Intermittent Ca2+ signals mediated by Orai1 regulate basal T cell motility
Figures
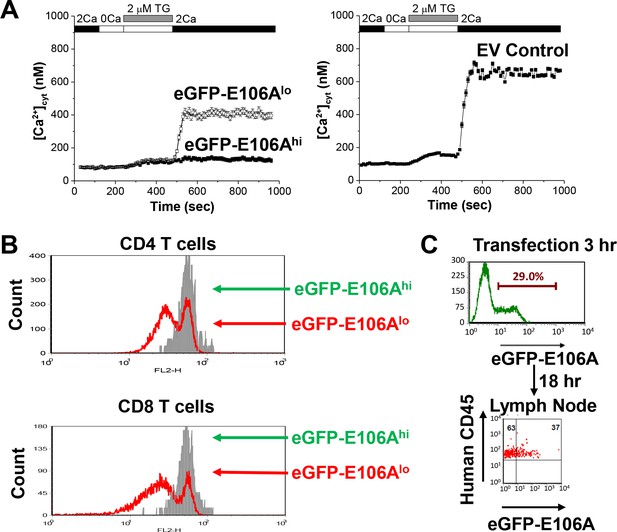
Effects of expressing Orai1-E106A on human T cells.
(A) Averaged thapsigargin-induced Ca2+ entry, measured by fura-2, in activated human CD4+ T cells transfected with eGFP-Orai1-E106A (left) or empty vector control (EV, right, n = 133 cells); eGFP-E106A transfected cells were grouped into two populations, either eGFP-E106Ahi with high eGFP fluorescence (solid squares, n = 43 cells) or eGFP-E106Alo with no detectable eGFP fluorescence (empty squares, n = 115 cells); bars represent SEM. (B) Primary human CD4+ and CD8+ T cells were transfected with eGFP-E106A, then uniformly labeled with the fluorescent cell tracker dye CMTMR and co-cultured with SEB-pulsed primary human dendritic cells from the same donor; proliferation was assessed after 72 hr by CMTMR dilution as measured by flow cytometry. (C) Human CD3+ T cells were transfected with eGFP-E106A and expression level was measured 3 hr post-transfection before adoptive transfer into reconstituted NOD.SCI.β2 mice; cells were recovered from lymph nodes 18 hr later and eGFP fluorescence was used to measure homing to lymph nodes.
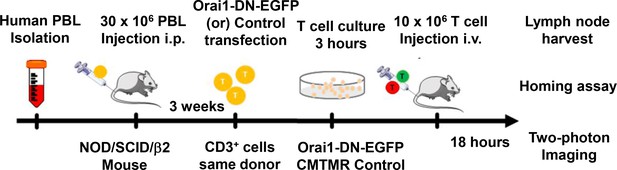
Protocol for homing and two-photon imaging of transfected human CD3+ T cells in reconstituted NOD.SCID.β 2 mouse lymph node.
https://doi.org/10.7554/eLife.27827.004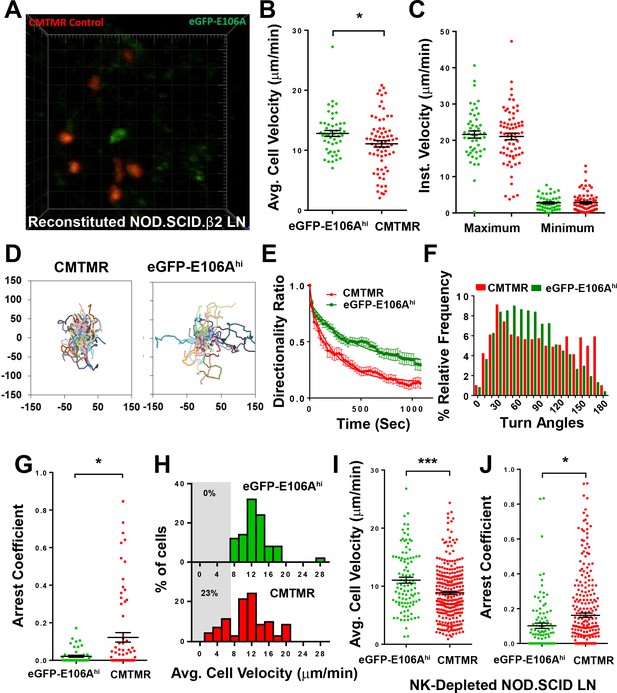
Orai1 block increases human T cell motility within reconstituted NOD.SCID.β2 lymph nodes.
