An unexpected role for the yeast nucleotide exchange factor Sil1 as a reductant acting on the molecular chaperone BiP
Figures
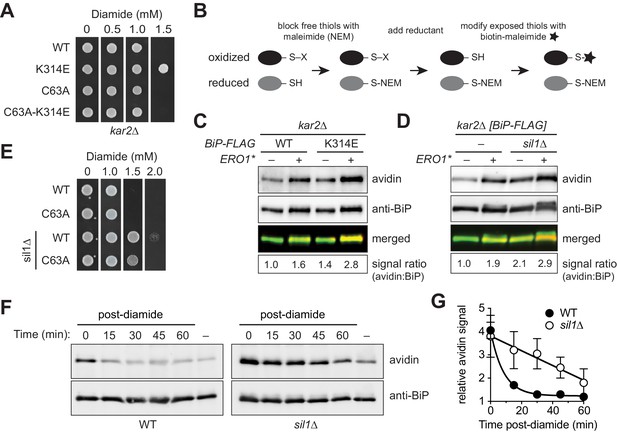
Sil1 regulates BiP oxidation state in cells.
(A) Yeast strains (CSY289, 290, 612, 689) were spotted onto SMM plates containing 0–1.5 mM diamide and incubated for 2 d at 30°C. (B) Schematic for the biotin-switch procedure. (C) Yeast strains deleted for endogenous BiP (kar2∆) containing plasmids encoding FLAG-tagged BiP were assayed for oxidized BiP levels using the biotin-switch protocol. Oxidative stress was generated by overexpression of Ero1*. BiP was immunoprecipitated, and total and oxidized BiP were detected by Western blotting. The relative levels of oxidized BiP are expressed as the ratio of the intensity of the avidin and anti-BiP signals. The signal ratio was set to 1.0 for wild-type cells grown in the absence of Ero1*. (D) Lysates were prepared from the indicated yeast after Ero1* induction. Oxidized BiP levels were detected and quantified as in C. (E) Yeast strains (CSY5, 275, 448, 449) were spotted onto YPD plates containing 0–2.0 mM diamide and were incubated for 2 d at 30°C. (F) Cells were treated with 5 mM diamide for 15 min, diamide was removed, and cells were returned to 30°C until harvest. Oxidized BiP levels were determined as in C. (G) Plot of the averaged quantified data ± SEM from F and a second independent experiment using the same protocol. For each strain, the signal ratio was set to 1.0 for cells grown without diamide.
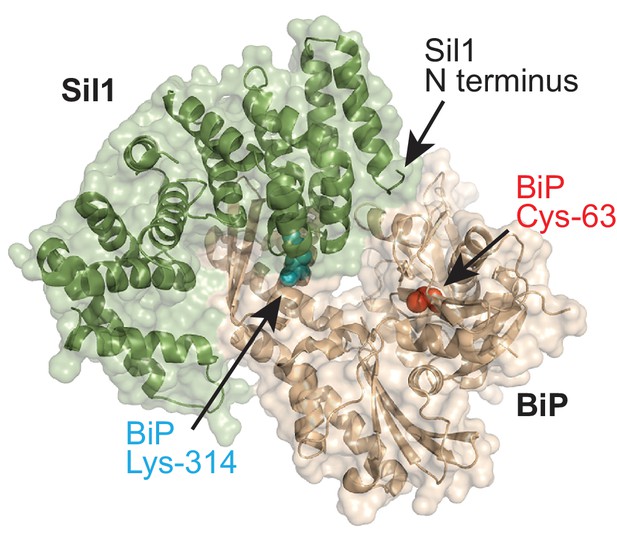
BiP-Sil1 structure.
Sil1-BiP complex structure (PBD ID: 3QML) encompassing the armadillo repeats from yeast Sil1 (residues 113–421) and the yeast BiP ATPase domain (residues 43–426) (Yan et al., 2011). BiP is shown in wheat; Sil1 is colored green. The redox-active BiP cysteine is shown as a red sphere. BiP lysine-314 is shown as a cyan sphere.
