Learning induces the translin/trax RNase complex to express activin receptors for persistent memory
Figures
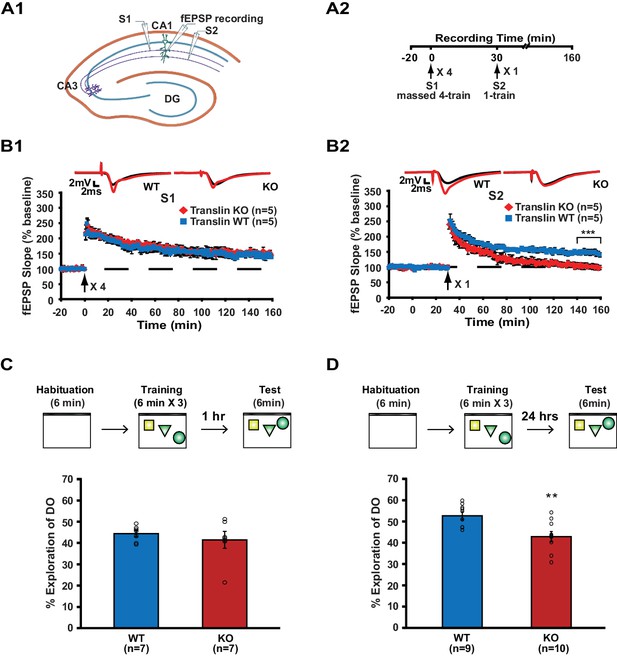
Translin KO mice display impaired synaptic tagging and long-term memory formation.
(A1) A schematic diagram showing the two independent inputs (S1 and S2) converging onto postsynaptic neurons in CA1. (A2) Experimental scheme. (B) Slices from translin KO mice showed unaltered 4-train long-lasting potentiation in pathway S1 (B1; two-way repeated-measures ANOVA, F(1,8) = 0.03, p=0.9), but displayed reduced persistent potentiation after 1-train stimulation in pathway S2 (B2); two-way repeated-measures ANOVA, F(1,8) = 36.5, p=0.0003). Average traces for the baseline (black) and the last 20 min (red) are shown above each graph. (C and D) Top, experimental scheme. Translin KO mice explored the displaced object (DO) significantly less than WT littermates 24 hr after training (C; t-test, p=0.004), while explored DO similar to WT littermates 1 hr after training (D; t-test, p=0.5). ** indicates p<0.005. *** indicates p<0.0005. n, number of mice. Data are represented as mean ±SEM.
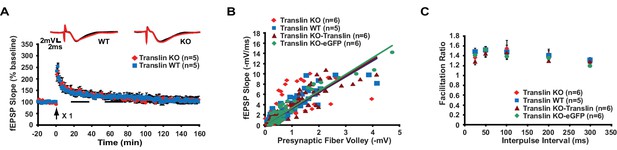
Short-lived LTP and basal synaptic transmission are unaltered in translin KO mice.
(A) One-train stimulation was delivered to hippocampal area CA1. Translin KO mice showed unaltered short-lasting LTP induced by 1-train stimulation compared to WT littermates (two-way repeated-measures ANOVA, F(1,8) = 0.07, p=0.8). Average traces for the baseline (black) and the last 20 min (red) are shown above each graph. (B and C) Data from translin KO mice injected with either translin- or eGFP-virus are supplements for Figure 7 and included for comparison with uninjected WT and KO mice. (B) The input-output relationship was similar across hippocampal slices from WT littermates, translin KO mice, and translin KO mice injected with either translin or eGFP virus (one-way ANOVA, F(3,22) = 0.5, p=0.7). (C) Slices from WT littermates, translin KO mice, and translin KO mice injected with translin or eGFP virus displayed similar PPF at different interpulse intervals (two-way repeated measures ANOVA, F(3,24) = 0.5, p=0.7). n, number of mice. Data are represented as mean ±SEM.
