Kinetics of the LOV domain of ZEITLUPE determine its circadian function in Arabidopsis
Figures
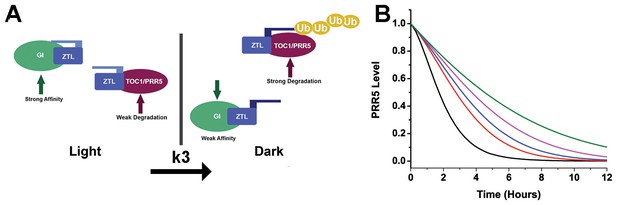
Models of ZTL photochemistry and regulation of circadian period.
(A) In the light, ZTL associates with both GI and degradation targets (PRR5/TOC1). During the day, the strong affinity for GI allows GI, ZTL, TOC1 and PRR5 levels to rise. Upon dusk, the adduct form of ZTL decays with a rate constant k3, leading to a conformational change. The conformation change decreases GI affinity and leads to ubiquitination of ZTL targets. (B) Modeling PRR5 degradation as a function of k3 (see Equation 1 and Materials and methods for model generation and parameter selection). Using k3 for WT (black), G80R (red), V48I (blue), G46S:G80R (magenta) and V48I:G80R (green) leads to predictable changes in PRR5 degradation patterns. (Figure 1—supplements 1, 2 and 3).
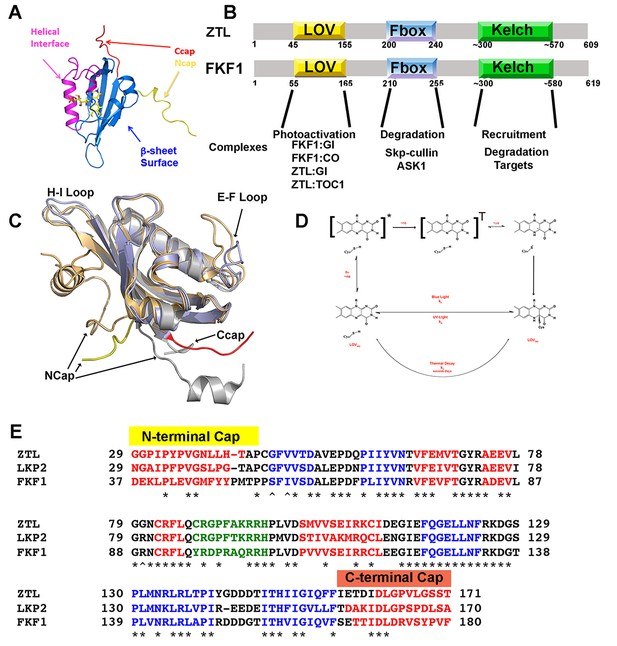
LOV chemistry and signal transduction.
(A) PAS/LOV domains signal through four elements. Multi-domain proteins reorient Ncap (yellow) and Ccap (red) elements to affect activity of signaling domains or recruit additional proteins to the PAS/LOV surface. The β-sheet (blue) and helical surface (magenta) act as protein:protein interactions motifs. (B) ZTL, and FKF1 domain architecture. The N-terminal LOV domain engages multiple targets. Light-activation then regulates Fbox activity to target proteins for degradation. (C) Superposition of VVD (light orange), AsLOV2 (grey) and ZTL. The ZTL LOV core (blue) is similar to existing LOV structures. Significant deviations exist at the Ncap (ZTL: Yellow), Ccap (ZTL: Red) and the E-F and H-I loops. These elements are believed to be important for signaling. (D) LOV proteins are activated by blue-light absorption to form an excited singlet state. The singlet rapidly undergoes intersystem crossing to generate a triplet intermediate. Proton coupled electron transfer from the conserved Cys residue and subsequent C4a adduct formation activates LOV proteins. The adduct state then decays in the dark or presence of UV-light. (E) Sequence alignment of ZTL family. Predicted helical regions (red), β-sheet (blue) and an extended E-F loop (green) are noted. * implies conserved, ∧ residues tuning ZTL and FKF1 kinetics. The Ncap and Ccap regions are labeled.
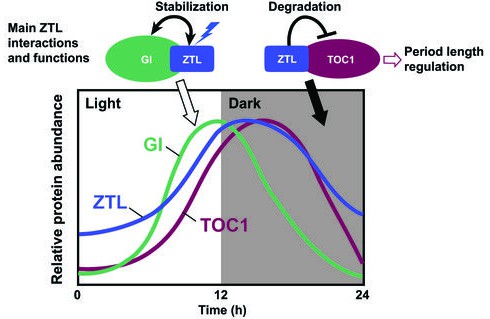
Schematic diagram of the daily protein abundance profile of ZTL and its function in the circadian clock.
ZTL function in the circadian clock involves interactions with two core clock proteins, GI and TOC1. The protein abundance patterns of ZTL, GI, and TOC1 proteins are based on previously published results (Más et al., 2003a; Kim et al., 2007). The circadian clock regulates the expression profile of GI protein with its peak near the end of the light period. The ZTL-GI interaction is enhanced when the ZTL LOV domain absorbs blue light. This interaction stabilizes both ZTL and GI proteins toward the end of the day, thus ZTL protein level also peaks at that time, although ZTL transcript levels are constitutive. TOC1 expression is also clock regulated with expression mainly during the nighttime. ZTL interacts with TOC1 in a light-independent manner, however ZTL mediated TOC1 degradation is enhanced at night. ZTL mutants lead to enhanced accumulation of TOC1 protein, leading to period lengthening of the circadian clock.