(A) Two-photon microscopy of migrating human T cells, showing eGFP-E106A transfected cells in green and CMTMR-labeled mock transfected cells in red, within intact mouse lymph node 18 hr after adoptive co-transfer of 5 × 106 of each cell type. (B) Average cell velocities of eGFP-E106Ahi (n = 50) versus CMTMR-labeled control (n = 71) T cells; bars represent mean ± SEM, data from independent experiments using four different donors (12.8 ± 0.5 µm/min vs. 11.1 ± 0.5 µm/min for E106Ahi vs CMTMR cells, p=0.0268). (C) Maximum and minimum cellular instantaneous velocities of eGFP-E106Ahi (green) versus CMTMR-labeled (red) control T cells. (Hodges-Lehmann median difference of −0.21 µm/min, −2.82 to 2.16 µm/min 95% confidence interval for maximum velocity and −0.36 µm/min, −1.02 to 0.36 µm/min 95% confidence interval for minimum velocity) (D) Superimposed tracks with their origins normalized to the starting point. Cells were tracked for more than 20 min. n = 111 (CMTMR), n = 58 (eGFP-E106Ahi) (E) Directionality ratio (displacement/distance) over elapsed time. For Orai1-blocked cells in green, tau = 397 s; vs CMTMR controls in red, tau = 238 s, n = 49 time points. (F) Histogram of turn angles in eGFP-E106Ahi (green) and CMTMR controls (red). Mean ± SEM, 74.5 ± 1.0 degrees for Orai1 blocked cells vs 86.5 ± 1.5 degrees for CMTMR controls, p=0.0001, two-tailed T test. (G) Arrest coefficients of eGFP-E106Ahi compared with CMTMR-labeled control T cells, defined as fraction of time with instantaneous velocity <2 µm/min. (For Orai1-blocked cells in green, 0.02 ± 0.01; vs. CMTMR controls in red, 0.12 ± 0.03, p=0.0406) (H) Frequency distribution of average cell velocities for eGFP-E106Ahi (top) and CMTMR-labeled control T cells (bottom), cells with average velocity <7 µm/min are highlighted in gray; tick marks denote the center of every other bin. (I,J) Average cell velocities (I) and arrest coefficients (J) of eGFP-E106Ahi (green, n = 102) vs CMTMR-labeled control (red, n = 278) human T cells in NK-cell-depleted immunodeficient mouse lymph nodes. Average cell velocities: 11.0 ± 0.5 µm/min vs. 8.8 ± 0.3 µm/min, p=0.0004; Arrest coefficients: 0.10 ± 0.02 vs. 0.16 ± 0.01, p=0.0516 for E106Ahi vs CMTMT cells; bars represent mean ± SEM, data from independent experiments using eight different donors, ***p<0.005.
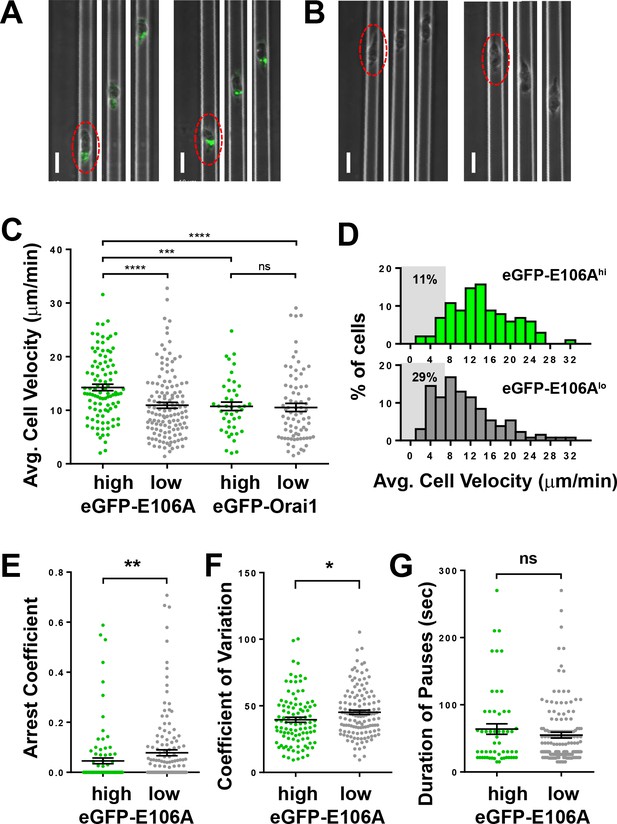
Orai1 block reduces frequency of pausing during human T cell motility in vitro.
