Human cardiac fibroblasts adaptive responses to controlled combined mechanical strain and oxygen changes in vitro
Figures
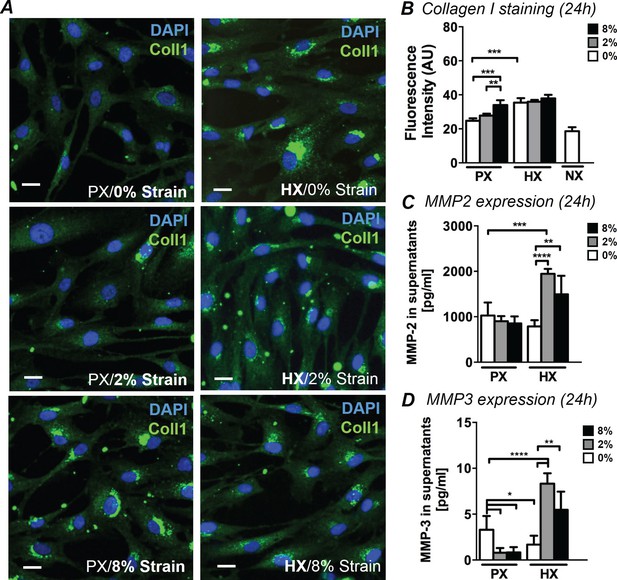
Analyses of early ECM adaptive responses by CFs subject to combined environmental stimulation.
(A) Representative images of CFs fixed after 24 hr stimulations and stained for collagen I (green) and DAPI (blue). Scale bars = 20 μm. An intense perinuclear staining is observed in CFs stimulated either by 8% mechanical strain or by HX. (B) Quantitative fluorescence intensity analyses on Collagen I staining plotted as graph. Collagen I staining is increased by HX, 8% strain or NX conditions. Data collected from one cell donor, two independent experiments, minimum number of experimental replicates n = 4, technical replicates (multiple images per replicate) n = 3. Detected amounts of MMP-2 (C) and MMP-3 (D) in supernatants of CFs. Both MMP-2 and MMP-3 expression is significantly increased in combined HX and mechanical strain stimulation. Protein secretion data collected from one cell donor, two independent experiments, minimum number of experimental replicates n = 4, technical replicates n = 2. White histograms correspond to 0% strain conditions, grey histograms to 2% strain conditions and black histograms to 8% strain conditions. Two-way ANOVA test was performed for all groups. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001. One outlier measurement in MMP-3 expression was detected by performing Grubb’s test (α = 0.05; p<0.05) and removed from the analysis.
-
Figure 1—source data 1
Collagen intensity levels.
- https://doi.org/10.7554/eLife.22847.004
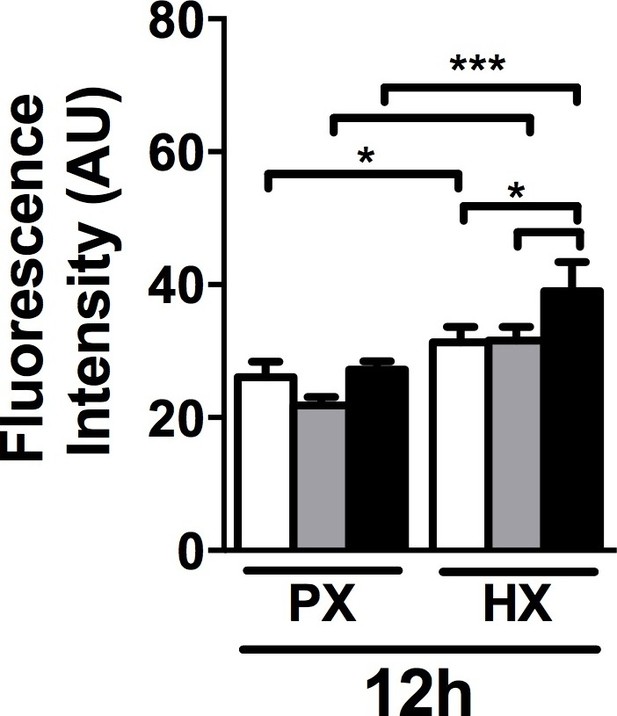
Quantitative fluorescence intensity analyses on collagen I staining performed on CFs fixed after 12 hr of combined stimulation.
