The role of PDF neurons in setting the preferred temperature before dawn in Drosophila
Figures
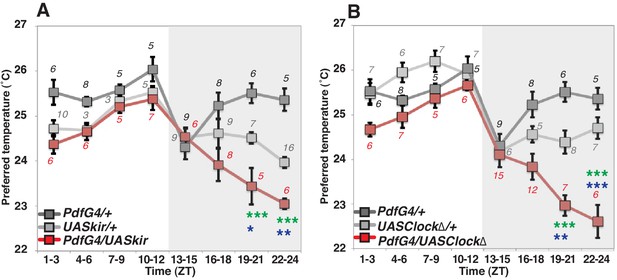
PDF neurons are required for preferred temperature before dawn.
(A) TPR of Pdf-Gal4/UAS-Kir2.1 (red), Pdf-Gal4/+ (dark gray) and UAS-Kir2.1/+ (light gray) flies over 24 hr. Numbers represent the number of assays. Pdf-Gal4 drives sLNvs and lLNvs. (B) TPR of Pdf-Gal4/UAS-Clock∆ (red), Pdf-Gal4/+ (dark gray) and UAS-Clock∆/+ (light gray) flies over 24 hr. Preferred temperatures were calculated using the distribution of flies by temperature preference behavior. Data are shown as the mean preferred temperature in each time zone (ZT1-3, 4–6, 7–9, 10–12, 13–15, 16–18, 19–21 and 22–24.) The pre-dawn time is ZT19-21 and ZT22-24. Zeitgeber Time (ZT; 12 hr light/dark cycle; ZT0 is lights-ON, ZT12 is lights-OFF). Numbers represent the number of assays. The preferred temperatures among Gal4/UAS, Gal4/+ and UAS/+ flies in each time zone were analyzed using One-way ANOVA and Tukey-Kramer tests (Supplementary file 1). Stars indicate p values of Tukey-Kramer tests when Gal4/UAS are statistically different from both Gal4/+ (stars in green) and UAS/+ (stars in blue). ****p<0.0001, **p<0.01 or *p<0.05.
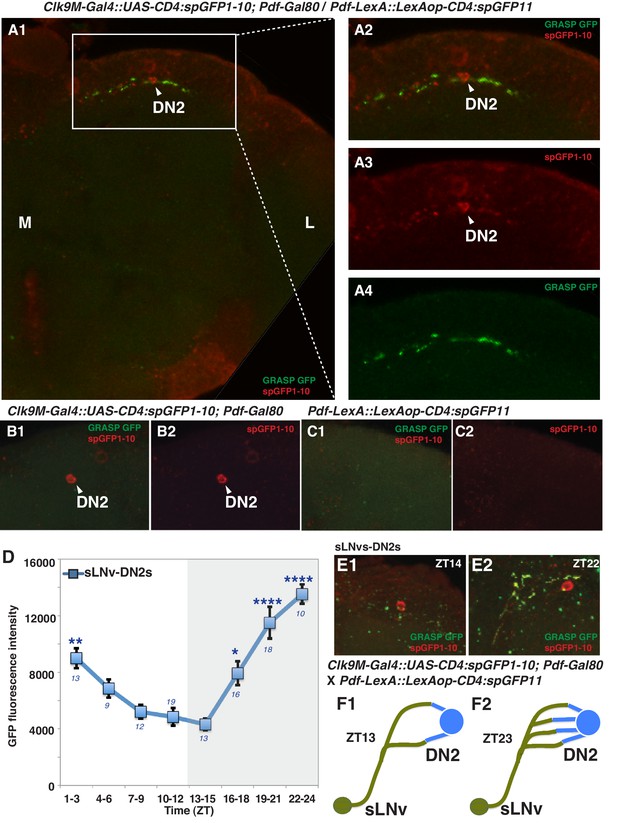
sLNvs contact DN2s, and the number of contacts peaks before dawn.
(A1) GRASP between DN2s and sLNvs. The terminal area of sLNvs in dorsal lateral brain was magnified. The reconstituted GFP signals (green) were only detected in the dorsal protocerebrum, but not in the sLNvs soma nor the other region of their projections. Clk9M-Gal4::UAS-CD4:spGFP1-10; Pdf-Gal80 and Pdf-LexA::LexAop-CD4:spGFP11 flies were used to express split-GFP1-10 in DN2s, and split-GFP11 in LNvs, respectively. When the two fly lines were crossed, a reconstituted GFP signal (green) was detected (A4). The soma and projection of DN2s (red) were detected by anti-GFP (A3). The merged image of A3 and A4 (A2). M: medial, L: lateral. (B,C) Neither of the split GFP fragments alone in DN2s or sLNvs had reconstituted GFP fluorescence signals (no green signal in B and C). (D) Comparisons of GRASP signals between sLNvs and DN2s (blue) throughout the course of the day. GFP fluorescence intensity was measured and the mean values were plotted. Numbers represent the number of GRASP experiments. One-way ANOVA and Tukey-Kramer tests compared GFP fluorescence intensity of each time zone in sLNv-DN2s (blue). The intensity of all the time zones were compared with ZT 13–15 (blue). One-way ANOVA; p<0.0001, F (7, 72)=5.226. One-way ANOVA; ****p<0.0001, F (7, 102)=16.09. (E) A representative image of GRASP between sLNvs and DN2s at ZT14 (E1) and ZT22 (E2). Clk9M-Gal4::UAS-CD4:spGFP1-10; Pdf-Gal80 and Pdf-LexA::LexAop-CD4:spGFP11 flies were used to express split-GFP1-10 in DN2s and split-GFP11 in LNvs, respectively. (F) A schematic of the relationship between sLNvs and DN2s at ZT13 and ZT23. The number of the sLNv-DN2 contacts dramatically fluctuates throughout the day and peak before dawn (ZT22-24) (Figure 2D). At ZT22-24, there are the greatest number of contacts between sLNvs and DN2s.
