Crosstalk within a functional INO80 complex dimer regulates nucleosome sliding
Figures
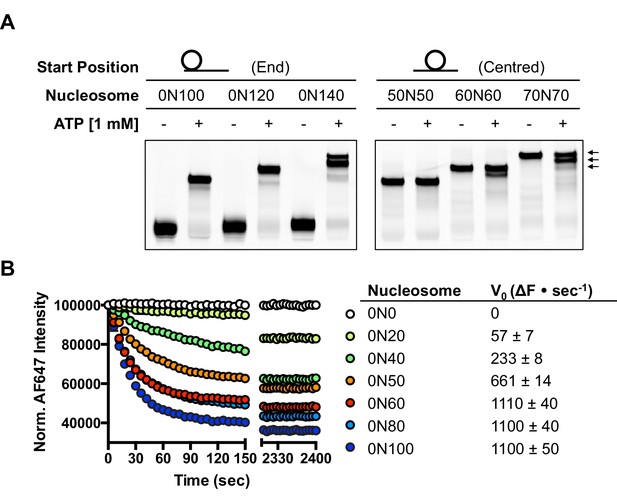
hINO80 positions nucleosomes 50 base pairs from DNA ends.
(A) Nucleosomes positioned either at the end (left panel) or centre (right panel) of DNA fragments were incubated with hINO80. As reported previously (Udugama et al., 2011) overhangs of 100 bp or less resulted in centrally positioned nucleosomes, while those positioned at the centre remained centrally located. (This property, as well as others (nucleosome binding affinity, sliding activity and ATPase), are also observed for recombinant human INO80 complex containing a full length Ino80 subunit and additional (non-conserved) subunits [Figure 1—figure supplement 1]). Increasing the overhang from 100 to 120 bp produced two bands corresponding to a centrally positioned nucleosome and 10 bp from the centre. Increasing to a 140 bp overhang gave three bands – centred, 10 bp from centre and 20 bp from centre (marked with arrows). Incubation of the appropriate centrally positioned nucleosomes gave similar products. (B) (left) FRET-based sliding assay for nucleosomes with different flanking DNA lengths, (right) initial rates of sliding for nucleosomes with different flanking DNA lengths. Comparison with gel-based assay is presented in Figure 1—figure supplement 2.
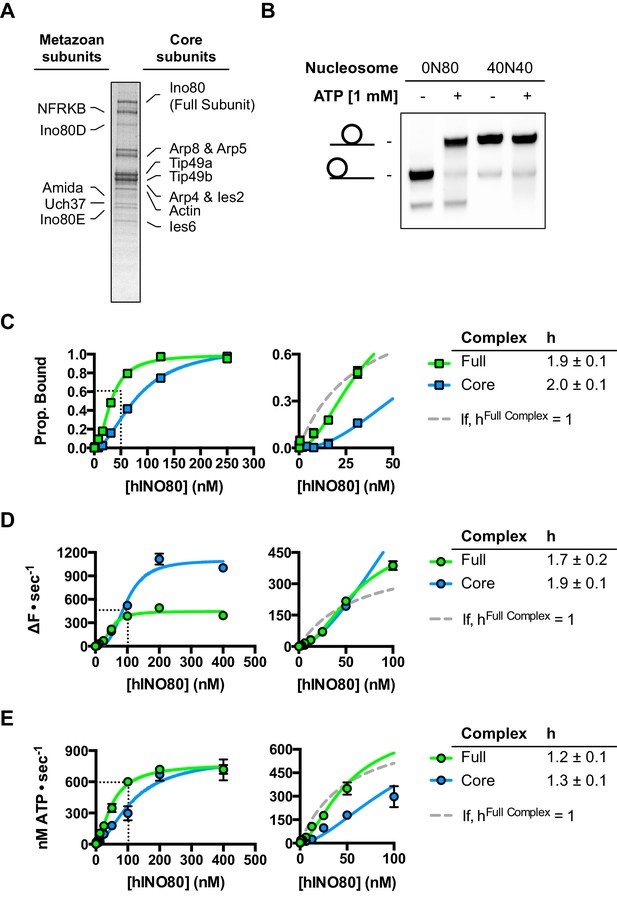
Characterisation of full length recombinant human INO80 complex.
(A) Coomassie-stained SDS gel of full length recombinant human INO80 complex. Subunit identities were confirmed by Western blotting. All subunits identified by Chen et al. (2011) are purified with the complex with the exception of MCRS1. Yields and quality of the complex are not as good as for hINO80 core complex (Figure 1—figure supplement 2) so the main experiments in this study were characterised with the hINO80 core complex which has similar properties to the full complex (Willhoft et al., 2016 and panels (B) - (E) below.). (B) Full human INO80 complex centres mono nucleosomes on short DNA fragments. (C) Full and core human INO80 complexes have similar binding affinity for nucleosomes. Data shown are MST experiments with 0N100 nucleosomes. For these data (and in panels D and E), the dotted line is the fit with h = 1 for the full complex. (D) Full and core human INO80 complexes also show similar cooperativity in sliding 0N100 nucleosomes although the full complex shows a lower activity. (E) Full and core human INO80 complexes also show similar cooperativity and activity in 0N100 nucleosome stimulated ATPase.