Collagen I intensity is increased by HX condition with a more prominent effect when HX was combined to 8% cyclic strain. Data collected from one cell donor, two independent experiments, minimum number of experimental replicates n = 4, technical replicates (multiple images per replicate) n = 3. White histograms correspond to 0% strain conditions, grey histograms to 2% strain conditions and black histograms to 8% strain conditions. Two-way ANOVA test was performed. *p<0.05, ***p<0.001.
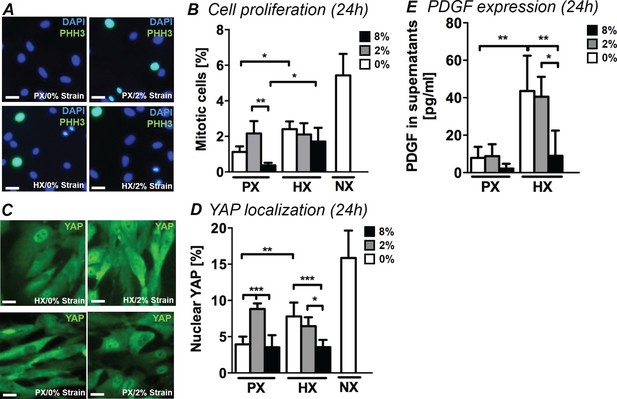
Analyses of CFs proliferation under combined environmental stimulation.
(A) Representative images of CFs fixed after 24 hr stimulations and stained for PHH3 (green) and DAPI (blue). Scale bars = 50 μm. (B) Quantitative analyses of mitotic cell fraction plotted as graph. Cell mitosis is increased by either HX alone and by 2% strain at PX. (C) Representative images of CFs fixed after 24 hr stimulations and stained for YAP (green) and DAPI (blue). Scale bars = 50 μm. (D) Quantitative analyses of cells presenting nuclear YAP plotted as graph. YAP translocation into nuclei is increased by either HX or by 2% strain at PX. Data in Panels B and D were collected from one cell donor, two independent experiments, minimum number of experimental replicates n = 4, technical replicates (multiple images per replicate) n = 3. (E) Detected amounts of PDGF in supernatants of CFs plotted as graph. PDGF expression is significantly increased in HX alone and HX combined with mechanical strain stimulation (2% strain). Protein secretion data collected from one cell donor, two independent experiments, minimum number of experimental replicates n = 4, technical replicates n = 2. White histograms correspond to 0% strain conditions, grey histograms to 2% strain conditions and black histograms to 8% strain conditions. Two-way ANOVA test was performed for mitotic cells and nuclear YAP, whereas Kruskal-Wallis test was performed on PDGF expression. *p<0.05, **p<0.01, ***p<0.001.
-
Figure 2—source data 1
Mitotic cells and YAP localization data.