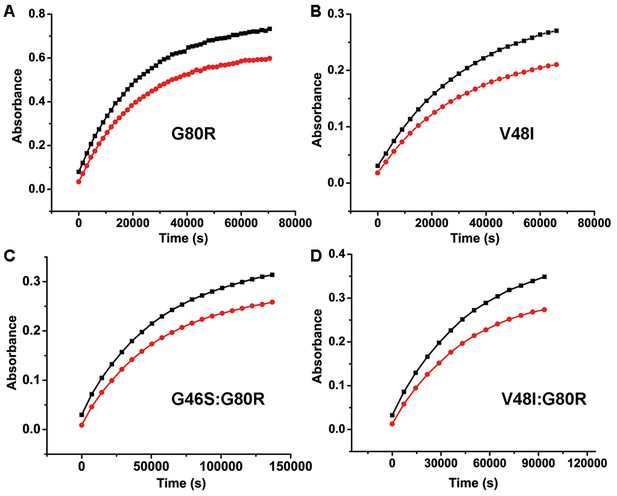
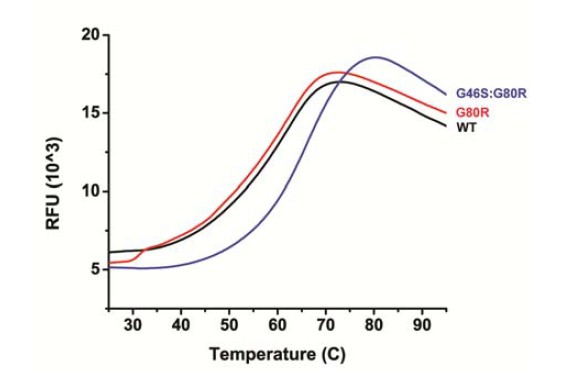
Kinetics of ZTL Variants (A–D) Kinetics of ZTL adduct decay are determined from the absorbance recovery at 450 (black) and 478 nm (red).
G80R; τ = 6.6 hr (A) and V48I; τ = 10.7 hr (B) recover on similar timescales. The G46S:G80R; τ = 21 hr (C) variant recovers with a 2-fold slower time constant. A V48I:G80R variant recovers τ >65 hr (see Table 1 for more information). Due to the long time constant for V48I:G80R, full recovery cannot be observed without the presence of an imidazole base catalyst (Pudasaini and Zoltowski, 2013). The recovery data shown (D) is under base catalyzed conditions (150 mM imidazole) to allow full recovery.
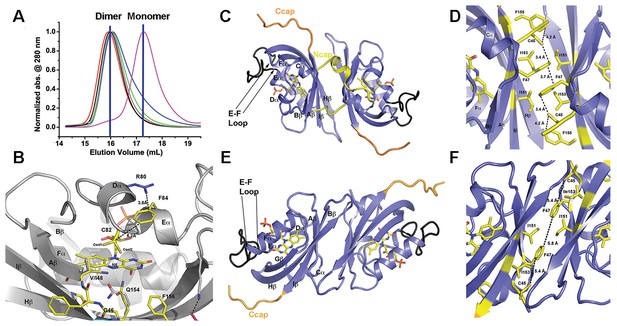
Structural analysis and LOV dimer formation in ZTL.
(A) G80R (dark-state, black; light-state (grey), WT (red), V48I:G80R (green), G46S:G80R (blue) all elute as dimers with apparent MWs of 38–41 kDa compared to the expected monomer of 16 kDa. Multi-Angle-Light-Scattering (MALS) confirms dimer formation in WT 29–165 (absolute MW 33 ± 2 kDa) and 16–165 constructs (See Figure 2—figure supplement 1). Introduction of an I151R abolishes dimer formation (magenta; apparent MW = 22 kDa). (B) Structure of ZTL active site (yellow) and residues involved in structural or kinetic modulation of signaling (purple). R80 within Dα forms a π-cation interaction with F84 directly above the photoreactive C82, resulting in steric stabilization of adduct formation. The observed steric stabilization through the π-cation interaction is consistent with the longer photocycle in G80R. V48I positions the additional methyl group into a pocket adjacent to N5, C82 and Q154. Comparisons of AsLOV2 structures (white; buried conformation), dark-state ZTL (yellow; exposed conformation) demonstrates that V48I can impact the position of Q154 between buried and exposed conformations. Movement of Q154 correlates with movement of F156 in Iβ. (C–F) ZTL monomers are defined by an antiparallel β-sheet flanked by a series of α-helices (Cα, Dα, Eα, Fα). The helices cradle the photoreactive FMN adjacent to C82 (Eα helix). ZTL contains a 9-residue insert linking the E-F helices that accommodates the adenine ring of FAD in some LOV proteins (black) (Zoltowski et al., 2007). N- and C-terminal extensions (Ncap/Ccap; yellow) are largely disordered; however, a short helix within the Ncap reaches across a dimer interface in some molecules to form contacts between the Cα and Dα helices. Two dimer interfaces are formed through the β-scaffold in ZTL. The compact dimer (C,D) differs from the elongated dimer (E,F) by a 2.0 Å translation along the β-sheet. Key residues in the dimer interface are shown in yellow. The translation disrupts a network of sulfur-π and π-π interactions involving C45 and F47, centered around I151. (Figure 2—figure supplements 1 and 2).