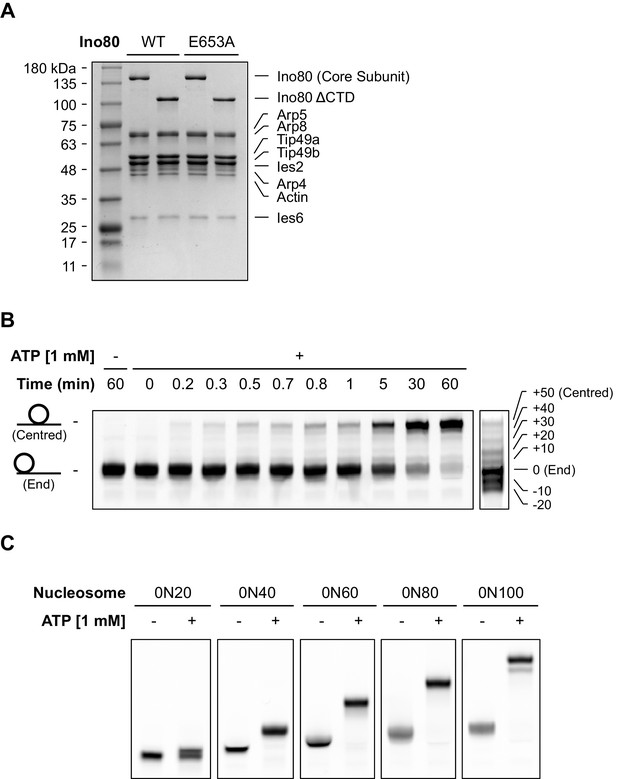
Characterisation of hINO80 complex.
(A) Coomassie-stained SDS gel of protein complexes used for these studies. (B) At a 1:10 INO80:nucleosome ratio, the sliding of 0N100 nucleosomes by hINO80 is highly processive, with appearance of centred nucleosomes very early in the time course and long before most of the nucleosomes have been shifted at all from the ends. This suggests a small proportion of the enzyme is in a highly active state. The data in Figure 3 show that the proportion of this increases as the ratio to nucleosomes increases that we interpret to be two hINO80 complexes binding to each nucleosome. At early time points at low ratios of hINO80:nucleosomes it is evident that discrete bands appear at intermediate positions between the end- and centrally-positioned nucleosomes. Using different length overhangs, we observed one intermediate band for each additional 20 bp of overhang, corresponding to a shift towards the centre of 10 bp each time. Hence, the bands are spaced approximately 10 bp apart. At first, we thought this might represent a step size for the protein as suggested by studies on other remodellers, but we observe a similar banding pattern if we heat a sample of our nucleosomes in the absence of enzyme revealing that this is an intrinsic property of the nucleosomes themselves rather than of the enzyme catalysed reaction (see right end panel). We prepare end-positioned nucleosomes by reconstitution on the strongly positioning Widom 601 sequence (Lowary and Widom, 1998). The most plausible explanation for the ‘intermediate’ bands is, therefore, that phasing of the DNA sequence within the Widom 601 sequence favours positions every 10 bp (i.e. one turn of the duplex) to bring regions into phase that favour being placed on the inside or outside of the nucleosome wrap, a property of DNA sequences that has been recognised for over three decades (Drew and Travers, 1985). The intermediate bands we observe could therefore correspond to either a ‘step-size’ or simply pausing of nucleosomes at preferred sites during translocation by multiple smaller steps between these sites. Since nucleosomes wrap just under 150 bp of duplex around the core, and INO80 slides mono-nucleosomes to the centre of DNA fragments (Udugama et al., 2011), it would require an overhang of around 300 bp to completely shift an end-positioned nucleosome away from the influence of the Widom 601 sequence. (C) hINO80 can centre nucleosomes with short overhangs. Nucleosomes with 20, 40, 60, 80 and 100 bp overhangs all shift towards the centre.
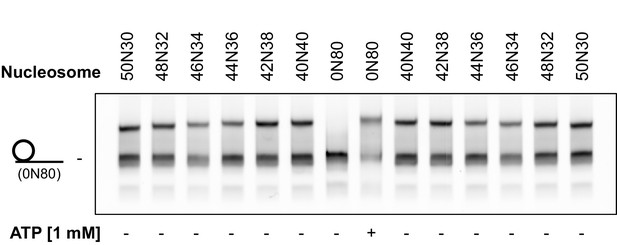
Nucleosomes shifted by hINO80 are located close to the centre of a short DNA fragment.
Specific nucleosome markers were prepared by ligating different length overhangs on either side of nucleosomes prepared on a WIDOM 601 sequence as described in the Methods. Sliding reactions with 0N80 nucleosomes were carried out in the presence or absence of ATP. The 0N80 nucleosome was added to each of the marker lanes as an additional standard to show even running of the gel. The sliding assay was incubated for 1 hr at 37°C as described in Materials and methods. The spread of product bands is similar to that of the markers and the gel allows differential mobility to be detected between marker nucleosomes differing by just 2 bp from the centre showing the spread of products to be within a few base pairs of the central location.
