Chlamydia interfere with an interaction between the mannose-6-phosphate receptor and sorting nexins to counteract host restriction
Figures
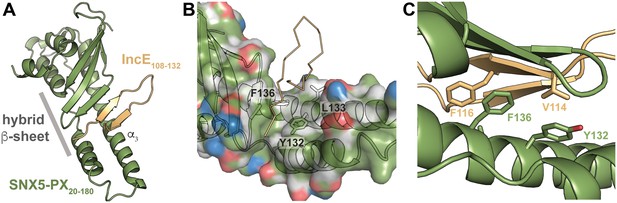
IncE binds in a hydrophobic groove on SNX5-PX, extending the SNX5-PX β-sheet.
(A) The structure of SNX5-PX20-180 (green) bound to IncE108-132 (gold). Binding of IncE to SNX5 leads to the formation of a hybrid β-sheet. (B) Surface and ribbon representation of SNX5-PX showing the hydrophobic binding groove. Atoms are colored according to the scheme described in (Hagemans et al., 2015) to highlight hydrophobic surfaces. Carbon atoms not bound to nitrogen or oxygen atoms are colored grey, oxygens carrying the negative charges in glutamate and aspartate are red and nitrogens carrying the positive charges in lysine and arginine are blue, while all remaining atoms are green. The IncE interacting residues are shown as sticks and labeled. IncE is displayed as gold ribbon. (C) Close-up view of the SNX5-PX20-180:IncE108-132 binding surface with interacting residues shown as sticks.
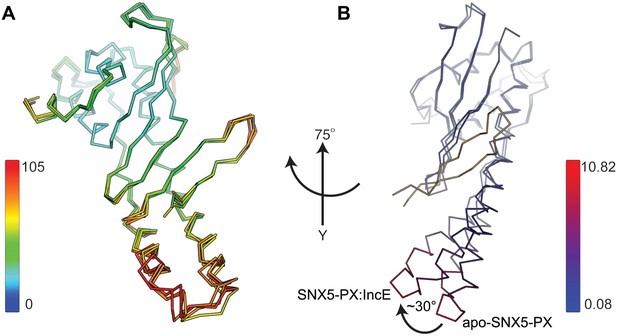
Structural analysis of the SNX5-PX:IncE complex.
(A) Overlay of the four SNX5-PX:IncE complexes present in the asymmetric unit. The B-factors of the Cα atoms are indicated by colors along a spectrum as shown in the figure. (B) Overlay of the SNX5-PX:IncE complex with the apo-SNX5-PX core structure (PDB ID 3HPC). The structures are colored by their root-mean-square deviation (RMSD) indicated by a color gradient from blue to red. The average RMSD is 1.77 Å. Molecules are rotated 75° around the y-axis compared to panel A.
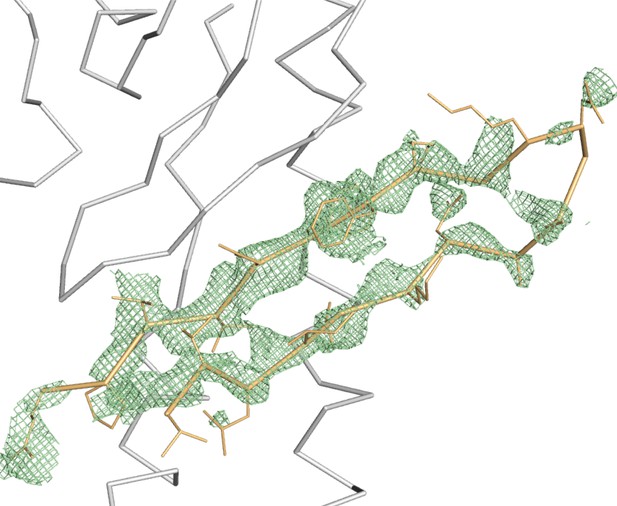
Difference density map reveals IncE108-132.
