Genome mining unearths a hybrid nonribosomal peptide synthetase-like-pteridine synthase biosynthetic gene cluster
Figures
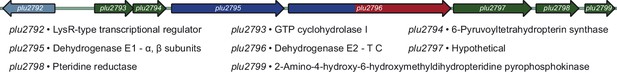
The pepteridine biosynthetic locus.
Green, pteridine synthesis genes; Blue, pyruvate dehydrogenase-like genes; Red, NRPS-like genes. T, thiolation domain; C, condensation domain.
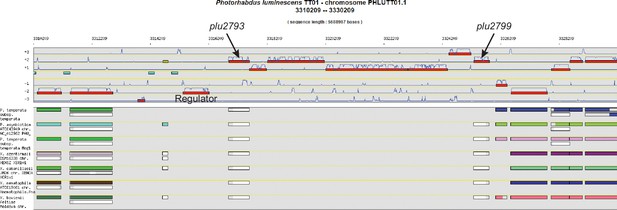
Genome synteny analysis using MicroScope.
Genome synteny analysis revealed a genomic island in P. luminescens TT01 (note absence of biosynthetic genes in phylogenetically related species, lower panel) harboring a number of enzymes possessing homology to pteridine and NRPS biosynthetic machineries (+1, +2, and +3 reading frames, upper panel). The regulator in the genomic island was not included in our design, as the pathway was placed under the control of a T7 promoter.
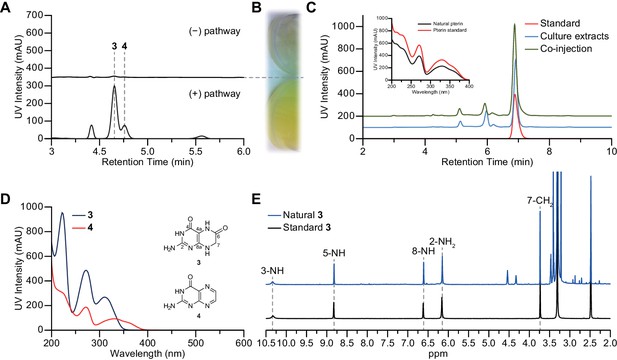
Production of 7,8-dihydroxanthopterin (3) and pterin (4) by heterologous expression.
(A) HPLC traces (310 nm) of butanol extracts showing compounds 3 and 4 being over-produced in the heterologous expression strain. (B) The edge of a culture flask growing E. coli BAP1 carrying an empty vector pET28a (top), and a culture flask growing E. coli BAP1 carrying the wild-type pepteridine pathway (bottom). (C) HPLC traces (310 nm) from HPLC co-injection with authentic pterin (4), and UV absorption spectral comparison between natural and authentic pterin (inset). (D) UV-vis absorption spectra of compounds 3 and 4. (E) 1H NMR comparison of natural (blue) and standard (black) 3 in DMSO-d6.
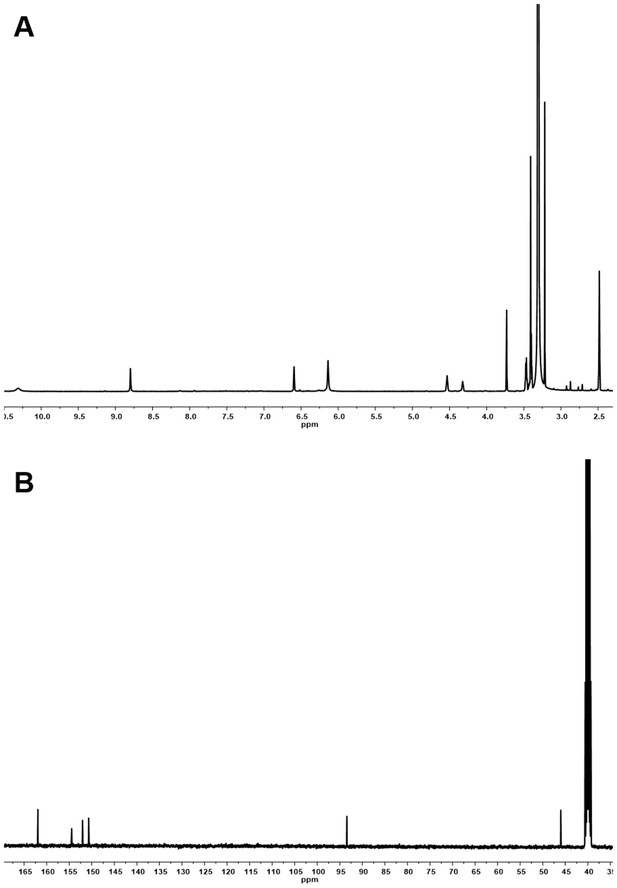
1H and 13C NMR spectra of compound 3.
(A) 1H NMR spectrum of compound 3 in DMSO-d6. (B) 13C NMR spectrum of compound 3 in DMSO-d6.