- https://doi.org/10.7554/eLife.22847.007
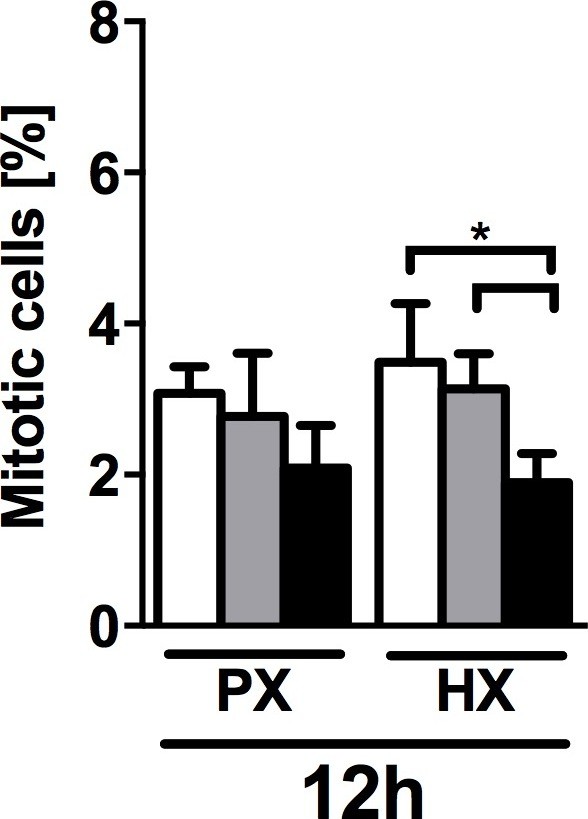
Quantification of mitotic cells (PHH3+/DAPI) performed on CFs fixed after 12 hr of combined stimulation (PX, HX and mechanical strain).
Cell mitosis is negatively regulated by 8% cyclic strain combined with HX condition. Data collected from one cell donor, two independent experiments, minimum number of experimental replicates n = 4, technical replicates (multiple images per replicate) n = 3. White histograms correspond to 0% strain conditions, grey histograms to 2% strain conditions and black histograms to 8% strain conditions. Two-way ANOVA test was performed. *p<0.05.
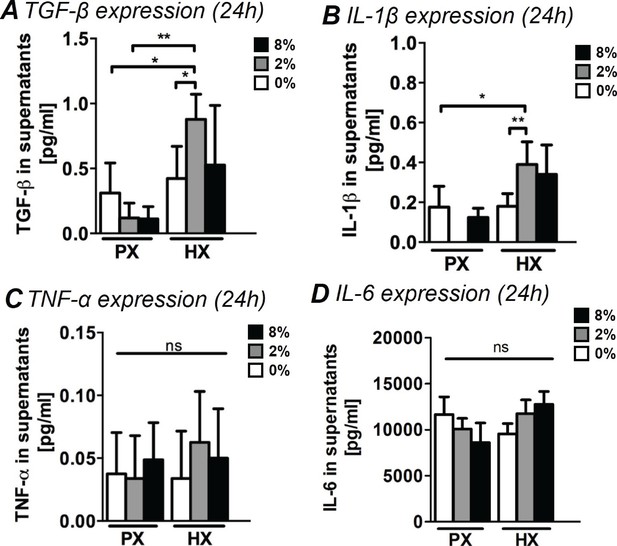
Analyses of pro-fibrotic and pro-inflammatory cytokines in supernatants of CFs.
(A) TGF-β expression in supernatants. TGF-β expression resulted significantly increased by the combination of HX and mechanical strain (2% strain). (B) Expression of IL-1β in CFs supernatants. IL-1β expression resulted significantly upregulated by the combination of HX and mechanical strain (2% strain). (C) TNF-α expression in supernatants. Extremely low amounts were detected and environmental stimulations do not induce significant differences in the expression. (D) IL-6 expression in supernatants. IL-6 resulted abundantly expressed by CFs with no statistically significant differences induced by environmental stimulations. Protein secretion data from each marker was collected from one cell donor, two independent experiments, minimum number of experimental replicates n = 4, technical replicates n = 2. White histograms correspond to 0% strain conditions, grey histograms to 2% strain conditions and black histograms to 8% strain conditions. Kruskal-Wallis tests were performed for all groups except for IL-6 analyzed with Two-way ANOVA tests. *p<0.05, **p<0.01, ns = non-significant.
-
Figure 3—source data 1
Protein expression data.