(A,B) Confocal microscopy of eGFP-E106A transfected human CD4+ T cells in microfabricated channels 7 µm high by 8 µm wide, showing two individual eGFP-E106Ahi T cells (A) and two eGFP-E106Alo T cells (B), each circled in red in the first frame; individual images taken 1 min apart, scale bar = 10 µm. (C) Comparison of average cell velocities of eGFP-E106A transfected T cells (eGFP-E106Ahi cells in green, n = 102; eGFP-E106Alo cells in gray, n = 131; 14.2 ± 0.6 µm/min vs. 10.9 ± 0.5 µm/min, p<0.0001 for E106Ahi vs E106Alo cells) and eGFP-Orai1 transfected control T cells (eGFP-Orai1hi cells in green, n = 43; eGFP-Orai1lo cells in gray, n = 76; 10.7 ± 0.8 µm/min vs. 10.5 ± 0.8 µm/min for Orai1hi vs Orai1lo cells; Hodges-Lehmann median difference of −0.84 µm/min, −2.96 to 1.28 µm/min 95% CI); bars represent mean ± SEM, data from independent experiments using five different donors. (D) Frequency distribution of average cell velocities of eGFP-E106Ahi (top) and eGFP-E106Alo (bottom) human T cells, cells with average velocity <7 µm/min are highlighted in gray; tick marks denote the center of every other bin. (E) Arrest coefficients of eGFP-E106Ahi vs eGFP-E106Alo human T cells, defined as fraction of time each individual cell had an instantaneous velocity <2 µm/min (0.05 ± 0.01 vs. 0.08 ± 0.01 for E106Ahi vs E106Alo cells, p=0.0015). (F) Variance in velocity of eGFP-E106Ahi vs eGFP-E106Alo human T cells, coefficient of variation is calculated by standard deviation divided by the mean of instantaneous velocity for each individual cell (39.5 ± 1.9% vs. 45.1 ± 1.6% for E106Ahi vs E106Alo cells, p=0.0138). (G) Duration of pauses for eGFP-E106Ahi vs eGFP-E106Alo human T cells (Hodges-Lehmann median difference of 0 s, −8.43 to 4.71 s 95% CI for E106Ahi vs E106Alo cells); bars represent mean ± SEM, *p<0.05, **p<0.01, ***p<0.005, ****p<0.001.
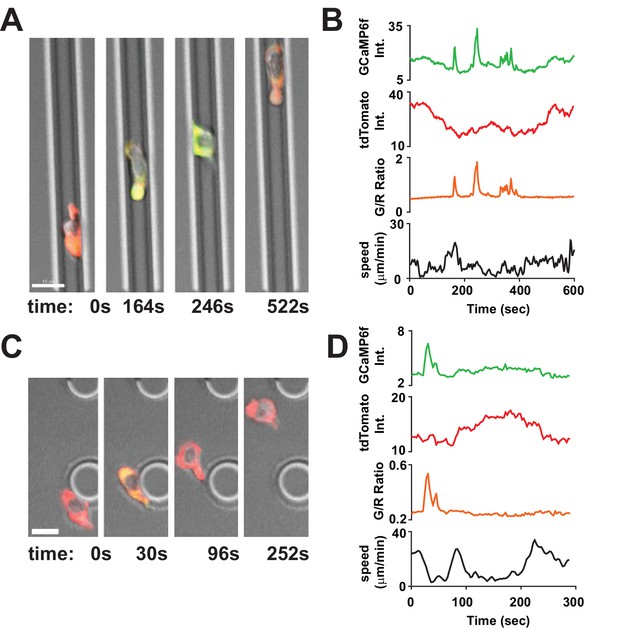
Tracking Ca2+signals in human T cells in vitro with Salsa6f.
