EMC1-dependent stabilization drives membrane penetration of a partially destabilized non-enveloped virus
Figures
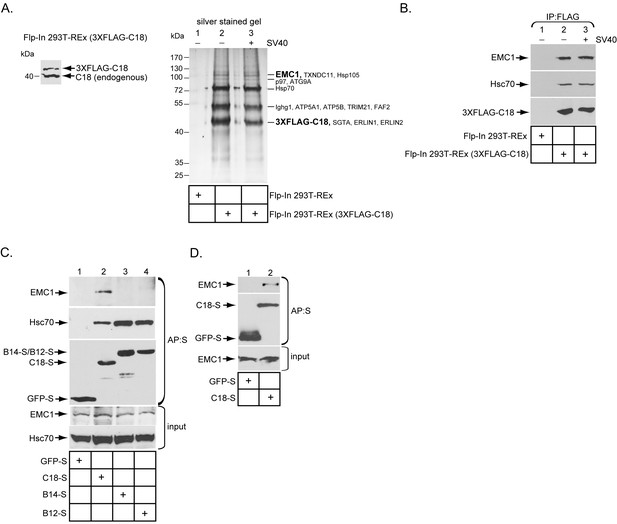
The ER membrane J-protein C18 binds to the EMC1 transmembrane protein.
(A) Expression of 3XFLAG-C18 and endogenous C18 from the Flp-In 293T-REx (3XFLAG-C18) cell line was analyzed by SDS-PAGE followed by immunoblotting with an antibody against C18. 3XFLAG-C18 was immunopurified from induced Flp-In 293T-REx cells infected with SV40 MOI ~50 (‘+’) or mock infected (‘−’). Lysates from the parental cells were used as a negative control. Bound proteins were eluted by 3X FLAG peptide, and subjected to SDS-PAGE followed by silver staining. Bands (indicated on the right) were excised and subjected to mass spectrometry analysis. Protein identities of the bands are listed on the right side of the gel. (B) Samples prepared as (A) were immunoblotted using the indicated antibodies. (C) HEK 293T cells were transfected with the indicated constructs, and after one day of transfection, cells were lysed with 1% Triton X-100 followed by affinity purification with S-agarose beads. The eluted samples were subjected to SDS-PAGE followed by immunoblotting using the indicated antibodies. (D) COS-7 cells were transfected with the indicated constructs, and after two days of transfection, cells were lysed with 1% DBC followed by affinity purification with S-agarose beads. The affinity purified samples were subjected to immunoblotting using the indicated antibodies. Please see Table 1 for the shot –gun mass spectroscopy data.
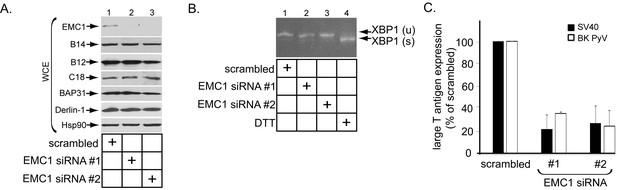
EMC1 is essential in supporting SV40 and BK PyV infection.
(A) CV-1 cells were transfected with either negative control siRNA (scrambled), EMC1 siRNA#1, or #2. Two days after transfection, cells were lysed with 1% Triton X-100 and the resulting whole cell extract (WCE) was subjected to SDS-PAGE followed by immunoblotting using the indicated antibodies. (B) CV-1 cells were transfected with either scrambled, EMC1 siRNA#1, or #2 for two days. RNA was isolated from the cells, and RT-PCR was performed to identify splicing of the XBP1 mRNA. Cells treated with DTT served as a positive control. (C) CV-1 cells transfected with either scrambled siRNA, or EMC1 siRNA#1 or #2 for 48 hr were infected with either SV40 (MOI ~0.5) for 20 hr (black bars) or BK PyV (MOI ~0.5) for 48 hr (open bars), fixed, and then stained for TAg. Infection was scored using immunofluorescence microscopy. Data are normalized to the scrambled siRNA condition. Values represent the mean ± SD (n ≥ 3). Statistically significant differences (p<0.05) were observed under all EMC1 knockdown conditions when compared to scrambled. Role of EMC subunits 2–10 in SV40 infection is shown in Figure 2—figure supplement 1.