The refined Fo-Fc density map after molecular replacement, contoured at 2σ, shows positive density not present in the molecular replacement model. SNX5-PX20-180 and the final model of the IncE peptide are shown as grey and gold ribbons, respectively.
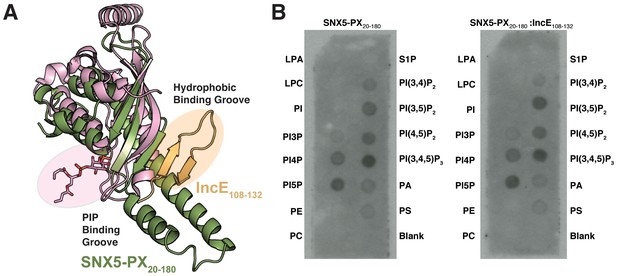
The hydrophobic binding groove of SNX5-PX is distant from the PIP binding groove.
(A) Overlay of the SNX5-PX:IncE structure with the crystal structure of the PX domain from p40phox bound to phosphatidylinositol 3-phosphate (PDB ID 1H6H) (Bravo et al., 2001). SNX5-PX and IncE are colored green and gold, respectively. The PX domain from p40phox is colored pink and the phosphatidylinositol 3-phosphate is shown as sticks with carbons colored pink. (B) IncE does not affect SNX5 binding to immobilized lipids. Purified SNX5-PX20-180 alone or in complex with IncE108-120 was incubated with membranes containing various phospholipids as described in Methods. SNX5 binding was detected using anti-SNX5 antibody. A blot representative of two independent biological experiments is shown. LPA, lyosophophatidic acid. LPC, lysophosphocholine. PtdIns, phosphatidylinositol. PI(3)P, phosphatidylinositol (3) phosphate. PI(4)P, phosphatidylinositol (4) phosphate. PI(5)P, phosphatidylinositol (5) phosphate. PE, phosphatidylethanolamine. PC, phosphatidylcholine. S1P, sphingosine 1-phosphate. PI(3,4)P2, phosphatidylinositol (3,4) bisphosphate. PI(3,5)P2, phosphatidylinositol (3,5) bisphosphate. PI(4,5)P2, phosphatidylinositol (4,5) bisphosphate. PI(3,4,5)P3, phosphatidylinositol (3,4,5) triphosphate. PA, phosphatidic acid. PS, phosphatidylserine.
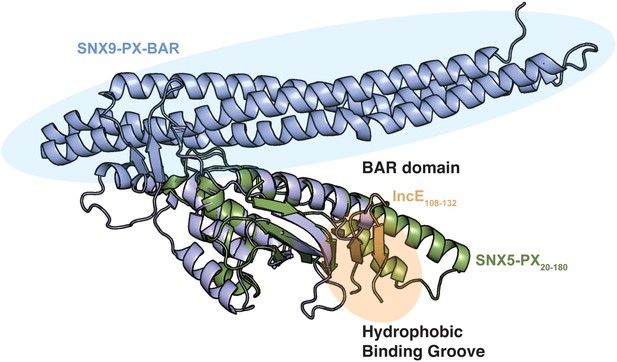
Overlay of the SNX5-PX:IncE structure with the structure of the SNX9-PX-BAR domain.
The color scheme for SNX5 and IncE is the same as in Figure 1. SNX9 (PDB ID 2RAK) (Pylypenko et al., 2007) is colored blue. The structure of SNX5-PX:IncE is rotated 90˚ around the x-axis compared to Figure 1—figure supplement 3A.