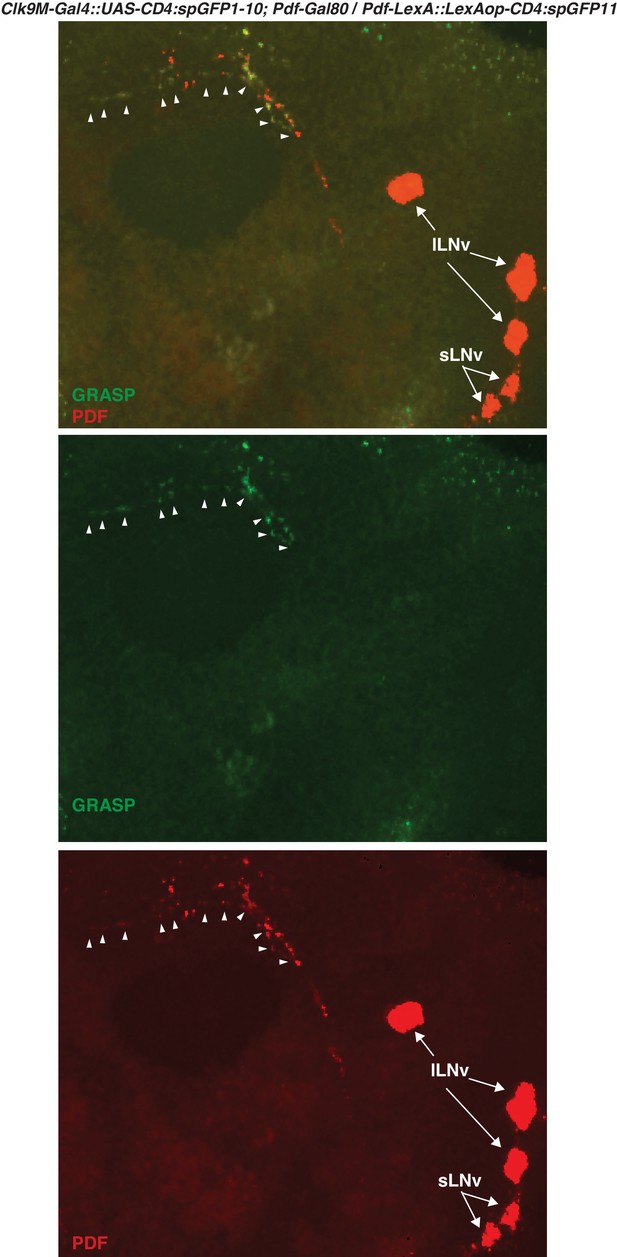
GRASP between DN2s and sLNvs.
Clk9M-Gal4::UAS-CD4:spGFP1-10; Pdf-Gal80 and Pdf-LexA::LexAop-CD4:spGFP11 flies were crossed. The reconstituted GFP signals (green, arrowheads) were only detected in the dorsal protocerebrum, but not in the sLNvs soma nor the other region of their projections. The somas of LNvs (red, arrows) were labeled by anti-PDF (red).
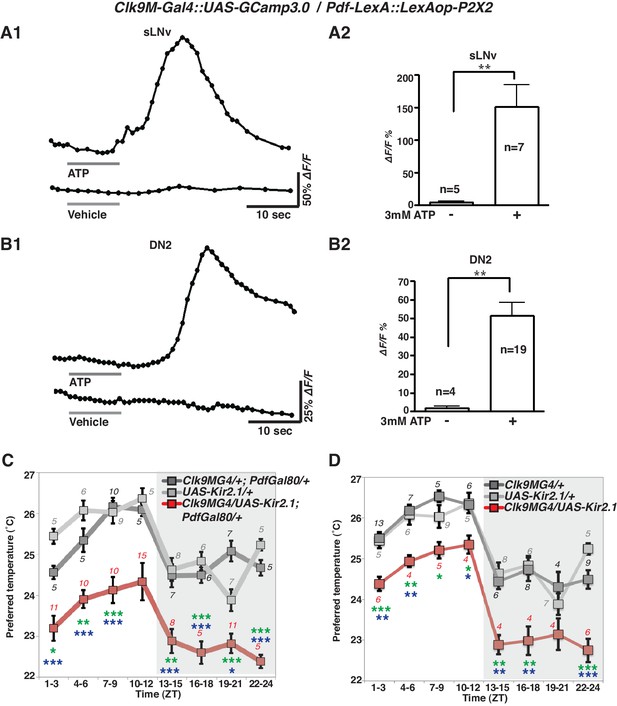
sLNvs activate DN2s, and the loss of DN2 activation results in a lower temperature preference.