- https://doi.org/10.7554/eLife.22847.010
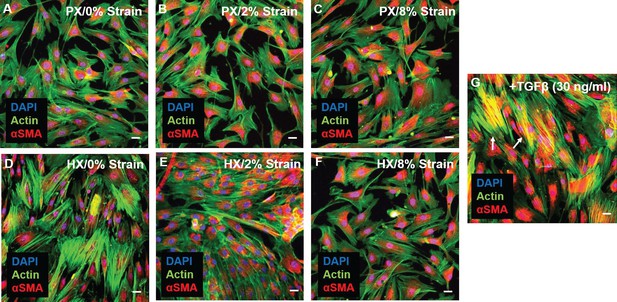
Representative images of CFs fixed after 24 hr combined stimulations (A–F) and stained for actin (green) and αSMA (red), nuclei were counterstained with DAPI (blue).
Scale bars = 20 μm. αSMA is similarly expressed in all conditions and diffused in perinuclear and cytoplasmic localization. (G) Positive control for αSMA immunofluorescence through stimulation with 30 ng/ml TGF-β: CFs exhibit co-localization of αSMA staining with actin stress fibers (resulting in yellow signal) denoting differentiation into myofibroblasts (white arrows). Representative images were collected from a pool of images obtained from one cell donor, one independent experiment, minimum number of experimental replicates n = 4, technical replicates n = 3.
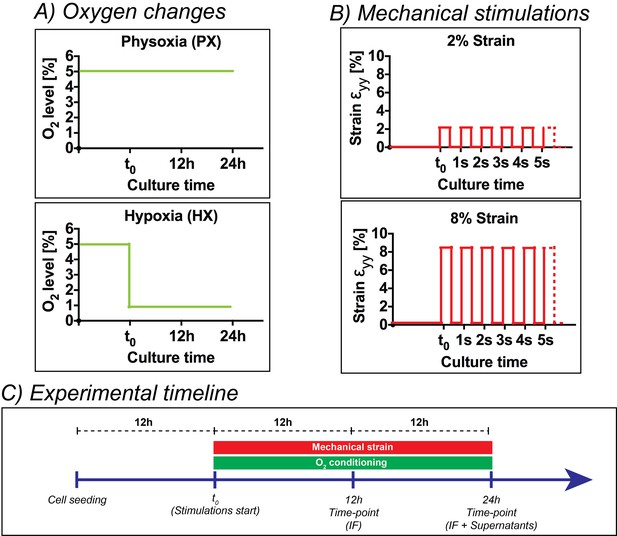
Overview of the experimental design of the present work.
(A) Oxygen changes reproduced in the present work: an incubation at PX; an abrupt reduction to HX environments (1% O2) maintained for 24 hr. (B) Mechanical stimulation regimes investigated in the present work: a static control at constant 0% strain; a cyclic mechanical strain stimulation at 2% strain and 1 Hz frequency; a cyclic mechanical strain stimulation at 8% strain and 1 Hz frequency. (C) Experimental timeline of the experiments performed: cells were seeded on microdevices and allowed to adhere to the culture membranes for 12 hr at PX and 0% strain before starting the combination of environmental stimuli. After 12 hr from the beginning of stimulations, we fixed samples for immunofluorescence analyses and after 24 hr we fixed samples for immunofluorescence analyses and collected supernatants for protein expression quantifications.
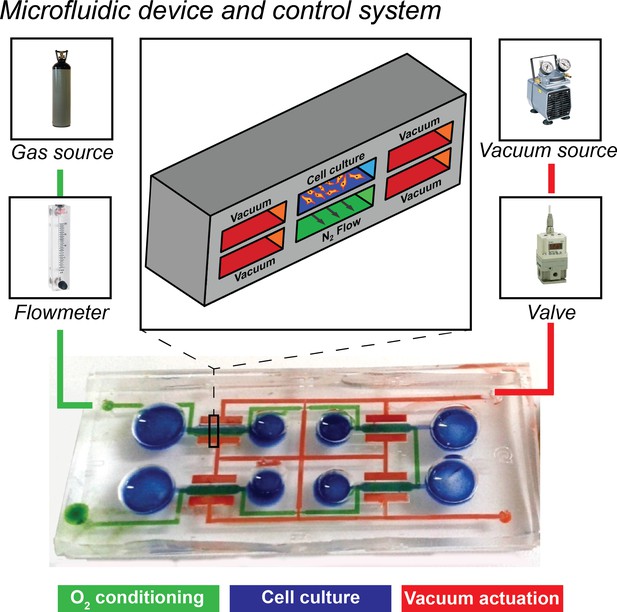
Overview of the microfluidic platform employed in the present work together with control system.