(A,C) Confocal microscopy of Salsa6f transfected human CD4+ T cells in ICAM-1 coated microchannels 7 µm high by 8 µm wide (A, Video 1) and open space (C, Video 2), showing merged red (tdTomato), green (GCaMP6f), and DIC channels; circular structures shown in (C) are support pillars part of the PDMS chamber; scale bar = 10 µm, time = s. (B,D) Total intensity tracings of GCaMP6f (green) and tdTomato (red) fluorescence, G/R ratio (orange), and speed (black), for corresponding T cells shown in (A) and (C); data representative of independent experiments from three different donors.
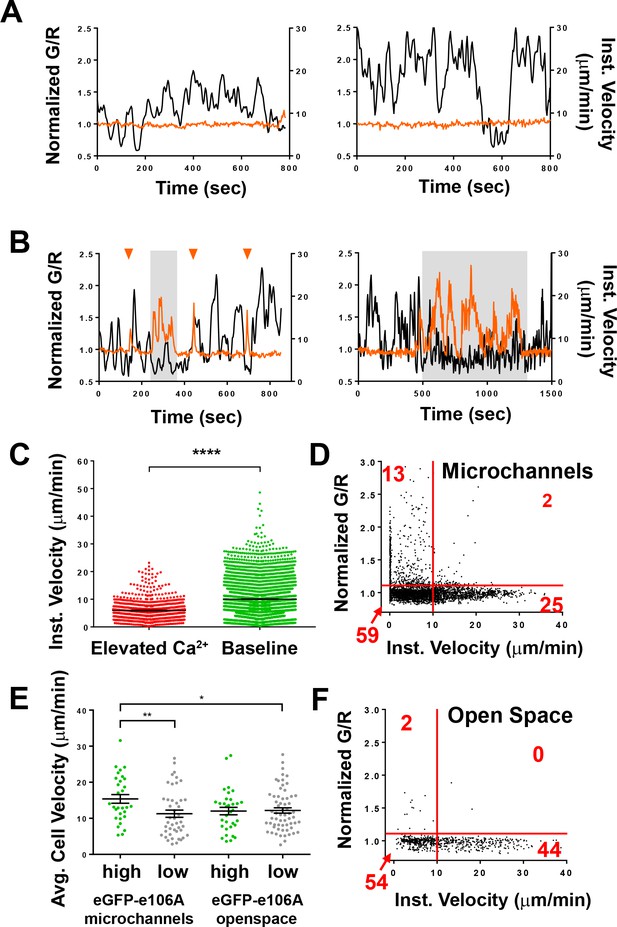
Spontaneous Ca2+signals during human T cell motility in vitro are correlated with reduced velocity.
(A,B) Sample tracks from Salsa6f-transfected human T cells in microchannels, with intracellular Ca2+ levels as G/R ratios for each time point, normalized to zero-time (orange), overlaid with instantaneous cell velocity (black), cells in (A) have stable Ca2+ levels, cells in (B) show brief Ca2+ transients (arrowheads) or sustained Ca2+ signaling (gray highlights). (C) Instantaneous velocity of Salsa6f-transfected human T cells in microchannels during elevated cytosolic Ca2+ levels (red) and during basal Ca2+ levels (green); n = 22 cells, data from independent experiments using three different donors; ****p<0.001. (D) Scatter plot of Salsa transfected human T cells in microchannels, instantaneous cell velocity versus normalized G/R ratio for each individual time point analyzed; red numbers in each quadrant show percent of time points, split by 1.10 normalized G/R ratio and 10 µm/min; n = 4081 points. (E) Mean track velocity of eGFP-E106A-transfected human T cells, comparing eGFP-E106Ahi (green) and eGFP-E106Alo T cells (gray) in confined microchannels vs open space; n = 30, 44, 33, and 62 cells, respectively (15.4 ± 1.2 µm/min vs. 11.3 ± 1.0 µm/min for E106Ahi vs E106Alo cells in microchannels; p=0.0099 and 12.0 ± 1.0 µm/min vs. 12.2 ± 0.7 µm/min for E106Ahi vs E106Alo cells in open space; Hodges-Lehmann median difference of 0.15 µm/min, −2.46 to 2.40 µm/min 95% CI). Bars represent mean ± SEM, data from independent experiments using two different donors, *p<0.05, **p<0.01. (F) Scatter plot of Salsa transfected human T cells in open space, instantaneous cell velocity versus GCaMP6f/tdTomato R/R0 for each individual time point analyzed; red numbers in each quadrant show percent of cells, split by 1.10 normalized G/R ratio and 10 µm/min; n = 723 points.