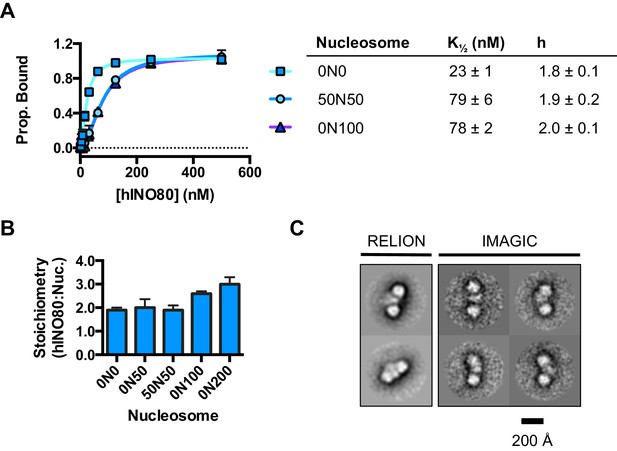
hINO80 binds nucleosomes cooperatively.
(A) Equilibrium binding between hINO80 and nucleosomes determined by MST using 20 nM nucleosome ligands. The data were fitted to a cooperative binding curve (left) to determine K1/2 saturation points and Hill coefficients (h) (right). Raw data and analysis are presented in Figure 2—figure supplement 1. (B) Stoichiometry of hINO80 binding to a variety of nucleosomes. In all experiments, the nucleosome concentration used was 500 nM. The 0N100, 0N0 and 50N50 nucleosomes each bind two hINO80 complexes. However, lengthening of the overhang provides an adventitious binding site until, at 200 bp, three hINO80 complexes can be accommodated per nucleosome. Data analysis is presented in Figure 2—figure supplement 2. (C) Negative stain electron microscopy of hINO80 complexed with 50N50 nucleosomes. The 2D classes show evidence for two hINO80 complexes flanking a nucleosome. RELION classes (shown on the left) were used to select principal views and contained most of the particles, from which smaller groups were selected for classes in IMAGIC (shown on the right). The IMAGIC classes contained fewer particles but which were more closely aligned in view so show more detail.
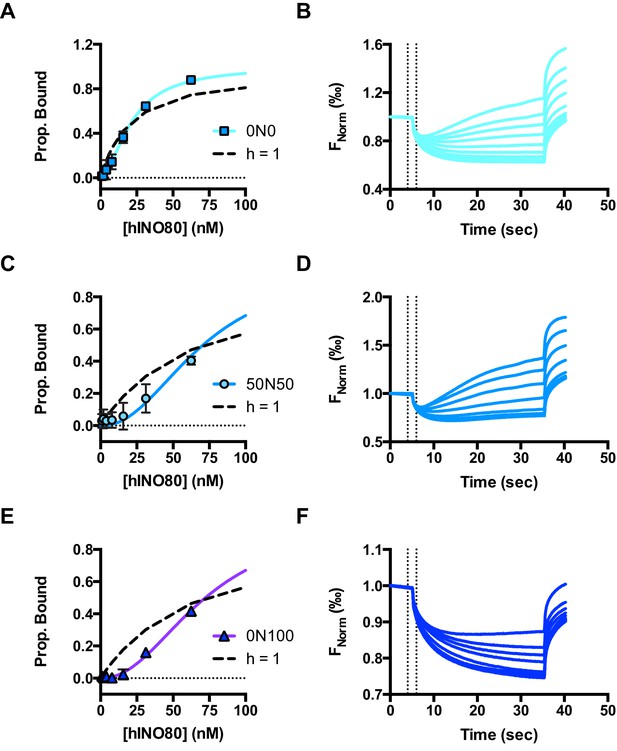
Raw data and fits for hINO80:nucleosome interactions shown in Figure 2A.
(A) Fit of MST binding data at low hINO80 concentrations to emphasise the deviation from fit to a non-cooperative (hyperbolic) curve. These data are for 0N0 nucleosomes. (B) Representative raw MST data traces taken from one series from a group of replicates for 0N0 nucleosome. The dotted lines delineate the region used for analysis. (C) Fit of MST binding data at low hINO80 concentrations to emphasise the deviation from fit to a non-cooperative (hyperbolic) curve. These data are for 50N50 nucleosomes. (D) Representative raw MST data traces taken from one series from a group of replicates for 50N50 nucleosome. The dotted lines delineate the region used for analysis. (E) Fit of MST binding data at low hINO80 concentrations to emphasise the deviation from fit to a non-cooperative (hyperbolic) curve. These data are for 0N100 nucleosomes. (F) Representative raw MST data traces taken from one series from a group of replicates for 0N100 nucleosome. The dotted lines delineate the region used for analysis.
