Co-targeting the tumor endothelium and P-selectin-expressing glioblastoma cells leads to a remarkable therapeutic outcome
Figures
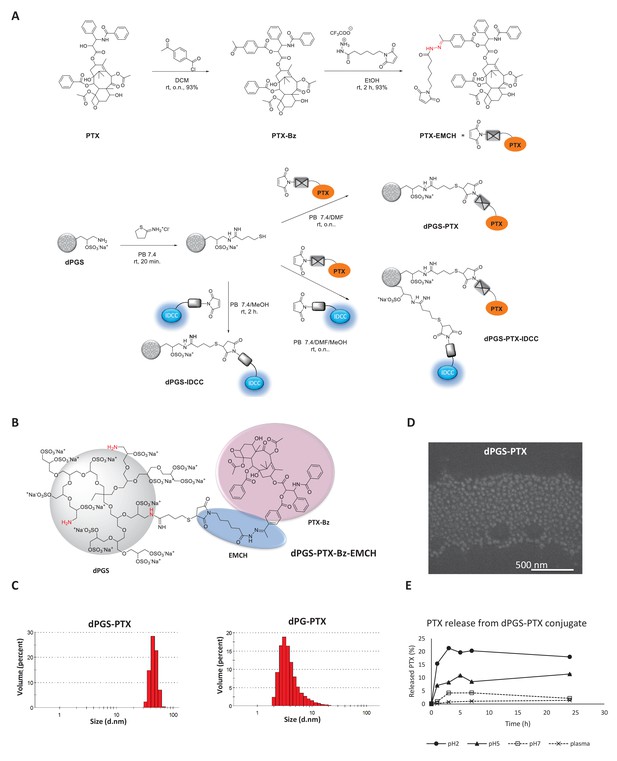
Synthesis and physicochemical characterization of dPG-PTX nanoconjugates.
(A) Scheme depicting the synthesis of PTX-EMCH and dPGS conjugates. (B) Chemical structure of dPGS-PTX. (C) Size distribution of dPGS-PTX and dPG-PTX nanoconjugates, obtained by zetasizer ZS. Data is representative of 3 individual experiments. (D) SEM image of dPGS-PTX nanoconjugate as obtained by Quanta 200 FEG Environmental SEM. (E) Analysis of PTX cumulative release (%) from dPGS-PTX at different pH values and in human plasma, as obtained from HPLC measurements. Data represent mean ± s.d. of three independent experiments.
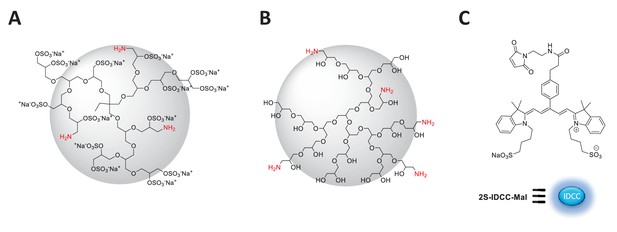
Idealized chemical structure of dendritic nanocarriers based on polyglycerol.
(A) dPGS amine bearing 90% surface sulfate groups and 5% amine groups (B) dPG amine bearing 10% amine groups and 90% neutral hydroxyl groups. (C) Chemical structure of 2S-IDCC-maleimide dye.
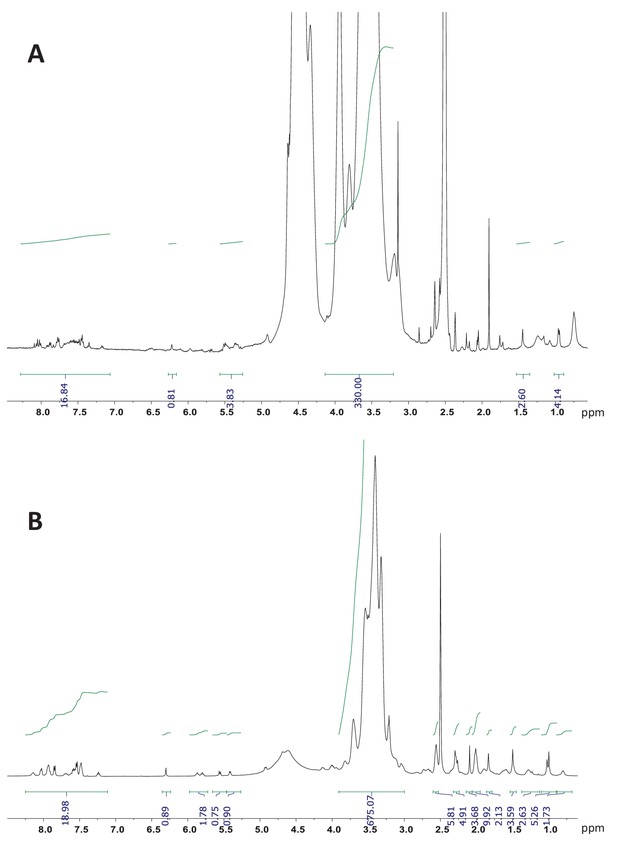
1H NMR (700 MHz, DMSO-d6) of dPGS-PTX and dPG-PTX nanoconjugates.