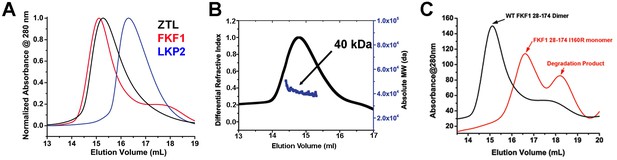
Dimerization of ZTL/FKF1/LKP2 LOV domains.
(A) ZTL (16-165; apparent MW = 57 kDa; absolute MW from MALS ~40 kDa, panel B) and FKF1 (28–174; apparent MW from SEC 62 kDa; absolute MW from MALS ~42 kDa) elute as dimers on SEC as confirmed by multi-angle light scattering (B). LKP2 (16–165 apparent MW = 33 kDa) adopts a much smaller hydrodynamic radius indicating distinct differences in oligomeric structure and/or affinity. ZTL 16–165 was used to allow comparison of similar length constructs and allow use of a stable FKF1 construct characterized previously (Nakasako et al., 2005), (B) Multi-angle light scattering of ZTL 16–165. The expected monomer MW was ~18 kDa, indicating ZTL exists as a constitutive dimer. (C) Introduction of I160R variants (equivalent to ZTL I151R) in FKF1 renders the protein monomeric. It is also more susceptible to proteolysis.
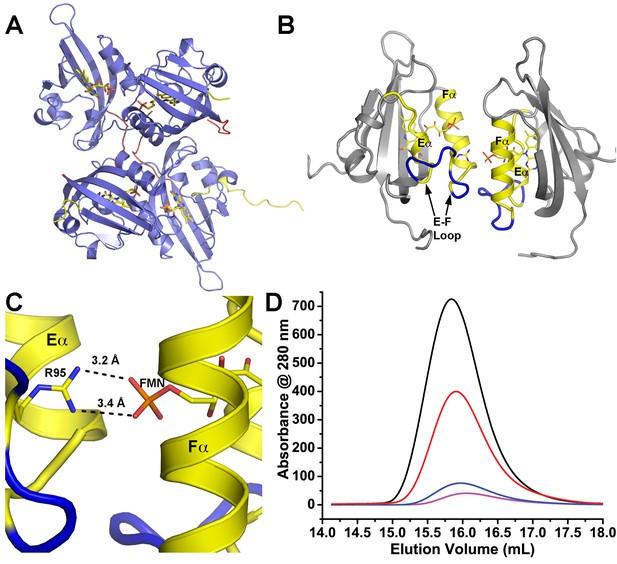
Dark-state structure of ZTL 29–165 and helical dimer.
(A) ZTL crystallizes as a tetramer, where the helical interfaces are buried within the crystal lattice. (B) A parallel helical dimer is observed in the crystal lattice that is characterized by slightly asymmetric contacts between the E and F helices (yellow) and associated loops (blue). (C) Contacts between R95 in the E-F loop (blue) and the phosphate of the bound FMN of the neighboring molecule stabilize the helical dimer. (C) Introduction of a R95A variant does not affect in vitro dimerization of ZTL 29–165. The elution profile of R95A is not concentration dependent and reflects a dissociation constant unable to be determined by SEC (Kd <0.2 µM). Traces depicted and apparent MWs are: dark-state (Black- 41.3 kDa at 130 µM and Blue- 39.6 kDa at 13 µM), light-state (Red-40.8 kDa at 130 µM and Magenta- 37.8 at 13 µM).
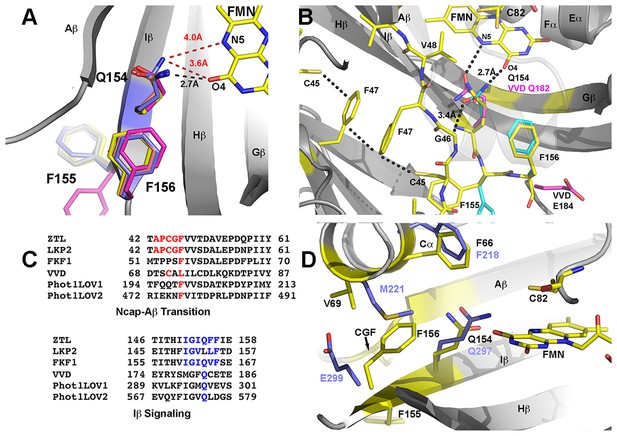
Q154 links Ncap, Ccap and helical elements.
(A) Q154 exists in multiple conformations in WT ZTL structures. They differ in interactions with the active site flavin. An exposed conformation forms strong H-bonds to the O4 position (black dotted line). A buried conformation forms weaker interactions (3.6 Å) with O4 that leads to closer interactions at N5 (4.0 Å; red dotted line). The altered conformation is coupled to movement of F156 into the active site and multiple conformations of F155, forming a QFF motif. The altered conformations define ZTL signaling as distinctly different than existing LOV structures. (B) The unusual orientations of Q154 differ from other LOV proteins that typically show strong interactions near N5 (VVD; magenta). The heterogeneous conformations of Q154 directly abut G46 in a CGF motif allowing formation of the sulfur-π and π-π interactions involving C45 and F47. F156 then adopts a buried conformation in contrast to the equivalent residue in VVD (E184) (C) Sequence alignments of LOV proteins depict conserved elements within the CGF motif (red) and QFF motif (blue) in Arabidopsis thaliana ZTL, LKP2, FKF1, phototropin 1 LOV1 and LOV2. Sequence conservation indicates divergent signaling mechanisms within the ZTL/FKF1 family compared to existing LOV allostery models. (D) Comparisons of ZTL (yellow, black lettering) and Arabidopsis thaliana phototropin 1 LOV1 (PDB: 2Z6C; blue). The altered conformation of Q154 draws F156 into the active site. The buried conformation of F156, leads to movement of Cα (F66, V69). (Figure 3—figure supplement 1)..