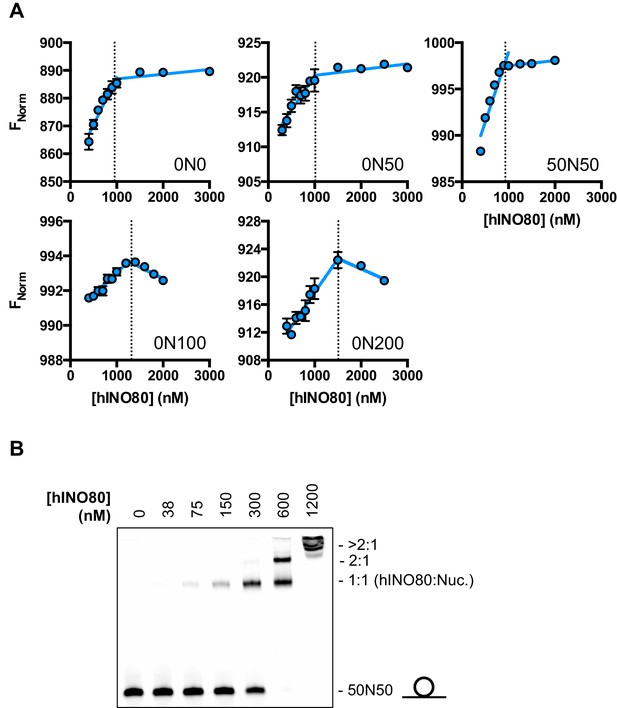
Characterisation of the hINO80:nucleosome interaction.
(A) Raw data and data fit for stoichiometry measurements by MST. All experiments were carried out with 500 nM nucleosome. The reducing FNorm values with longer overhangs are likely due to adventitious binding to the overhang rather than the nucleosome as suggested by the non-stoichiometric values for binding that increase as the overhang lengthens. (B) Gel mobility band shift showing formation of two distinct hINO80:nucleosome complexes. At high concentrations (>600 nM) the complexes aggregate and run at the top of the gel. The centrally positioned nucleosome (50N50) concentration was 300 nM.
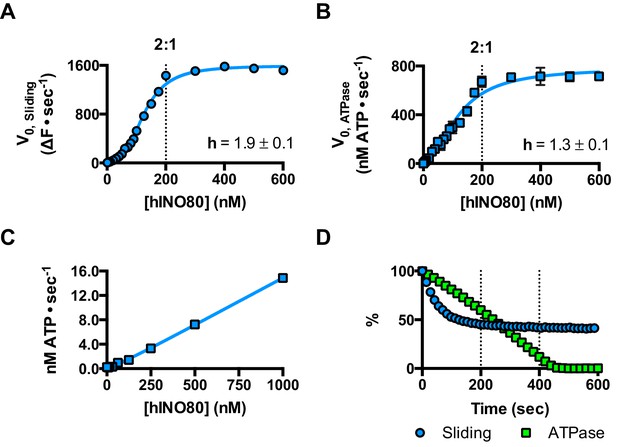
hINO80 operates as a functional dimer.
(A) Sliding activity of nucleosomes increases sigmoidally (h = 1.9 ± 0.1) (Figure 3—figure supplement 1) with respect to hINO80 concentration, peaking at a ratio of two hINO80 complexes per 0N100 nucleosome. Experiments were done at a nucleosome concentration of 100 nM. Experiments at 300 nM nucleosome show similar activity profile again peaking at a ratio of 2:1 hINO80:nucleosome (Figure 3—figure supplement 1). (B) ATPase activity follows a similar profile as sliding activity, reaching a maximum at the same ratio of hINO80:nucleosome, but with a lower cooperativity (h = 1.3 ± 0.1) (Figure 3—figure supplement 1). (C) ATPase activity of hINO80 shows a linear dependence on protein concentration in the absence of nucleosomes. (D) Normalised fluorescence traces corresponding to sliding and ATPase activities of hINO80, at a 2:1 ratio of complex to 0N100 nucleosomes. ATPase activity (determined from the slope) shows a similar rate even after sliding has reached completion (<200 s). Further analysis of these data with 50N50 nucleosomes is presented in Figure 3—figure supplement 2.
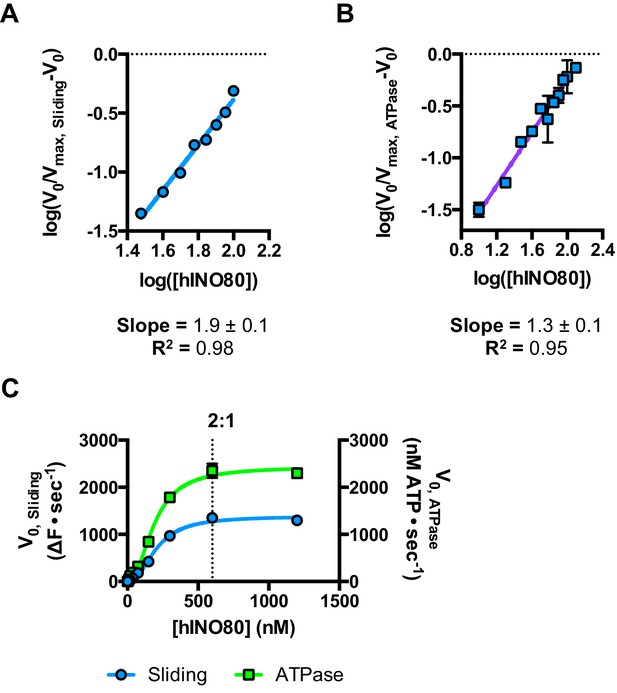
Hill plots and stoichiometry for sliding and ATPase.