A microdevice is filled with color dyes to highlight the microfluidic circuits. Blue channels represent the four cell culture chambers, green channel represents the lower circuit flowing below all four culture chambers meant for oxygen conditioning, red channel represent vacuum actuation circuit for application of mechanical strain to cell cultures. A 3D sketch of the cross section of a single culture chamber is shown with the same color code (upper inset). Elements of the control system are also highlighted: vacuum system is actuated by connecting a vacuum source to a switching valve regulating the amount of cyclic vacuum to be delivered to the devices; oxygen conditioning circuit is actuated by connecting a nitrogen gas source to a flowmeter to regulate the flow of deoxygenating gas through the lower conditioning channel (green).
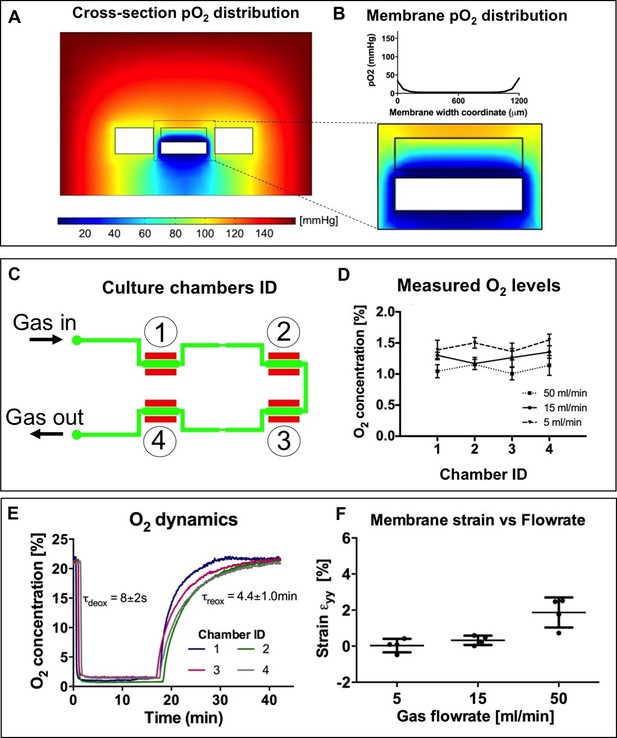
Numerical modeling and experimental characterization of oxygen conditioning system included in the microdevices employed for the present work.
(A) Spatial distribution of oxygen tension in a culture chamber cross-section upon application of a fixed oxygen tension (0 mmHg) at the lower channel boundaries simulating the flow of deoxygenating nitrogen gas. The side vacuum chamber boundaries are also set at a fixed oxygen tension to simulate cyclic vacuum application. Considering the membrane region, where cells are cultured (B), the oxygen tension is uniformly distributed with limited border effects near the channel walls. (C) Outline of the lower layer of the microdevices showing conditioning gas inlets and outlets together with numbered culture chambers. (D) Experimental measurements of near-membrane oxygen concentrations performed by means of a needle-based oxygen sensor on all four culture chambers of n = 4 microdevices upon application of varying flowrates (5, 15 and 50 ml/min) of gas in the lower conditioning channel. Oxygen levels reach approximately 1% uniformly throughout all four chambers of the microdevices. (E) Experimental measurements of near-membrane oxygen concentrations performed on all four culture chambers of n = 4 microdevice with respect to time (one measurement every 3 s). Deoxygenation time constant, representing the application of conditioning gas flowrate, results in the order of seconds; reoxygenation time constant, representing the interruption of conditioning gas flowrate, results in the order of minutes. (F) Impact of conditioning gas flowrates on the resulting membrane strain when no vacuum is applied. No membrane strain is observed for 5 ml/min or 15 ml/min flowrates while 50 ml/min gas flowrates cause membrane strain of approximately 2%.