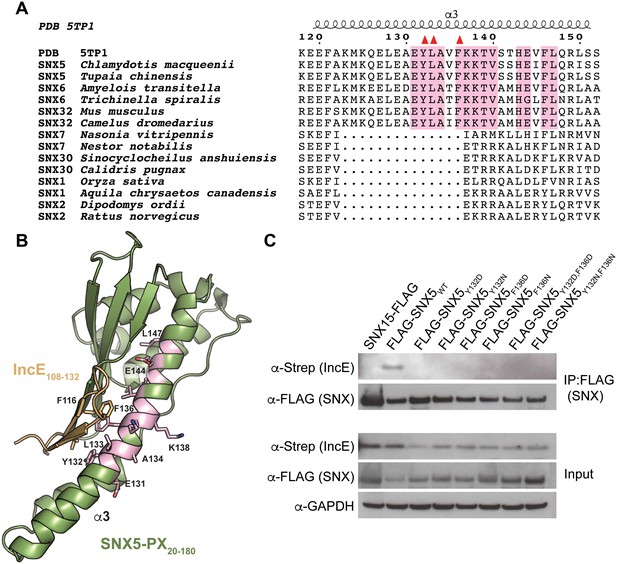
The IncE binding residues are highly conserved among the SNX5/6/32 family of proteins and implicated in protein-protein interactions.
(A) Alignment of selected SNX family members showing the conservation of the IncE-interacting residues Y132, L133 and F136 (red triangles) in SNX5/6/32 family members and their divergence in related SNX-BAR proteins. The only invariant residues in all SNX5/6/32 sequences are highlighted in pink. (B) Structural representation of the invariant residues (pink) shown in A. (C) HEK293T cells were transiently co-transfected with the indicated full-length FLAG-tagged SNXWT constructs and with Strep-tagged IncE101-132. Lysates were immunoprecipitated with anti-FLAG beads and immunoblotted with the indicated antibodies. Input represents 1% of lysates. SNX15 serves as a negative control. The data shown is representative of two independent biological experiments.
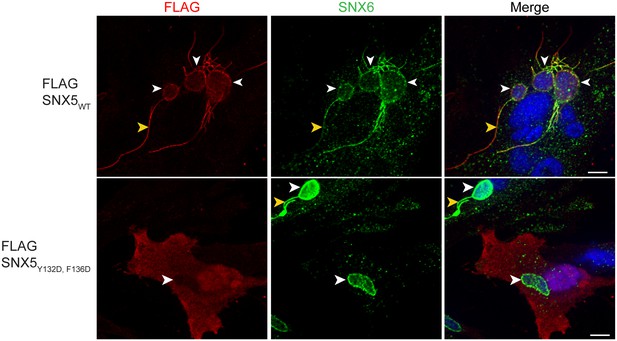
The hydrophobic binding groove of SNX5-PX is required for recruitment of SNX5 to C. trachomatis inclusions.
HeLa cells transiently expressing full-length FLAG-SNX5WT or FLAG-SNX5Y132D,F136D were infected with C. trachomatis for 24 hr and analyzed by confocal microscopy for the localization of FLAG-SNX5 and endogenous SNX6. Shown are single z slices. White arrowheads point to inclusions. Yellow arrowheads point to tubules. Scale bar, 10 µm. The data shown is representative of two biological replicates.
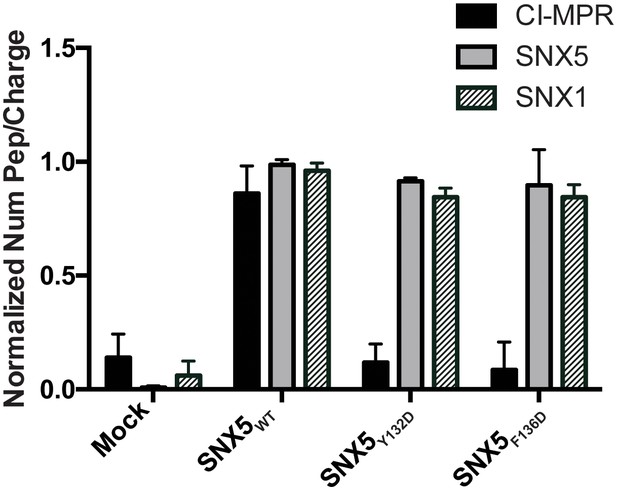
Comparative AP-MS for transfected SNX5WT, SNX5Y132D, and SNX5F136D.