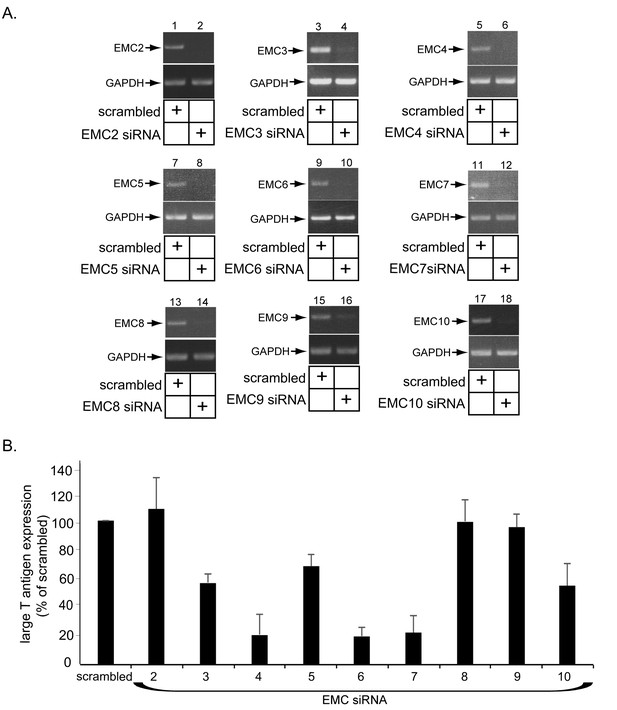
Role of EMC subunits 2–9 in SV40 infection.
(A) CV-1 cells were transfected with either the negative control siRNA (scrambled) or the indicated siRNA directed against the different EMC subunits. RT-PCR analyses were performed to reveal the extent of knockdown of the mRNA message. GAPDH serves as the loading control. (B) Cells in B were incubated with SV40 and the extent of infection analyzed as in Figure 2C.
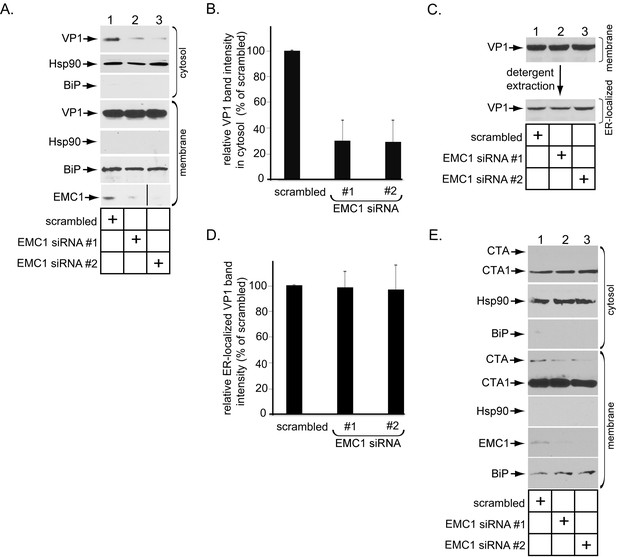
EMC1 promotes SV40 ER-to-cytosol membrane penetration.