(A) dPGS-PTX. (B) dPG-PTX. The integration of the aromatic peaks of PTX-Bz (19 hr) between 8.2–7.0 and the peaks corresponding to the dendritic backbone gives a molar ratio drug/conjugate of approximately 1.
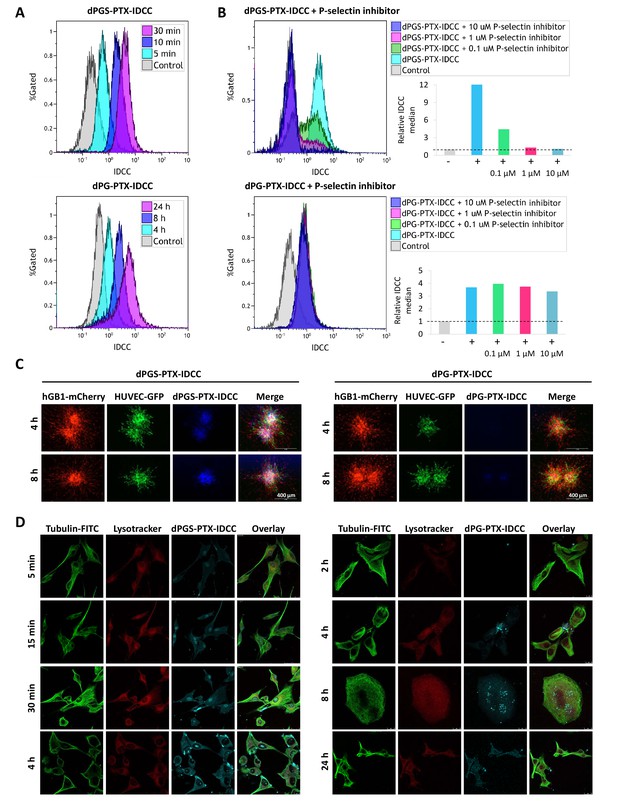
Targeted sulfated conjugate dPGS-PTX-IDCC efficiently internalizes into human glioblastoma cells and spheroids via P-selectin.
(A) Flow cytometry analysis of the cellular uptake of dPGS-PTX-IDCC following 5, 10 and 30 min incubation and dPG-PTX-IDCC following 4, 8 and 24 hr incubation in patient-derived glioblastoma cells (hGB1). (B) Flow cytometry analysis of the cellular uptake of dPGS-PTX-IDCC and dPG-PTX-IDCC following treatment with 0.1, 1 and 10 µM P-selectin inhibitor. (C) Internalization of IDCC-labeled dendritic conjugates (blue) into tumor spheroids composed of mCherry-labeled hGB1 cells (red), GFP-labeled HUVEC (green) and astrocytes. (D) Cellular uptake and intracellular trafficking of dPGS-PTX-IDCC and dPG-PTX-IDCC conjugates in U-87 MG cells. The figure depicts representative confocal images of IDCC-labeled dendritic conjugates (cyan), tubulin (green) and the lysosome (red).