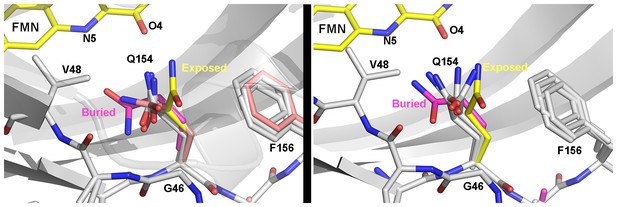
Heterogeneous orientations of Q154.
WT (left) and G80R (right) conformations of Q154 compared to buried conformations in VVD (magenta; PDBID 2PD7) and exposed conformations in ZTL V48I:G80R (yellow). In both WT and G80R, Q154 samples orientations covering the range between exposed and buried conformations. One orientation in WT (salmon) lies close to the buried conformation. These heterogeneous conformations may contribute to ZTL retaining light and dark-state functions.
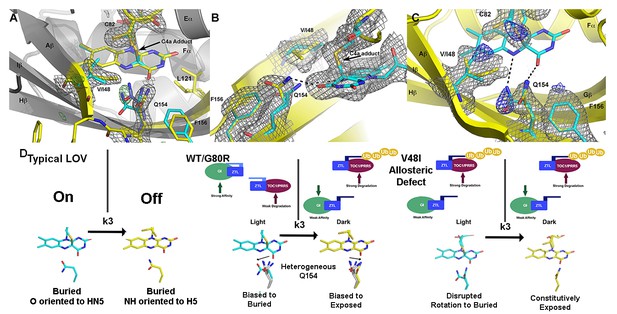
Structural effects on ZTL chemistry and signaling.
(A) Comparison of dark-state V48I:G80R (yellow), light-state V48I:G80R (cyan) and dark-state AtLOV1 phototropin 1 (purple: PDB ID: 2Z6C) molecules. 2Fo-Fc (2.0 σ grey mesh) and Fo-Fc (3.0 σ green mesh) are depicted for dark-state V48I:G80R. Lack of density for an adduct is consistent with minimal population of the light-state species. Density shows clear selection of the I48 methyl group towards Q154. Some residual density is present in the buried conformation of Q154 that indicates either partial occupancy of the site in the dark, or residual light-state conformations. The buried conformation correlates with the orientation of Q154 in all other LOV structures as depicted by AsLOV2. Electron density for FMN is excluded to allow clear observation of electron density for active site side-chains. (B) Light state crystal structure of V48I:G80R, 2Fo-Fc data shown at 2.0 σ show (grey mesh) clear electron density for C4a adduct formation. (C) Rotated view of the active site of the light-state ZTL molecule. Modeling of Q154 in the exposed conformation (yellow; panel C) results in Fo-Fc (blue mesh; panel C) density at 3.0 σ for the light-state molecule. Electron density for the FMN is excluded for clarity. The data confirms rotation of Q154 to a buried conformation (cyan) following adduct formation. Rotation of Q154 is coupled to rearrangement of V/I48. In V48I, the additional methyl groups blocks rotation, partially inhibiting population of the buried conformation of Q154 (cyan conformation). (D) Predicted divergent ZTL model of allostery and signal transduction based on the integrated structural, mutational and in vivo data. Orientations of WT/G80R Q154 are derived from the dark-state G80R ZTL structure with buried and exposed conformations shown for reference from V48I:G80R. WT/G80R and V48I:G80R deviate from typical LOV models (derived from VVD: light PDBID 3RH8, dark PDBID 2PD7) on the position of Q154. WT/G80R ZTL retains a heterogeneous orientation of Q154. We propose that Q154 is heterogeneous regardless of lighting conditions, but biased towards the buried conformation in the light. At dusk adduct decay, with rate constant k3, causes the Q154 conformation to be biased towards an exposed conformation, accelerating ubquitination of protein targets. For V48I, I48 selects the exposed conformation in the dark and leads to only partial burying of Q154 (shown in C, (D), leading to constitutively high ubquitination activity that mimics the dark-state of WT-ZTL. (Figure 4—figure supplements 1 and 2).
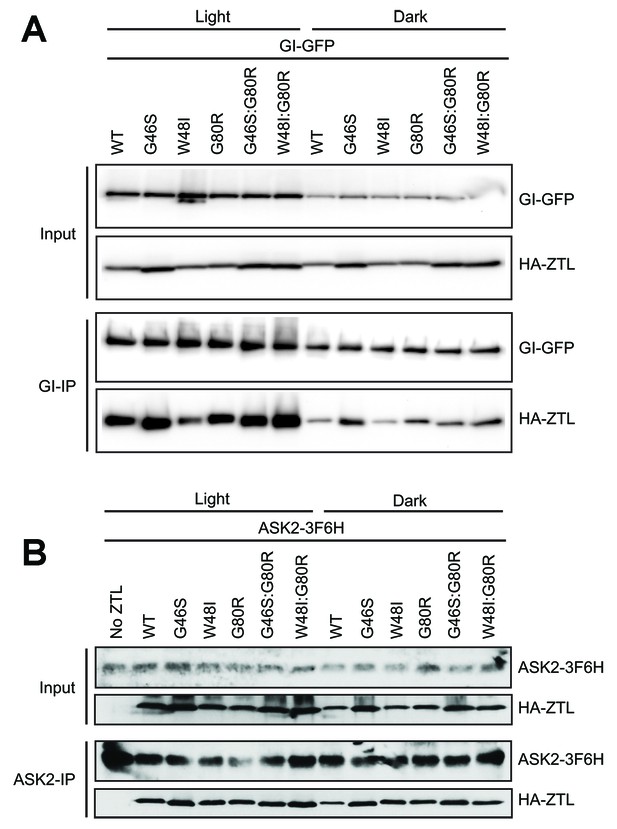
Effect of ZTL variants on protein complex formation.
