Parallel evolution of influenza across multiple spatiotemporal scales
Figures
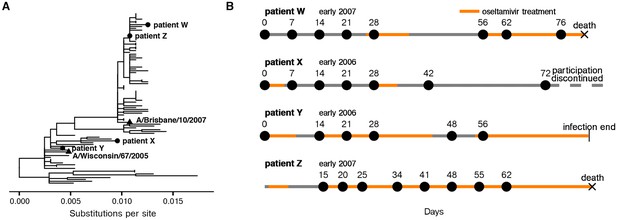
Long-term H3N2 influenza infections in four immunocompromised patients.
(A) Phylogenetic relationship between initial patient consensus sequences and 63 unique circulating influenza strains collected in the USA from 2004 to 2007, as inferred from the HA gene. (B) Overview of patient influenza infections and treatments. Periods of oseltamivir treatment are shown in orange. Dates of sequenced nasal wash samples are calculated relative to the first influenza-positive nasal wash. Low-quality samples are not shown here and were excluded from downstream analysis. Materials and methods and Figure 1—figure supplement 1 give full clinical histories.
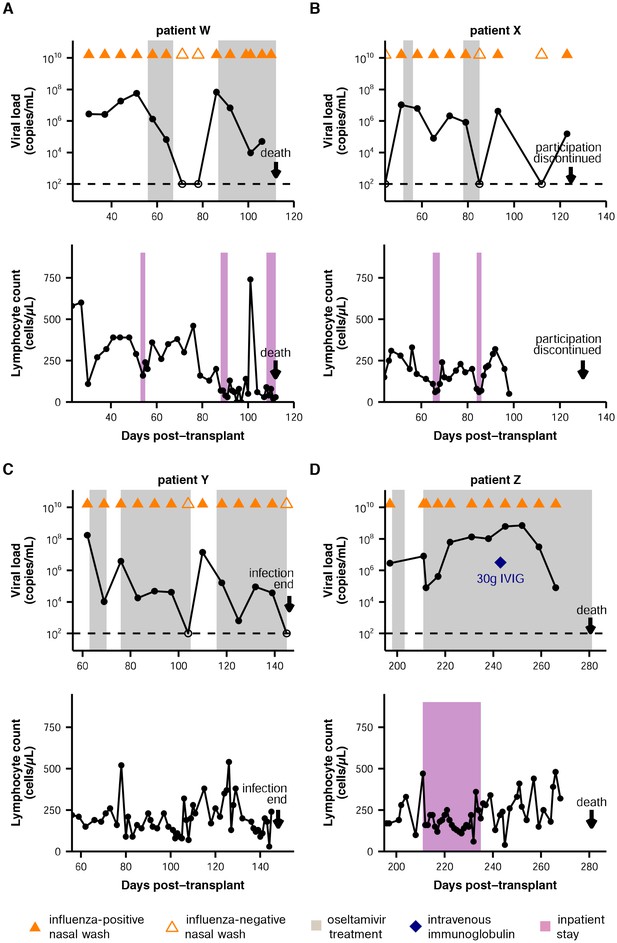
Summary of patient infections.
Influenza viral load over time, as quantified by qRT-PCR, and lymphocyte counts over time are shown for all patients. For the plots of viral load, nasal wash samples are marked with triangles, with influenza-positive nasal washes indicated by solid coloring. Oseltamivir treatment is indicated by gray shading. Intravenous immunoglobulin treatment is marked with a blue diamond. The limit of detection is marked with a dashed line. For the plots of lymphocyte counts, inpatient stays are indicated in purple.
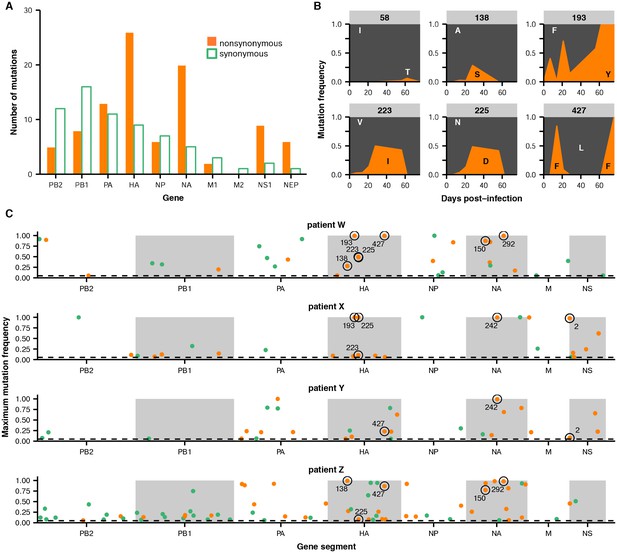
Within-host influenza variants.