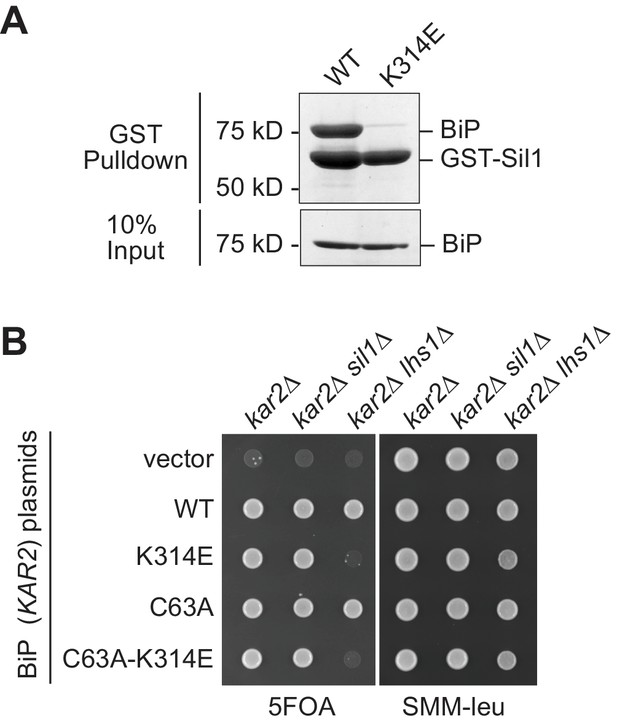
A BiP K314E mutation disrupts Sil1 binding.
(A) GST-Sil1, bound to glutathione-agarose beads, was incubated with recombinant wild-type BiP or mutant BiP-K314E proteins for 1 hr. Bound proteins were eluted from the agarose beads with SDS, and proteins were resolved by SDS-PAGE and visualized using a Coomassie blue stain. (B) CSY214 (kar2∆ [pCS623]), CSY594 (kar2∆ sil1∆ [pCS623]), and CSY595 (kar2∆ lhs1∆ [pCS623]) strains were transformed with LEU2-marked plasmids coding for the indicated BiP alleles (pCS681, pKP37, pCS685, pKP97) or empty vector. Transformants were spotted onto SMM plates containing 5-FOA (to select against wild-type BiP plasmid pCS623) or SMM minus leucine plates (to confirm successful transformation with the LEU2-marked plasmid). A sil1∆ lhs1∆ yeast strain is inviable; the observed inviability of the BiP-K314E mutant in the lhs1∆ strain background is consistent with an inability of the BiP-K314E mutant to associate with Sil1, which phenocopies a sil1 null strain.
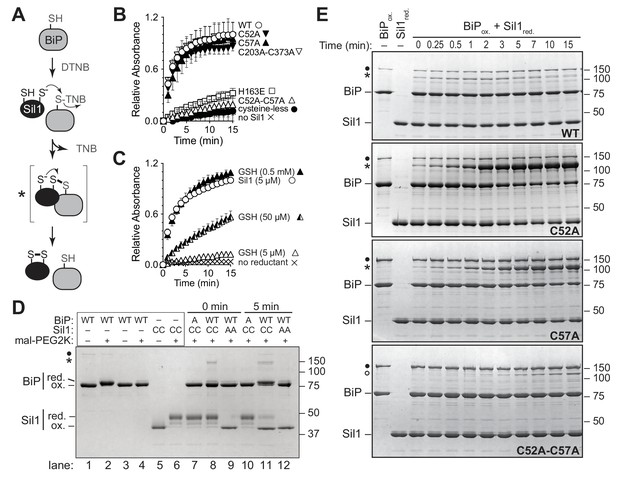
Sil1 N-terminal cysteines facilitate reduction of oxidized recombinant BiP in vitro.
(A) Schematic for monitoring Sil1 activity as a BiP-cysteine reductant. (B) Reduction of recombinant BiP oxidized by DTNB (BiP-TNB), in the presence of recombinant wild-type or mutant Sil1, was monitored by following the change in absorbance at 412 nm (indicative of TNB release) over time. (C) BiP-TNB reduction by reduced glutathione (GSH) or Sil1 was monitored as in B. Data in B and C represent the mean values from three independent experiments; error bars depict the range. (D) Thiol-disulfide exchange between recombinant wild-type BiP (WT) or a cysteine-less BiP-C63A mutant (A) and Sil1-C203A-C373A (with the N-terminal cysteines; CC) or a cysteine-less Sil1 (lacking the N-terminal cysteines; AA) was monitored by following the presence or absence of free thiols in both proteins. BiP and/or Sil1 were incubated for the indicated times, and reactions were quenched with the addition of the thiol-modifying agent mal-PEG2K. Samples were separated by non-reducing SDS-PAGE and visualized with Coomassie blue. Lanes 1–6 are shown as mobility controls. Lanes 1 and 2 contain BiP that was not reacted with DTNB; all other lanes include BiP incubated with DNTB prior to the addition of Sil1, and later mal-PEG2K. Labels indicate disulfide-linked BiP-Sil1 (asterisk) and BiP-BiP (filled circle) species. (E) Coomassie-stained gels follow recombinant BiP-TNB reaction with wild-type and mutant Sil1 proteins. Samples were quenched at the indicated times with N-ethylmaleimide (NEM) and separated by non-reducing SDS-PAGE. A BiP-Sil1 mixed-disulfide species (asterisk) accumulates with Sil1-C52A and Sil1-C57A. A second BiP-Sil1 species maintained in the absence of the N-terminal Sil1 cysteine pair is noted with an open circle.