(A) CV-1 cells transfected for 24 hr with the indicated siRNAs were infected with SV40 at MOI ~5, harvested 15 hpi, and subjected to the ER-to-cytosol membrane transport assay (see Materials and methods). Cytosolic Hsp90 and ER-resident BiP were used as markers for the cytosol and membrane fractions, respectively. In this assay, the amount of samples loaded in both the cytosol and membrane fractions represent 10% of the total amount in the respective fractions, and they were immunoblotted in parallel with the same exposure time. (The black line indicates that lane 3 of the EMC1 immunoblot was sliced from the same film used for lanes 1 and 2). (B) Relative VP1 band intensities from the cytosol fraction from (A) were determined using ImageJ (NIH). Data are normalized to scrambled siRNA. Values represent the mean ± SD of three independent experiments, and are statistically significant (p<0.05). (C) To isolate ER-localized SV40, the membrane fraction in (A) was solubilized in 1% Triton X-100, and the extracted material subjected to SDS-PAGE followed by immunoblotting with the indicated antibodies. (D) Relative VP1 band intensities from (C) were determined using ImageJ (NIH). Data are normalized to scrambled siRNA. Values represent the mean ± SD of three independent experiments. (E) CV-1 cells transfected with the indicated siRNAs were incubated with cholera toxin for 90 min. Cells were harvested, fractionated as in (A), and the cytosol and membrane fractions analyzed by using the indicated antibodies. We showed SV40-induced EMC1 foci in Figure 3—figure supplement 1.
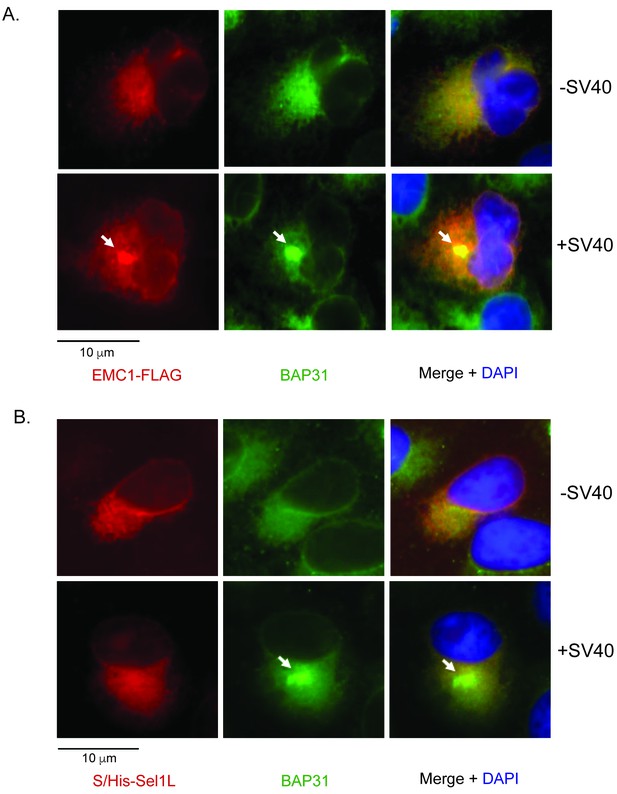
SV40 induces EMC1 to form foci.
(A) CV-1 cells expressing EMC1-FLAG incubated with or without SV40 for 9 hr were stained for FLAG or an antibody against BAP31. (B) CV-1 cells expressing S/His-Sel1L incubated with or without SV40 for 9 hr were stained for S or an antibody against BAP31.
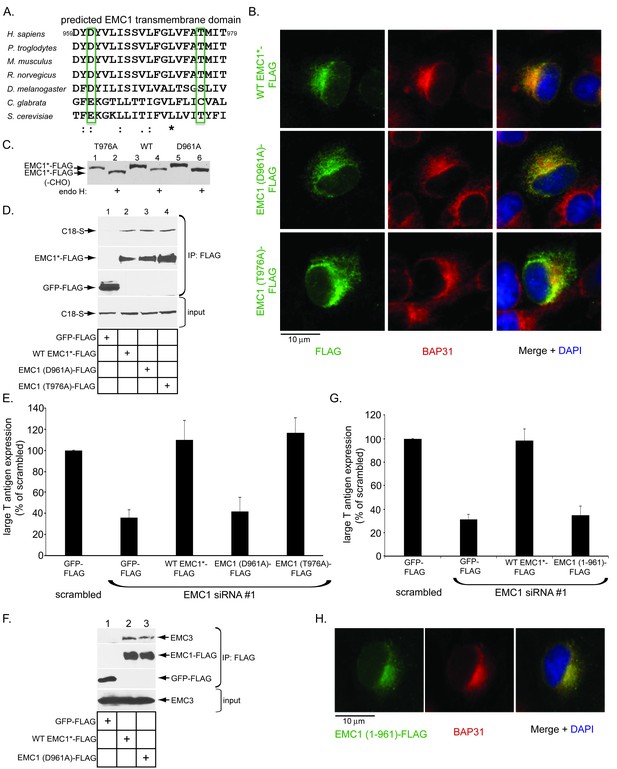
The predicted EMC1 transmembrane domain residue D961 plays a critical role during SV40 infection.