-
Figure 5—figure supplement 2—source data 1
O2 concentrations data.
- https://doi.org/10.7554/eLife.22847.018
Tables
Summary of the main cellular responses observed in experiments of combined environmental stimulation. Cellular responses were included in the corresponding stimulation condition if the measured parameter was significantly different from the parameter at reference condition PX(5% O2)/0% strain (single arrow ↑) and if more than two-fold significant difference was observed compared to PX(5% O2)/0% strain condition (double arrow ↑↑). Different background cell color are added as a function of the number of significant cellular responses observed (yellow: 2 or less changes; orange: up to 5 changes; red: more than 5 changes).
| 0% Strain | 2% Strain | 8% Strain | |
|---|---|---|---|
| HX (1% O2) | ↑↑ Mitosis ↑↑ Nuclear YAP ↑↑ PDGF ↑ Collagen ↓ MMP-3 | ↑ Mitosis ↑ Nuclear YAP ↑↑ PDGF ↑ Collagen ↑↑ MMP-2–3 ↑↑TGF-β ↑ IL-1β | ↑ Collagen ↑ MMP-2 |
| PX (5% O2) | Reference condition | ↑ Mitosis ↑↑ Nuclear YAP ↓ MMP-3 | ↑ Collagen ↓ MMP-3 |
Summary of literature studies investigating early CFs responses under standard pro-fibrotic and inflammatory stimulation. Quantitative fold-changes of selected markers (fibrotic vs. control condition) are reported for studies that carried out protein expression assays or proliferation assays at early stimulation timepoints and compared to our on-chip model (grey background). The on-chip model well correlates to standard in vitro models in terms of ECM remodeling markers (Collagen 1, MMPs), cell proliferation and pro-fibrotic or inflammatory cytokynes (TGF-β and IL-1β). The on-chip model did not elicit production of IL-6 and TNF-α as compared to standard in vitro models. Assays: WB=Western Blot; ELISA= Enzyme-linked immunosorbent assay; HP= Hydroxyproline assay; CCK-8= Cell-counting kit eight proliferation assay; FACS= Fluorescence-activated cell sorting; Count= Standard cell counting; BrdU= Bromodeoxyuridine proliferation assay. Chemical stimuli: Ang II= Angiotensin II; TGF-β= Transforming Growth Factor-β; IL-1α/IL-1β= Interleukin-1α/1β; TNF-α= Tumor Necrosis Factor-α. Source data file linked to Table 2 shows this comparison data plotted in a chart.