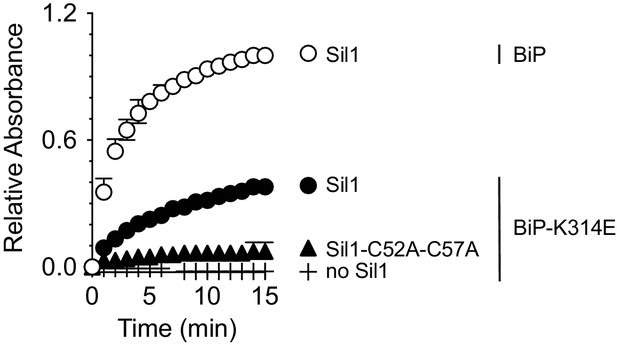
Oxidized BiP-K314E is a relatively poor substrate for Sil1.
Recombinant wild-type BiP or BiP-K314E were oxidized with DTNB, and the ability of wild-type Sil1 or mutant Sil1 (C52A-C57A) to reduce the oxidized BiP proteins was monitored by following the change in absorbance at 412 nm over time. Data represent the mean values from three independent experiments; error bars depict the range. Equivalent maximal absorbance values (equal amounts of TNB release) were observed when the reductant DTT was added in excess to the oxidized wild-type BiP or BiP-K314E proteins (data not shown), confirming that wild-type BiP and BiP-K314E were both equally oxidized by DTNB.
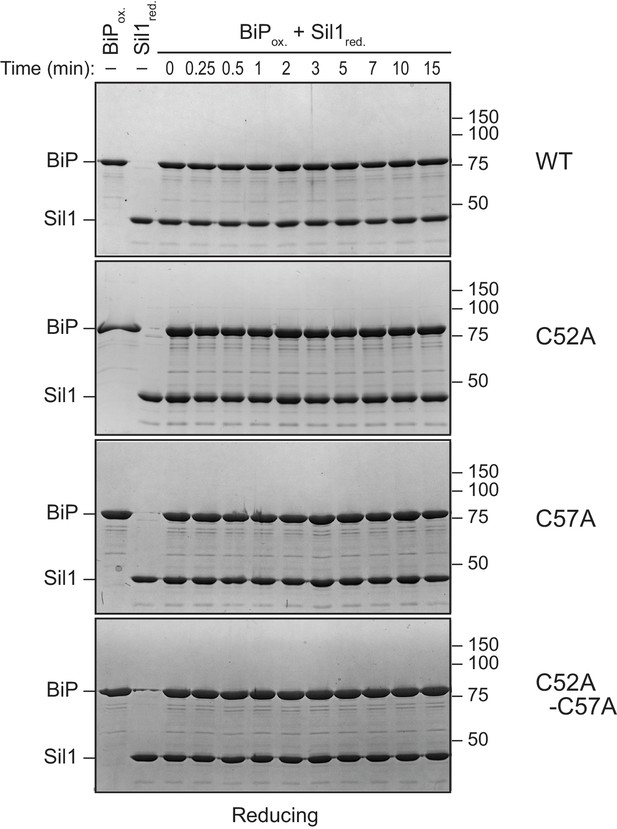
High molecular weight protein species are resolved by reducing SDS-PAGE.
Samples from Figure 2E were treated with BME (5%) prior to electrophoresis, and proteins were visualized with a Coomassie blue stain. The disappearance of the high molecular weight bands present under non-reducing conditions is consistent with the designation of these bands in Figure 2E as disulfide-bonded BiP-BiP (filled circle) and BiP-Sil1 (asterisk) species.
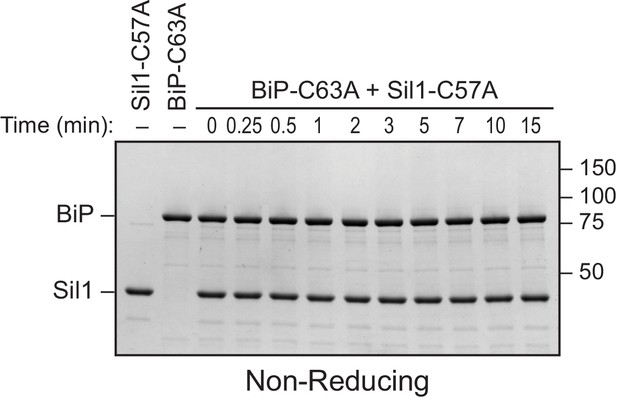
BiP's cysteine is required to form the disulfide-bonded species observed with Sil1-C57A.