(A) Multiple sequence alignment of the predicted transmembrane domain of EMC1 from human (H. sapiens), chimpanzee (P. troglodytes), mouse (M. musculus), rat (R. norvegicus), fly (D. melanogaster), haploid yeast (C. glabrata), and baker’s yeast (S. cerevisiae). Highlighted amino acids (in green rectangles) represent highly conserved charged or polar residues. (B) CV-1 cells expressing the indicated constructs were stained with an anti-FLAG antibody. BAP31 was used as an ER marker. DAPI positions the nucleus. (C) WT EMC1*-FLAG, EMC1 (D961A)-FLAG, or EMC1 (T976A)-FLAG expressing cells were lysed and treated with endoglycosidase H followed by SDS-PAGE and immunoblotting using a FLAG antibody. (D) Cell lysates generated from HEK 293T cells expressing the indicated constructs were subjected to immunoprecipitation using anti-FLAG antibody coated beads. The bound samples were analyzed by immunoblotting using anti-FLAG and anti-S antibodies. (E) CV-1 cells were transfected with scrambled or the indicated siRNA for 24 hr prior to transfection with the indicated FLAG-tagged constructs for 24 hr. Cells were then infected with SV40 (MOI ~0.5) for 20 hr, fixed, and stained with anti-FLAG and anti-large T antigen antibodies. The percentages of T antigen positive cells were determined only in FLAG-expressing cells by using immunofluorescence microscopy. Values represent means ± SD from three independent experiments. (F) COS-7 cells were transfected with the indicated constructs for 48 hr prior to cell lysis using a buffer containing 1% DBC followed by immunoprecipitation with anti-FLAG antibody. The bound materials were analyzed by immunoblotting using the indicated antibodies. (G) As in E, except cells expressing EMC1 (1-961)-FLAG were also analyzed. (H) As in B, except EMC1 (1-961)-FLAG was used. WT EMC1*-FLAG represents siRNA-resistant EMC1, and all EMC1 mutants were generated using this construct as the template.
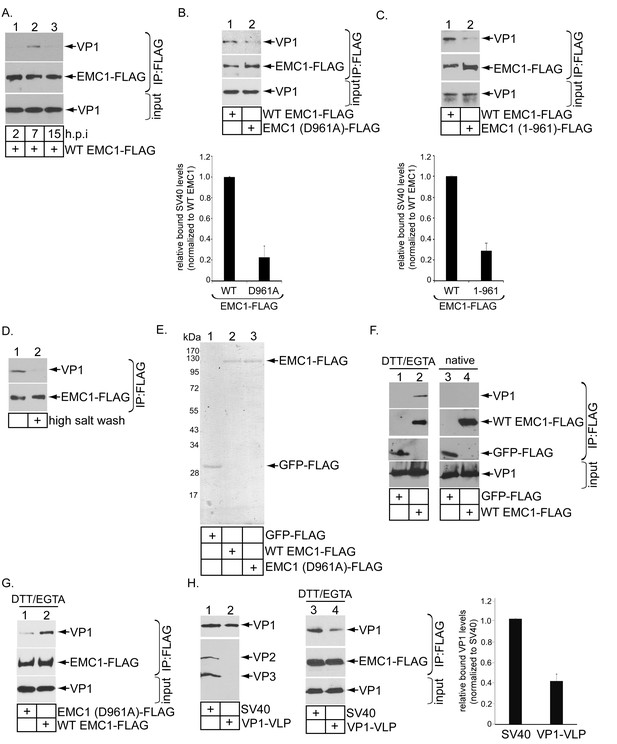
EMC1 binds to SV40 during infection and in vitro.