(A) Number of nonsynonymous (orange) and synonymous (green) variants in each influenza gene. We identified within-host viral mutations that reached a frequency of at least 5% in two independent sequencing replicates from any patient sample. (B) Frequencies over time for all HA mutations in patient W. Each subplot represents a site in HA and is labeled by codon number. Ancestral identities are colored in gray and mutant ones in orange. (C) Maximum frequencies reached by all nonsynonymous (orange) and synonymous (green) mutations in each patient. Mutations circled in black emerged independently in multiple patients and are labeled by codon number. The dotted line indicates the minimum frequency threshold of 5%. Materials and methods and Figure 2—figure supplement 1 describe procedures used for variant calling and quality control. Figure 2—figure supplements 2–5 give full frequency trajectories for all mutations in all patients. Figure 2—figure supplement 6 shows mutations in HA and NA on their respective crystal structures. Figure 2—figure supplement 7 describes permutation tests that assess the significance of the observed parallelism between patients.
-
Figure 2—source data 1
Primers used for viral deep sequencing.
5’ primer tail sequence is indicated in plain text, homology to the U12/U13 regions in bold text, and gene-specific sequence in bold underlined text. Primers were modified from the universal primers described in (Hoffmann et al., 2001) to account for the A/G polymorphism at the U4 site of the U12 region.
- https://doi.org/10.7554/eLife.26875.006
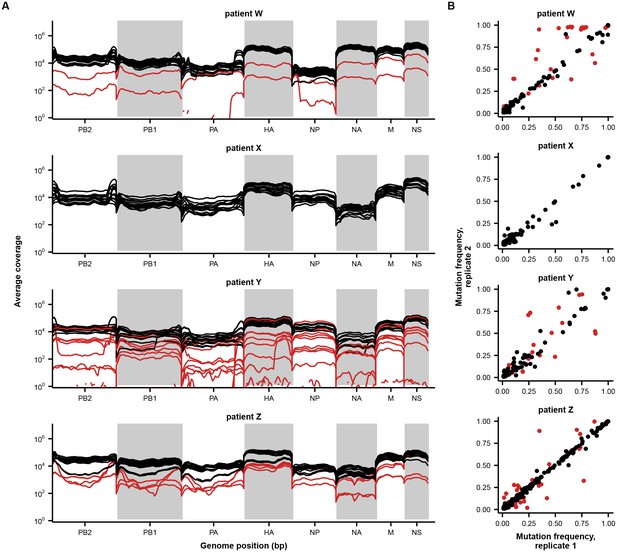
Sample quality controls.
(A) Sequencing coverage across the genome. Average sequencing coverage for each sample and library replicate is shown in 50 bp bins across the genome. Each line indicates a separate library replicate. Low-quality samples are colored in red and were not included in downstream analyses. (B) Correlation between mutation frequencies in replicate sequencing libraries. Mutations in low-quality samples are colored in red and were excluded from downstream analyses. These samples were not shown in Figure 1B. A mutation was called if a base other than the initial consensus base reached a frequency of at least 1% with a total coverage of at least 200 reads in both replicates. Note that this variant calling threshold is more lenient than the 5% threshold used to call variants in downstream analyses; this more lenient threshold identifies more variants to get a better measurement of replicability. Samples were excluded from downstream analyses if the average difference between mutation frequencies exceeded 0.05. Replicate libraries were prepared beginning with independent reverse-transcription reactions.
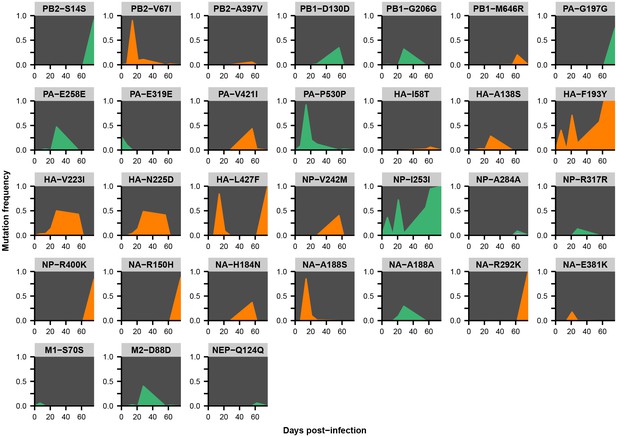
Within-host variants in patient W.