Reduced recombinant Sil1-C57A was mixed with an equal concentration of a cysteine-less BiP protein (BiP-C63A), which was mock treated with DTNB. At the indicated times, sample buffer containing NEM was added to quench any reactive thiols as in Figure 2E. Proteins were resolved by non-reducing SDS-PAGE, and the gel was stained with Coomassie blue. In contrast to the strong disulfide-bonded species observed upon mixing Sil1-C57A mutant and wild-type BiP, no high molecular weight bands were observed at any point during the reaction of Sil1-C57A and cysteine-less BiP. These data are consistent with the identification of the strong high molecular weight disulfide-bonded species in Figure 2E (denoted with an asterisk) as a BiP-Sil1 mixed-disulfide pair. No high molecular weight species were observed for the BiP-C63A protein reacted with DTNB, consistent with the identification of the band observed with wild-type BiP after treatment with DTNB as a BiP-BiP disulfide bonded species (Figure 2E, filled circle).
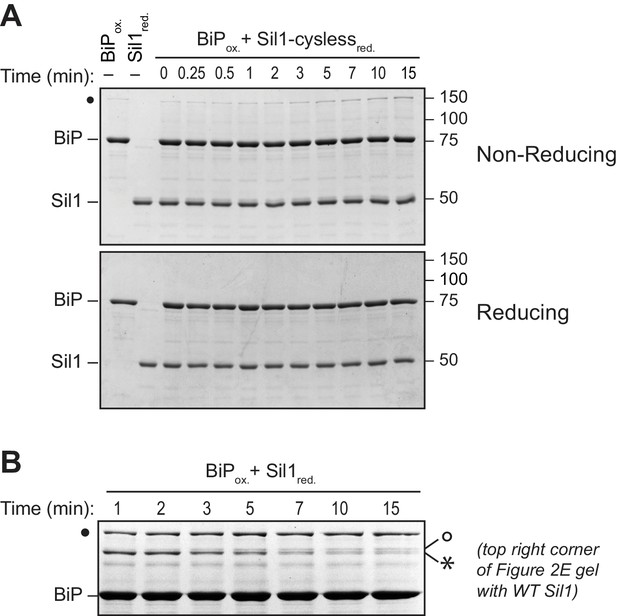
High molecular weight species observed under non-reducing conditions are absent when BiP is incubated with a cysteine-less Sil1 mutant.
(A) BiP-TNB was reacted with an equal concentration of cysteine-less Sil1 (C52A-C57A-C203A-C373A) pretreated with DTT. Reactions were quenched at the indicated time with sample buffer containing NEM as in Figure 2E. Proteins were resolved under non-reducing or reducing SDS-PAGE and visualized with a Coomassie blue stain. The only redox-dependent high molecular weight band observed with these proteins is the BiP-BiP disulfide-bonded species (filled circle), which is present even in the absence of Sil1. (B) An enlarged image of the upper right corner of the top gel from Figure 2E, which shows the reaction of reduced wild-type Sil1 and BiP-TNB. A transient doublet of disulfide-bonded proteins is visible during the timecourse. The bottom doublet band (asterisk) appears earlier and wanes at later time points, which is consistent with a transient interaction between BiP and the N-terminal Sil1 cysteine pair. The appearance of the upper double band (open circle) later during the timecourse mirrors the appearance of a disulfide-bonded species at later times of incubation between BiP-TNB and an N-terminal cysteine Sil1 mutant (Sil1-C52A-C57A). We anticipate that these bands (open circle) reflect a limited association between BiP and the cysteines in Sil1 present in the armadillo repeats, which is consistent with the absence of these bands when reactions are carried out with cysteine-less Sil1 (panel A) or cysteine-less BiP proteins (Figure 2—figure supplement 3).