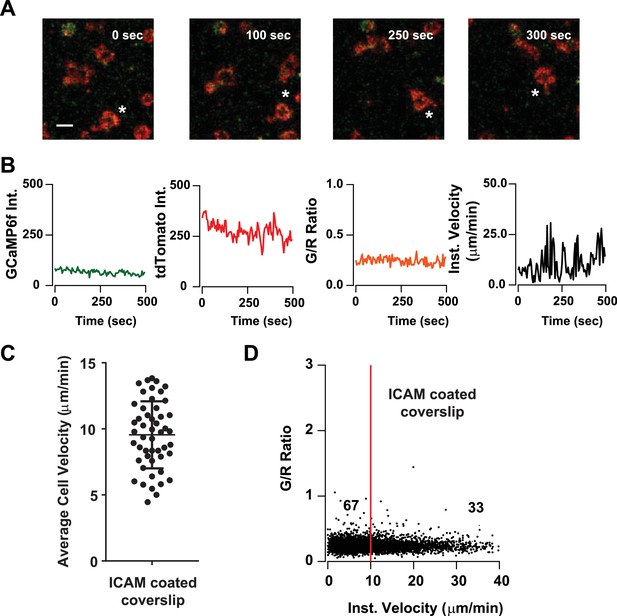
Tracking cell motility and Ca2+signals in Cd4-Salsa6f T cells on ICAM coated coverslips.
(A) Confocal microscopy of Cd4-Salsa6f (Het) Th1 cells on open-field ICAM-1-coated coverslips showing merged red (tdTomato) and green (GCaMP6f) channels at three different time points. Scale bar 10 μm. * indicates movement of one cell. (B) Recordings of GCaMP6f (green), tdTomato (red) fluorescence, G/R ratio (orange), and speed (black), for corresponding T cell shown in (A). (C) Average cell velocities of Cd4-Salsa6f Th1 cells on 5 μg/ml ICAM-1-coated coverslips (n = 52 cells, two independent experiments). (D) Scatter plot showing instantaneous cell velocity versus GCaMP6f/tdTomato (G/R) ratio for each individual time point analyzed; numbers show percent of time points, split by 10 µm/min, as in Figure 8D,F; n = 5289 points.
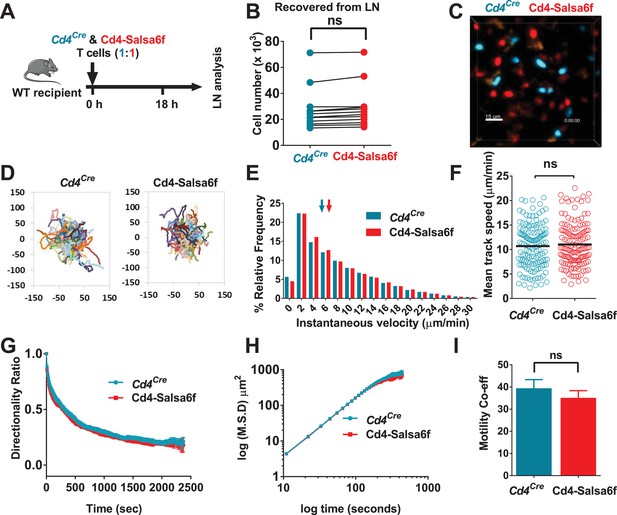
Motility of Salsa6f T cells in lymph node following adoptive transfer.