Mutation frequencies over the course of the infection are shown at all variable sites in patient W. Ancestral identities are colored in gray, derived nonsynonymous identities in orange, and derived synonymous identities in green. Sites were called as variant if a base other than the initial consensus reached a frequency of at least 5% in both replicate libraries of at least one sequenced time point. The frequencies shown here are the average of the two replicates. Mutations located in the overlap of M1/M2 or NS1/NEP genes are displayed twice with the appropriate annotation for each gene.
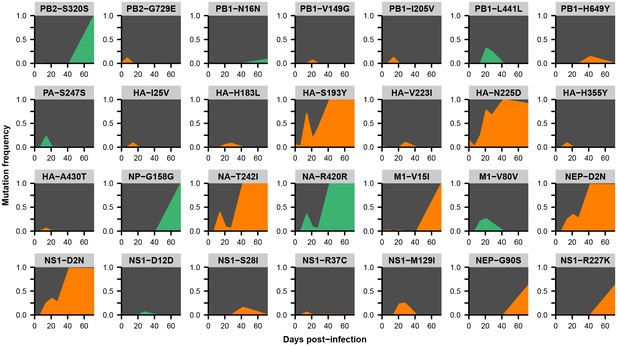
Within-host variants in patient X.
Mutation frequencies over the course of the infection are shown at all variable sites in patient X. Ancestral identities are colored in gray, derived nonsynonymous identities in orange, and derived synonymous identities in green. Sites were called as variant if a base other than the initial consensus reached a frequency of at least 5% in both replicate libraries of at least one sequenced time point. The frequencies shown here are the average of the two replicates. Mutations located in the overlap of M1/M2 or NS1/NEP genes are displayed twice with the appropriate annotation for each gene.
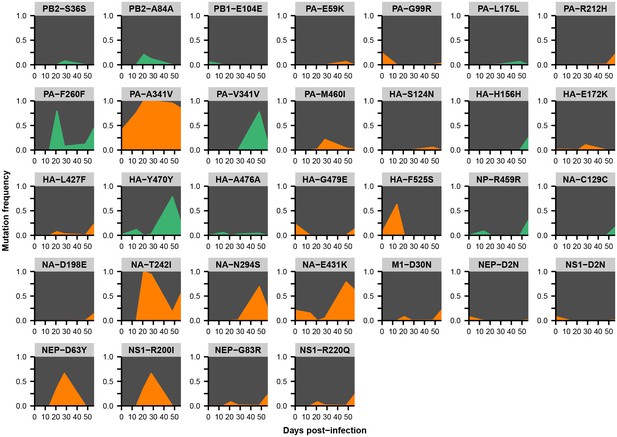
Within-host variants in patient Y.
Mutation frequencies over the course of the infection are shown at all variable sites in patient Y. Ancestral identities are colored in gray, derived nonsynonymous identities in orange, and derived synonymous identities in green. Sites were called as variant if a base other than the initial consensus reached a frequency of at least 5% in both replicate libraries of at least one sequenced time point. The frequencies shown here are the average of the two replicates. Mutations located in the overlap of M1/M2 or NS1/NEP genes are displayed twice with the appropriate annotation for each gene.