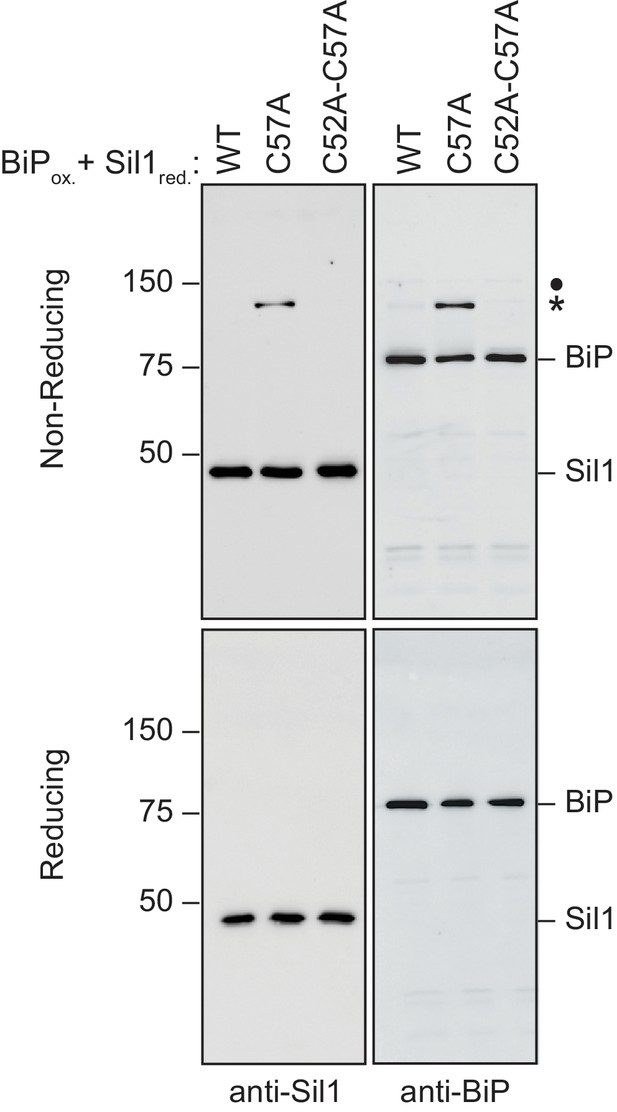
High molecular weight species observed under non-reducing conditions (and enhanced when Sil1 contains a single N-terminal cysteine) contain both Sil1 and BiP.
Reduced Sil1 (WT), Sil1-C57A, Sil1-C52A-C57A, and BiP-TNB were prepared, and BiP-TNB and the indicated Sil1 proteins were reacted, as in Figure 2E. Reactions were quenched 5 min after mixing of the BiP-TNB and Sil1 proteins by the addition of sample buffer containing NEM. Samples were separated by non-reducing or reducing SDS-PAGE; for reducing conditions, BME (5%) was added prior to electrophoresis. Samples were transferred to nitrocellulose, and Sil1 and BiP were detected by immunoblotting with rabbit anti-Sil1 or anti-BiP antibodies. Non-reducing and reducing samples were run on a single gel, and the membrane was cut after transfer prior to the probing with the individual antibodies. Labels indicate disulfide-linked BiP-Sil1 (asterisk) and a weakly detected BiP-BiP dimer (filled circle).
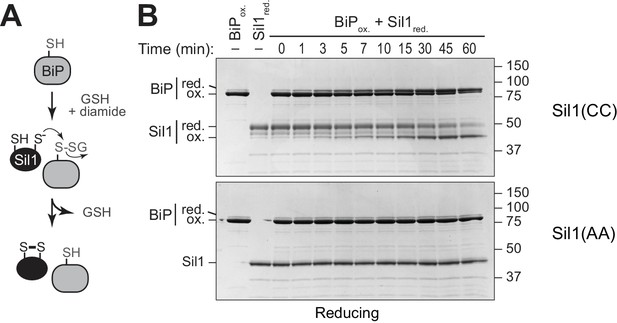
Sil1 can reduce glutathionylated BiP.
(A) Schematic for formation and reduction of glutathionylated BiP. (B) Glutathionylated BiP was prepared as described in the Materials and methods. Reduction of glutathionylated BiP by Sil1-C203A-C373A (with the N-terminal cysteines; CC) or a cysteine-less Sil1 (lacking the N-terminal cysteines; AA) was monitored by following the presence or absence of free protein thiols. At the indicated times, reactions were quenched with the addition of the thiol-modifying agent mal-PEG2K, which irreversibly reacts with reduced thiols. Samples were separated by reducing SDS-PAGE, and visualized with Coomassie blue. Proteins with free thiols that become modified with mal-PEG2K show a decreased electrophoretic mobility relative to proteins with oxidized cysteines that do not react with mal-PEG2K; the oxidized and reduced forms of each protein are indicated.
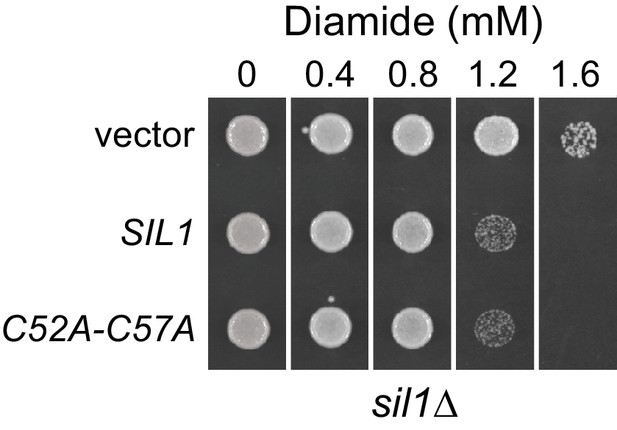
A yeast strain expressing a Sil1 mutant that lacks reducing activity (Sil1-C52A-C57A) does not show an increased resistance to diamide.