(A) In vivo interactions between GI and ZTL variants were detected by transient co-expression in N. benthamiana under continuous light or dark conditions. WT and ZTL variants all show enhanced interactions with GI in the presence of light. Ncap variants lead to altered GI interactions. V48I disrupts light-state GI interactions, consistent with altered conformational changes. G46S enhances dark-state binding to GI due to partial dark-state activation. These identify the Ncap as the interaction surface for GI. (B) Effect of ZTL LOV domain mutations on ASK2. Proteins were precipitated with anti-FLAG (B) or protein A antibody, and the presence of ZTL variants was detected by anti-HA antibody. All variants indicate that ASK2 interacts with ZTL regardless of lighting conditions.
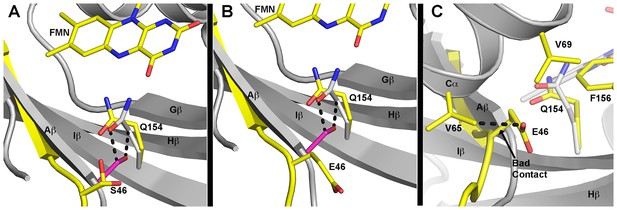
Predicted differences in G46 mutants.
(A–C) Predicted models of G46 mutations and hypothetical effect based on WT ZTL structures. (A) The WT ZTL (grey cartoon) structure cannot accommodate a sidechain at position 46. Addition of a Ser residue (G46S) at this position leads to steric clashes (2.2 Å) between the Cβ position and the exposed conformation (grey) of Q154. Additional clashes are present from the OH group (dashed lines). G46S can be accommodated by very modest movement of Aβ (yellow) and rotation of Q154 to the buried position (yellow). Such reorientation favors the putative light-state conformation of Q154. S46 can adopt to orientations that do not clash with any residues in the ZTL structure (shown). One orientation H-bonds with Q154. Such interactions should not disrupt the ZTL fold. (B) The equivalent orientation of the Glu sidechain in a G46E that minimizes steric clashes. (C) A G46E mutation requires rotation of the Glu sidechain out of the active site pocket. The orientation with minimum steric clashes retains a van der Waals contact (2.5 Å) with V65 in Cα (dashed line). In addition, charged E46 is forced into a hydrophobic pocked lined by V65, V69 and F156. We predict these combined interactions destabilize the LOV fold in G46E.
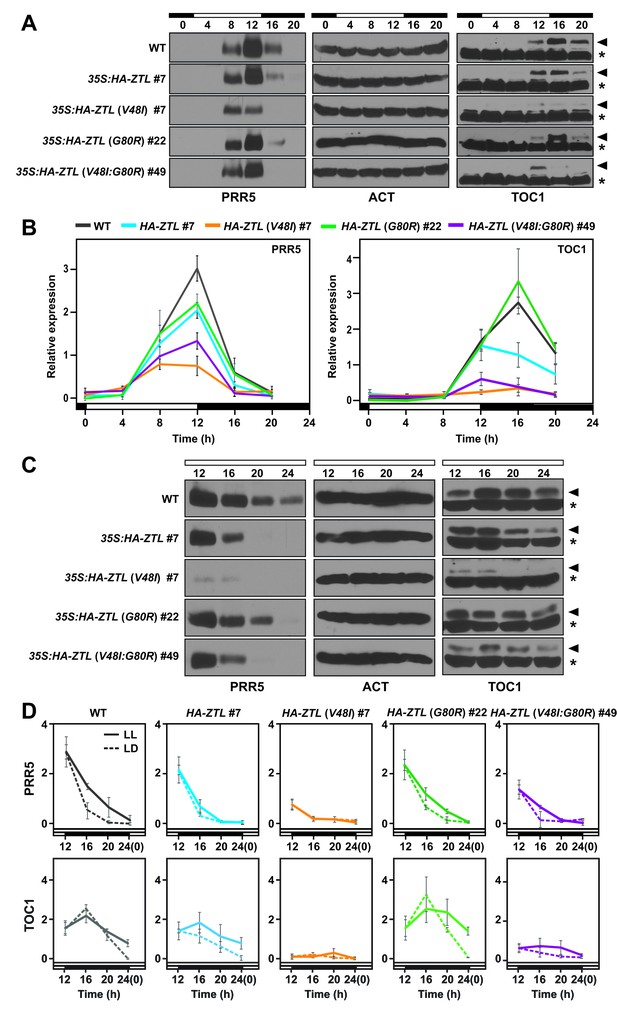
Diurnal and circadian expression profiles of PRR5 and TOC1 proteins in ZTL variant overexpressors.