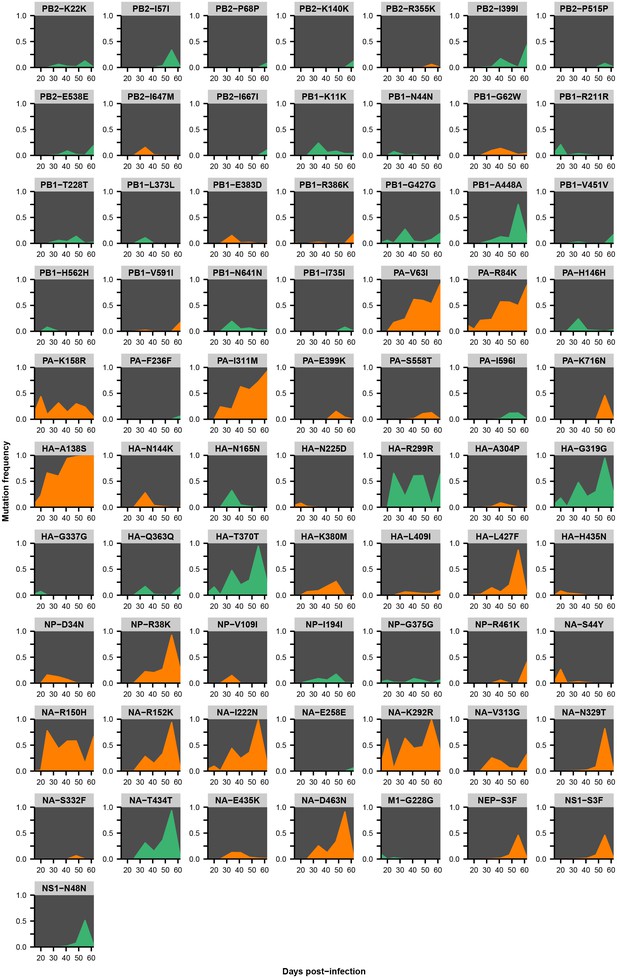
Within-host variants in patient Z.
Mutation frequencies over the course of the infection are shown at all variable sites in patient Z. Ancestral identities are colored in gray, derived nonsynonymous identities in orange, and derived synonymous identities in green. Sites were called as variant if a base other than the initial consensus reached a frequency of at least 5% in both replicate libraries of at least one sequenced time point. The frequencies shown here are the average of the two replicates. Mutations located in the overlap of M1/M2 or NS1/NEP genes are displayed twice with the appropriate annotation for each gene.
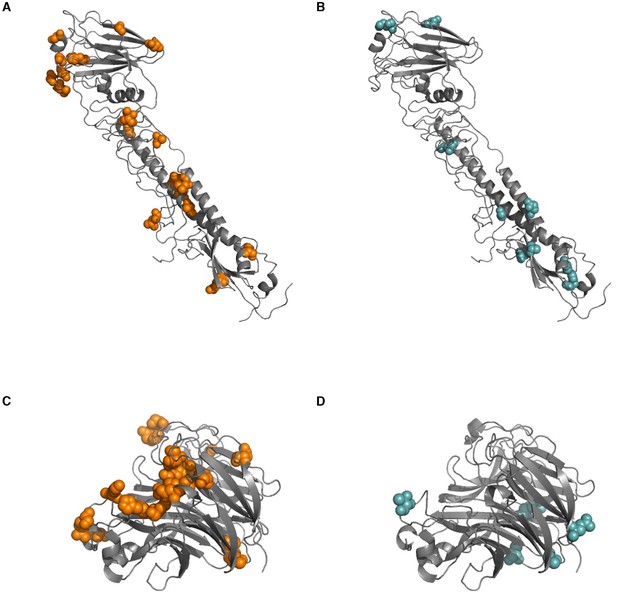
Sites of within-host mutation.
Sites of (A) nonsynonymous and (B) synonymous within-host mutations are shown on an HA crystal structure (PDB 4HMG [Weis et al., 1990]). Sites of (C) nonsynonymous and (D) synonymous within-host mutation are shown on an NA crystal structure (PDB 2BAT [Varghese et al., 1992]). All sites of synonymous and nonsynonymous mutation are shown here, in contrast to Figure 4A, which only shows sites in HA at which nonsynonymous mutations arise in multiple patients in our study.
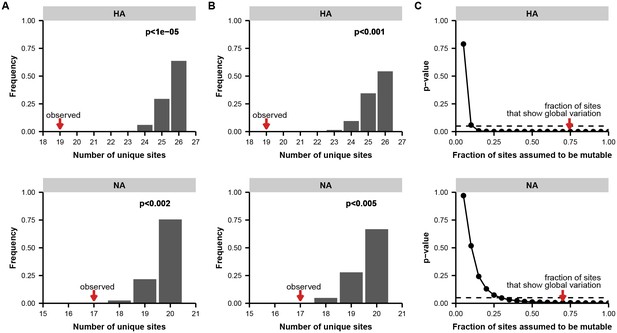
Permutation tests for parallel evolution between patients.