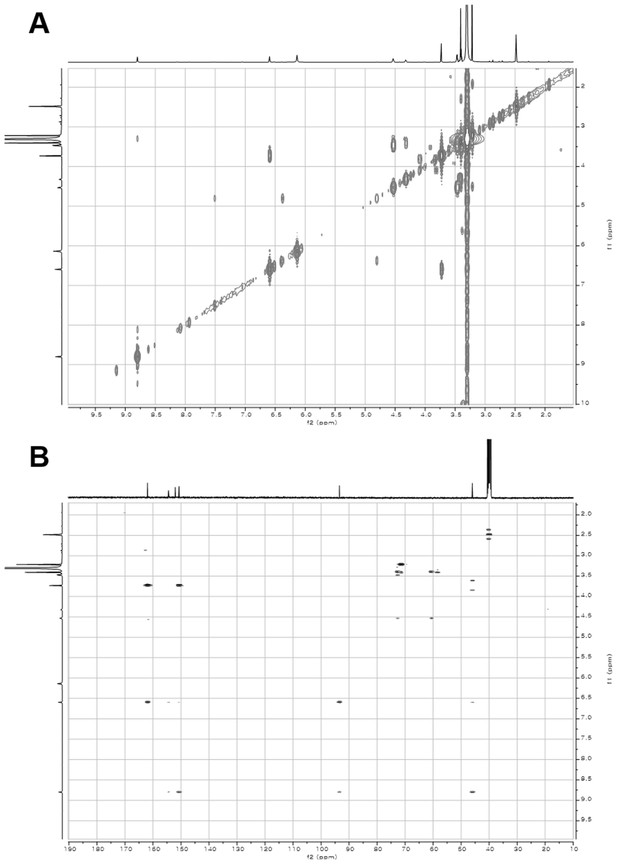
2D- (gCOSY and gHMBCAD) NMR spectra of compound 3.
(A) gCOSY NMR spectrum of compound 3 in DMSO-d6. (B) gHMBCAD NMR spectrum of compound 3 in DMSO-d6.
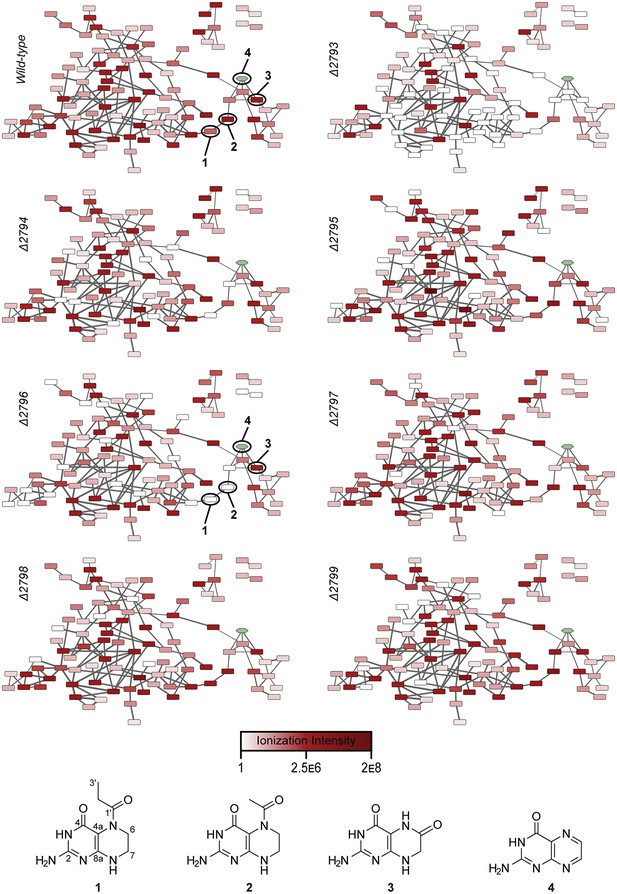
Relative abundances of wild-type pathway-dependent metabolites in wild-type and mutant pepteridine pathways.
The average ionization intensity is depicted for each molecular feature under a given genetic condition (wild-type and Δplu2793 through Δplu2799). Alterations in abundance are correlated with changes in nodal color intensity among genetic constructs, allowing visual assessment of product distributions for a given mutation. See Figure 3—figure supplement 2 for information on mass of pathway-dependent metabolites. Structural characterization of compounds 1 and 2 is shown in Figure 3—figure supplements 4–10, Table 2, and the Materials and methods section.
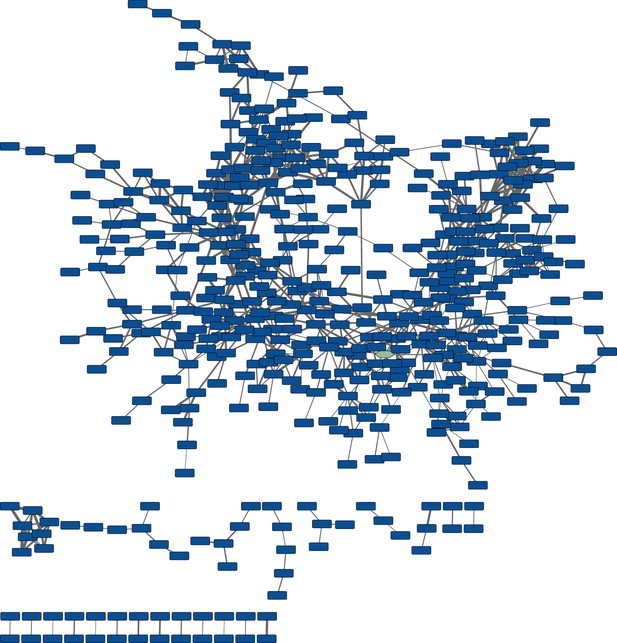

Untargeted molecular networking of culture extracts from cells harboring the wild-type pepteridine biosynthetic pathway.
The node with the green hue represents pterin (4) networked into the untargeted wild-type map. This network includes abundant molecules from the pathway in addition to primary metabolites and media components.