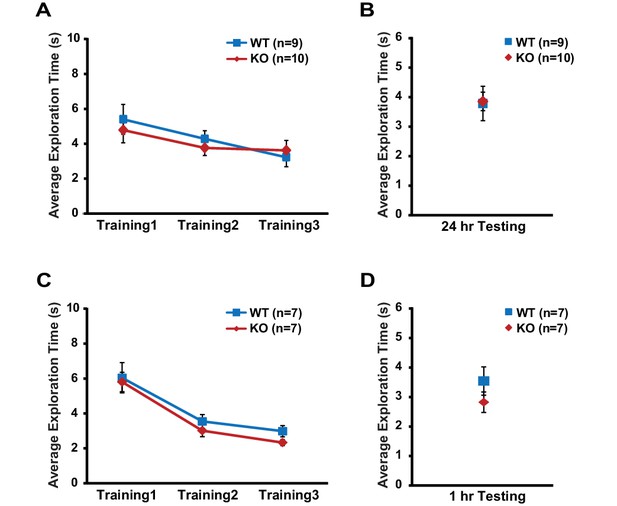
Exploratory behavior during object-location memory task is unaltered in translin KO mice.
Compared to WT littermates, translin KO mice displayed unaltered total object exploration time across training sessions (two-way repeated measures ANOVA, (A); F(1,17) = 0.2, p=0.7, 24 hr task, (C) F(1,12) = 1.1, p=0.3, 1 hr task), and during testing (one-way ANOVA, (B); F(1,17) = 0.01, p=0.9. (D); F(1,12) = 1.3, p=0.3). n, number of mice. Data are represented as mean ±SEM.
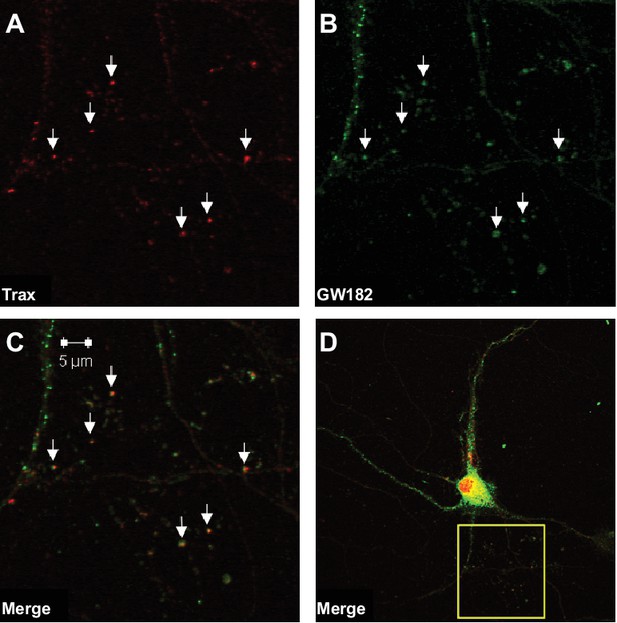
Trax co-localizes with P-bodies.
Immunostaining images display a rat hippocampal neuron stained for trax (A red) and GW182, a marker of P-bodies (B) green). (C) Merged image. (D) The box indicates the area depicted at higher magnification in the other three panels. Arrowheads indicate puncta that are co-labelled by trax and GW182.
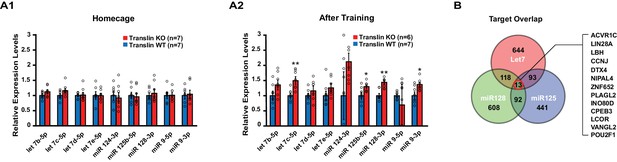
Translin KO mice display increased levels of several microRNAs targeting the activin A receptor type 1C (ACVR1C) 30 min following training in the object location memory task.