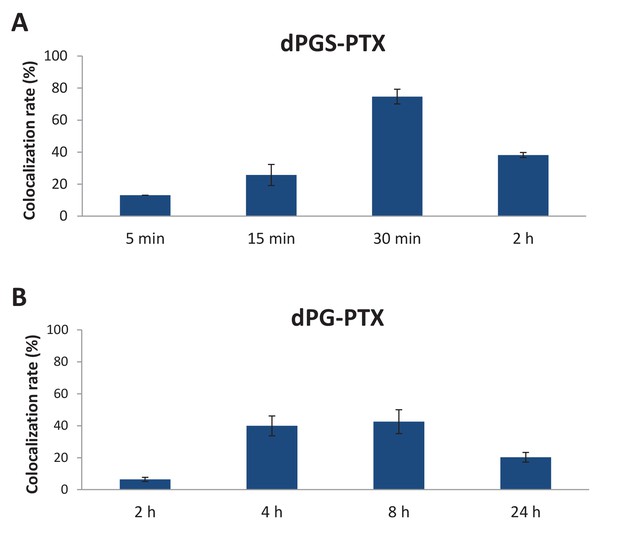
Targeted sulfated conjugate dPGS-PTX colocalizes with tubulin efficiently and rapidly.
(A) Quantification of dPGS-PTX-IDCC colocalization with tubulin. (B) Quantification of dPG-PTX-IDCC colocalization with tubulin. Data represent mean ± s.d.
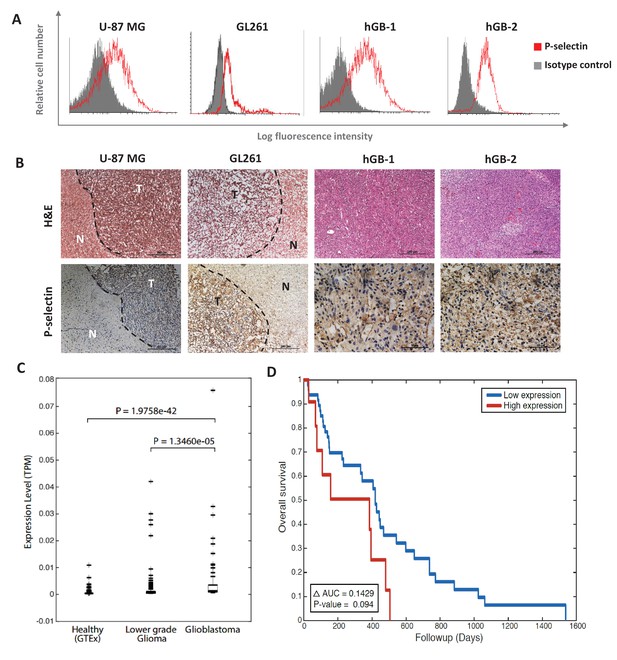
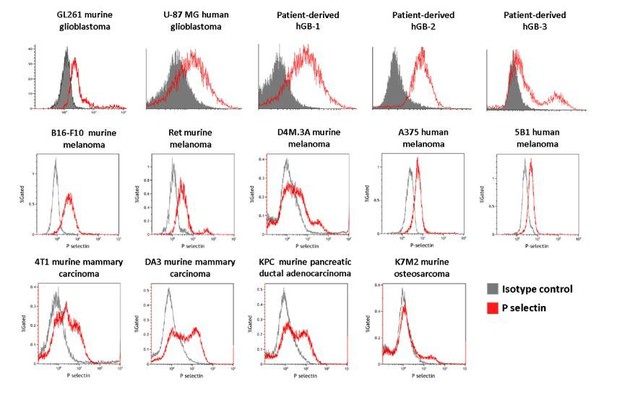
P-selectin is expressed in human and murine glioblastoma.
(A) Flow cytometry analysis of P-selectin expression in U-87 MG, GL261 and two patient-derived glioblastoma cell lines. Images are representative of 3 individual experiments. (B) Representative H&E staining and immunohistochemistry staining for P-selectin expression in intracranial U-87 MG and GL261 tumors and in glioblastoma patient specimens. Positive staining is shown in brown. hGB-1 and hGB-2: human glioblastoma patient-derived cells and tissues. T: Tumor; N: Normal. (C) Comparison of P-selectin expression in healthy brain, lower grade gliomas and glioblastoma from data obtained from the Genotype-Tissue Expression (GTEx) collection and from The Cancer Genome Atlas (TCGA). (D) Kaplan-Meier survival curves obtained from TCGA data of glioblastoma patients with high and low P-selectin expression (using 63 samples with top and bottom 10% of SELP expression).