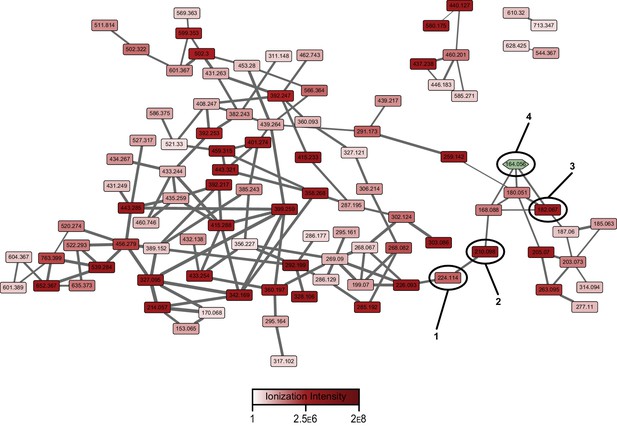
Pathway-targeted molecular networking of the wild-type pepteridine biosynthetic pathway.
Comparative metabolomic analysis between control and experimental culture samples enabled targeting (MS2) of molecular features dependent on the presence of the biosynthetic pathway.
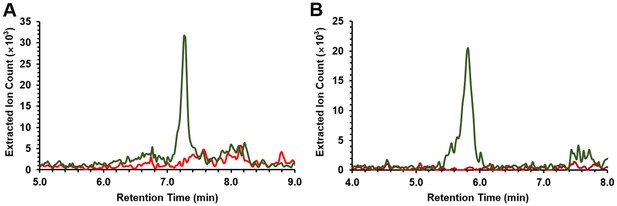
LC/MS Extracted Ion Count (EIC) chromatograms of compounds 1 (A) and 2 (B) from butanol extracts of the pepteridine heterologous expression strain.
LC/MS traces were extracted with m/z 224 corresponding to compound 1 (A) and m/z 210 corresponding to compound 2 (B) from butanol extracts of the E. coli BAP1 culture broths carrying the wild-type pepteridine pathway (green) or the empty vector (pET28a, red).
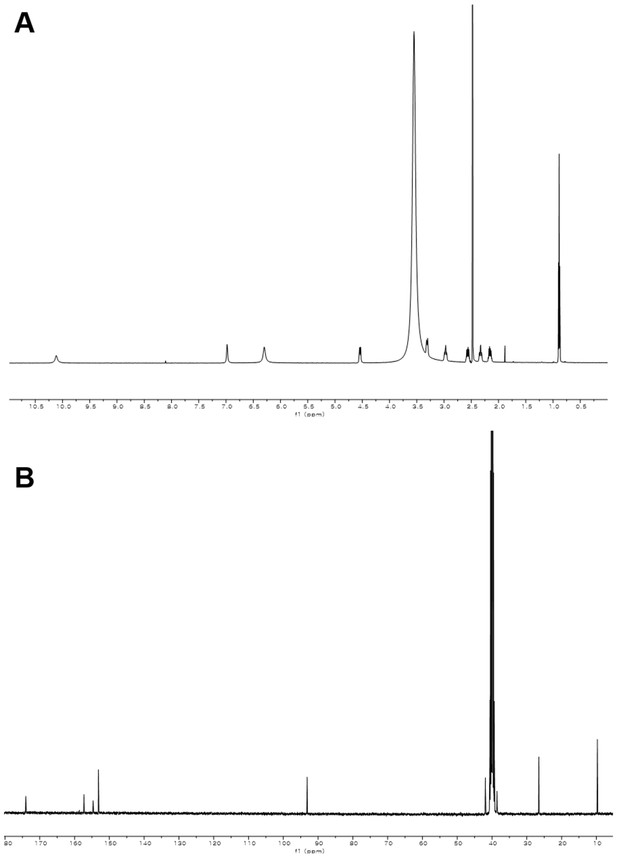
1H and 13C NMR spectra of compound 1.
(A) 1H NMR spectrum of compound 1 in DMSO-d6. (B) 13C NMR spectrum of compound 1 in DMSO-d6.
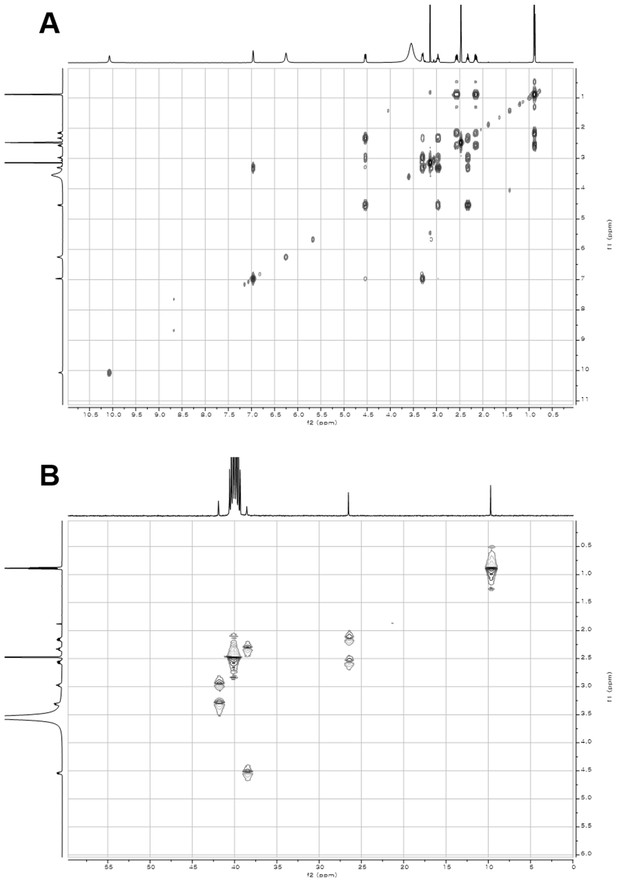
2D- (gCOSY and gHSQCAD) NMR spectra of compound 1.