Real-time RT-PCR was performed on hippocampal samples to validate candidate microRNAs. (A) Left, no significant difference was observed between translin KO and WT hippocampus in homecage condition. Right, the levels of let-7c-5p (t-test, p=0.003), miR-125b-5p (t-test, p=0.04), miR-128–3p (t-test, p=0.001) and miR-9–3p (t-test, p=0.03) were increased 30 min after training in translin KO hippocampus relative to WT hippocampus. (B) Venn diagram shows 13 overlapping targets of Let-7, miR-128 and miR-125. The 13 target genes are listed in a descending order of prediction context scores. ACVR1C has the highest context score. * indicates p<0.05, ** indicates p<0.005. Data are represented as mean ±SEM.
-
Figure 3—source data 1
Common target genes of the conserved mRNA targets of let-7, miR-125 and miR-128.
(Descending order of aggregate context scores).
- https://doi.org/10.7554/eLife.27872.007
-
Figure 3—source data 2
Conserved mRNA targets of let-7 between humans and mice.
Conserved mRNA targets of miR-125 between humans and mice. Conserved mRNA targets of miR-128 between humans & mice.
- https://doi.org/10.7554/eLife.27872.008
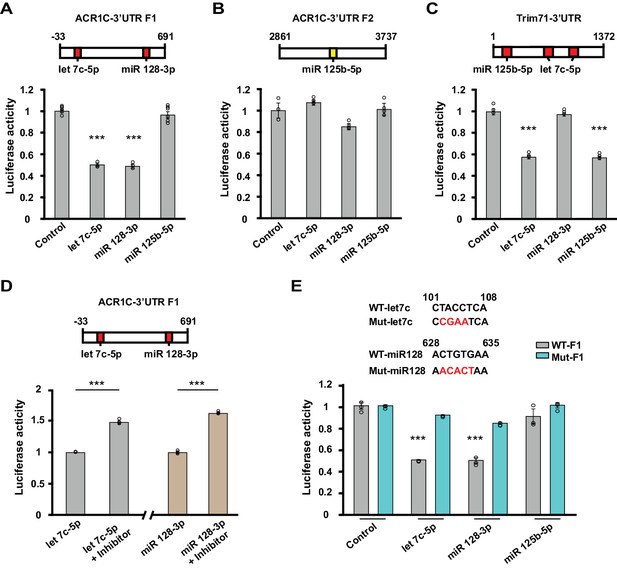
Let-7c-5p and miR-128–3p suppress luciferase activity of ACVR1C-3’UTR.
(A–D) Top, schematic diagrams illustrating the location of target sites for selected microRNAs in 3’UTR fragments. Values shown above the boxes indicate nucleotide positions relative to the start of the 3’UTR. Red indicates target sites of given microRNAs with high context percentile scores (> 80th percentile), while yellow indicates a lower context score. Bottom, bar graphs illustrate normalized luciferase activity (relative to control) from cells transfected with the listed microRNA mimics at a concentration of 2 nM (n = 3–6 per group). (A) The luciferase activity of the F1 reporter construct was reduced by either let-7c-5p (target position: 101–108) or miR-128–3 p (target position: 628–635) treatment (one-way ANOVA F(3,14) = 136.7, p<0.0001; Dunnett’s post hoc test control vs let-7c-5p or miR-128–3p: p=0.00000004 or 0.00000004, respectively). (B) The luciferase activity of the F2 reporter construct was unaffected by miR-125–5p (target position: 3248–3254) (one-way ANOVA F(3,8) = 5.59, p=0.023; Dunnett’s post hoc test control vs let-7c-5p, miR-128–3 p or miR-125–5 p: p=0.5, p=0.07, or p=0.9, respectively). (C) Treatment with either miR-125–5 p (target position: 79–86) or let-7c-5p (target position: 166–173 and 262–269) decreased the luciferase activity of the positive control TRIM71 reporter construct (one-way ANOVA F(3,8) = 217.799, p<0.0001; Dunnett’s post hoc test control vs let-7c-5p or miR-125–5 p: p=0.0000002 or p=0.0000002, respectively). (D) Left, luciferase activity was enhanced when cells were co-transfected with Let-7c-5p (2 nM) and its inhibitor (LNA-anti-let7c-5p, 25 pmol) compared to Let-7c-5p treatment alone (t-test, p=0.00003). Right, luciferase activity was increased when cells were co-transfected with miR128-3p (2 nM) and its inhibitor (LNA-anti- miR128-3p,25 pmol) compared to miR128-3p treatment alone (t-test, p=0.000006). (E) Top, schematics of WT and mutant sequences. Bottom, the effect of Let-7c-5p or miR128-3p treatment on reducing the luciferase activity of the WT F1 construct was prevented in the mutant F1 construct (let7c-5p: WT vs. Mut, t-test, p=0.00000005; miR 128–3p: WT vs. Mut, t-test, p=0.0001). *** indicates p<0.0005. Data are represented as mean ±SEM.