(A1) A representative graph of sLNv activation via P2X2 expression in sLNvs. Both P2X2 and GCaMP3.0 were expressed in sLNvs by using Clk9M-Gal4::UAS-GCaMP3.0 / Pdf-LexA:: LexAop-P2X2 flies. The representative trace of GCaMP fluorescence in a sLNv neuron showed a great increase by application of 3 mM ATP into the bath (upper line), but it showed no response to the vehicle control (bottom line). ATP was present in the bath until the end of calcium imaging acquisition. (B1) A representative graph of DN2 activation via P2X2 expression in sLNvs. GCaMP3.0 and P2X2 were expressed in DN2s and sLNvs, respectively, by using Clk9M-Gal4::UAS-GCaMP3.0 / Pdf-LexA:: LexAop-P2X2. The representative trace of GCaMP fluorescence in DN2 neurons showed an excitation following the activation of P2X2 in sLNvs, which are activated by 3 mM ATP perfusion into bath (upper line), but it showed no response to the vehicle control (bottom line). (A2, B2) The bar graph shows mean maximum GCaMP fluorescence increase in sLNvs (A2) and DN2s (B2) to bath-applied ATP and vehicle. Unpaired t-test between ATP and vehicle. Numbers represent the number of experiments. (C) TPR of Clk9M-Gal4/UAS-Kir2.1; Pdf-Gal80/+ (red), Clk9M-Gal4/+; Pdf-Gal80/+ (dark gray) and UAS-Kir2.1/+ (light gray) flies over 24 hr. Data are shown as the mean preferred temperature. Numbers represent the number of assays. Stars indicate p values of Tukey-Kramer tests when Gal4/UAS are statistically different from both Gal4/+ (stars in green) and UAS/+ (stars in blue). ****p<0.0001, **p<0.01 or *p<0.05. One-way ANOVA ZT1-3; p=0.0002, F (2, 20)=14.79. ZT4-6; p<0.0001, F (2, 20)=21.88. ZT7-9; p<0.0001, F (2, 28)=17.75. ZT10-12; p=0.014, F (2, 24)=5.211. ZT13-15; p=0.0004, F (2, 22)=11.99. ZT16-18; p<0.0001, F (2, 16)=26.03. ZT19-21; p<0.0001, F (2, 24)=19.61. ZT22-24; p<0.0001, F (2, 14)=83.46. (D) TPR of Clk9M-Gal4/UAS-Kir2.1 (red), Clk9M-Gal4/+ (dark gray) and UAS-Kir2.1/+ (light gray) flies over 24 hr. Data are shown as the mean preferred temperature. Numbers represent the number of assays. Stars indicate p values of Tukey-Kramer tests when Gal4/UAS are statistically different from both Gal4/+ (stars in green) and UAS/+ (stars in blue). ****p<0.0001, **p<0.01 or *p<0.05. One-way ANOVA ZT1-3; p=0.0005, F (2, 21)=11.36. ZT4-6; p=0.0012, F (2, 14)=11.26. ZT7-9; p=0.0163, F (2, 16)=5.387. ZT10-12; p=0.0069, F (2, 12)=7.741. ZT13-15; p=0.0012, F (2, 15)=10.91. ZT16-18; p=0.0019, F (2, 15)=9.791. ZT19-21; p=0.1172, F (2, 12)=2.577. ZT22-24; p<0.0001, F (2, 17)=24.33.
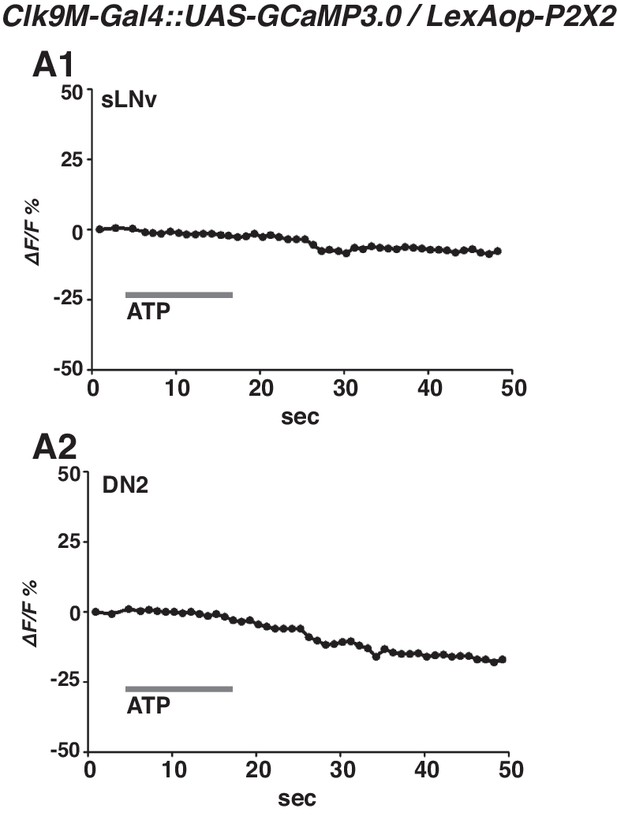
Lack of P2 × 2 expression in sLNvs leads to no responses to ATP application in both sLNvs and DN2s.