(A) gCOSY NMR spectrum of compound 1 in DMSO-d6. (B) gHSQCAD NMR spectrum of compound 1 in DMSO-d6.
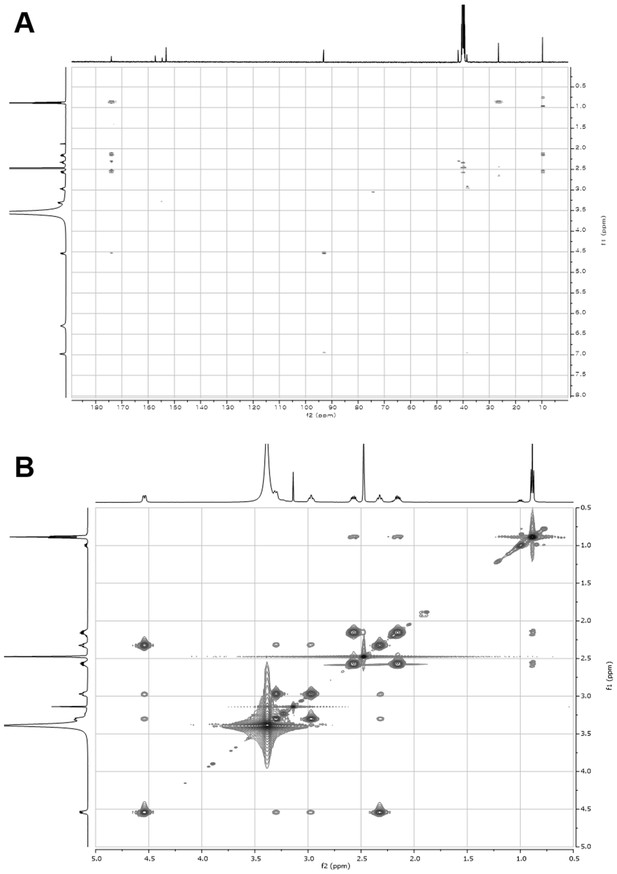
2D- (gHMBCAD and NOESY) NMR spectra of compound 1.
(A) gHMBCAD NMR spectrum of compound 1 in DMSO-d6. (B) NOESY NMR spectrum of compound 1 in DMSO-d6.
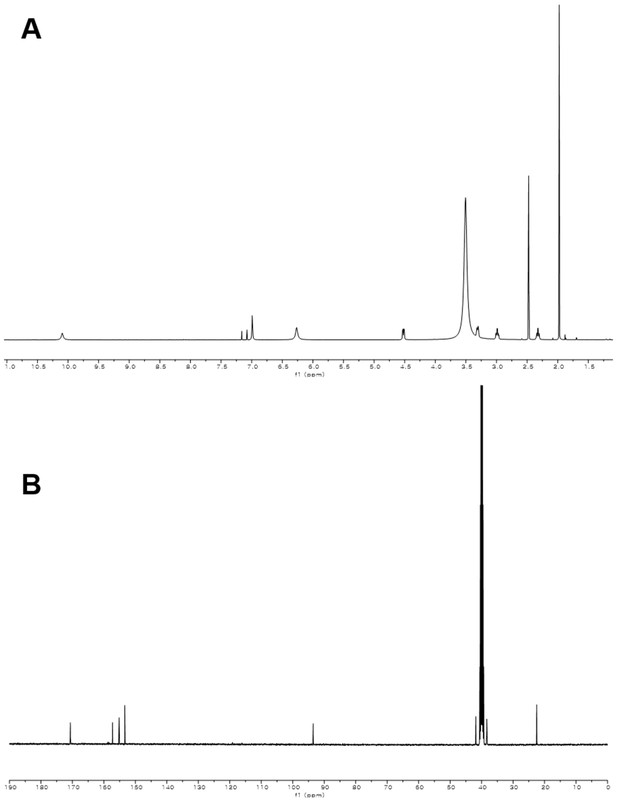
1H and 13C NMR spectra of compound 2.
(A) 1H NMR spectrum of compound 2 in DMSO-d6. (B) 13C NMR spectrum of compound 2 in DMSO-d6.