The Y-axis quantifies the number of peptides/charge detected by LC-MS/MS for CI-MPR, SNX5, and SNX1 that affinity purified with the indicated full-length SNX5 proteins. Mock refers to untransfected cells. The graph is normalized to SNX5WT. Whereas SNX1 is equally well represented in the affinity-purified lysates from WT and mutant SNX5, the representation of CI-MPR peptides differed significantly between SNX5WT and each of the SNX5 mutants. Data are mean +/− SD from three independent biological experiments.
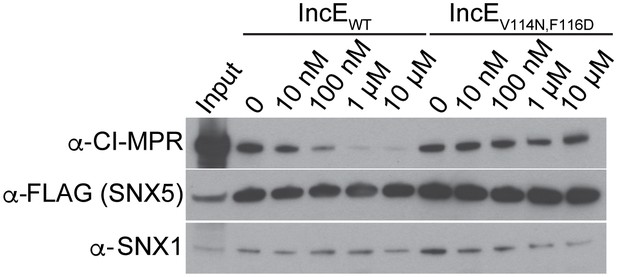

IncE108-132 interferes with CI-MPR binding to SNX5-associated complexes.
Lysates from HEK293T cells transiently expressing full-length FLAG-SNX5WT were incubated with anti-FLAG beads in the presence of the indicated concentrations of wild type or mutant IncE108-132 under non-equilibrium conditions. Eluates were immunoblotted with the indicated antibodies. Input represents 1% of lysates. WT but not mutant IncE interferes with binding of CI-MPR to FLAG-SNX5-containing complexes. The data shown is representative of three biological experiments.
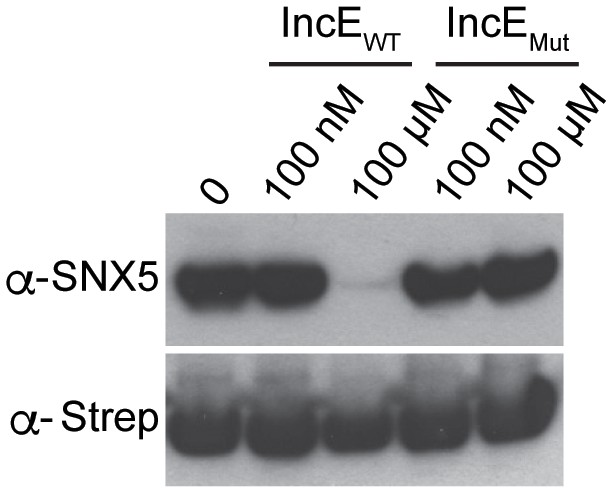
IncE peptide interferes with IncE101-132 binding to SNX5-PX20-180 in vitro.
Purified 6xHis-MBP-IncE101-132-Strep was immobilized to Strep-Tactin beads and incubated with purified 8xHis-SNX5-PX20-180 in the presence of the indicated concentrations of WT IncE108-132 or IncEV114N,F116D (Mut) under non-equilibrium conditions. Eluates were immunoblotted with the indicated antibodies. WT but not mutant IncE peptide interferes with binding of IncE to SNX5-PX.