A sil1∆ strain (CSY448) transformed with the plasmids pJW7 (SIL1), pCS876 (sil1-C52A-C57A), or an empty vector was spotted onto SMM plates containing 0–1.6 mM diamide and incubated for 2 d at 30°C.
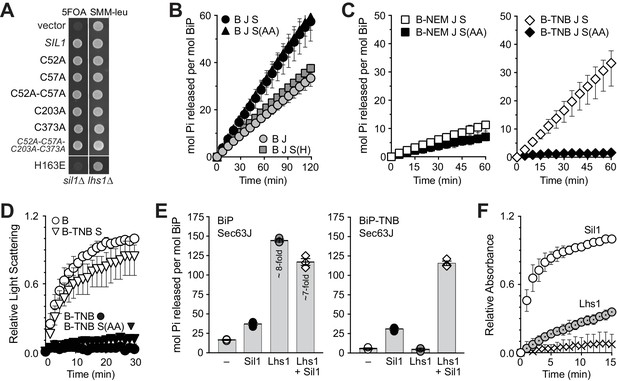
Sil1 regulates BiP activity both as a nucleotide exchange factor (NEF) and a reductant.
(A) Sil1 cysteine mutants maintain nucleotide exchange activity. A sil1∆ lhs1∆ strain covered with an URA3-marked SIL1 plasmid was transformed with LEU2-marked plasmids encoding the indicated Sil1 proteins. Their ability to substitute for wild-type Sil1 was assessed after counter-selection of the URA3-plasmid on 5-FOA. (B) BiP (B) ATPase activity was monitored by following the accumulation of free phosphate in the presence of Sec63J (J) and Sil1 (S), Sil1-C52A-C57A (S(AA)) or Sil1-H163E (S(H)). (C) ATP hydrolysis rates of BiP oxidized with NEM (B-NEM) or DNTB (B-TNB) was monitored in combination with J, S, or S(AA). In B and C, mean values of triplicate experiments are shown; error bars depict the range. (D) Aggregation of denatured rhodanese was assayed by monitoring light scattering (associated with aggregation) over time. Denatured rhodanese was diluted away from denaturant into buffer containing BiP or BiP-TNB that had been pre-incubated in the presence or absence of reduced, recombinant Sil1. Mean values of three independent experiments are shown; error bars depict the range. (E) The accumulation of free phosphate 15 min (left panel) or 30 min (right panel) post-ATP addition was determined for reduced BiP and BiP-TNB incubated with Sil1 (1:1 ratio) and/or Lhs1 (1:0.2 ratio) plus J-protein. Data show the mean rate of phosphate release ± SEM of three independent experiments. (F) Reduction of BiP-TNB by Sil1 or Lhs1 was monitored spectroscopically as in Figure 2. Mean values of four independent experiments are shown; error bars depict the range.
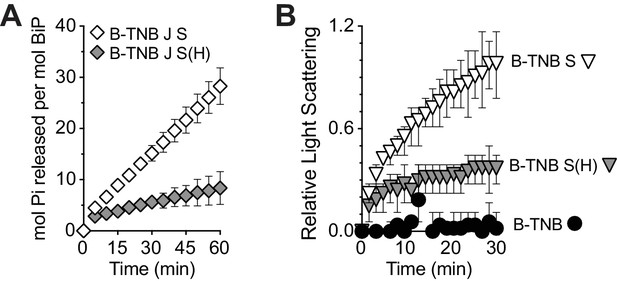
Sil1-H163E only modestly reverses the decreased ATPase and increased holdase activities associated with oxidized BiP.
(A) ATP hydrolysis rates of BiP oxidized with DNTB (B-TNB) were monitored in combination with Sec63J (J) and Sil1 (S) or Sil1-H163E (S(H)). Mean values of duplicate experiments are shown; error bars depict the range. (B). Aggregation of denatured rhodanese was assayed as in Figure 4D. Denatured rhodanese was diluted away from denaturant into buffer containing BiP-TNB that had been pre-incubated in the presence or absence of reduced, recombinant wild-type Sil1 or Sil1-H163E mutant. Mean values of three independent experiments are shown; error bars depict the range.
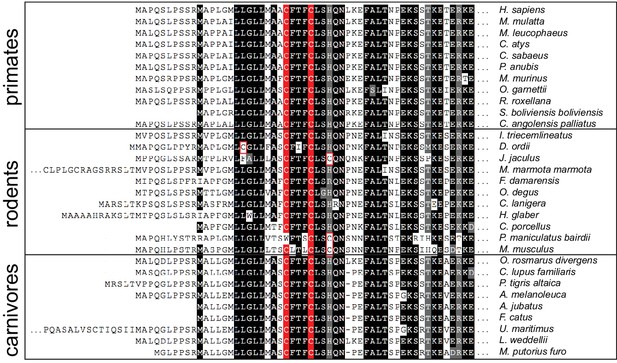

Mammalian SIL1 orthologs contain a conserved pair of cysteines within the N-terminal region.