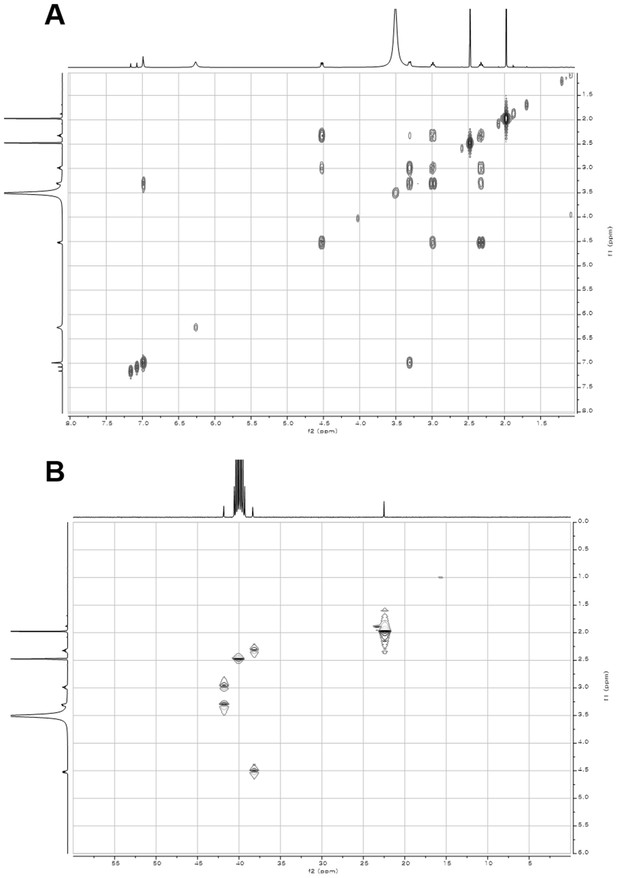
2D- (gCOSY and gHSQCAD) NMR spectra of compound 2.
(A) gCOSY NMR spectrum of compound 2 in DMSO-d6. (B) gHSQCAD NMR spectrum of compound 2 in DMSO-d6.
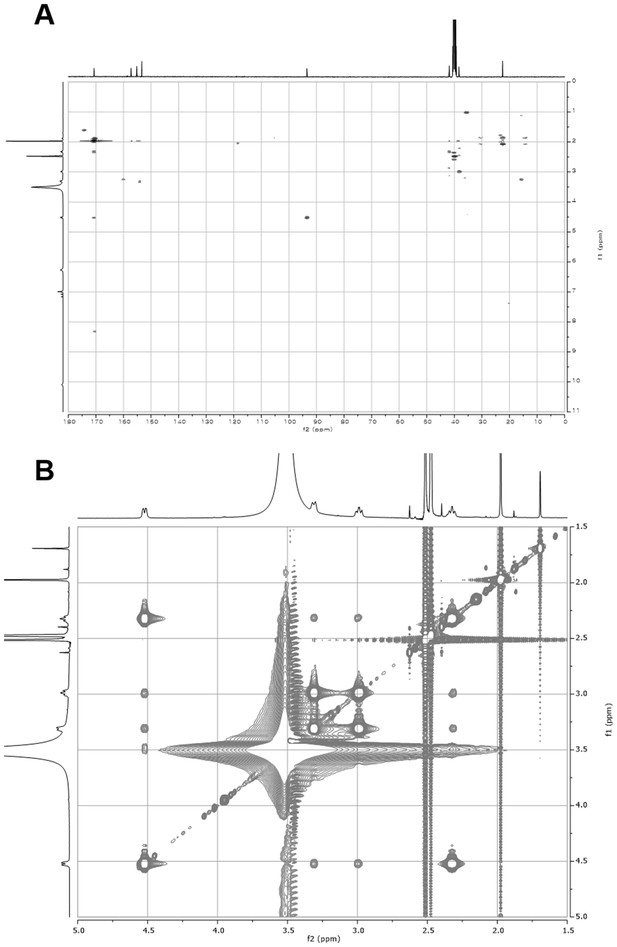
2D- (gHMBCAD and NOESY) NMR spectra of compound 2.
(A) gHMBCAD NMR spectrum of compound 2 in DMSO-d6. (B) NOESY NMR spectrum of compound 2 in DMSO-d6.
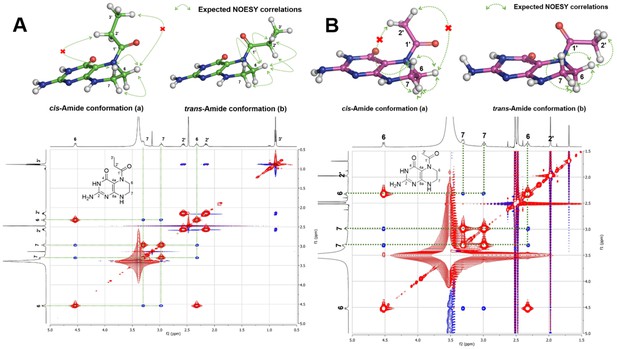
Amide conformation of compounds 1 and 2.
(A) Plausible cis- (a) and trans- (b) amide conformations of compound 1 (top) and interpretation of 2D NOESY spectral data of compound 1 (bottom), (B) Plausible cis- (a) and trans- (b) amide conformations of compound 2 (top) and interpretation of 2D NOESY spectral data of compound 2 (bottom).
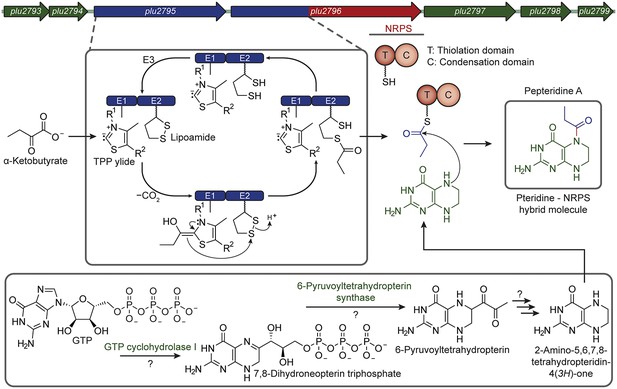
Proposed pepteridine biosynthesis.