Collagen Expression | ||||||
| Ref. | Species | Assay | Stimulus | Culture Time | Change vs. control (standard model) | Change vs. control (24 hr on-chip model) |
| (Xiao et al., 2016) | Mouse | WB | Ang II | 24 hr | ↑1.5-fold | ↑1.45-fold |
| (Guo et al., 2016) | Rat | WB | TGF-β | 48 hr | ↑2.2-fold | |
| (Pan et al., 2013) | Rat | WB | TGF-β | 24 hr | ↑1.5-fold | |
| (Peng et al., 2010) | Human | HP | TGF-β | 48 hr | ↑1.6-fold | |
| (Li et al., 2015) | Rat | WB | Ang II | 24 hr | ↑2.1-fold | |
MMP-2 Expression | ||||||
| Ref. | Species | Assay | Stimulus | Culture Time | Change vs. control (standard model) | Change vs. control (24 hr on-chip model) |
| (Xiao et al., 2016) | Mouse | WB | Ang II | 24 hr | ↑1.8-fold | ↑2-fold |
| (Rhaleb et al., 2013) | Rat | WB | IL-1β | 72 hr | ↑2-fold | |
| (Brown et al., 2007) | Rat | WB | IL-1 | 48 hr | ↑2.2-fold | |
| (Li et al., 2015) | Rat | WB | Ang II | 24 hr | ↑2-fold | |
| MMP-3 Expression | ||||||
| Ref. | Species | Assay | Stimulus | Culture Time | Change vs. control (standard model) | Change vs. control (24 hr on-chip model) |
| (van Nieuwenhoven et al., 2013 ) | Human | WB | IL-1α | 24 hr | ↑2-fold | ↑2.7-fold |
| (Brown et al., 2007) | Rat | WB | IL-1 | 48 hr | ↑2-fold | |
| Cell proliferation | ||||||
| Ref. | Species | Assay | Stimulus | Culture Time | Change vs. control (standard model) | Change vs. control (24 hr on-chip model) |
| (Xiao et al., 2016) | Mouse | CCK-8 | Ang II | 24 hr | ↑2-fold | ↑1.9-fold |
| (Guo et al., 2016) | Rat | FACS | TGF-β | 48 hr | ↑2-fold | |
| (Porter et al., 2004) | Human | Count | TNF-α | 4 days | ↑1.5-fold | |
| (Dobaczewski et al., 2010) | Mouse | BrdU | TGF-β | 24 hr | ↓2-fold | |
| (Vivar et al., 2016) | Rat | FACS | TGF-β | 72 hr | ↓2-fold | |
| (Ai et al., 2015) | Rat | CCK-8 | Ang II | 48 hr | ↑1.8-fold | |
| (Li et al., 2015) | Rat | MTT | Ang II | 24 hr | ↑2-fold | |
| TGF- β Expression | ||||||
| Ref. | Species | Assay | Stimulus | Culture Time | Change vs. control (standard model) | Change vs. control (24 hr on-chip model) |
| (Gu et al., 2012) | Mouse | WB | Ang II | 24 hr | ↑2-fold | ↑2.7-fold |
| (Xiao et al., 2016) | Mouse | WB | Ang II | 24 hr | ↑1.8-fold | |
| (Ai et al., 2015) | Rat | ELISA | Ang II | 48 hr | ↑2.5-fold | |
| (Li et al., 2015) | Rat | WB | Ang II | 24 hr | ↑1.9-fold | |
| IL-1β Expression | ||||||
| Ref. | Species | Assay | Stimulus | Culture Time | Change vs. control (standard model) | Change vs. control (24 hr on-chip model) |
| (Turner et al., 2009) | Human | ELISA | IL-1α | 24 hr | ↑8-fold | ↑2.2-fold |
| (Turner et al., 2009) | Human | ELISA | TNF-α | 24 hr | ↑2-fold | |
| TNF-α Expression | ||||||
| Ref. | Species | Assay | Stimulus | Culture Time | Change vs. control (standard model) | Change vs. control (24 hr on-chip model) |
| (Turner et al., 2009) | Human | ELISA | IL-1α | 24 hr | ↑4-fold | ns |
| (Humeres et al., 2016) | Rat | Luminex | TGF-β | 72 hr | ↓4-fold | |
| (Yokoyama et al., 1999) | Rat | ELISA | Ang II | 6h | ↑5-fold | |
| IL-6 Expression | ||||||
| Ref. | Species | Assay | Stimulus | Culture Time | Change vs. control (standard model) | Change vs. control (24 hr on-chip model) |
| (Turner et al., 2009) | Human | ELISA | IL-1α | 24 hr | ↑19-fold | ns |
| (Turner et al., 2009) | Human | ELISA | TNF-α | 24 hr | ↑3.5-fold | |
| (Turner et al., 2007) | Human | ELISA | TNF-α | 24 hr | ↑2.8-fold | |
-
Table 2—source data 1
Data from Table 2 plotted as chart.
- https://doi.org/10.7554/eLife.22847.014