Cd4Cre and Cd4-Salsa6f (Hom) cells are shown in teal and in red, respectively. (A) Experimental design to characterize homing and motility of Cd4-Salsa6f cells. CTV-labeled Cd4Cre cells and CTY-labeled Cd4-Salsa6f cells (1:1) were adoptively transferred into wildtype mice, 18 hr prior to LN harvesting. (B) Paired numbers of CTV+ and CTY+ cells recovered from lymph nodes (p=0.65, Mann Whitney test). (C) Representative median filtered, maximum intensity projection image showing simultaneously imaged Cd4Cre and Cd4-Salsa6f cells the lymph node, scale bar = 30 µm. See Video 3. (D) Superimposed tracks with their origins normalized to the starting point. Cells were tracked for more than 20 min. n = 140. (E) Frequency distribution of instantaneous velocities; arrows indicate median, tick marks at the center of every other bin (n > 14,800, three independent experiments). (F) Scatter plot showing mean track speed, black bars indicate overall mean values (11.1 ± 0.4 and 10.7 ± 0.4 µm/min, for Cd4Cre and Cd4-Salsa6f cells respectively, p=0.69; n = 140). (G) Directionality ratio (displacement/distance) over elapsed time (tau = 461 s for Cd4Cre in teal; tau = 474 s for Cd4-Salsa6f in red. n = 217 time points). (H) MSD vs time, plotted on a log-log scale. (I) Measured motility coefficient from 140 tracks (35.1 ± 3.2 vs 39.4 ± 3.9 µm2/min for Cd4Cre and Cd4-Salsa6f cells, p=0.65).
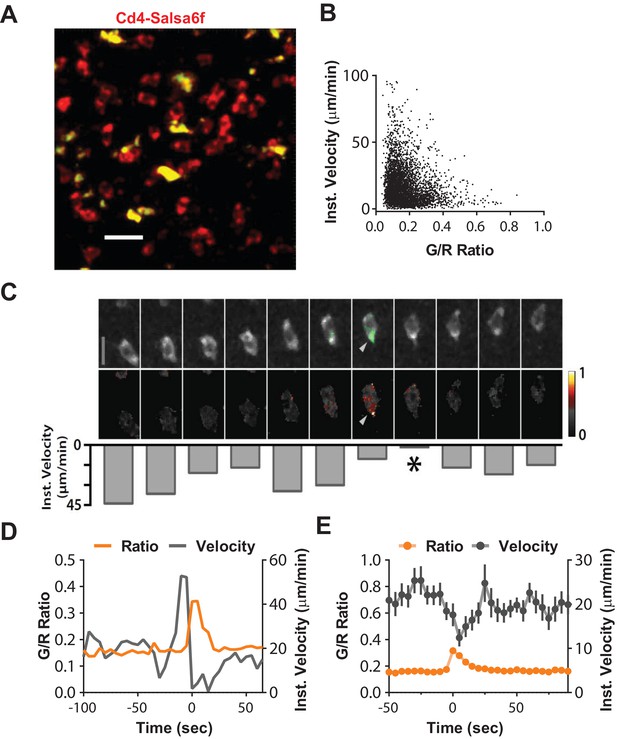
Suppression of motility during spontaneous Ca2+transients.
(A) Median filtered, maximum intensity projection showing cytosolic labeling (exclusion of Salsa6f from the nucleus) in adoptively transferred Cd4-Salsa6f (Hom) cells (red) in the lymph node of wild-type recipients. Autofluorescent structures appear as yellow bodies. Scale bar = 20 µm. See Video 4. (B) Scatterplot of instantaneous 3D velocity vs ratio of GCaMP6f (green) to tdTomato (red) fluorescence intensity (r = −0.24, Spearman's rank correlation, p<0.0001, n = 4490 pairs). (C) Image sequence showing a migrating T cell and calcium transient from (A). Top row: TdTomato signal is shown in grayscale, overlaid with GCaMP6f signals in green. Scale bar = 10 µm. (Video 5). Center row: Heat map of Green/Red ratios matched to corresponding images in the top row. Arrows indicate local Ca2+ transient. Bottom row: inverted bar graph showing corresponding instantaneous 3D velocities. Asterisk marks a pause in cell motility. (D) Representative track from Cd4-Salsa6f (Hom) T cells in lymph nodes, showing intracellular Ca2+ levels measured by G/R ratio (orange) on left Y-axis and instantaneous 3D velocity (gray) on right Y-axis. (E) Averaged time course of the instantaneous 3D velocity (gray trace, right Y-axis) aligned by the corresponding rise in Salsa6f G/R ratio (orange, left Y-axis). The velocity minimum at time = 5 s-is significantly lower than a baseline from −30 to −10 s (p<0.0001 two-tailed T-test, n = 39 cells).