α-Ketobutyrate is processed by the atypical dehydrogenase-NRPS-like complex to generate a propionyl-loaded carrier protein (blue). Pteridine enzymes interact with the metabolism of the host to generate tetrahydropterin (green). The NRPS C domain couples these substrates to form the cis-amide bond (red) as illustrated for 1.
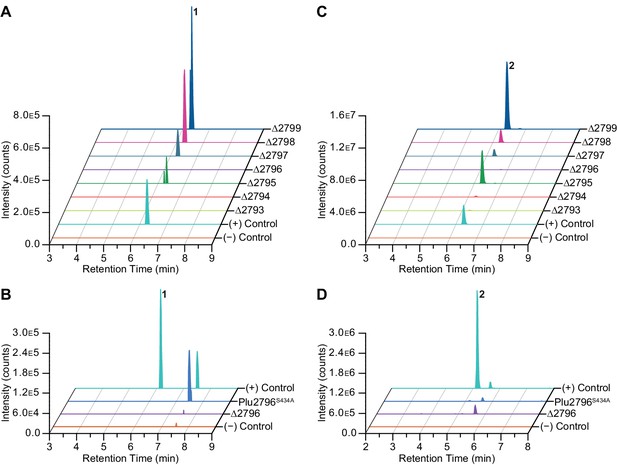
Gene deletion and NRPS inactivation analyses on 1 (A–B) and 2 (C–D).
(A–B) Extracted ion chromatograms of 1 for wild-type and mutants are shown. (C–D) Extracted ion chromatograms of 2 for wild-type and mutants are shown.
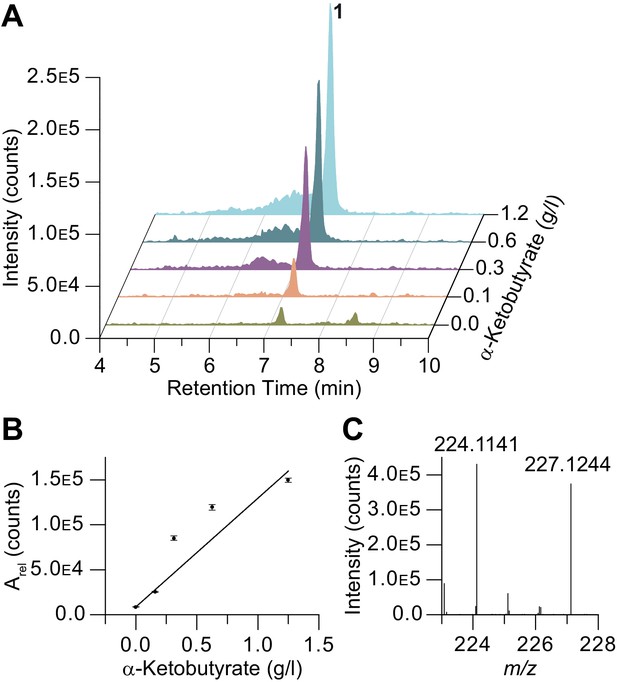
α-Ketobutyrate feeding studies on 1.
(A–B) Production enhancement of 1 in α-ketobutyrate supplementation studies (Arel, relative integration value). (C) 13C4-α-ketobutyrate supplementation leads to 13C3 incorporation in 1 consistent with the proposed biosynthesis.
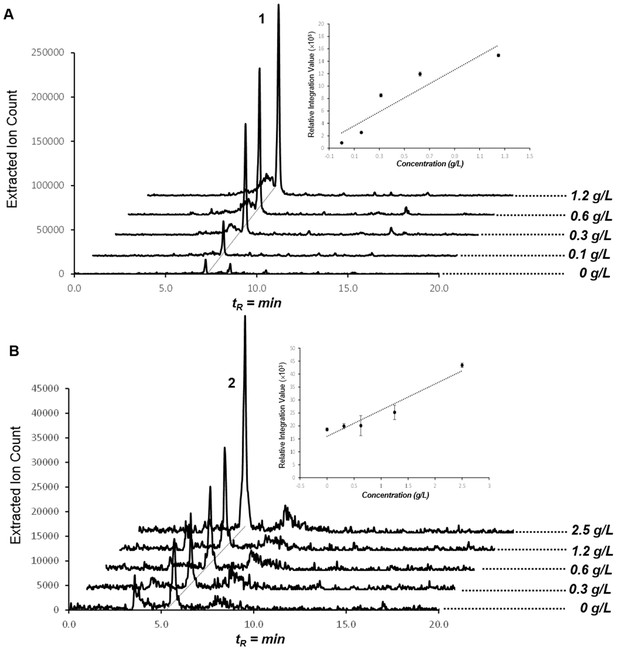
Production of compounds 1 and 2 in dose-dependent α-ketobutyrate (A) and pyruvate (B) feeding studies.
HPLC/MS traces were extracted with m/z 224 corresponding to compound 1 (A) and m/z 210 corresponding to compound 2 (B) from butanol extracts of the culture broths fed with varying concentrations of α-ketobutyrate (A) and pyruvate (B). Dose-response plots for 1 and 2 were determined using extracted peak integration values.