(A) Distribution of unique variable sites when sites are drawn at random along the full length of the indicated gene, matching the number of variants empirically observed in each patient. These simulations test a simple null model in which each site in the gene is equally likely to mutate. We calculated the number of unique sites of mutation in each simulation as a metric of parallelism: fewer unique sites means that more parallel mutation has occurred. The p-value indicates the proportion of 100,000 simulations in which the number of unique sites is less than or equal to what is empirically observed. (B) Distribution of overlapping sites for simulations as described in (A), but with a constrained null model in which the fraction of sites considered mutable is the fraction that shows at least two instances of nonsynonymous mutation in the global H3N2 influenza population between 2000 and 2015 (see Materials and methods) to account for constraints on protein evolution. For both HA and NA, the observed overlap of mutations between patients is statistically significant under this set of constraints. (C) p-values as described in (A), calculated across a range of constraints on the fraction of mutable sites. The constrained null model indicated in (B) is indicated with a red arrow. For both HA and NA, the observed parallelism is statistically significant at a threshold of 0.05 unless it is assumed that fewer than 15% of the sites in HA or 35% of the sites in NA are mutable.
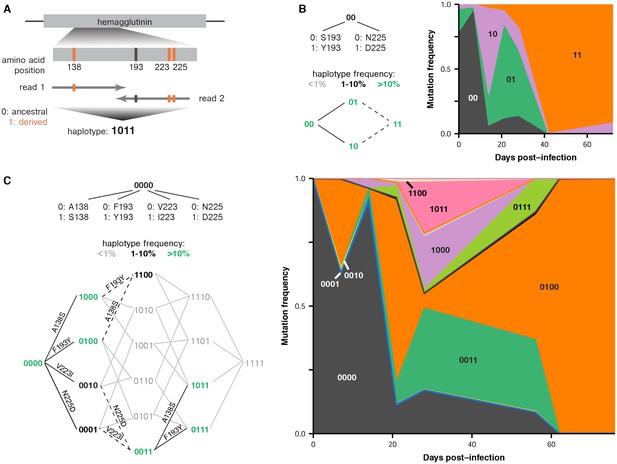
Parallel emergence of the same mutations within single infected hosts.
(A) Method for inferring partial haplotypes from short-read sequencing data. We identified paired-end reads that spanned multiple sites of interest along a gene and determined whether the read carried the ancestral or derived allele at each site. (B) Frequencies of haplotypes at HA sites 193 and 225 in patient X. Evolutionary paths from the ancestral to double-mutant state are shown, with haplotypes colored according to their maximum frequency during the infection. Solid black lines connect pairs of haplotypes that are both present at a frequency of above 1% and that unambiguously occurred through the indicated mutation. Dashed lines indicate that multiple mutations could have produced a particular haplotype. Gray lines indicate that a mutation did not arise at a detectable frequency on a particular haplotype background. (C) Frequencies of haplotypes at HA sites 138, 193, 223, and 225 in patient W. Figure 3—figure supplement 1 estimates the rate of PCR recombination as described in Materials and methods. Figure 3—figure supplement 2 and 3 show the number of paired-end reads that spanned the mutations in the haplotypes in patients X and W.
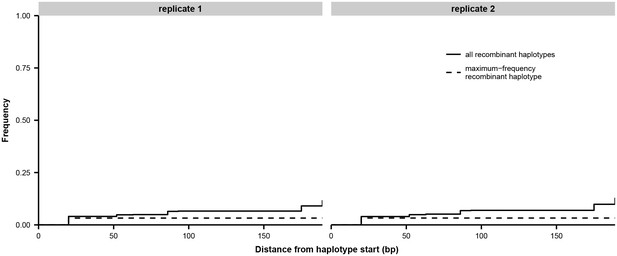
Estimate of PCR recombination rate.
The total frequency of all recombinant haplotypes and the frequency of the single most common recombinant haplotype are shown as a function of distance from the haplotype start. We mixed equal volumes of extracted RNA from the first influenza-positive nasal washes from patients W and X and prepared replicate libraries for sequencing as described in Materials and methods. In the absence of PCR recombination, all haplotypes should consist entirely of bases from one or the other original sample: for instance, 00000000 or 11111111. Based on sequencing of the unmixed samples from patients W and X, we identified eight sites of fixed differences within a 200 bp region of the HA gene, and we inferred haplotypes in the mixture sample at these eight sites. Beginning at the first haplotype site, we tallied the proportion of haplotypes that had experienced recombination by each successive site in the haplotype. We did not seek to distinguish between PCR recombination and sequencing errors: the haplotypes 00100000 and 00111111 were both recorded as having experienced recombination by the third haplotype site. The maximum-frequency recombinant haplotype never exceeded 3.5% of the total haplotypes.