An alignment of protein sequences from SIL1 orthologs found in primates, rodents, and carnivores reveals a conserved cysteine pair in the N-terminal region. Sequences are shaded at 90% identity. Cysteines conserved between 90% of the chosen orthologs are highlighted in red; additional cysteine residues are outlined in red.
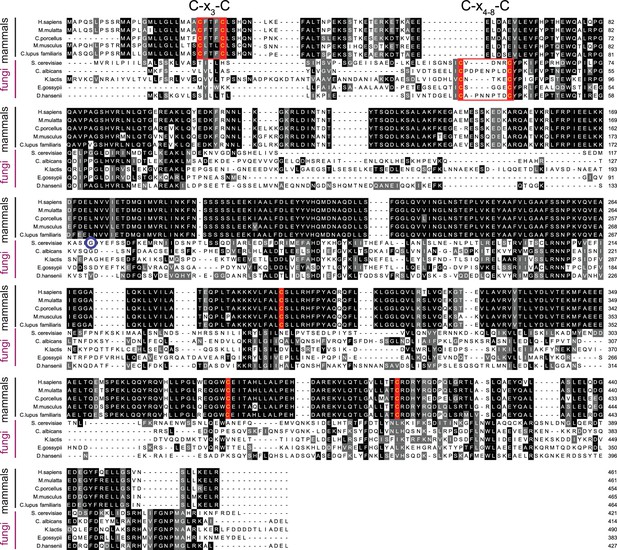

Mammalian and fungal Sil1 proteins contain a conserved N-terminal cysteine pair.
Alignment of orthologous Sil1 protein sequences from five mammalian and five fungal species. Protein sequences were aligned using ClustalW (Larkin et al., 2007) and formatted using Multiple Align Show (Stothard, 2000) with a consensus-shading setting of 50% identity (black) or similarity (grey). Cysteines showing a 50% conservation between orthologs are highlighted in red text (instead of black). The N-terminal redox-active cysteine pair identified in S. cerevisiae Sil1 and shared between fungal orthologs is boxed in red; conserved N-terminal mammalian cysteines are also boxed in red. A blue circle notes S. cerevisiae residue 113, the most N-terminal residue from the Sil1 structure construct (Figure 1—figure supplement 1). Note the limited sequence identity observed between the fungal and mammalian Sil1 orthologs, and also among the fungal Sil1 orthologs.
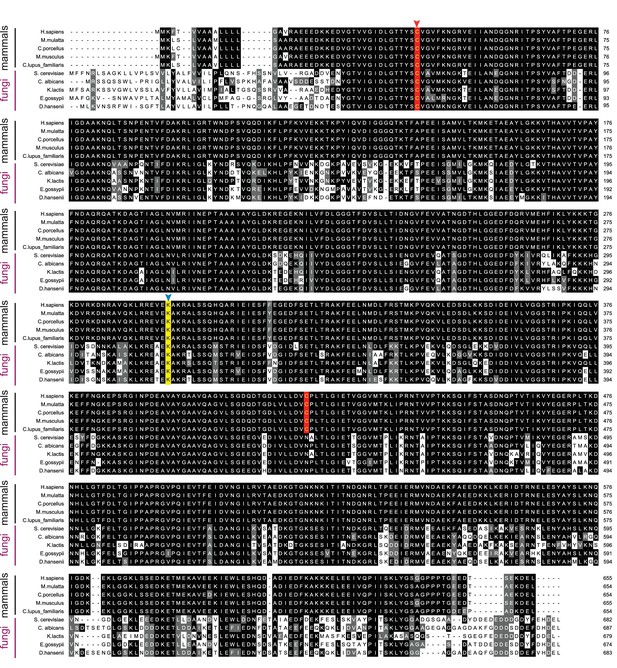
Sequence alignment of BiP orthologs.
Alignment of BiP ortholog protein sequences from five mammalian and five fungal species. Sil1 proteins from the same species are displayed in Figure 5—figure supplement 1. Protein sequences were aligned and formatted as described for Figure 5—figure supplement 1. In contrast to the low degree of sequence conservation observed between Sil1 orthologs, a high degree of sequence conservation is seen among the BiP orthologs. A red arrow highlights the conserved redox active cysteine within the BiP ATPase domain. A blue arrow and yellow background indicate the position of lysine-314 in yeast BiP, which is conserved across species.