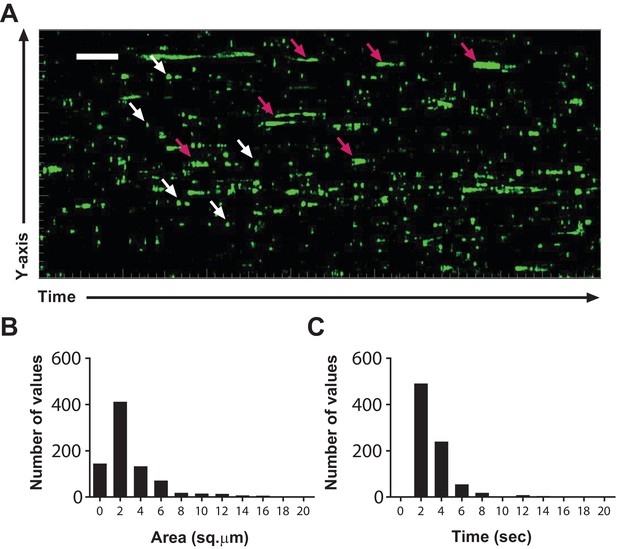
T cell Ca2+transients in the steady-state lymph node.
(A) Calcium history map of steady-state lymph node. Maximum intensity YT projection of 1200 processed green channel time points showing localized sparkles (white arrows) and cell-wide global Ca2+ transients (magenta arrows). Scale bar = 50 µm along Y axis, 50 s along T axis. See Video 6. (B) Frequency distribution of the area of local Ca2+ signals. (C) Frequency distribution of the duration of local Ca2+ signals.
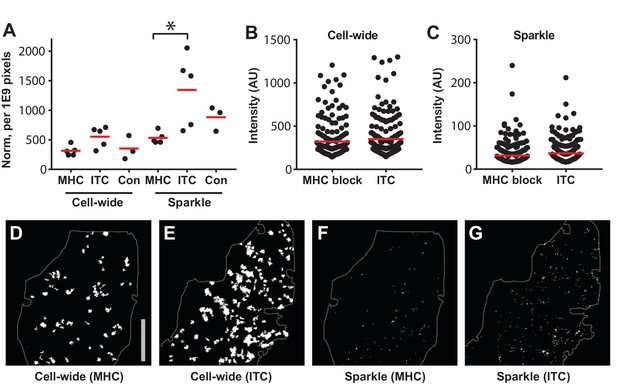
MHC block and Ca2+transients in steady state lymph nodes.
(A) The frequency of cell-wide and local (sparkles) Ca2+ transients in CD-Salsa6f (Hom) lymph nodes 48 hr after injection of MHC class I and II blocking antibodies (MHC), isotype control antibody (ITC), or no antibody (Con). Red bars indicate mean values. For MHC-blocked compared to ITC, the relative event frequencies were: for cell-wide, 314 ± 38 vs 553 ± 77, mean ± SEM, p=0.06; for sparkles, 532 ± 44 vs 1343 ± 272, mean ± SEM, p=0.02, Mann-Whitney test. (B,C) Integrated green channel intensities of Ca2+ transients normalized to SD of green channel for cell-wide events (B) and for sparkles (C). Red bars indicate mean values. For MHC-blocked vs ITC, the relative amplitudes were: for cell-wide (B), 321 ± 14 vs 350 ± 15, mean ± SEM; for sparkles (C), 32 ± 2 vs 37 ± 2, mean ± SEM. (D–G) Representative thresholded images showing cell-wide and local Ca2+ transients, 48 hr after treatment with anti MHC I and II or ITC antibody. The area of the imaging field analyzed is indicated. Scale bar = 100 μm.
Videos
Salsa6f-transfected human T cell in confined microchannel.
Merged red (tdTomato), green (GCaMP6f), and DIC channels; scale bar = 10 µm, time shown in hr:min:s. This video corresponds to Figure 4A.