(A) PRR5 and TOC1 protein levels were analyzed in WT, 35S: HA-ZTL, 35S: HA-ZTL (V48I), 35S: HA-ZTL (G80R) and 35S: HA-ZTL (V48I:G80R) under 12L/12D conditions. Actin (ACT) was used as a loading control for PRR5. Arrowhead indicates the band corresponding to TOC1 protein, while an asterisk indicates a nonspecific cross-reacting band, which is used as a loading control. (B) Relative expression level of PRR5 and TOC1 were determined in WT, 35S: HA-ZTL, 35S: HA-ZTL (V48I), 35S: HA-ZTL (G80R) and 35S: HA-ZTL (V48I:G80R) under 12L/12D conditions. Actin and the TOC1 nonspecific bands were used for normalizing protein loadings for quantification of PRR5 and TOC1. The data represent the averages ±SEM obtained from three biological replicates. (C) PRR5 and TOC1 protein levels were analyzed in WT, 35S: HA-ZTL, 35S: HA-ZTL (V48I), 35S: HA-ZTL (G80R) and 35S: HA-ZTL (V48I:G80R) during the subjective night under constant light conditions. (D) Relative levels of PRR5 and TOC1 proteins were determined in WT, 35S: HA-ZTL, 35S: HA-ZTL (V48I), 35S: HA-ZTL (G80R) and 35S: HA-ZTL (V48I:G80R). Dashed lines represent protein levels under 12L/12D conditions, while solid lines represent protein levels under constant light conditions. The data represent the averages ±SEM obtained from three biological replicates. (Figure —figure supplement 1 and 2).
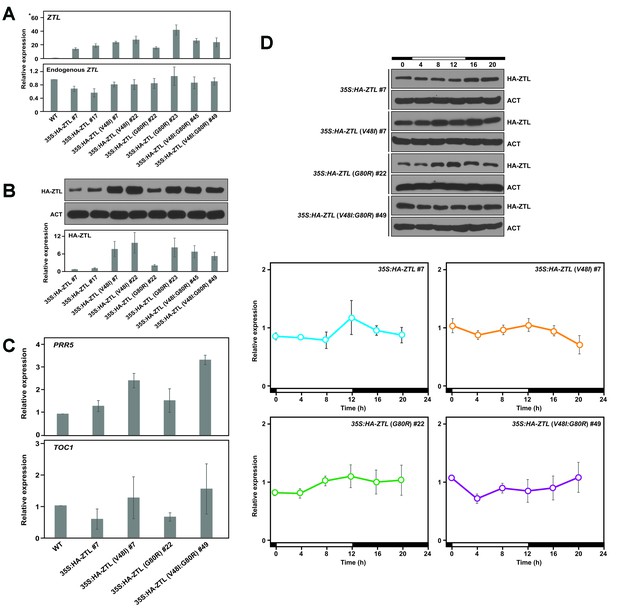
Relative expression levels of clock proteins.
Expression levels of total ZTL and endogenous ZTL transcripts (A), and also ZTL protein (B) were analyzed in WT, 35S: HA-ZTL, 35S: HA-ZTL (V48I), 35S: HA-ZTL (G80R) and 35S: HA-ZTL (V48I:G80R) plants harvested at ZT8 under 12L/12D conditions. (A) Relative expression levels of ZTL were determined using IPP2 as the internal control. (B) Representative blot images of HA-ZTL and ACT are shown. Relative expression levels of ZTL protein were determined from three biological replicates. (C) Expression levels of PRR5 and TOC1 transcripts were analyzed in WT, 35S: HA-ZTL, 35S: HA-ZTL (V48I), 35S: HA-ZTL (G80R) and 35S: HA-ZTL (V48I:G80R) plants harvested at ZT8 under 12L/12D conditions. Relative expression levels of PRR5 and TOC1 were determined using IPP2 as the internal control. (D) Diurnal expression levels of ZTL protein were analyzed in 35S: HA-ZTL, 35S: HA-ZTL (V48I), 35S: HA-ZTL (G80R) and 35S: HA-ZTL (V48I:G80R) plants under 12L/12D conditions. Representative blot images of HA-ZTL and ACT are shown. Relative expression levels of ZTL protein were determined using ACT as the internal control. All data represent averages ±SEM obtained from three biological replicates.
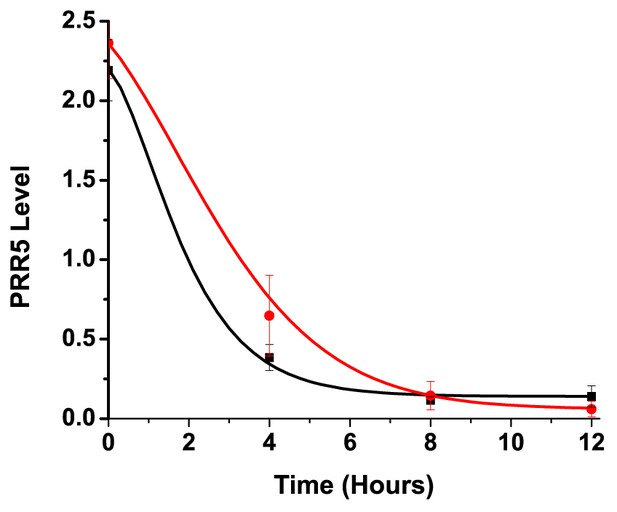
Comparison of model to PRR5 degradation in vivo.
Comparison of PRR5 degradation model (Equation 1 and Equation S10) with experimental data. The red curve (simulated for G80R) shows reasonable precision in predicting the observed delay in PRR5 degradation for G80R data (red circles) compared to simulated WT (black) and WT data (black squares). See methods for model generation and parameter estimation.