Tables
Plasmids.
| Plasmid | Description | Marker | Source |
|---|---|---|---|
| pCS623 | KAR2 | CEN URA3 | (Wang et al., 2014) |
| pCS757 | KAR2-FLAG | CEN LEU2 | (Wang et al., 2014) |
| pCS878 | kar2-K314E-FLAG | CEN LEU2 | This study |
| pCS452 | PGAL1-ERO1*-myc | CEN URA3 | (Sevier et al., 2007) |
| pCS681 | KAR2 | CEN LEU2 | (Wang et al., 2014) |
| pCS685 | kar2-C63A | CEN LEU2 | (Wang et al., 2014) |
| pKP37 | kar2-K314E | CEN LEU2 | This study |
| pKP97 | kar2-C63A-K314E | CEN LEU2 | This study |
| pJW7 | SIL1 | CEN URA3 | This study |
| pCS876 | sil1-C52A-C57A | CEN URA3 | This study |
| pHS116 | SIL1 | CEN LEU2 | This study |
| pKS20 | sil1-C52A | CEN LEU2 | This study |
| pKS21 | sil1-C57A | CEN LEU2 | This study |
| pKS24 | sil1-C52A-C57A | CEN LEU2 | This study |
| pKS23 | sil1-C203A | CEN LEU2 | This study |
| pKS22 | sil1-C373A | CEN LEU2 | This study |
| pCS923 | sil1-C52A-C57A-C203A-C373A | CEN LEU2 | This study |
| pCS925 | sil1-H163E | CEN LEU2 | This study |
| pCS817 | His6-kar2-(42-682) | KAN | (Wang et al., 2014) |
| pCS818 | His6-kar2-(42-682)-C63A | KAN | (Wang et al., 2014) |
| pKP85 | His6-kar2-(42-682)-K314E | KAN | This study |
| pCS637 | sil1-(20-407)-His6 | AMP | This study |
| pCS870 | sil1-(20-407)-C52A-His6 | AMP | This study |
| pCS871 | sil1-(20-407)-C57A-His6 | AMP | This study |
| pCS875 | sil1-(20-407)-H163E-His6 | AMP | This study |
| pCS877 | sil1-(20-407)-C52A-C57A-His6 | AMP | This study |
| pCS948 | sil1-(20-407)-C203A-C373A-His6 | AMP | This study |
| pCS895 | sil1-(20-407)-C52A-C57A-C203A-C373A- His6 | AMP | This study |
| pKP52 | GST-sil1-(22-406) | AMP | This study |
| pHS130 | His6-lhs1-(21-877)-StrepII | KAN | (Xu et al., 2016) |
| pCS675 | GST-sec63J-(121-221) | AMP | (Wang et al., 2014) |
Strains.
| Strain | Genotype | Source |
|---|---|---|
| CSY5 | MATa GAL2 ura3-52 leu2-3,112 | (Wang et al., 2014) |
| CSY214 | MATa GAL2 ura3-52 leu2-3,112 kar2∆::KanMX [pCS623] | (Wang et al., 2014) |
| CSY275 | MATa GAL2 ura3-52 leu2-3,112 kar2-C63A | (Wang et al., 2014) |
| CSY278 | MATa GAL2 ura3-52 leu2-3,112 kar2-C63A can1::PGAL1-ERO1*-myc | (Wang et al., 2014) |
| CSY289 | MATa GAL2 ura3-52 leu2-3,112 kar2∆::KanMX [pCS681] | (Wang et al., 2014) |
| CSY290 | MATa GAL2 ura3-52 leu2-3,112 kar2∆::KanMX [pCS685] | (Wang et al., 2014) |
| CSY612 | MATa GAL2 ura3-52 leu2-3,112 kar2∆::KanMX [pKP37] | This study |
| CSY689 | MATa GAL2 ura3-52 leu2-3,112 kar2∆::KanMX [pKP97] | This study |
| CSY622 | MATa GAL2 ura3-52 leu2-3,112 kar2∆::KanMX [pCS757] | This study |
| CSY646 | MATa GAL2 ura3-52 leu2-3,112 kar2∆::KanMX [pCS878] | This study |
| CSY448 | MATalpha GAL2 ura3 leu2 sil1∆::KanMX | This study |
| CSY449 | MATalpha GAL2 ura3 leu2 lys2∆0 kar2-C63A sil1∆::KanMX | This study |
| CSY581 | MATalpha GAL2 ura3 leu2 lhs1∆::KanMX | This study |
| CSY594 | MATa GAL2 ura3 leu2 sil1∆::KanMX kar2∆::KanMX [pCS623] | This study |
| CSY595 | MATa GAL2 ura3 leu2 lhs1∆::KanMX kar2∆::KanMX [pCS623] | This study |
| CSY625 | MATa GAL2 ura3 leu2 sil1∆::KanMX kar2∆::KanMX [pCS757] | This study |
| CSY647 | MATa GAL2 ura3 leu2 sil1∆::KanMX lhs1∆::KanMX [pJW7] | This study |