(A) Representative graphs of GCaMP fluorescence in sLNv (A1) and DN2 (A2) using Clk9M-Gal4::UAS-GCaMP3.0 / LexAop-P2X2 flies. GCaMP3.0 were expressed both in sLNvs and DN2s. Without Pdf-LexA to drive P2X2 in sLNvs, no responses were detected in the somas of either sLNvs (A1) or DN2s (A2).
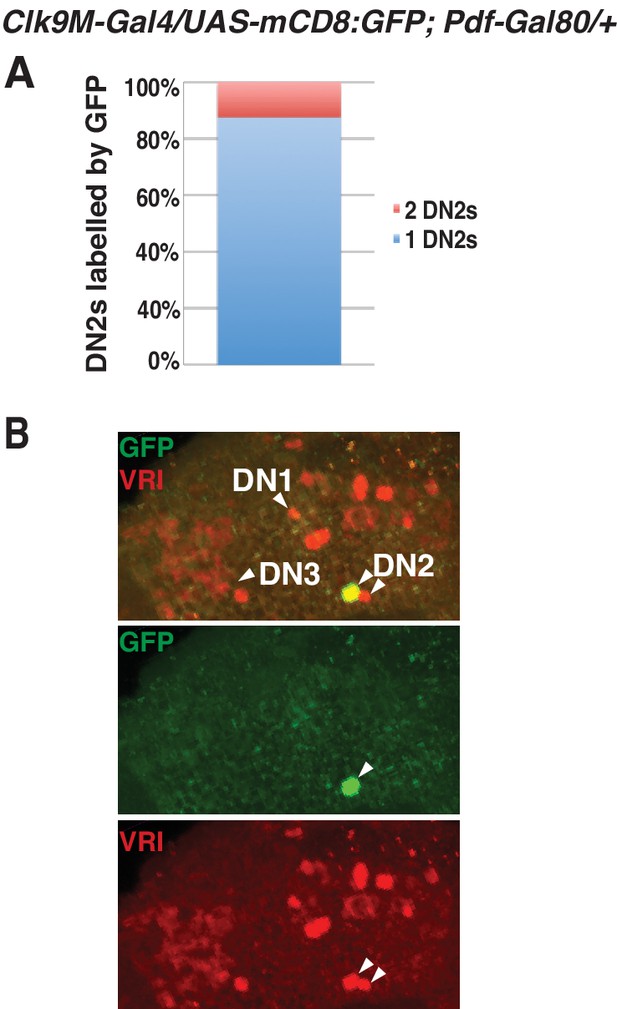
The DN2 driver is only expressed in one of the two sets of DN2s.
(A) Clk9M-Gal4;Pdf-Gal80 is expressed in one of the two sets of DN2s. While less than 20% flies had both sets of DN2s labeled with GFP (red), more than 80% flies had only one of the sets of DN2s labeled (blue). (B) A representative brain image of Clk9M-Gal4/UAS-mCD8:GFP; Pdf-Gal80/+. While VRI is expressed in two sets of DN2s, GFP only drives in one DN2 group.
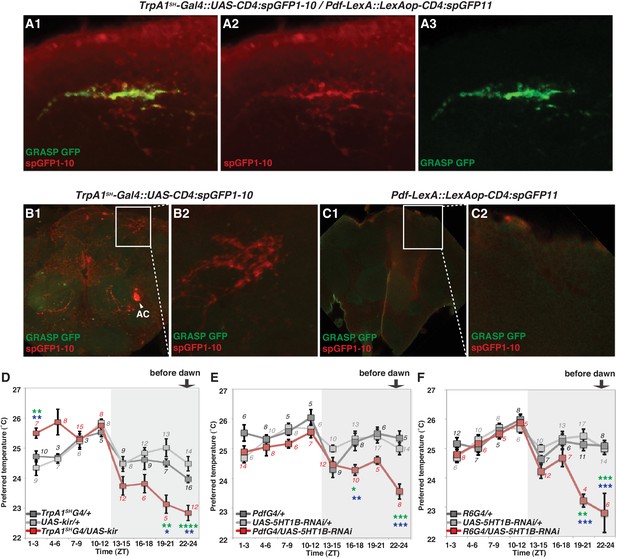
TrpA1SH-Gal4 and Pdf-Gal4 expressing neurons contact and are involved in regulating temperature preference before dawn.