(A) CV-1 cells expressing WT EMC1-FLAG and subsequently infected with SV40 for the indicated time period were harvested, and the resulting lysate subjected to immunoprecipitation using anti-FLAG antibody coated agarose beads. The bound materials were subjected to SDS-PAGE followed by immunoblotting with the indicated antibodies. (B) Cells expressing WT EMC1-FLAG or EMC1 (D961A)-FLAG were infected with SV40 for 7 hr. Cells were processed and analyzed as in (A). (right graph) Quantification of the relative levels of SV40 bound to WT EMC1 and EMC1 (D961A). Data are normalized to WT EMC1. Values represent means ± SD from three independent experiments. (C) As in B, except EMC1 (1-961)-FLAG was used. (D) The SV40-EMC1 interaction was analyzed at 7 h.p.i. as in (A), except the precipitated material was divided into two sets, with one set subjected to high salt (0.5 M NaCl) wash. The remaining bound material was analyzed by SDS-PAGE and immunoblotting as above. (E) The indicated FLAG-tagged proteins were expressed in and purified from HEK 293T cells, and their purity analyzed by SDS-PAGE followed by Coomassie staining. (F) GFP-FLAG or WT EMC1-FLAG was incubated with either DTT/EGTA treated-SV40 or intact untreated SV40. The FLAG-tagged proteins were precipitated using anti-FLAG antibody coated agarose beads, and the bound materials subjected to SDS-PAGE followed by immunoblotting using the indicated antibodies. (G) WT or EMC1 (D961A)-FLAG was incubated with DTT/EGTA treated-SV40, and the samples analyzed as in (F). (H) (lanes 1–2) VP1-VLP and WT SV40 were analyzed for the presence of VP1, VP2, and VP3. (lanes 3–4) DTT/EGTA-treated SV40 or VP1-VLP was incubated with EMC1-FLAG. The extent of interaction was analyzed as in F. The EMC1-bound VP1 level was quantified in the right graph.
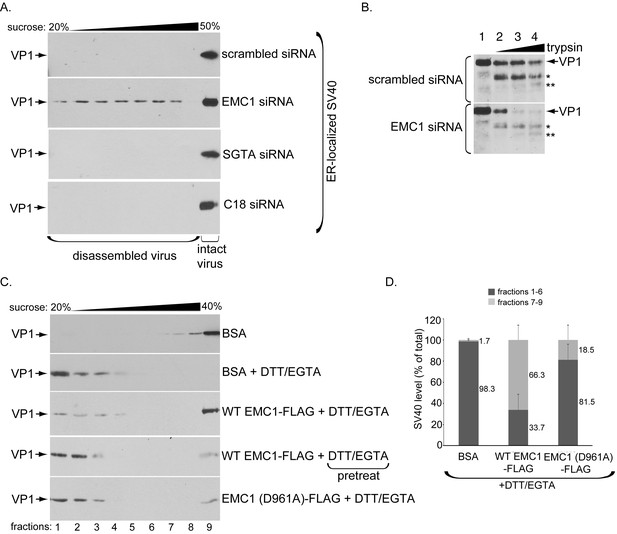
EMC1 stabilizes ER membrane-penetrating SV40.
(A) Extracts containing ER-localized SV40 derived from CV-1 cells treated with the indicated siRNAs were layered on top of a discontinuous sucrose gradient (20–50% sucrose), and centrifuged. Fractions were collected from the top of the gradient and analyzed for presence of SV40 by immunoblotting using VP1 antibodies. (B) Extracts containing ER-localized SV40 derived from CV-1 cells treated with the indicated siRNAs were subjected to limited proteolysis using increasing trypsin concentrations. The samples were then subjected to SDS-PAGE followed by immunoblotting using polyclonal VP1 antibodies. * and ** denote degraded VP1 monomers. (C) Purified SV40 was incubated with either BSA, BSA + DTT/EGTA, WT EMC1-FLAG + DTT/EGTA, or EMC1 (D961A)-FLAG + DTT/EGTA. Alternatively, SV40 was pre-incubated with DTT/EGTA followed by addition of WT EMC1-FLAG. Samples were layered on top of a discontinuous sucrose gradient (20–40% sucrose) and centrifuged. Fractions were analyzed as in (A). (D) The amount of VP1 in fractions 1–6 (corresponding to disassembled SV40) and fractions 7–9 (corresponding to largely intact SV40) in samples incubated with BSA, WT EMC1-FLAG, or EMC1 (D961A)-FLAG in the presence of DTT/EGTA in C were quantified.
