Structural basis for inhibition of erythrocyte invasion by antibodies to Plasmodium falciparum protein CyRPA
Figures
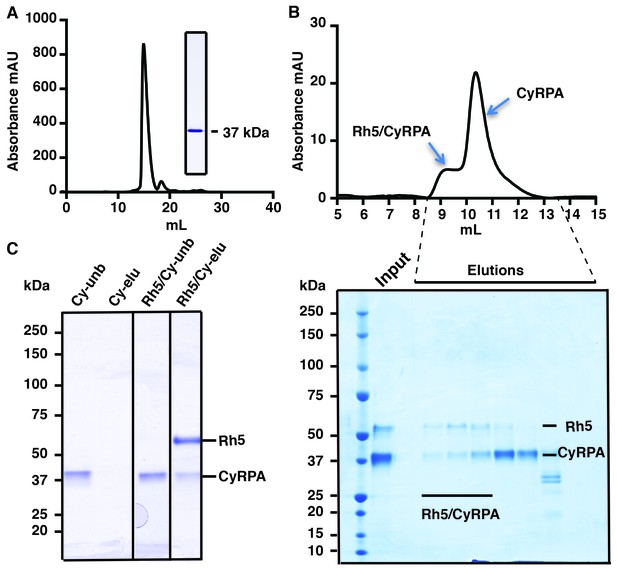
Production of functional recombinant CyRPA.
(A) Purified recombinant CyRPA was analysed by size exclusion chromatography and SDS-PAGE. (B) Formation of the PfRh5/CyRPA complex was monitored by size exclusion chromatography, CyRPA formed a complex with Rh5 and the complex was eluted in the peak labelled as Rh5/CyRPA. The chromatographic profiles are shown (top panel) and fractions eluted from the column (lower panel) were analysed by SDS-PAGE. (C) Immunoprecipitation of FLAG-PfRh5 after incubation with CyRPA. Cy-unb, CyRPA alone; Cy-eluted, elution from anti-FLAG antibody beads of CyRPA alone; Rh5/Cy-unb, FLAG-PfRh5 and excess CyRPA incubated and unbound protein analysed; Rh5/Cy-elu, FLAG-PfRh5 and CyRPA eluted from anti-FLAG antibody beads.
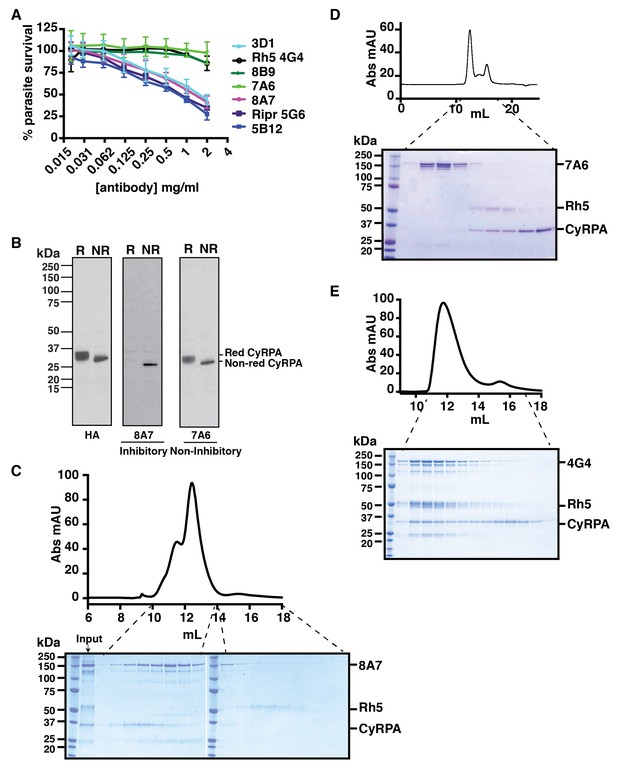
Anti-CyRPA monoclonal antibodies inhibit P. falciparum growth and interaction of PfRh5.
(A) In vitro growth inhibition (GIA) assays were performed to assess the abilities of the monoclonal antibodies raised against recombinant CyRPA to block P. falciparum parasite growth in human erythrocytes. Assays were performed twice in triplicate and error bars denote SD of the mean of 6 values. Anti-Rh5 mAb 4G4 (not inhibitory) and anti-Ripr mAb 5G6 (inhibitory) are included as GIA controls. (B) Immunoblot of inhibitory (8A7) and non-inhibitory (7A6) monoclonal antibodies against proteins from CyRPA-HA tagged transgenic P. falciparum schizonts in reduced (R) and non-reduced (NR) condition. The P. falciparum parasites expressed haemagglutinin-tagged CyRPA protein. (C) Monoclonal antibody 8A7 blocked binding of PfRh5 to CyRPA. PfRh5 and CyRPA were incubated together with 8A7 and the complex formation monitored by size exclusion chromatography and SDS-PAGE analysis under non-reducing conditions. Monoclonal antibody 8A7 bound to CyRPA, preventing the PfRh5/CyRPA complex formation. CyRPA was co-eluted with 8A7 in the earlier fractions and the free Rh5 eluted in the later fractions. (D) Monoclonal antibody 7A6 did not inhibit the formation of the PfRh5/CyRPA complex as monitored by size exclusion chromatography and SDS-PAGE analysis. This antibody did not bind to native CyRPA indicating the linear epitope is not surface exposed. (E) Non-inhibitory anti-PfRh5 monoclonal antibody 4G4 did not significantly inhibit the formation of the PfRh5/CyRPA complex. The PfRh5/CyRPA complex bound to antibody 4G4 and was eluted together with 4G4 as a tri-molecule complex in the earlier fractions.