(A) Hill plot for sliding activity data shown in Figure 3. (B) Hill plot for ATPase activity shown in Figure 3. (C) Protein concentration dependence of sliding and ATPase activities at 300 nM nucleosome concentration (compare to 100 nM in Figure 3). The stoichiometry remains 2:1.
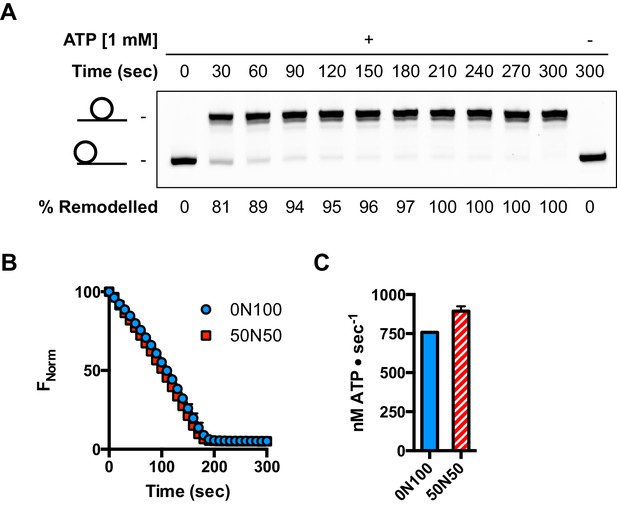
ATPase during and after sliding.
(A) Gel analysis of sliding reaction at 2:1 ratio of hINO80:nucleosome (0N100). Samples from appropriate time points taken from an assay under the same conditions as Figure 3D. Analysis on band intensities shows the reaction to be >90% complete after 90 s. (B) Time course of ATPase activity at saturating concentrations of 50N50 and 0N100 nucleosomes. (C) Nucleosome-stimulated ATPase activity with saturating concentration of 0N100 and 50N50 nucleosomes is similar.
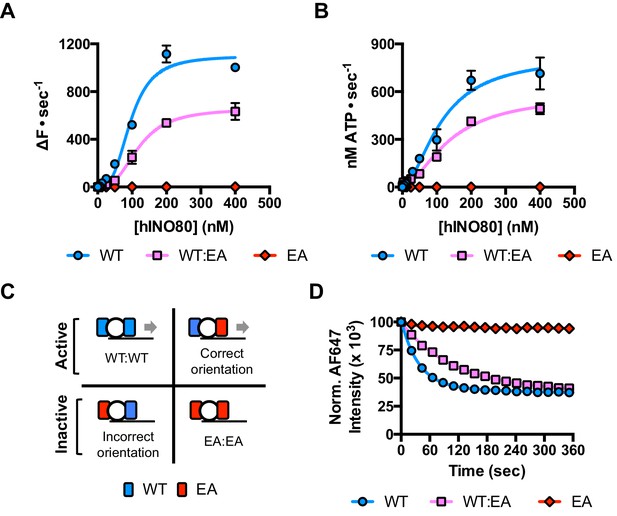
A single ATPase is both necessary and sufficient for sliding.
(A) Concentration dependence of sliding activity for wild type hINO80 (WT), ATPase dead mutant (EA), and an equimolar mixture of the two complexes (WT:EA) at a nucleosome (0N100) concentration of 100 nM. Binding and stoichiometry experiments are shown in Figure 4—figure supplement 1. (B) ATPase activity under the same conditions as (A). (C) Punnett square showing distribution of dimer species in a 1:1 ratio mixture. (D) Time course of sliding reactions at 2:1 hINO80:nucleosome.
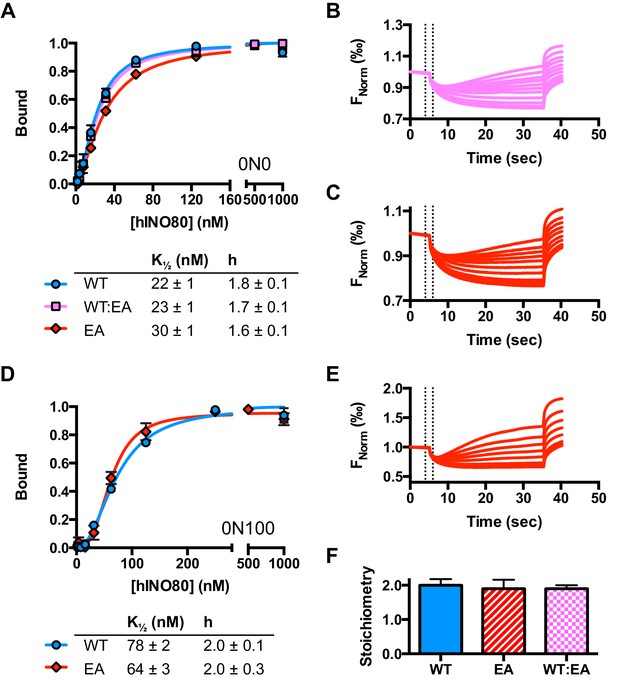
Characterisation of the E653A hINO80 mutant.