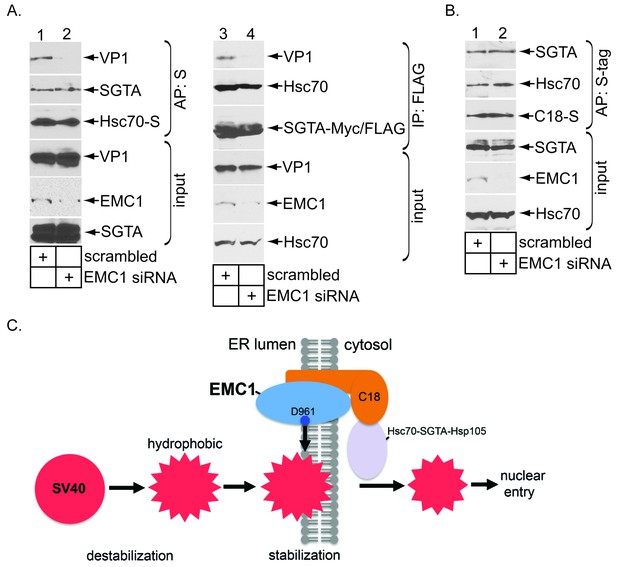
EMC1 depletion impairs SV40’s engagement with the cytosolic extraction machinery.
(A) (lanes 1–2) Extracts from SV40-infected CV-1 cells expressing Hsc70-S and treated with the indicated siRNAs were subjected to affinity purification using S-agarose beads. The bound materials were subjected to immunoblotting using the indicated antibodies. (lanes 3–4) As in lanes 1–2, except COS-7 cells were expressing SGTA-Myc/FLAG. (B) Extracts from COS-7 cells treated with the indicated siRNAs were subjected to affinity purification using S-agarose beads. The bound materials were subjected to immunoblotting using the indicated antibodies. (C) A model depicting how EMC1 stabilizes the partially destabilized SV40 in the ER membrane. When SV40 reaches the ER from the plasma membrane, ER factors destabilize SV40, exposing its internal hydrophobic proteins VP2 and VP3. This generates a partially destabilized hydrophobic viral particle that binds to and integrates into the ER lipid bilayer. Within the bilayer, we propose that EMC1 deploys its D961 residue to engage and stabilize this partially destabilized virus, thereby preventing it from premature disassembly. EMC1-dependent stabilization enables the cytosolic Hsc70-SGTA-Hsp105 machinery to extract the virus into the cytosol in order to complete the membrane transport process.
Tables
Shot-gun mass spectrometry analyses of EMC subunits that bound to C18. 'Shot-gun' mass spectrometry analyses of the FLAG-precipitated material derived from either 3XFLAG-C18 expressing cells or the control parental cells not expressing 3XFLAG-C18 in Figure 1A. Only the amounts of unique peptides corresponding to the 10 EMC subunits, as well as C18, are shown.
| unique peptide (Flp-In 293T-REx) | unique peptide (Flp-In 293T-REx 3XFLAG-C18) | |
|---|---|---|
| C18 | 0 | 16 |
| EMC1 | 0 | 7 |
| EMC2 | 3 | 4 |
| EMC3 | 0 | 5 |
| EMC4 | 0 | 2 |
| EMC5 | 0 | 0 |
| EMC6 | 0 | 1 |
| EMC7 | 0 | 0 |
| EMC8 | 0 | 0 |
| EMC9 | 0 | 0 |
| EMC10 | 0 | 1 |