(A) GRASP between ACs and sLNvs. TrpA1SH-Gal4::UAS-CD4:spGFP1-10 and Pdf-LexA::LexAop-CD4:spGFP11 flies were used to express split-GFP1-10 in ACs and split-GFP11 in LNvs, respectively. When these fly lines were crossed, a reconstituted GFP signal (green) was detected only at the distal terminus (A3). The axons of ACs (red) were labeled by anti-GFP-Cy5 (A2), which can detect split-GFP1-10 expressed in AC neurons. The merged image of A2 and A3 (A1). (B,C) Neither of the split GFP fragments alone in ACs or sLNvs had a reconstituted GFP fluorescence signal (green). (B2, C2): magnified images of B1 and C1, respectively. (D) TPR of TrpA1SH-Gal4/UAS-Kir2.1 (red), TrpA1SH-Gal4/+ (dark gray) and UAS-Kir2.1/+ (light gray) flies over 24 hr. (E) TPR of Pdf-Gal4/UAS-5HT1B-RNAi (red), Pdf-Gal4/+ (dark gray) and UAS-5HT1B-RNAi /+ (light gray) flies over 24 hr. (F) TPR of R6-Gal4/UAS-5HT1B-RNAi (red), R6-Gal4/+ (dark gray) and UAS-5HT1B-RNAi /+ (light gray) flies over 24 hr. R6-Gal4 drives sLNvs, but not lLNvs. The preferred temperatures among Gal4/UAS, Gal4/+ and UAS/+ flies in the each time zone were analyzed using one-way ANOVA and Tukey-Kramer tests (Supplementary file 1). Stars indicate p values of Tukey-Kramer tests when Gal4/UAS are statistically different from both Gal4/+ (stars in green) and UAS/+ (stars in blue). ****p<0.0001, ***p<0.001, **p<0.01 or *p<0.05.
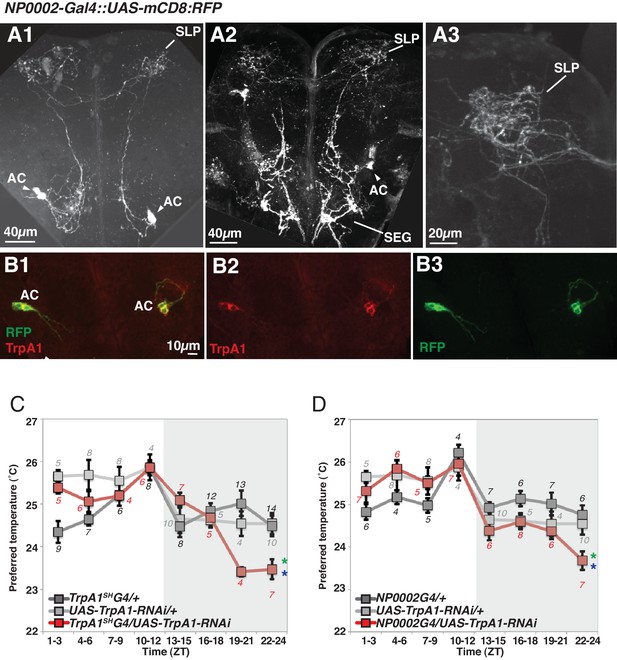
TRPA1 knockdown in ACs causes lower temperature preference before dawn.
(A–B) NP0002-Gal4 is expressed in AC neurons (A). Expression of NP0002-Gal4/ UAS-mCD8-RFP (green) in the fly brain (B). NP0002-Gal4/ UAS-mCD8-RFP expression (green) was overlapped in AC neurons labeled by TRPA1 antibody staining (red). (C–D) TRPA1 knockdown using TrpA1SH-Gal4 (C) and NP0002-Gal4 (D). TPRs of TrpA1SH-Gal4/ UAS-TrpA1-RNAi (red), TrpA1SH-Gal4/+ (dark gray) and UAS-TrpA1-RNAi /+ (light gray) flies over 24 hr (C). TPRs of NP0002-Gal4/ UAS-TrpA1-RNAi (red), NP0002-Gal4/+ (dark gray) and UAS-TrpA1-RNAi /+ (light gray) flies over 24 hr (D). Numbers represent the number of assays. The preferred temperatures among Gal4/UAS, Gal4/+ and UAS/+ flies in the each time zone was analyzed using One-way ANOVA and Tukey-Kramer tests (Supplementary file 1). Stars indicate p values of Tukey-Kramer tests when Gal4/UAS are statistically different from both Gal4/+ (stars in green) and UAS/+ (stars in blue). *p<0.05. SLP: superior lateral protocerebrum, SEG: subesophageal ganglion.
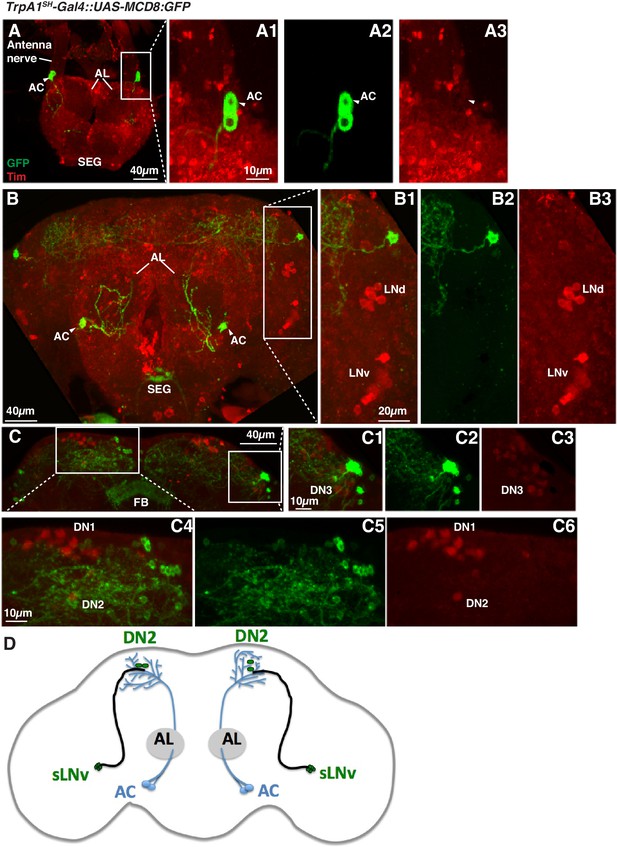
TrpA1SH-Gal4 expressing cells do not overlap with clock neurons in the brain.