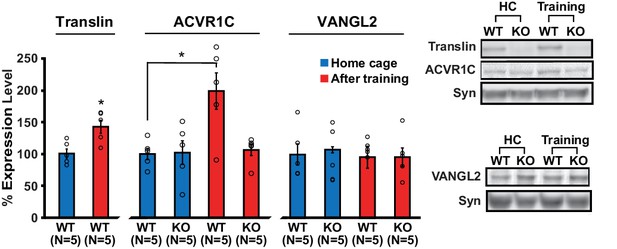
Loss of translin prevents elevation of ACVR1C protein levels 30 min following training in the object location memory task.
Training induced an increase in translin levels in hippocampal synaptosomes (t-test, p=0.01), while the loss of translin prevented a training-induced upregulation of synaptosomal ACVR1C levels in the hippocampus (two-way ANOVA: genotype, F(1,16) = 5.2, p=0.03; training, F(1,16) = 6.8, p=0.02; genotype X training, F(1,16) = 5.7, p=0.03; Dunnett’s post hoc test, WThomecage vs. WTtraining: p=0.01, WThomecage vs. KOtraining: p=0.9). However, synaptosomal vang-like 2 (VANGL2) levels were unaffected by either genotype or training (two-way ANOVA: genotype, F(1,16) = 0.06, p=0.8; training, F(1,16) = 0.3, p=0.6; genotype X training, F(1,16) = 0.06, p=0.8). Syn: synaptophysin. HC: homecage. * indicates p<0.05. n, number of mice. Data are represented as mean ±SEM.
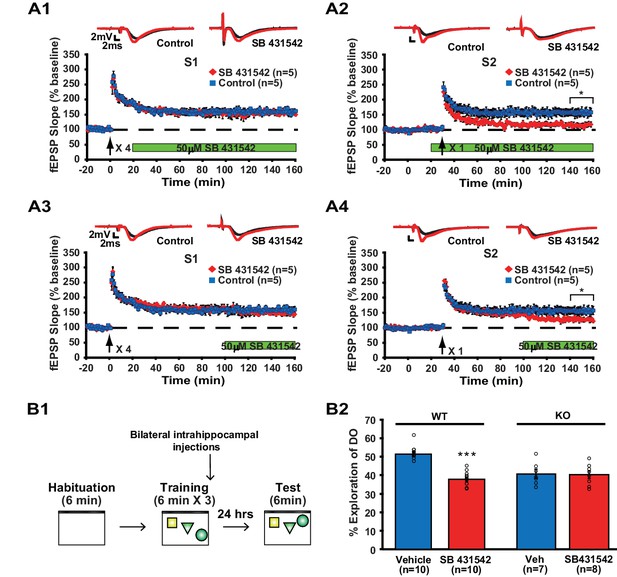
Inhibition of ACVR1C impairs synaptic tagging and long-term memory in WT mice, not translin KO mice.