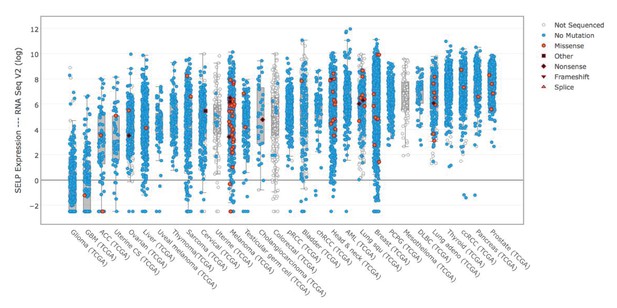
P-selectin (SELP) is differentially expressed across various cancer types.
Comparison of P-selectin expression in various cancer types from data obtained from the TCGA data portal (sorted by the median).
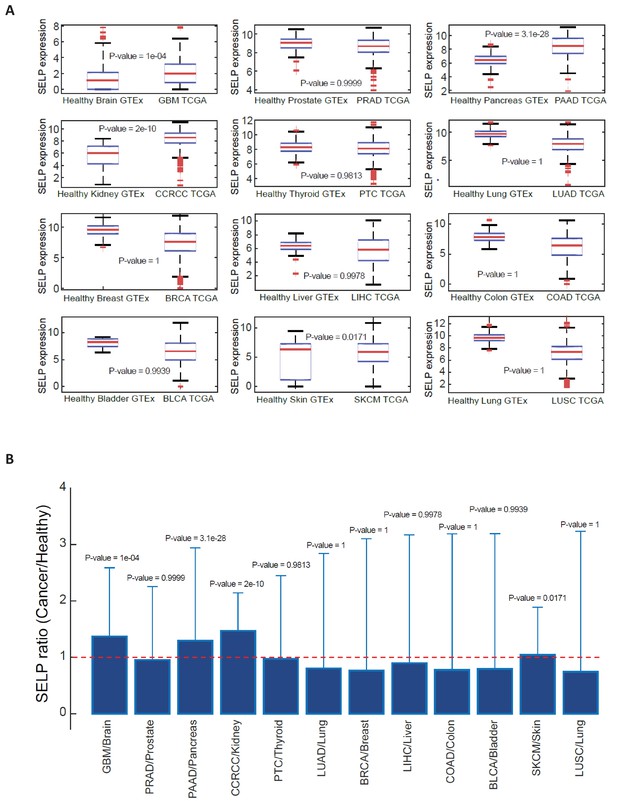
SELP expression in healthy and cancerous tissues.
Data was obtained from XENA database (http://xena.ucsc.edu/public-hubs/), which normalizes gene expression from TCGA and GTEx together. (A) For each tissue type, the distribution of SELP gene in healthy versus cancerous tissue was compared via a Wilcoxon one-sided rank sum test, and the boxplots are presenting the distribution of SELP in each tissue. (B) The bar plots are showing the ratio of mean expression in cancerous vs. healthy in each tissue type, and the error bars represent the standard deviation over all sample points.
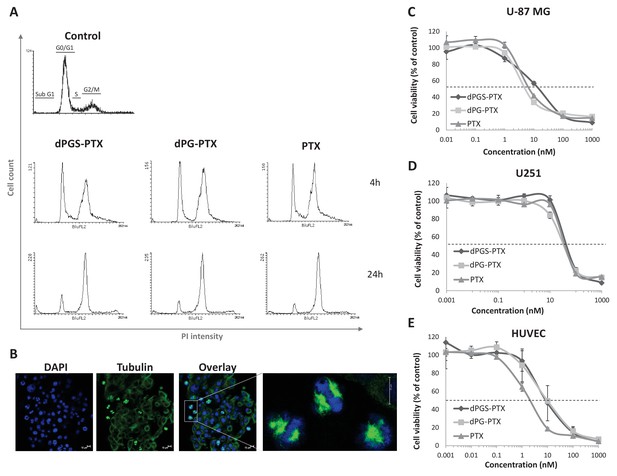
The anti-proliferative activity of PTX is retained following conjugation with dPG or dPGS nanocarriers.