(A) Equilibrium binding to 0N0 nucleosomes of wild type (WT), E653A mutant (EA) and a 1:1 mixture (WT:EA) of hINO80 complexes. Analysis of WT data is shown in Figure 2 and Figure 2—figure supplement 1. (B) Representative raw MST data traces taken from one series from a group of replicates for 0N0 nucleosome with the WT:EA mixture. The dotted lines delineate the region used for analysis. (C) Representative raw MST data traces taken from one series from a group of replicates for 0N0 nucleosome with the EA complex alone. The dotted lines delineate the region used for analysis. (D) Equilibrium binding to 0N100 nucleosomes as above. The affinity for nucleosomes (0N0 and 0N100) was indistinguishable with every combination tested. (E) Representative raw MST data traces taken from one series from a group of replicates for 0N100 nucleosome with the EA complex alone. The dotted lines delineate the region used for analysis. (F) Stoichiometry of binding to 0N0 nucleosomes of wild type (WT), E653A mutant (EA) and a 1:1 mixture (WT:EA) of hINO80 complexes.
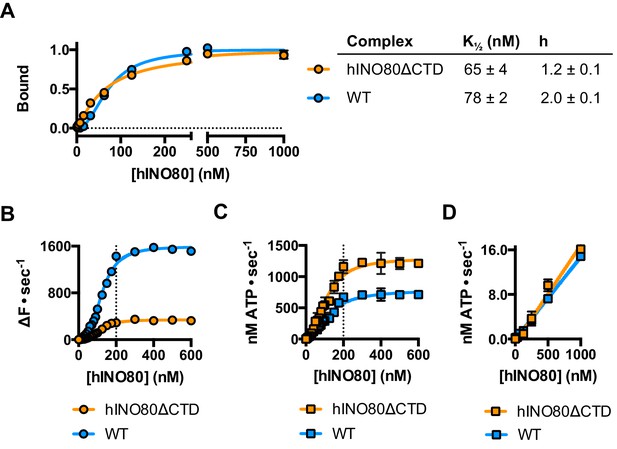
hINO80ΔCTD is deficient in ATPase coupling.
(A) Comparison of nucleosome affinity of hINO80 and hINO80ΔCTD. The nucleosome was 0N100 at a concentration of 20 nM. Hill coefficients and K1/2 values are tabulated on the right. Raw MST data are presented in Figure 5—figure supplement 1. (B) Comparison of protein concentration dependence for sliding activity for hINO80 and hINO80ΔCTD complexes. The maximum rate of sliding for hINO80ΔCTD is approximately 5-fold lower than for hINO80 but is achieved at a 2:1 ratio of complex:nucleosome in both cases. Analysis of the data is presented in Figure 5—figure supplement 1. (C) Comparison of protein concentration dependence for nucleosome-stimulated ATPase activity for hINO80 and hINO80ΔCTD complex. The maximum ATPase rate for hINO80ΔCTD is approximately 2-fold higher than for hINO80 but is still attained at a 2:1 ratio of complex:nucleosome (0N100) in both cases. Analysis of the data is presented in Figure 5—figure supplement 1. (D) Dependence of ATPase activity on protein concentration for hINO80 and hINO80ΔCTD in the absence of nucleosomes.
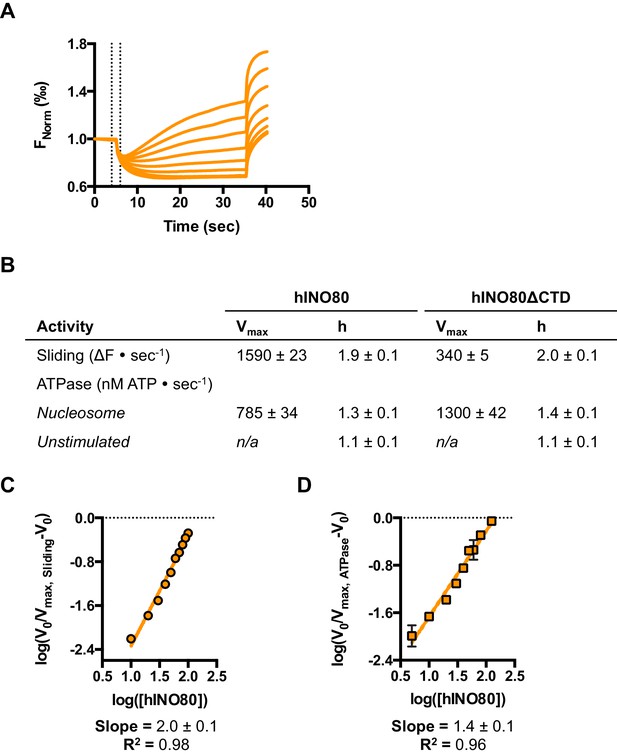
Characterisation of hINO80ΔCTD.