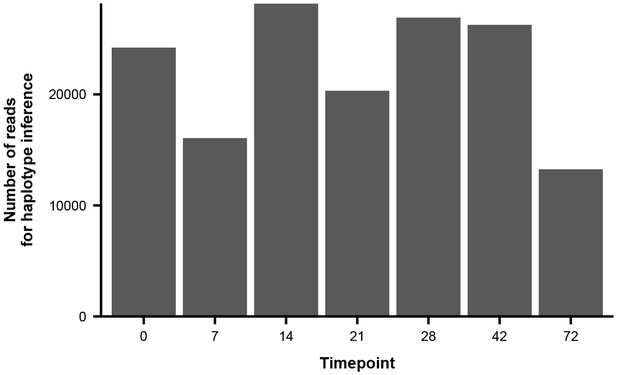
Number of paired-end reads used to infer haplotype dynamics in patient X.
Each bar represents the average number of paired-end reads that spanned both variable sites of interest in Figure 3B across the two sequencing replicates.
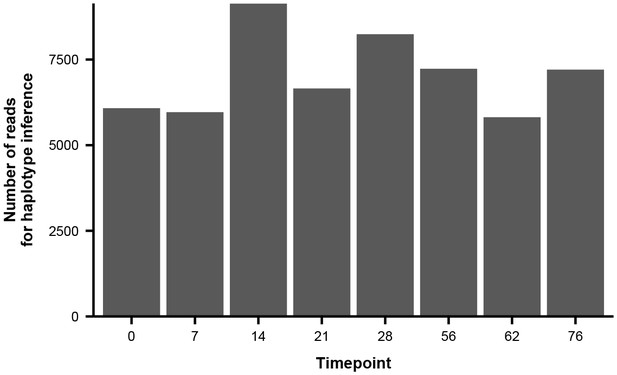
Number of paired-end reads used to infer haplotype dynamics in patient W.
Each bar represents the average number of paired-end reads that spanned the four variable sites of interest in Figure 3C across the two sequencing replicates.
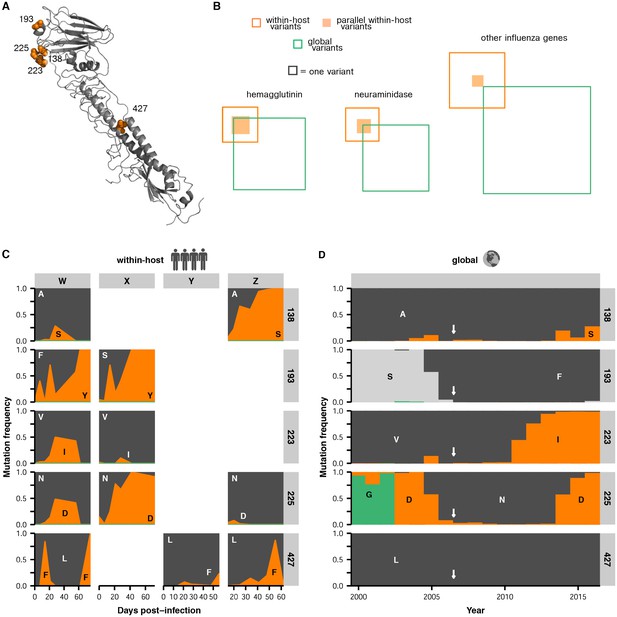
Parallel mutations at within-host and global scales.
(A) Sites of parallel within-host mutation plotted on an HA crystal structure (PDB 4HMG [Weis et al., 1990]). (B) Overlap of within-host (orange) and global (green) variable sites in HA, NA, and all other influenza genes. Sites at which mutations arise in more than one patient are indicated in solid orange. We defined global variable sites as those at which a variant reached a frequency of at least 10% in a given year after 2000 in the GISAID database of global influenza sequences (Bogner et al., 2006). Numbers of within-host and global mutations are given in Figure 4—source data 1. (C) Mutation frequencies over time within individual patients for parallel within-host mutations in HA. Ancestral identities are colored in gray and mutant ones in orange. (D) Global variant frequencies between 2000 and 2015 in H3N2 influenza at sites of parallel within-host mutation in HA. The approximate timing of the patient infections (2006–2007) is indicated by a white arrow. Figure 4—figure supplement 1 displays variant frequencies for all sites of parallel mutation at the within-host and global scales. Figure 4—figure supplement 2 describes permutation tests that assess the significance of the overlap in mutations at the within-host and global scales.