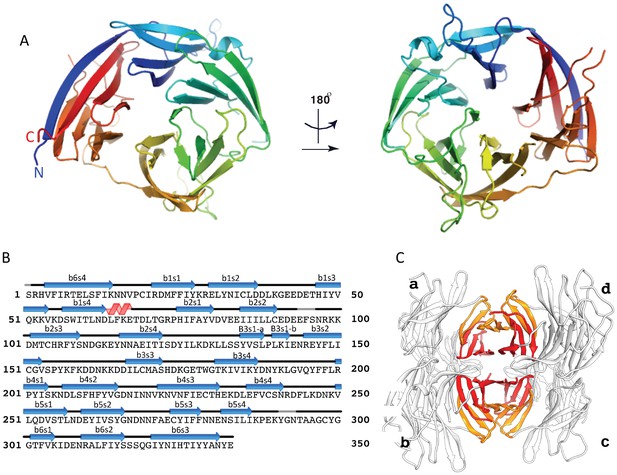
The crystal structure of the uncomplexed CyRPA.
(A) Orthogonal view of the ribbon representation of CyRPA, colored in rainbow fashion from the N-terminus (blue) through to the C- terminus (red). (B) Amino acid sequence and the secondary structure of CyRPA, showing the location of the 24 canonical strands of the six-bladed β-propeller [labelled ‘βmsn’ where the index n denotes the strand and the index m the sheet (n = 1,..,4; m = 1,..,6)]. (C) Assembly of the 222 pseudo-symmetric tetramer of CyRPA within the crystallographic asymmetric unit, showing the formation of the extended 5–5 and 6–6 sheets (orange and red, respectively). Each monomer of CyRPA in the asymmetric unit is labelled a, b, c, d.
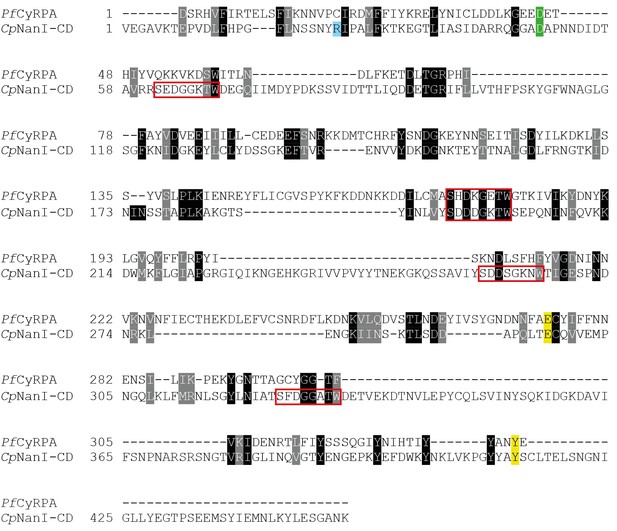
Comparison of CyRPA with C.
perfringens NanI sialidase. Clustal Omega sequence alignment of P. falciparum CyRPA (without signal peptide) and the catalytic domain of Clostridium perfringens NanI sialidase (Newstead et al., 2008). These sequences share 16% identity and 33% similarity. Conserved sialidase features include Asp box motifs (red boxes), catalytic acid/base residue (green), catalytic nucleophile residue (yellow) and the acidic residue that modulates the nucleophile’s pKa (blue).
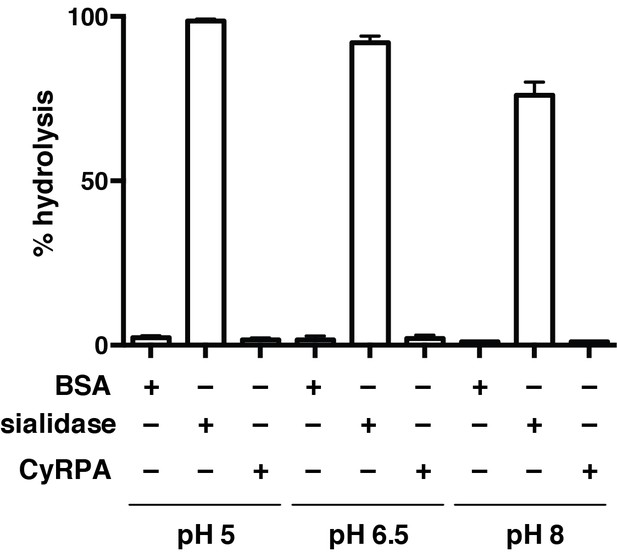
Recombinant CyRPA does not have sialidase activity.