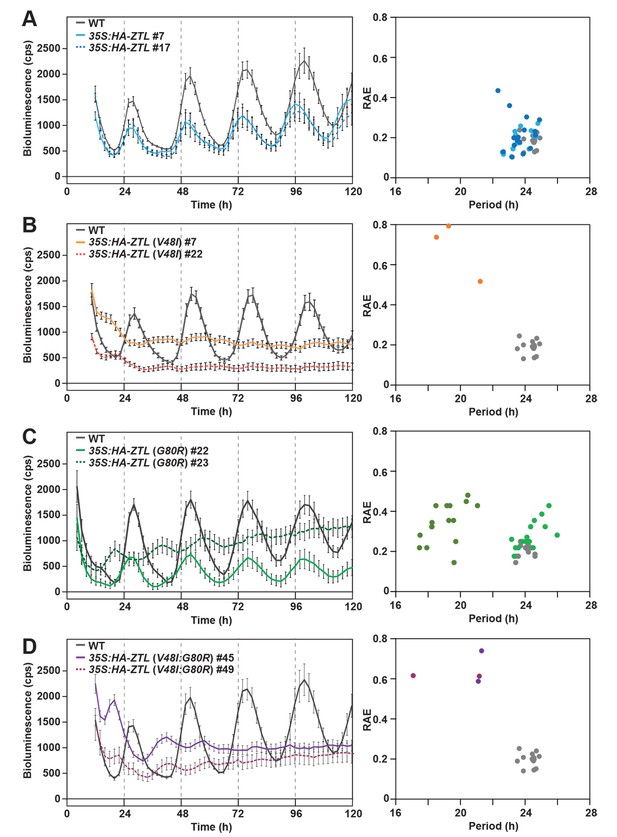
Circadian clock phenotypes of ZTL variant overexpressors.
(A–D) CCA1:LUC activity was analyzed in WT, 35S:HA-ZTL (A), 35S:HA-ZTL (V48I) (B), 35S:HA-ZTL (G80R) (C) and 35S:HA-ZTL (V48I:G80R) (D) lines under continuous light conditions. CCA1:LUC traces represent the averages ±SEM of the results obtained from eight individual seedlings. Period length estimation and relative amplification errors of 16 individual measurements are shown.
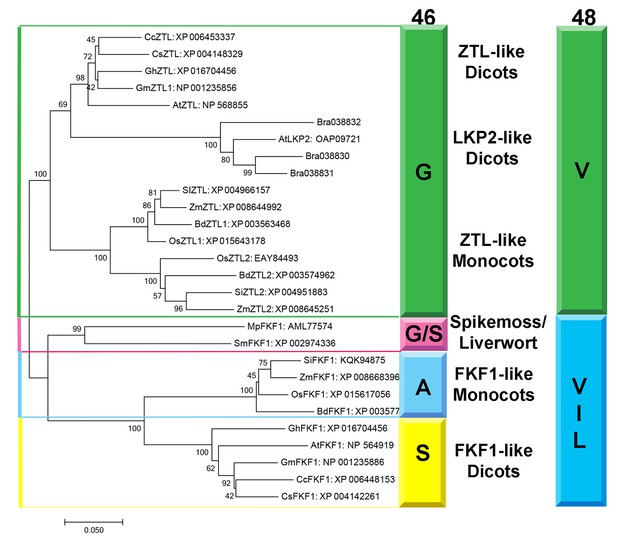
Phylogenetic Analysis of FKF1/ZTL family members in plants.
Residue identity at position 46 (Colored Bar) distinguishes ZTL-like, LKP2-like and FKF1-like proteins consistent with evolutionary diversification of signaling mechanisms. LKP2 is isolated to a clade containing Brassica rapa members that all contain a Q154L substitution. Al ZTL members contain G46 which is necessary to promote the alternative conformation of Q154. Spikemoss and liverwort FKF1’s are isolated indicating a possible intermediate function. Accession numbers for all sequences are shown after the protein name. The evolutionary history was inferred using the Minimum Evolution method (Rzhetsky and Nei, 1992). The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (500 replicates) are shown next to the branches (Felsenstein, 1985). The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Poisson correction method (Zuckerkandl and Pauling, 1965) and are in the units of the number of amino acid substitutions per site. The ME tree was searched using the Close-Neighbor-Interchange (CNI) algorithm (Nei and Kumar, 2000) at a search level of 1. The Neighbor-joining algorithm (Saitou and Nei, 1987) was used to generate the initial tree. The analysis involved 28 amino acid sequences. All positions containing gaps and missing data were eliminated. There were a total of 535 positions in the final dataset. Evolutionary analyses were conducted in MEGA7 (68). (Figure 7—figure supplement 1).

Sequence Alignment of FKF1/ZTL family members in plants.
ZTL/FKF1/LKP2 homologs from dicots; (Arabidopsis thaliana (At), Brassica rapa (Bra), Glycine max (Gm), Cucumis sativus (Cs), Citrus clementine (Cc), Gossypium hirsutum (Gh), Avena sativa (As), monocots; Oryza sativa japonica (Os), Setaria italica (Si), Zea mays (Zm), Brachypodium distachyon (Bd), Liverwort; Marchantia polymorpha (Mp) and Spikemoss; Selaginella moellendorffii (Sm). ZTL and FKF1 cluster in reference to conserved CGF and QFF motifs. See Figure 7 for corresponding accession numbers. All ZTL proteins conserve G46 and V48 (blue). In FKF1 the position corresponding to G46 contain Ala (FKF1-like monocots) or Ser (FKF1-like dicots) (red); Phototropins contain an Asn at the equivalent position (green). * denote conserved residues through all proteins. All proteins conserve the canonical GXNCRFLQ motif (magenta) as well as residues leading into the E-F loop. The FKF1 species differ in the residues immediately following the LOV consensus sequence in the beginning of the E-F loop (ZTL: C87 and G89; FKF1: F87 and D89). Liverwort and spikemoss sequences diverge containing elements consistent with both ZTL and FKF1 (G46 and ZTL E-F loop but I48 for Mp; S46 and F87 but G89 for Sm), indicating an evolutionary transition. The QFF (blue) motif is more divergent. All ZTL/FKF1 contain a Phe at position 156 that occupies an alternative buried position compared to solvent exposed hydrophilic residues in other LOV proteins.