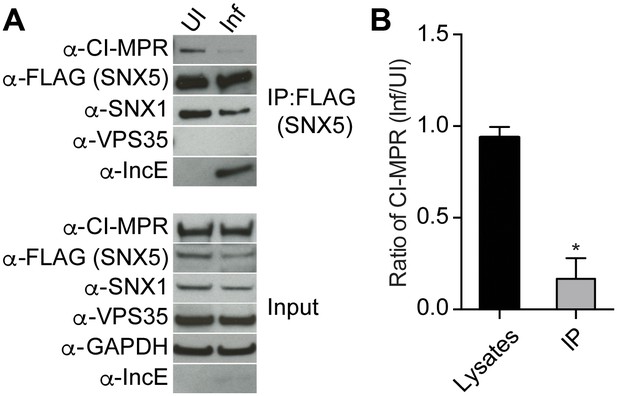
C. trachomatis infection interferes with CI-MPR binding to SNX5 complexes.
(A) HeLa cells were transiently transfected with full-length FLAG-SNX5WT for 24 hr and then were infected (Inf) or left uninfected (UI) with C. trachomatis for 24 hr. FLAG-SNX5WT was immunoprecipitated and eluates were immunoblotted with the indicated antibodies. Input is 1% of lysates used for immunoprecipitation. (B) Quantitation of the ratio of CI-MPR from infected to uninfected cells in lysates or in eluates from FLAG-SNX5 immunoprecipitations (IP). Data are mean +/− SD from three independent biological experiments. *p=0.0004 compared to lysates, unpaired t-test.
Tables
SNX5 protein-protein interactions and their dependence on the IncE binding groove. Shown is the average of the spectral counts from the listed proteins detected in the affinity purified lysates prepared (in triplicate) from untransfected cells (mock) or from cells transfected with full-length SNX5WT, SNX5F136D, or SNX5Y132D mutants. Only interactions exhibiting a BFDR ≤ 0.1 when compared to affinity purified lysates from untransfected cells are listed. The proteins that have a SAINT score (Teo et al., 2014) of 1 (the highest probability possible) are highlighted in green and the SAINT score is shown in parentheses. Of the proteins found to interact with SNX5WT, CI-MPR and IGF1R show a difference in their SAINT score between the WT and the mutants, suggesting that the mutations block the protein-protein interaction. (ND, not determined due to lack of spectral counts in data set).
| ProtID | Name | Mock | SNX5 WT | SNX5 F136D | SNX5 Y132D |
|---|---|---|---|---|---|
| Q9Y5X3 | SNX5_HUMAN | 4.33 | 134 (1) | 118.33 (1) | 121.67 (1) |
| Q13596 | SNX1_HUMAN | 0 | 70 (1) | 64 (1) | 64 (1) |
| O60749 | SNX2_HUMAN | 1 | 105.5 (1) | 101 (1) | 99.67 (1) |
| P11717 | CI-MPR_HUMAN | 0 | 12.67 (1) | 0.33 (0) | 0 (0) |
| P08069 | IGF1R_HUMAN | 0 | 8.67 (1) | ND | ND |
| P30048 | PRDX3_HUMAN | 0.33 | 4.67 (0.65) | 0.33 (0) | 1.33 (0.07) |
| Q15773 | MLF2_HUMAN | 0 | 1.67 (0.33) | 5.33 (0.98) | 2.67 (0.56) |
Additional files
-
Supplementary file 1
MaxQuant output for affinity purification experiments described in Figure 3.
Files are organized by the ‘RawFile’ names and describe LC-MS/MS runs recorded from three biological replicates. The file names are as follows: mock interactions (RawFiles FU20160603-24, FU20160603-32 and FU20160603-02), WT SNX5 interactions (RawFiles FU20160603-22, FU20160603-26m, and FU20160603-04) SNX5Y132D interactions (RawFiles FU20160603-20, FU20160603-34, and FU20160603-06) and SNX5F136D (RawFiles FU20160603-18, FU20160603-28, and FU20160603-08).
- https://doi.org/10.7554/eLife.22709.014
-
Supplementary file 2
Summary of results from SNX5 affinity purification and mass spectrometry.
Columns C-N show the number of peptide-charge state pairs (Num Pep/Charge) and columns O-Z show the sum of all peptide intensities (Intensity) for each protein detected in each biological replicate as indicated. Proteins are identified by UniProt accession numbers; entries with multiple UniProt accession numbers represent those identified by peptide sequences shared by multiple protein sequences.
- https://doi.org/10.7554/eLife.22709.015
-
Supplementary file 3
Data collection and refinement statistics.
- https://doi.org/10.7554/eLife.22709.016
-
Supplementary file 4
List of primers.
- https://doi.org/10.7554/eLife.22709.017