(A) Flow cytometry analysis of cell cycle following treatment with dPG-PTX, dPGS-PTX or free PTX at different time points. Images are representative of 3 individual experiments. (B) Representative confocal images of cells treated with dPGS-PTX for 8 hr. The nucleus is stained with DAPI (blue) and tubulin is stained with FITC-labeled antibody (green). (C–E) U-87 MG and U251 human glioblastoma cells and human umbilical vein endothelial cells (HUVEC) were incubated with serial concentrations of dPG-PTX, dPGS-PTX or free PTX for 72 hr and growth inhibition was evaluated. Data represent mean ± s.d. of triplicate wells. Graphs are representative of 3 individual experiments.
-
Figure 4—source data 1
Raw data of U-87 MG, U251 and HUVEC proliferation assays.
- https://doi.org/10.7554/eLife.25281.013
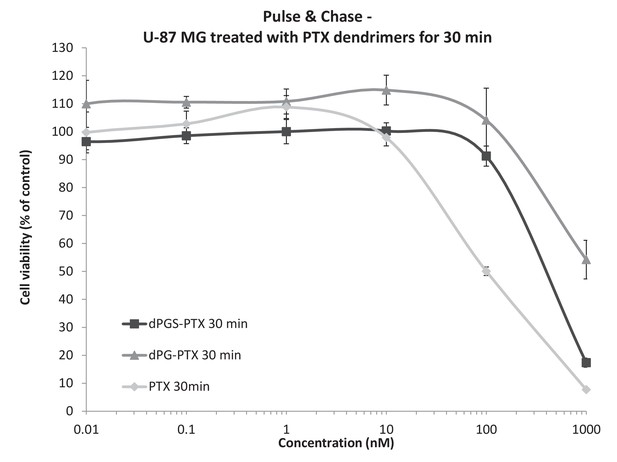
Differences in the in vitro activity of the dendritic conjugates are attributed to differences in their internalization kinetics.
U-87 MG cells were incubated with serial concentrations of dPG-PTX, dPGS-PTX or free PTX for 30 min, washed, left to grow for additional 72 hr and analyzed for growth inhibition. Data represent mean ± s.d. of triplicate wells. Graph is representative of 3 individual experiments.
-
Figure 4—figure supplement 1—source data 1
Raw data of U-87 MG proliferation assay following short-term exposure to treatments.
- https://doi.org/10.7554/eLife.25281.014
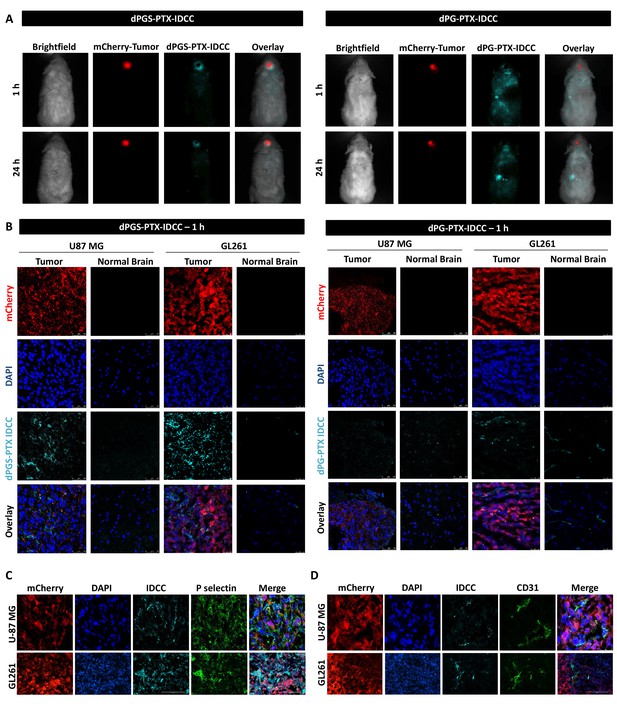
dPGS-PTX preferably accumulates in intracranial tumors.
(A) Representative non-invasive fluorescence images of mice bearing mCherry-labeled U-87 MG tumors (red) 1 hr and 24 hr following intravenous injection of IDCC-labeled dendritic conjugates (cyan) (n = 2). (B). Representative confocal images of brain sections of mice bearing U-87 MG or GL261 tumors 1 hr following intravenous injection of the dendritic conjugates (n = 2–3). Images depict DAPI-stained nucleus (blue), mCherry-labeled tumor cells (red) and IDCC-labeled dendritic conjugates (cyan). (C–D) Representative immunohistochemical staining for P-selectin (C) or CD31 (D) in U-87 MG and GL261 tumors following administration of dPGS-PTX-IDCC. Positive staining is shown in green (n = 3).