(A) Representative raw MST data traces taken from one series from a group of replicates for 0N100 nucleosome with the hINO80ΔCTD complex. The dotted lines delineate the region used for analysis. (B) Tabulated maximum rates and Hill coefficients for sliding and ATPase activities of hINO80 and hINO80ΔCTD complexes. (C) Hill plot for sliding activity data shown in Figure 5. (D) Hill plot for ATPase activity shown in Figure 5.
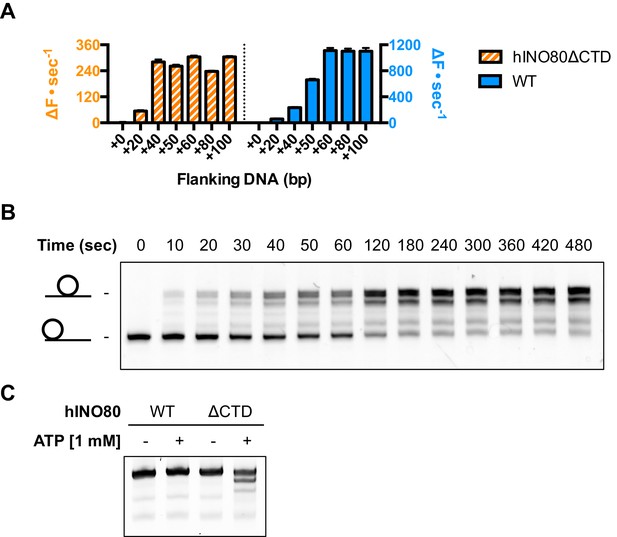
hINO80ΔCTD has altered end sensing.
(A) Initial sliding rates for hINO80 and hINO80ΔCTD on nucleosomes with different length overhangs. Full length hINO80 complex (right) senses DNA ends up to 60 bp while that of the hINO80ΔCTD complex (left) is reduced to under 40 bp. FRET-based assay data are shown in Figure 6—figure supplement 1. (B) hINO80ΔCTD has a reduced ability to centre nucleosomes. Although the majority of nucleosomes still reach the centre, the spread is much greater than for hINO80 (Figure 1 and Figure 3—figure supplement 2). (C) hINO80ΔCTD slides 50N50 nucleosomes away from the central location. These are unaltered by hINO80.
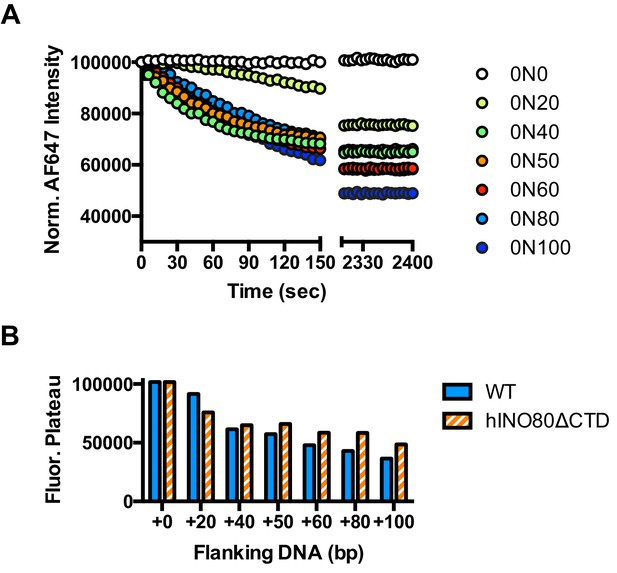
Flanking DNA length dependence of hINO80ΔCTD.
(A) FRET-based sliding assay for nucleosomes with different flanking DNA lengths with hINO80ΔCTD at 2:1 ratio of complex:nucleosome. (B) Plot of final fluorescence end points for reactions in (A). The different end points compared to hINO80 reflect the increased spread of positions on the DNA fragments for hINO80ΔCTD catalysed reactions.
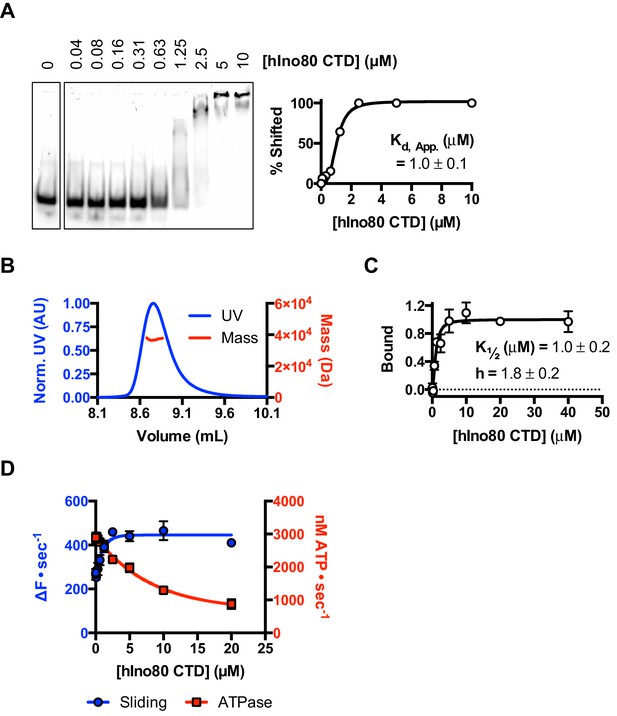
The CTD of hIno80 is a cooperative DNA binding protein.