(A) In hippocampal slices from WT mice, treatment with SB431542 (50 µM), an inhibitor of ACVR1C, starting 10 min before S2 stimulation blocked persistent potentiation in pathway S2 (A2; two-way repeated-measures ANOVA, F(1,8) = 13.9, p=0.007), without affecting 4-train long-lasting potentiation in pathway S1 (A1; two-way repeated-measures ANOVA, F(1,8) = 0.3, p=0.6). Treatment with SB431542 during the last 1 hr of recording impaired the maintenance of S2 potentiation (A4; two-way repeated-measures ANOVA, F(1,8) = 6.1, p=0.039), but not 4-train long-lasting potentiation in pathway S1 (A3; two-way repeated-measures ANOVA, F(1,8) = 1.4, p=0.3). (B1) SB431542 or vehicle was bilaterally injected into the hippocampus immediately after training in the object-location memory task. (B2) While WT mice injected with SB431542 (1 µM) explored the DO significantly less than the vehicle-treated group, translin KO mice injected with SB431542 explored the DO at a level similar to the vehicle-treated group 24 hr after training (two-way ANOVA: genotype, F(1,31) = 9, p=0.005; treatment, F(1,31) = 18.9, p=0.0001; genotype X treatment, F(1,31) = 16.4, p=0.0003; Tukey’s post-hoc, WTvehicle vs. WTSB431542: p<0.0001, KOvehicle vs. KOSB431542: p=0.9). n, number of mice. * indicates p<0.05, *** indicates p<0.00005. Data are represented as mean ±SEM.
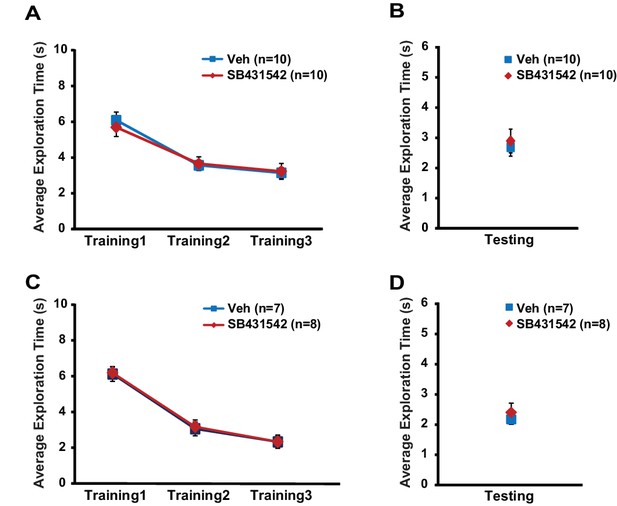
Related to Figure 6.
Exploratory behavior during object-location memory task is unaltered in mice injected with SB431542 into the hippocampus. WT mice subject to post-training intrahippocampal infusion of SB431542 or vehicle exhibited similar exploratory behavior across training sessions (A; two-way repeated measures ANOVA, F(1,18) = 0.039, p=0.8) and during testing session (B; t-test, p=0.7). Translin KO mice injected with SB431542 or vehicle after training showed similar total object exploration time across training sessions (C, two-way repeated measures ANOVA, F(1,13) = 0.03, p=0.9), and during testing (D, t-test, p=0.5). n, number of mice. Data are represented as mean ±SEM.