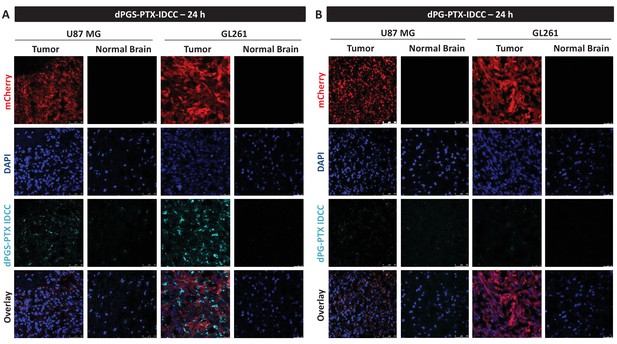
dPGS-PTX preferably accumulates in intracranial tumors at 24 hr.
Representative confocal images of brain sections of mice bearing U-87 MG or GL261 tumors 24 hr following intravenous injection of the dendritic conjugates (n = 2–3). Images depict DAPI-stained nucleus (blue), mCherry-labeled tumor cells (red) and IDCC-labeled dendritic conjugates (cyan). (A) dPGS-PTX-IDCC 24 hr following injection. (B) dPG-PTX-IDCC 24 hr following injection.
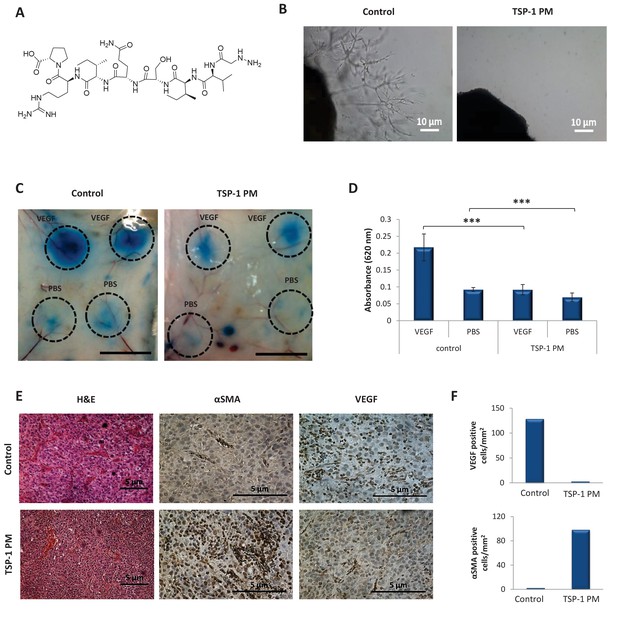
TSP-1 PM reduces the angiogenic potential of glioblastoma.
(A) Chemical structure of ABT-898, a TSP-1 mimetic peptide. (B) Sprouting of endothelial cells from mouse aortic following 8 days incubation with conditioned media collected from U-87 MG cells, in the absence or presence of TSP-1 PM. Images are representative of 3 individual experiments. (C) Assessment of vascular permeability by Miles assay following treatment with TSP-1 PM (n = 3). (D) Quantification of Evans Blue dye extracted from the skin. Data represent mean ± s.e.m. ***p<0.01. (E) Representative immunohistochemical staining of U-87 MG tumors (n = 3). Sections were stained with H&E or stained for αSMA and VEGF. (F) Quantification of αSMA or VEGF positive cells within U-87 MG tumors.