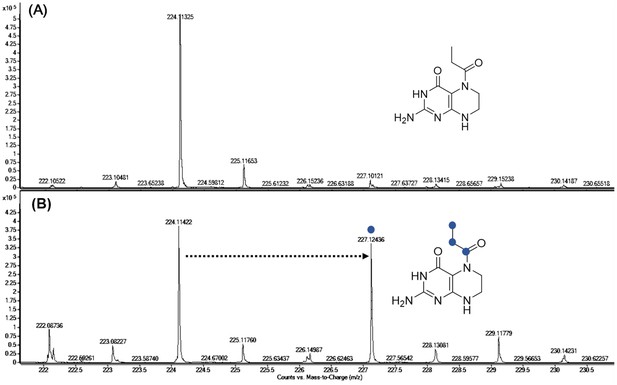
Characterization of 13C4-α-ketobutyrate incorporation in compound 1.
Incorporation of 13C4-α-ketobutyrate into compound 1 (B) led to a clear 3 Da mass shift of native 1 (A) via oxidative decarboxylation of α-ketobutyrate.
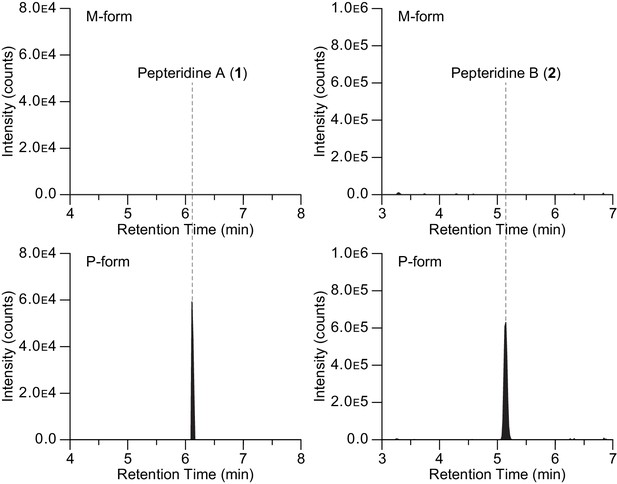
Extracted ion counts LC/HR-ESI-QTOF-MS analysis of pepteridines A (left panel) and B (right panel) from two genetically engineered P. luminescens strains locked in the phenotypic variants M-form (top) and P-form (bottom).
https://doi.org/10.7554/eLife.25229.026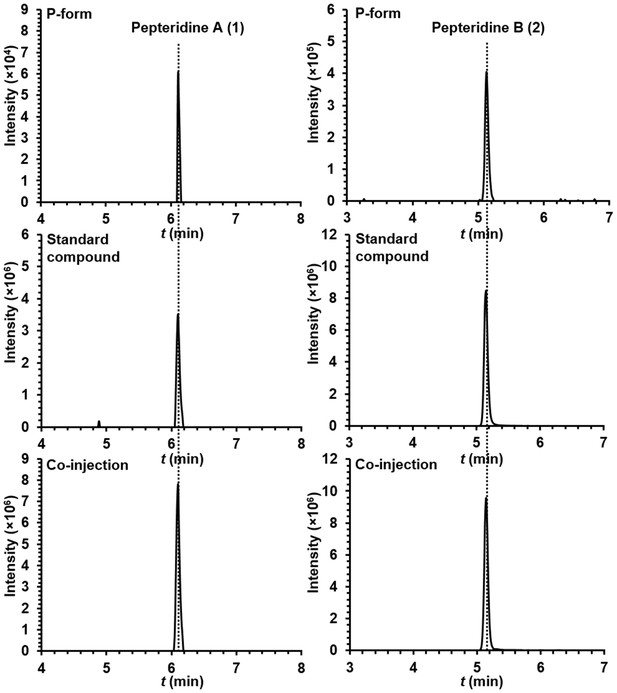
Extracted ion counts chromatograms from LC/HR-ESI-QTOF-MS analysis of pepteridines A (left panel) and B (right panel) from the butanol extracts of P-form culture broth (top), standard compounds (middle), and co-injection (bottom).
https://doi.org/10.7554/eLife.25229.027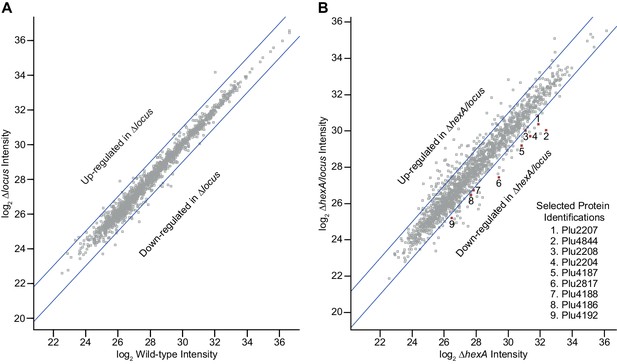
Quantitative proteomic analysis of a Δlocus strain in a wild-type background (A) and ΔhexA background (B).
https://doi.org/10.7554/eLife.25229.028-
Figure 8—source data 1
Proteins increased in WT vs. WTΔlocus strains by LC-MS/MS.
Intensities from label-free quantification calculated using MaxQuant, averaged for biological triplicate samples, then log2 transformed. Proteins presented exhibited ≥2 fold increased average intensity in WT compared with WTΔlocus. Two-tailed t-test performed using a cutoff of FDR = 0.01 and 2-fold signal difference, or p<0.05.
- https://doi.org/10.7554/eLife.25229.029
-
Figure 8—source data 2
Proteins increased in WTΔlocus vs. WT strains by LC-MS/MS.