Tables
Kinetics of thermal reversion for LOV constructs and variants at 296 K. Uncertainty is depicted as the standard deviation from three replicates.
| Construct | Time Constant, 1/k3 (hrs) |
|---|---|
| WT ZTL 29–165 | 1.4 ± 0.1 |
| G80R | 6.6 ± 0.1 |
| V48I | 10.7 ± 0.8 |
| G46S:G80R | 21 ± 3 |
| V48I:G80R | 65 hr<τ |
Data collection and refinement statistics (molecular replacement).
| WT ZTL Dark | G80R Dark | ZTL-Dark V48I:G80R | ZTL-Light V48I:G80R | |
|---|---|---|---|---|
| PDB ID Data collection | 5SVG | 5SVU | 5SVV | 5SVW |
| Space group | P3(1)21 | P3(1)21 | P3(1)21 | P3(1)21 |
| Cell dimensions | ||||
| a, b, c (Å) | 85.0, 85.0, 199.5 | 85.4, 85.4, 200.0 | 85.4, 85.4, 198.8 | 86.2, 86.2, 200.5 |
| abg (°) | 90, 90, 120 | 90, 90, 120 | 90, 90, 120 | 90, 90, 120 |
| Resolution (Å) * | 2.5 (2.59–2.5) | 2.6 (2.69–2.6) | 2.10 (2.18–2.1) | 2.29 (2.35–2.29) |
| Rsym or Rmerge | 7.6 (27.9) | 12.8 (23.4) | 5.6 (25.1) | 6.1 (19.9) |
| I / sI | 35.0 (11.1) | 55.4 (22.7) | 36.6 (12.6) | 19.8 (5.5) |
| Completeness (%) | 97.1 (98.6) | 97.9 (99.8) | 99.3 (99.0) | 87.0 (84.3) |
| Redundancy | 9.0 | 10.0 | 7.9 | 2.6 |
| Refinement | ||||
| Resolution (Å) | 2.5 | 2.6 | 2.1 | 2.3 |
| No. reflections | 28790 | 26153 | 49510 | 34582 |
| Rwork / Rfree | 17.8/24.0 | 16.4/23.2 | 16.3/20.0 | 16.2/22.8 |
| No. atoms | ||||
| Protein | 3964 | 3990 | 3986 | 3950 |
| Ligand/ion | 124 | 124 | 148 | 124 |
| Water | 142 | 139 | 343 | 269 |
| B-factors | ||||
| Protein | 47.3 | 46.8 | 34.6 | 37.5 |
| Ligand/ion | 35.2 | 35.4 | 27.6 | 27.5 |
| Water | 46.8 | 42.4 | 41.6 | 39.6 |
| R.m.s. deviations | ||||
| Bond lengths (Å) | 0.014 | 0.014 | 0.012 | 0.014 |
| Bond angles (°) | 1.58 | 1.61 | 1.46 | 1.65 |
| Ramachandran outliers | 2 (0.4%) | 3 (0.62%) | 1 (0.21%) | 1 (0.21%) |
-
*Highest-resolution shell is shown in parentheses.
Period length estimates of CCA1:LUC activity in WT and ZTL variants overexpression plants. See Figure 5—figure supplement 1 for expression levels.
| Genotype | Estimated Period length (hrs) | HA-ZTL protein abundance | *Estimated kdeg PRR5 LD (hrs−1) | *Estimated kdeg PRR5 LL (hrs−1) |
|---|---|---|---|---|
| WT | 24.36 ± 0.40 | ND | 0.3 ± 0.1 | 0.13 ± 0.1 |
| 35S:HA-ZTL #7 | 23.69 ± 1.23 | 1 | 0.5 ± 0.1 | 0.13 ± 0.1 |
| 35S:HA-ZTL #17 | 23.65 ± 0.72 | 1.5 ± 0.3 | ND | ND |
| 35S:HA-ZTL (V48I) #7 | ND | 11 ± 4 | 0.8 ± 0.1 | 0.5 ± 0.2 |
| 35S:HA-ZTL (V48I) #22 | ND | 14 ± 5 | ND | ND |
| 35S:HA-ZTL (G80R) #22 | 24.17 ± 1.08 | 3 ± 0.5 | 0.34 ± 0.05 | 0.14 ± 0.1 |
| 35S:HA-ZTL (G80R) #23 | 19.33 ± 1.06 | 12 ± 5 | ND | ND |
| 35S:HA-ZTL (V48I:G80R) #45 | ND | 10 ± 3 | 0.8 ± 0.2 | 0.3 ± 0.2 |
| 35S:HA-ZTL (V48I:G80R) #49 | ND | 7.6 ± 2 | ND | ND |
-
*Estimated kdeg values were extracted by fitting Equation S10 (below) to the PRR5 protein levels in vivo. For LD conditions, an average k is obtained by treating the system as only containing dark-state protein. Thus, the LD values are accurate as comparative terms between variants only.