(A) TrpA1SH-Gal4::UAS-mCD:GFP flies were stained with anti-GFP (green) and anti-Tim (red) (A1). AC neurons (arrow head) were labeled by GFP (A2) but not by the Tim antibody (A3). (B, C) TrpA1SH-Gal4 expressing cells did not overlap with lateral clock cells (LNv and LNd, (B) or dorsal clock cells (DN1, DN2 and DN3, (C). B1-3 and C1-6 are the magnified images of B and C, respectively. AL: antennal lobe, AN: antennal nerve, AC: AC neurons, FB: fan shaped body, SEG: subesophageal ganglion. (D) A schematic showing AC neurons’ projections as well as sLNv and DN2 clock neurons.
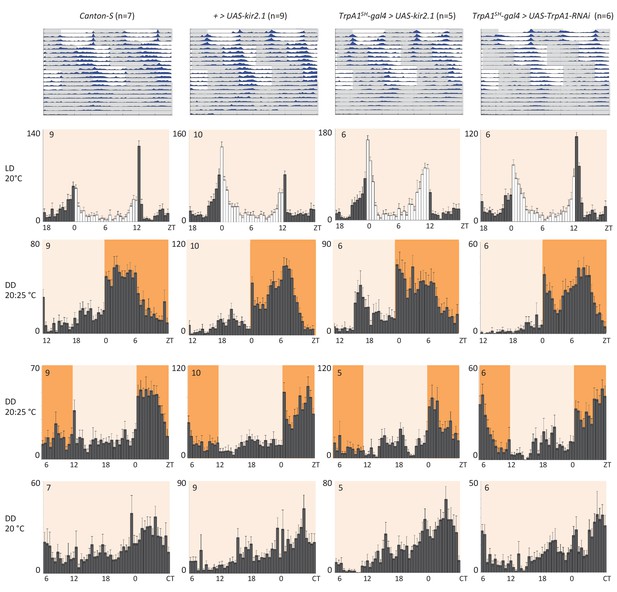
AC neuron silencing using the inwardly rectifying K+ channel Kir2.1 and TRPA1 knock down do not interfere with temperature entrainment.
Locomotor behavior of male flies of the genotypes indicated above each plot were analyzed in LD 20°C, followed by two 20°C: 25°C temperature cycles (TC) in DD, which were delayed by 6 hr compared to the previous regime. This was followed by five additional days in constant conditions (DD and 20°C). Top graphs show double-plotted average actograms, depicting behavioral activity throughout the experiment. White and grey areas depict light/warm and dark/cold periods respectively (actograms). Below, for the LD and TC parts the last 3 days and for the free-running part the first 3 days were averaged and plotted as histograms. White and grey bars indicate light and dark periods, respectively, while white background indicates 20°C periods and orange background 25°C periods (histograms). The number of animals analyzed is indicated in each histogram. The x-axis indicates time (hr) and y-axis indicates average total activity (number of beam crosses in 30 min).
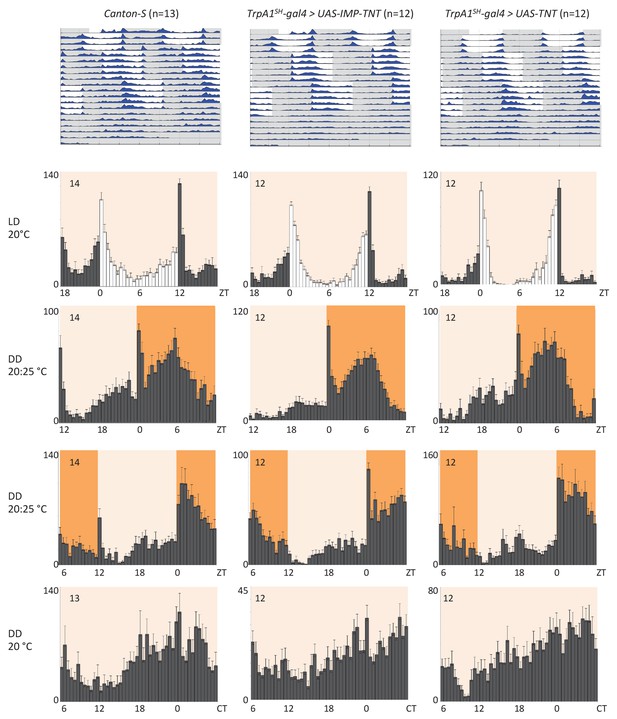
Blocking synaptic transmission of AC neurons using tetanus toxin light chain (TNT) does not interfere with temperature entrainment.