Intensities from label-free quantification calculated using MaxQuant, averaged for biological triplicate samples, then log2 transformed. Proteins presented exhibited ≥2 fold increased average intensity in WTΔlocus compared with WT. Two-tailed t-test performed using a cutoff of FDR = 0.01 and 2-fold signal difference, or p<0.05.
- https://doi.org/10.7554/eLife.25229.030
-
Figure 8—source data 3
Proteins increased in ΔhexA vs. ΔhexAΔlocus strains by LC-MS/MS.
Intensities from label-free quantification calculated using MaxQuant, averaged for biological triplicate samples, then log2 transformed. Proteins presented exhibited ≥2 fold increased average intensity in ΔhexA compared with ΔhexAΔlocus. Two-tailed t-test performed using a cutoff of FDR = 0.01 and 2-fold signal difference, or p<0.05 (+ indicates statistical significance).
- https://doi.org/10.7554/eLife.25229.031
-
Figure 8—source data 4
Proteins increased in ΔhexAΔlocus vs. ΔhexA strains by LC-MS/MS.
Intensities from label-free quantification calculated using MaxQuant, averaged for biological triplicate samples, then log2 transformed. Proteins presented exhibited ≥2 fold increased average intensity in ΔhexAΔlocus compared with ΔhexA. Two-tailed t-test performed using a cutoff of FDR = 0.01 and 2-fold intensity difference, or p<0.05 (+ indicates statistical significance).
- https://doi.org/10.7554/eLife.25229.032
-
Figure 8—source data 5
All proteins observed in WT and WTΔlocus strains by LC-MS/MS.
Intensities from label-free quantification calculated using MaxQuant and log2 transformed. Intensities for biological triplicate samples are presented. Average fold change shown for WT compared with WTΔlocus. Two-tailed t-test performed using a cutoff of FDR = 0.01 and 2-fold signal difference, or p<0.05 (+ indicates statistical significance).
- https://doi.org/10.7554/eLife.25229.033
-
Figure 8—source data 6
All proteins observed in ΔhexA and ΔhexAΔlocus strains by LC-MS/MS.
Intensities from label-free quantification calculated using MaxQuant and log2 transformed. Intensities for biological triplicate samples are presented. Average fold change shown for ΔhexA compared with ΔhexAΔlocus. Two-tailed t-test performed using a cutoff of FDR = 0.01 and 2-fold signal difference, or p<0.05 (+ indicates statistical significance).
- https://doi.org/10.7554/eLife.25229.034
Tables
Genetic dependency of molecular features.
| Gene | Dependent molecular features |
|---|---|
| plu2793 | 114 |
| plu2794 | 31 |
| plu2795 | 19 |
| plu2796 | 37 |
| plu2797 | 10 |
| plu2798 | 21 |
| plu2799 | 26 |
| plu2793/plu2796 | 12 |
-
Of the wild-type molecular features, this table denotes how many of them were deleted (i.e., not detected) in each mutant strain.
NMR spectral data of pepteridine A (1) and B (2) in DMSO-d6.
| Pepteridine A (1) | |||||
|---|---|---|---|---|---|
| No. | δCa | type | δHb | Mult (J in Hz) | HMBC |
| 1 | N | ||||
| 2 | 154.6 | C | |||
| 3 | NH | 10.07 | br s | ||
| 4 | 157.3 | C | |||
| 4a | 93.1 | C | |||
| 5 | N | ||||
| 6 | 38.5 | CH2 | 4.54 | dd (12.2, 3.6) | C-1′, C-4a |
| 2.33 | m | ||||
| 7 | 41.9 | CH2 | 3.30 | d (12.2) | |
| 2.97 | dt (12.0, 4.2) | C-6 | |||
| 8 | NH | 6.96 | br s | C-4a, C-6, C-7 | |
| 8a | 153.1 | C | |||
| 9 | N | ||||
| 1′ | 174.0 | C | |||
| 2′ | 26.5 | CH2 | 2.57 | dt (15.0, 7.4) | C-1′, C-3′ |
| 2.15 | dt (14.8, 7.3) | C-1′, C-3′ | |||
| 3′ | 9.7 | CH3 | 0.88 | t (7.4) | C-1′, C-2′ |
| NH2 | 6.25 | br s | |||
| Pepteridine B (2) | |||||
|---|---|---|---|---|---|
| No. | δCa | type | δHb | Mult (J in Hz) | HMBC |
| 1 | N | ||||
| 2 | 155.2 | C | |||
| 3 | NH | 10.04 | br s | ||
| 4 | 157.3 | C | |||
| 4a | 93.5 | C | |||
| 5 | N | ||||
| 6 | 38.3 | CH2 | 4.52 | dd (12.1, 3.5) | C-1′, C-4a |
| 2.32 | dt (11.7, 2.4) | ||||
| 7 | 41.8 | CH2 | 3.30 | d (12.0) | |
| 2.98 | dt (11.9, 4.1) | C-6 | |||
| 8 | NH | 6.96 | d (4.0) | ||
| 8a | 153.3 | C | |||
| 9 | N | ||||
| 1′ | 170.6 | C | |||
| 2′ | 22.5 | CH3 | 1.97 | s | C-1′ |
| NH2 | 6.22 | br s | |||
-
NMR spectra were recorded at b600 MHz for 1H NMR and a100 MHz for 13C NMR, respectively.
Additional files
-
Supplementary file 1
Molecular feature list and primers used.
(A) Molecular feature list dependent on the presence of the wild-type pathway. (B-D) Primers used.
- https://doi.org/10.7554/eLife.25229.035