-
Figure 4—source data 1
Overlap of mutations at the within-host and global scales.
- https://doi.org/10.7554/eLife.26875.019
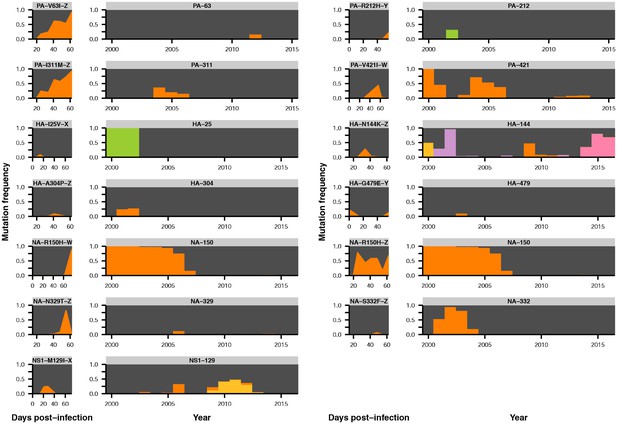
Parallel mutations at within-host and global scales.
Mutation frequencies over time are plotted within hosts and in the global population for sites that are variable across both scales. Parallel within-host mutations in HA are shown in Figure 4C and are omitted. This figure shows mutation frequencies for the remaining sites that were variable across scales. Within-host mutations are labeled by gene name, amino acid change, and patient ID. Ancestral identities are colored in gray and mutant ones in orange at sites of within-host mutation, and the same colors are applied to those amino acids in the global frequency plots where present. Global variant frequencies are shown twice for NA site 150 because mutations arise at that site in two independent patients.
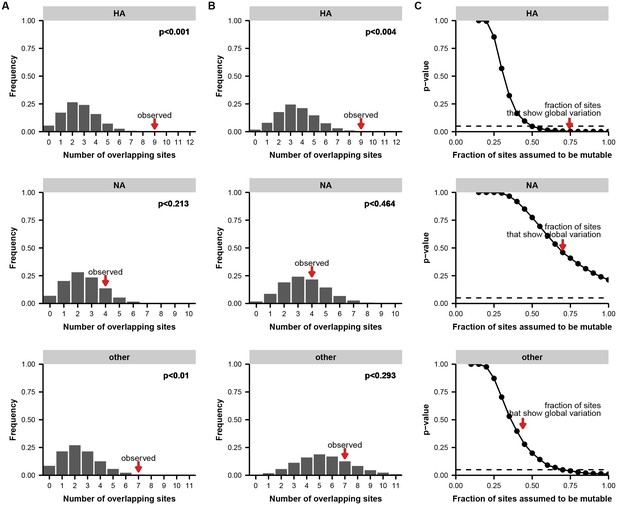
Permutation tests for parallel evolution across within-host and global scales.
(A) Distribution of overlapping sites when two sets of sites are drawn at random along the full length of the indicated gene or genes, matching the number of unique variable sites empirically observed in each patient and in the global influenza population (see Materials and methods). These simulations test a simple null model in which each site is equally like to mutate. We calculated the overlap between the two sets of sites in each simulation as a metric of parallelism: greater overlap means that more parallelism has occurred. The p-value indicates the proportion of 100,000 simulations in which the number of overlapping sites is greater than or equal to what is empirically observed. (B) Distribution of overlapping sites for simulations as described in (A), performed with a constrained null model in which the fraction of sites considered mutable is the fraction that shows at least two instances of nonsynonymous mutation in the global H3N2 influenza population between 2000 and 2015 (see Materials and methods) to account for constraints on protein evolution. For HA, but not NA or the other influenza genes in aggregate, the observed overlap of mutations at the within-host and global scales is statistically significant under this set of constraints. (C) p-values as described in (A), calculated across a range of constraints on the fraction of mutable sites. The constrained null model indicated in (B) is indicated with a red arrow. For HA, the observed parallelism is statistically significant at a threshold of 0.05 unless it is assumed that fewer than half the sites in the protein are mutable.
Additional files
-
Supplementary file 1
GISAID acknowledgement tables for global influenza sequences.
- https://doi.org/10.7554/eLife.26875.022
-
Supplementary file 2
Code and source data files for all analyses.
- https://doi.org/10.7554/eLife.26875.023