Locomotor behavior of male flies of the genotypes indicated above each plot (UAS-IMP-TNT: inactive toxin, UAS-TNT: active toxin) were analyzed in LD 20°C, followed by two 20°C: 25°C TC in DD, that were delayed by 6 hr compared to the previous regime. This was followed by five additional days in constant conditions (DD and 20°C). Top graphs show double-plotted average actograms, depicting behavioral activity throughout the experiment. White and grey areas depict light/warm and dark/cold periods respectively (actograms). Below, for the LD and TC parts the last 3 days and for the free-running part the first 3 days were averaged and plotted as histograms. White and grey bars indicate light and dark periods, respectively, while white background indicates 20°C periods and orange backgrounds 25°C periods (histograms). The number of animals analyzed is indicated in each histogram. The x-axis indicates time (hr) and y-axis indicates average total activity (number of beam crosses in 30 min).
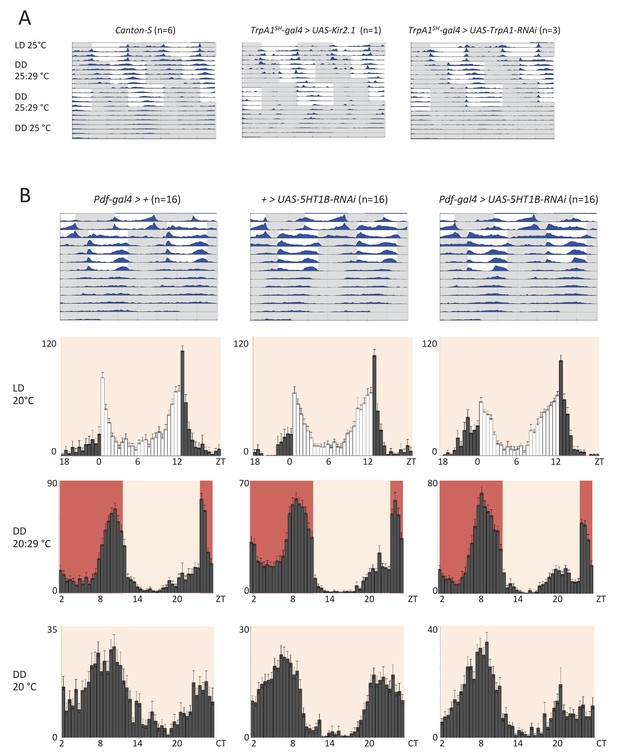
Silencing AC neurons does not interfere with synchronization to high temperature cycles (25°C: 29°C) (A) and down regulation of the serotonin receptor 5HT1B in Pdf neurons does not interfere with temperature entrainment (B).
(A) Locomotor behavior of male flies of the genotypes indicated above each plot were analyzed in LD 25°C, followed by two 25°C: 29°C TC in DD, that were delayed by 6 hr compared to the previous regime. This was followed by five additional days in constant conditions (DD and 25°C). Graphs show double-plotted average actograms, depicting behavioral activity throughout the experiment. White and grey areas depict light/warm and dark/cold periods respectively (actograms). Most of the flies with silenced AC neurons did not survive the experiment, presumably due to enhanced Kir2.1 and TNT expression in TrpA1SH-Gal4+ cells at the higher temperatures. Nevertheless, the few surviving flies showed normal entrainment to the temperature cycles. (B) Locomotor behavior of male flies of the genotypes indicated above each plot were analyzed in LD 20°C, followed by 5 days of an 8 hr advanced 20°C: 29°C TC in DD, followed by five additional days in constant conditions (DD and 20°C). Top graphs show double-plotted average actograms, depicting behavioral activity throughout the experiment. White and grey areas depict light/warm and dark/cold periods respectively (actograms). Below, the LD part, the last 3 days of the TC part, and the first 3 days of the free-running part, were averaged and plotted as histograms. White and grey bars indicate light and dark periods, respectively, while white backgrounds indicate 20°C periods and red backgrounds 29°C periods (histograms). The number of animals analyzed is indicated in each histogram. The x-axis indicates time (hr) and y-axis indicates average total activity (number of beam crosses in 30 min).
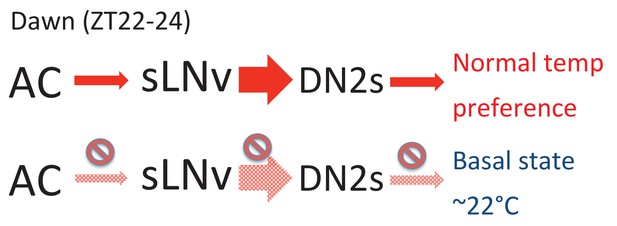
A model depicting the roles of AC, sLNv and DN2 neurons.
AC neurons detect ambient temperatures, and the information could be transmitted to sLNvs. At dawn (ZT22-24) sLNvs maximally activate DN2s, and the loss of DN2 activation results in flies preferring lower temperatures (basal state). Our data suggest a likely model in which sLNvs activate DN2s before dawn to drive proper temperature preference.
Tables
AC neuron silencing does not interfere with synchronization to high temperature cycles (25°C: 29°C) but reduces viability
Survival and synchronization of male flies exposed to the same temperature shift experiments as in Figure 5, except that the temperature cycled between 25°C and 29°C.