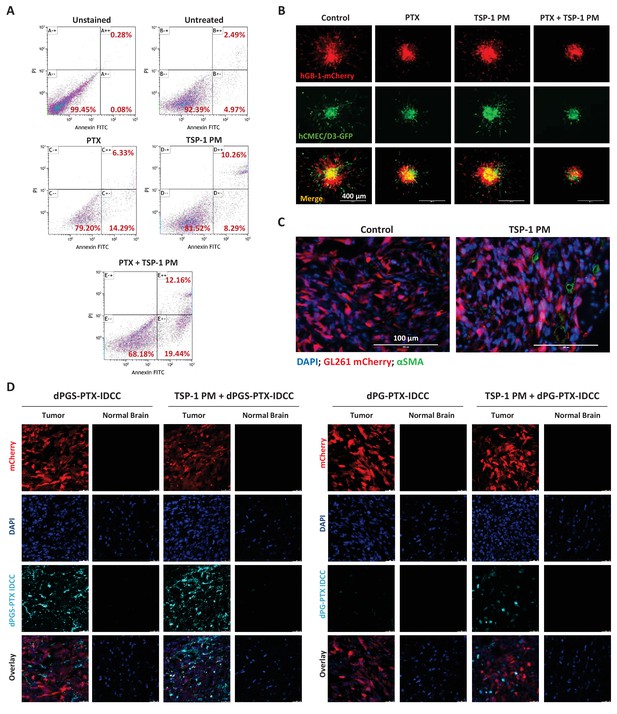
TSP-1 PM synergizes with dPGS-PTX to promote enhanced accumulation at the tumor site and cellular apoptosis.
(A) Flow cytometry analysis of apoptosis in HUVEC following treatment with 1 ng/ml TSP-1 PM, 10 nM PTX or their combination. (B) Representative images of 3D spheroid invasion following treatment with 1 ng/ml TSP-1 PM, 10 nM PTX or their combination. (C) Representative images of αSMA staining in GL261 tumors following treatment with saline (control) or 100 mg/kg TSP-1 PM. (D) Representative confocal images of brain sections 24 hr following intravenous injection of the dendritic conjugates, in the presence or in the absence of a 7 day pre-treatment with TSP-1 PM (100 mg/kg; QD IP). Images depict DAPI-stained nucleus (blue), mCherry-labeled tumor cells (red) and IDCC-labeled dendritic conjugates (cyan) (n = 3).
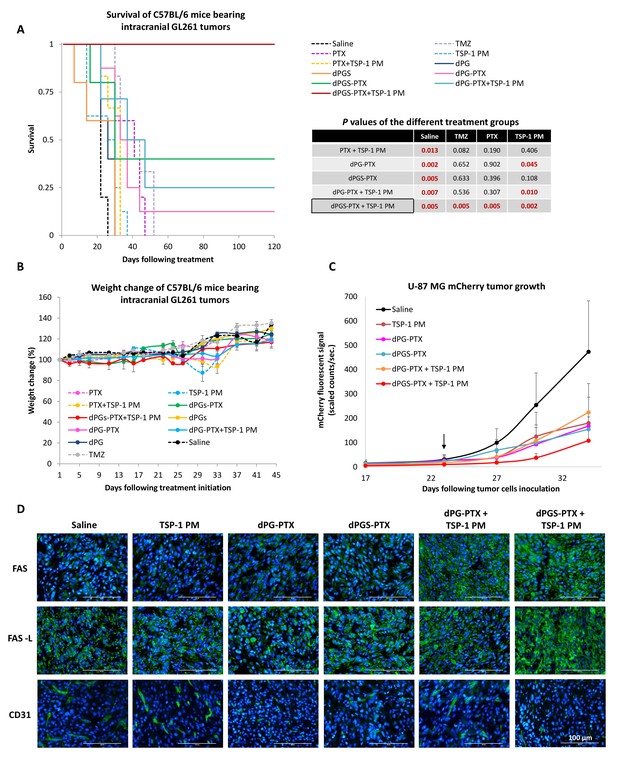
Combination therapy of dPGS-PTX with TSP-1 PM inhibits tumor growth and prolongs the survival of mice bearing intracranial glioblastoma.
(A) Kaplan-Meier curve for survival of mice bearing murine GL261 intracranial tumors following systemic injection of treatments (n = 4–8). Table shows statistical significance between different treatment groups. P values were determined using log rank test. (B) Body weight change, expressed as percent change from the day of treatment initiation. Data represent mean ± s.e.m. (n = 4–8). (C) Tumor growth of mCherry-labeled U-87 MG tumors following systemic administration of treatments (n = 3–8). Arrow points to time of treatment initiation. Data is represented as scaled counts/sec. of the mCherry fluorescent signal, detected by CRI Maestro imaging system. (D) Representative immunohistochemical images of U-87 MG tumors treated with TSP-1 PM, dPG-PTX, dPGS-PTX or their combinations. Tissues were stained for Fas/CD95, Fas-L/CD178 and CD31. Tumor cells are shown in blue (DAPI) and positive immunostaining is shown in green (n = 3). Data represent mean ± s.e.m.