Hydrolysis of 4-methylumbelliferyl N-acetyl-α-D-neuraminic acid (4MU-NeuNAc) in the presence of bacterial sialidase (neuraminidase) from Arthrobacter ureafaciens, CyRPA or BSA at different pH values. These values, obtained in triplicate, were normalized with respect to the fluorescence measured for 10 μM 4-methylumbelliferone in glycine buffer (0.9 M, pH 10) to determine the completeness of each reaction.
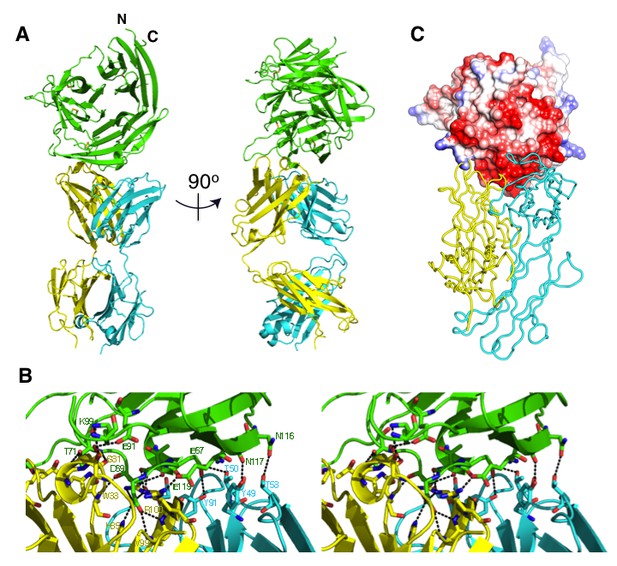
The crystal structure of the Fab 8A7 / CyRPA complex.
(A) Orthogonal views of the ribbon representation of the complex (CyRPA: green, Fab 8A7 light chain: cyan; Fab 8A7 heavy chain: yellow. (B) Hydrogen bond network within the interface between Fab 8A7 and CyRPA. (C) Surface of Fab 8A7 epitope of CyRPA colored according to surface potential (red negative, blue positive), with the Fab chain colored as in (A). Surface potential was computed using CHIMERA (Pettersen et al., 2004).
-
Figure 4—source data 1
Deduced hydrogen bonds between CyRPA and Fab 8A7.
Hydrogen bonds were computed using the program CONTACT within the CCP4 suite (Collaborative Computational Project N, 1994).
- https://doi.org/10.7554/eLife.21347.009

Amino acid sequence of anti-CyRPA monoclonal antibody 8A7 Fab fragment for both heavy and light chains.
https://doi.org/10.7554/eLife.21347.010Tables
Data collection and refinement statistics.
CyRPA | CyRPA/Fab | |
|---|---|---|
Wavelength (Å) | 0.954 | 0.954 |
Resolution range (Å) | 36.45–3.09 (3.20–3.09)* | 38.34–2.44 (2.53–2.44) |
Space group | P1 | P212121 |
Unit cell a, b, c (Å) α, β, γ (°) | 68.7, 83.56, 95.32 96.76, 104.11, 115.20 | 79.95, 87.38, 145.14 90, 90, 90 |
Total no. reflections | 121042 (10425) | 228716 (19565) |
Unique no. reflections | 32250 (2919) | 38347 (3695) |
Multiplicity | 3.8 (3.6) | 6.0 (5.3) |
Completeness (%) | 0.98 (0.89) | 0.99 (0.97) |
<I/σ(I)> | 10.84 (1.73) | 14.54 (2.02) |
Wilson B-factor (Å2) | 71.27 | 54.65 |
Rmerge | 0.12 (0.86) | 0.081 (1.033) |
Rmeas | 0.14 (1.01) | 0.089 (1.137) |
CC1/2 | 0.99 (0.67) | 0.998 (0.72) |
CC* | 1.00 (0.90) | 1.00 (0.915) |
No. reflections used in refinement | 32240 (2917) | 38345 (3695) |
No. reflections used for Rfree | 1584 (148) | 1891 (173) |
Rwork | 0.1871 (0.3620) | 0.1887 (0.2916) |
Rfree | 0.2297 (0.4022) | 0.2115 (0.3000) |
No. non-hydrogen atoms | 10627 | 6045 |
macromolecules | 10614 | 5990 |
Ligands | 12 | |
Protein residues | 1273 | 750 |
RMS bonds (Å) | 0.002 | 0.003 |
RMS angles (°) | 0.48 | 0.58 |
Ramachandran favored (%) | 95.2 | 97.4 |
Ramachandran allowed (%) | 4.8 | 2.6 |
Ramachandran outliers (%) | 0 | 0 |
Rotamer outliers (%) | 0.58 | 0.29 |
Clash score | 6.67 | 3.39 |
Average B-factor (Å2) | 82.87 | 68.03 |
Macromolecules | 82.91 | 68.15 |
Ligand | n.a. | 68.68 |
Solvent | 45.20 | 51.99 |
No. TLS groups | 22 | 10 |
-
*Statistics for the highest-resolution shell are shown in parentheses.