Salsa6f transfected human T cells in open microchamber, with merged red (tdTomato), green (GCaMP6f), and DIC channels, circular structures are support pillars part of the PDMS microchamber; scale bar = 10 µm, time shown in hr:min:s.
This video corresponds to Figure 4C.
Motility of Cd4-Salsa6f T cells in lymph node following adoptive transfer.
Cd4Cre and Cd4-Salsa6f cells and their trails are shown in teal and in red, respectively. Autofluorescent bodies appear as faint stationary yellow structures. Images were acquired at ~11 s interval. Playback speed = 50 frames per second; time shown in hr:min:sec. Video corresponds to Figure 6C.
Calcium signals in adoptively transferred Cd4-Salsa6f T cells.
Red signal from tdTomato expression in cytosol facilitates identification and tracking of cells; green GCaMP6f signal detects elevation of Ca2+. Autofluorescent structures appear as stationary yellow bodies. Movie is paused at frame 323, zoomed in to emphasize two examples of Ca2+ transients and an autofluorescent body. Images were acquired at 5 s interval. Major tick marks at 20 µm. Playback speed = 50 frames/s, time shown in hr:min:s. Video corresponds to Figure 7A.
A brief Ca2+signal filling the back of a moving adoptively transferred Salsa6f (Hom) T cell.
Left: composite of red tdTomato fluorescence pseudocolored grayscale with green GCaMP6f fluorescence. Right: Corresponding Green/Red ratios, masked to red channel as in Figure 7C. Images acquired at 1 frame every 5 s and 0.5 microns/pixel. Playback speed = 3 frames per second. 0.5 μm/pixel. Video corresponds to Figure 7C.
Calcium transients in steady state lymph nodes.
Cd4-Salsa6f (Hom) lymph node imaged at 0.5 s interval, processed to visualize Ca2+ transients (sparkles and cell-wide) in green. Red channel is turned off after beginning to facilitate viewing of Ca2+ transients. Autofluorescent structures appear as stationary green bodies. Playback speed = 100 frames/s. time shown in hr:min:s. Video corresponds to Figure 8A
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| recombinant DNA reagent | Salsa6f | Dong et al (doi: 10.7554/eLife. 32417) | ||
| strain, strain background (mouse) | Cd4-Salsa6f (Het), Cd4-Salsa6f (Hom) | Dong et al (doi: 10.7554/eLife. 32417) | ||
| strain, strain background (mouse) | NOD.Cg-PrkdcscidB2mtm1Unc/J (NOD.SCID.β2) | Jackson #002570 | ||
| strain, strain background (mouse) | NOD.CB17-Prkdcscid/J (NOD.SCID) | Jackson #001303 | ||
| strain, strain background (mouse) | Cd4Cre | Jackson #017336 | ||
| strain, strain background (mouse) | C57BL/6J | Jackson #000664 | ||
| transfected construct (synthetic) | eGFP-Orai1-E106A, eGFP-Orai1 | 23455504 | ||
| biological sample (human) | Primary T cells from healthy human subjects | UCI | IRB approved | |
| antibody | anti-MHC II (Clone Y3P), anti-MHC I (Clone AF6-88.5.5.3), IgG2a Isotype control (Clone: C1.18.4) | BioXCell | ||
| antibody | anti-NK cell antibody | Wako Chemicals | ||
| antibody | αCD3 and αCD28 | Tonbo Biosciences | ||
| peptide, recombinant protein | recombinant human ICAM | R&D Systems | ||
| peptide, recombinant protein | recombinant human IL-2 | BioLegend | ||
| commercial assay or kit | EasySep human T Cell Isolation Kit | Stemcell Technologies | ||
| commercial assay or kit | EasySep mouse T Cell Isolation Kit | Stemcell Technologies | ||
| commercial assay or kit | Nucleofection kit | Lonza | ||
| commercial assay or kit | Sylgard Elastomer 184 kit | Dow Corning | ||
| chemical compound, drug | Cell tracker CMTMR, CellTrace Yellow or CellTrace Violet | Life Technologies | ||
| software, algorithm | Imaris | Bitplane |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.27827.020





