
Igor Pro mean squared displacement TIRF analysis.
- https://doi.org/10.7554/eLife.21069.036