-
Figure 8—source data 1
Raw data of Kaplan-Meier survival curve.
- https://doi.org/10.7554/eLife.25281.021
-
Figure 8—source data 2
Raw data of the fluorescent signal of mCherry labeled U-87 MG intracranial tumors.
- https://doi.org/10.7554/eLife.25281.022
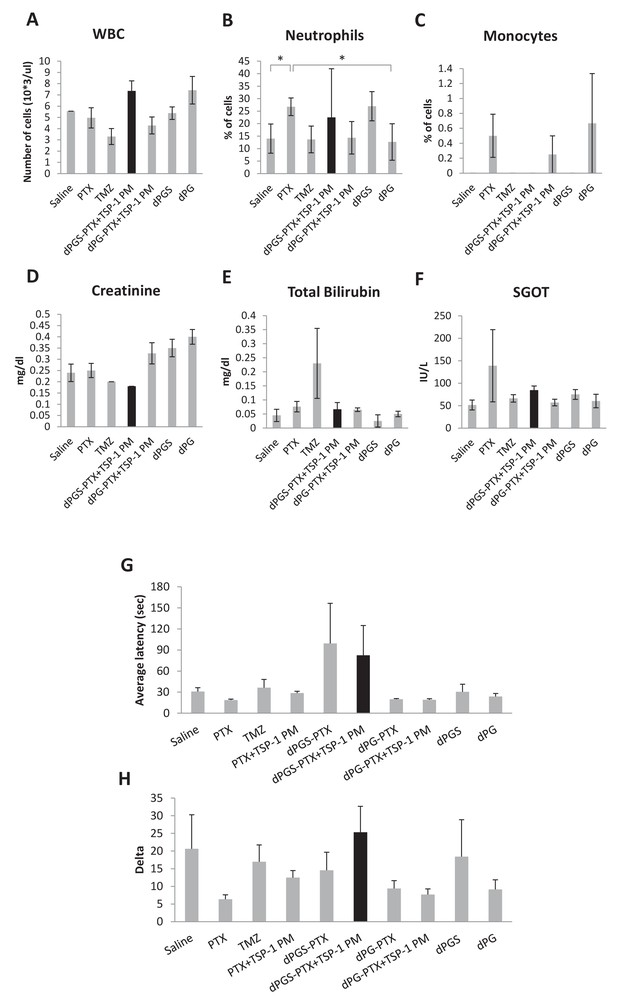
Combination treatment of dPGS-PTX and TSP-1 PM did not induce systemic toxicity.
(A–F) Blood was drawn from glioblastoma tumor-bearing mice (n = 5–8) following treatments and was analyzed for blood count (A–C) and biochemistry analysis (D–F). WBC, white blood cells; SGOT, serum glutamic oxaloacetic transaminase. (G) Average latency of mice to fall from the Rotarod. Data represent mean ± s.e.m. (n = 2–4). (H) Delta of latency to fall from a Rotarod between their first and last performance. Data represent mean ± s.e.m. (n = 2–4). *p<0.05.
-
Figure 8—figure supplement 1—source data 1
Raw data of blood count and biochemistry analysis.
- https://doi.org/10.7554/eLife.25281.023
-
Figure 8—figure supplement 1—source data 2
Raw data of mice Rotarod experiments.
- https://doi.org/10.7554/eLife.25281.024
Tables
Summary of physicochemical characterization of dendritic conjugates
https://doi.org/10.7554/eLife.25281.005| Compound | Molar ratio PTX/Conjugate | Molecular weight (Da) | Hydro-dynamic diameter (nm) | PDI | Diameter by SEM (nm) | Zeta potential (mV) |
|---|---|---|---|---|---|---|
| dPGS-PTX | ~1 | 12,807 | 45.9 ± 6.7 | 0.693 | 50 ± 20 | −47.1 ± 5.1 |
| dPG-PTX | ~1 | 11,207 | 4.2 ± 2.6 | 0.268 | NA | 16.5 ± 4.3 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.25281.025