(A) Gel mobility shift assay using 50 nM 50 bp duplex DNA with increasing concentrations of hIno80CTD. The gel (left) was digitised and band intensity plotted (right) to determine Kd apparent, which is approximately 1 μM. (B) SEC-MALS analysis of the hIno80CTD protein (SDS gel shown in Figure 7—figure supplement 1). The molecular weight was determined to be 38 ± 1.5 KDa compared to a calculated weight of a monomer of 35.3 KDa. (C) Binding of the hIno80CTD to a 50 bp DNA duplex determined by MST. K1/2 is 1.0 ± 0.2 μM with a Hill coefficient of 1.8 ± 0.2. (D) Effect of titrating in hIno80CTD into a reaction of hINO80ΔCTD (200 nM) with 100 nM 0N100 nucleosomes (compare to Figure 5). The ATPase rate decreases but initial sliding rate increases, demonstrating increased coupling of these activities. A Kd apparent for hIno80CTD calculated from the half maximal stimulation is approximately 1 μM. The binding event that regulates ATPase, however, has a lower K1/2 that cannot be determined accurately from these data but appears to be at least an order of magnitude weaker.
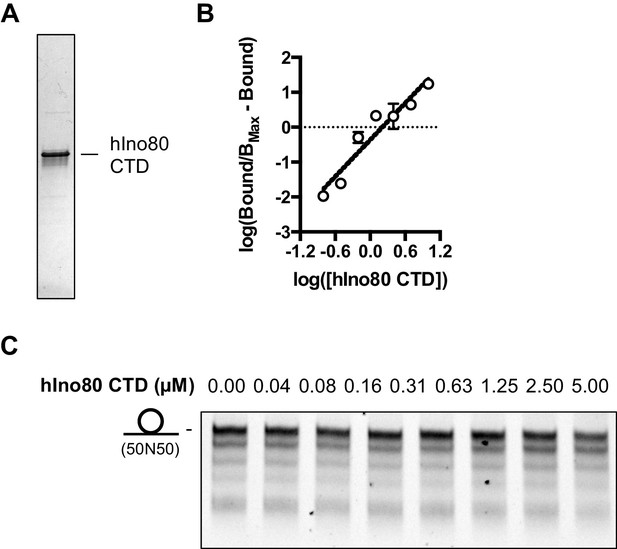
Characterisation of hIno80CTD protein.
(A) SDS gel of the purified Ino80CTD protein. (B) Hill plot for hIno80CTD binding data shown in Figure 7C. (C) Titration of hIno80CTD does not restore end sensing ability of the hINO80ΔCTD complex. Each lane is a sliding reaction carried out as described in the Materials and methods. This should be compared to the hINO80 data in Figure 6.
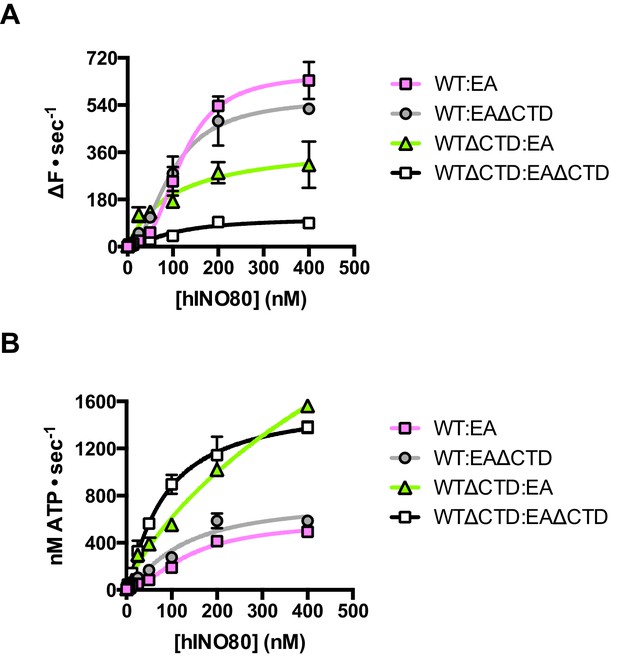
The contribution of the hIno80CTD in cis or trans.
(A) Concentration dependence of sliding activity for equimolar mixtures of different hINO80 complexes as indicated. Complexes are hINO80 (WT), hINO80ΔCTD (WTΔCTD), ATPase dead mutant (EA), and ATPase dead mutant lacking the CTD (EAΔCTD). Assays contained a nucleosome (0N100) concentration of 100 nM. The data are tabulated in Figure 8—source data 1. (B) ATPase activity under the same conditions as (A).
-
Figure 8—source data 1
Tabulated maximum rates and Hill coefficients for sliding and ATPase activities of mixtures of INO80 complexes as indicated.
- https://doi.org/10.7554/eLife.25782.021