Number of flies that survive and entrain in 29:25 TC protocol.
| Genotype | Total n | Survived until end experiment n | Entrain of survived n |
|---|---|---|---|
| Canton-S | 10 | 6 | 6* |
| TrpA1SH-gal4 > UAS-TeTxLC V1b (inactive) | 9 | 4 | 4 |
| TrpA1SH-gal4 > UAS-TeTxLC R3 (active) | 9 | 0 | |
| + > UAS-kir2.1 | 10 | 8 | 8 |
| TrpA1SH-gal4 > UAS-kir2.1 | 10 | 1 | 1* |
| TrpA1SH-gal4 > UAS-TrpA1 RNAi | 8 | 3 | 3* |
-
*actograms shown in Figure 5—figure supplement 2A.
Rhythm and temperature compensation analysis of control and TrpA1 loss-of-function mutant flies under free running conditions at different ambient temperatures.
| Genotype | Free run | n | % Rh | Period (h) ± SEM | RS ± SEM | Q10 |
|---|---|---|---|---|---|---|
| +/+ | 16°C | 9 | 64 | 24.67 ± 0.56 | 2.6 ± 0.20 | 0.97 |
| trpA11/+ | 16°C | 8 | 57 | 24.22 ± 0.57 | 2.9 ± 0.24 | 0.99 |
| +/trpA1w903* | 16°C | 12 | 75 | 23.83 ± 0.25 | 2.9 ± 0.28 | 1.01 |
| +/Df(3L)ED4415 | 16°C | 15 | 94 | 23.85 ± 0.19 | 3.2 ± 0.19 | 1.00 |
| trpA11/trpA11 | 16°C | 6 | 55 | 23.29 ± 0.38 | 2.0 ± 0.19 | 1.02 |
| trpA11/trpA1w903* | 16°C | 13 | 81 | 24.23 ± 0.17 | 3.4 ± 0.27 | 1.01 |
| trpA11/Df(3L)ED4415 | 16°C | 14 | 88 | 23.54 ± 0.25 | 2.9 ± 0.19 | 1.02 |
| +/+ | 29°C | 16 | 100 | 23.75 ± 0.06 | 4.8 ± 0.24 | |
| trpA11/+ | 29°C | 13 | 93 | 23.75 ± 0.12 | 3.9 ± 0.31 | |
| +/trpA1w903* | 29°C | 16 | 100 | 24.16 ± 0.09 | 3.5 ± 0.25 | |
| +/Df(3L)ED4415 | 29°C | 11 | 79 | 23.98 ± 0.07 | 4.1 ± 0.36 | |
| trpA11/trpA11 | 29°C | 7 | 78 | 23.57 ± 0.46 | 2.6 ± 0.26 | |
| trpA11/trpA1w903* | 29°C | 13 | 93 | 24.56 ± 0.11 | 3.3 ± 0.26 | |
| trpA11/Df(3L)ED4415 | 29°C | 14 | 88 | 24.25 ± 0.08 | 3.9 ± 0.31 |
Additional files
-
Supplementary file 1
One-way ANOVA and Tukey-Kramer tests showing a comparison of each genotype of flies (Gal4/UAS, Gal4/+ or UAS/+) within the same time zone.
The preferred temperatures among Gal4/UAS, Gal4/+ and UAS/+ flies were analyzed using One-way ANOVA and Tukey-Kramer tests. In each time zone, F and p values and degrees of freedom are shown. The comparison between Gal4/UAS and Gal4/+, Gal4/UAS and UAS/+ as well as Gal4/+ and UAS/+ are shown in green, blue and red, respectively (****p<0.0001, ***p<0.001, **p<0.01 or *p<0.05). Stars are shown in Figures 1 and 4D–F and Figure 4—figure supplement 1C–D when Gal4/UAS are statistically different from both Gal4/+ (stars in green) and UAS/+ (stars in blue). These time zones are highlighted in orange.
- https://doi.org/10.7554/eLife.23206.017
-
Supplementary file 2
One-way ANOVA and Tukey-Kramer tests showing a comparison of each control fly line (Pdf-Gal4/+, UAS-Kir/+, UAS-∆clock/+, UAS-5-HT1B-RNAi/+ and R6-Gal4 within the same time zone
The preferred temperatures among Pdf-Gal4/+, UAS-Kir/+, UAS-∆clock/+, UAS-5-HT1B-RNAi/+ and R6-Gal4 flies were analyzed using one-way ANOVA and Tukey-Kramer tests. In the each time zone, F and p values and degrees of freedom are shown. ***p<0.001, **p<0.01 or *p<0.05.
- https://doi.org/10.7554/eLife.23206.018
-
Supplementary file 3
One-way ANOVA and Tukey-Kramer tests showing TPR comparisons of each fly line
The daytime TPR of each fly line was analyzed using one-way ANOVA. The preferred temperatures of ZT1-3 were further compared to that of each time zone (ZT 4–6, 7–9 or 10–12) by Tukey-Kramer tests. F and p values and degrees of freedom are shown. ***p<0.001, **p<0.01 or *p<0.05
- https://doi.org/10.7554/eLife.23206.019