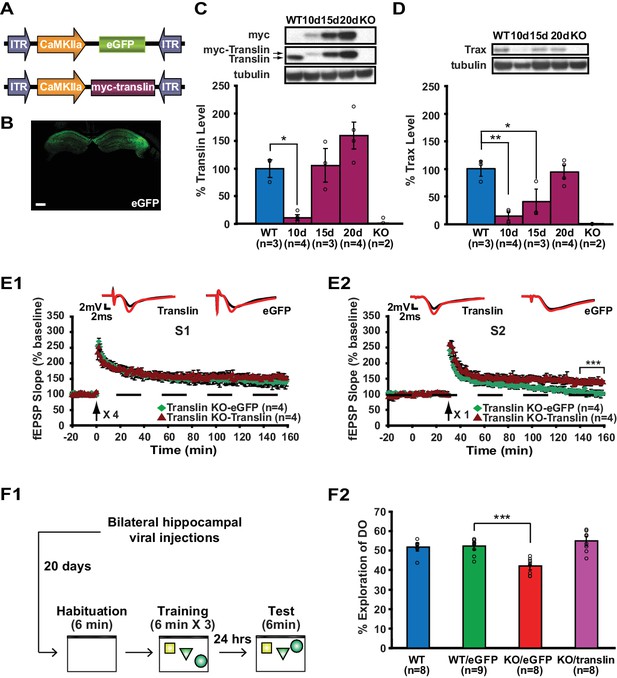
Viral restoration of translin/trax in hippocampal excitatory neurons rescues deficits in synaptic tagging and long-term memory in translin KO mice.
(A) Viral constructs designs. The expression of myc-tagged translin and eGFP is driven by the CamKII promotor. (B) A representative image of hippocampal eGFP expression from a coronal brain section of adult translin KO mice 20 days after viral injection. (C) The level of virally expressed myc-tagged translin reached WT levels 15 to 20 days after viral injection into the hippocampus of adult translin KO mice (one-way ANOVA, F(4,11) = 15.4, p=0.0002, Dunnett’s post hoc test, WT vs. KO15day: p=0.9, WT vs. KO20day: p=0.2). (D) Virally expressed translin restored trax to WT levels 20 days after injection (one-way ANOVA, F(4,11) = 14.7, p=0.0002, Dunnett’s post hoc test, WT vs. KO20day: p=0.9). (E) The viral restoration of translin levels in excitatory neurons of translin KO hippocampus reversed impaired persistent potentiation in pathway S2 (E2; two-way repeated-measures ANOVA, F(1,6) = 29.3, p=0.002) without affecting 4-train long-lasting potentiation in pathway S1 (E1; two-way repeated-measures ANOVA, F(1,6) = 2.5, p=0.2). (F1) A schematic diagram of the object-location memory task performed following bilateral hippocampal viral injections. (F2) Adult translin KO mice expressing eGFP explored the DO significantly less than WT littermates 24 hr after training. However, the behavior of adult translin KO mice expressing translin in excitatory neurons was similar to that of WT littermates or WT littermates expressing eGFP. Viral eGFP expression did not alter the behavior of WT littermates (one-way ANOVA, F(3,29) = 13.9, p=0.00001, Dunnett’s post hoc test, WTeGFP vs. KOeGFP: p=0.0001, WTeGFP vs. KOtranslin: p=0.5, WTeGFP vs. WT: p=0.9). n, number of mice. Scale bar: 20 µm. * indicates p<0.05, ** indicates p<0.005, *** indicates p<0.0005. Data are represented as mean ±SEM.
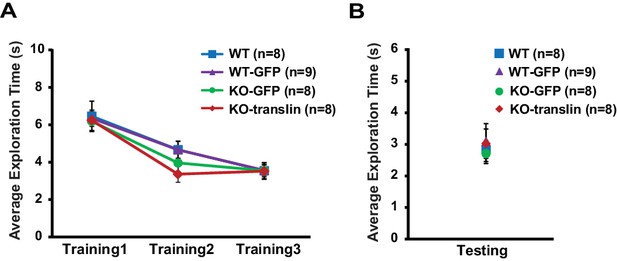
Viral expression does not alter exploratory behavior during object-location memory task in translin KO mice.
(A) Compared to non-injected WT littermates, translin KO mice injected with translin or eGFP virus and WT littermates injected with eGFP virus showed similar total object exploration time across training sessions (two-way repeated measures ANOVA, F(3,25) = 0.3, p=0.8). (B) Total object exploration time during testing was similar across groups (one-way ANOVA, F(3,29) = 0.2, p=0.9). n, number of mice. Error bars reflect S.E.M.
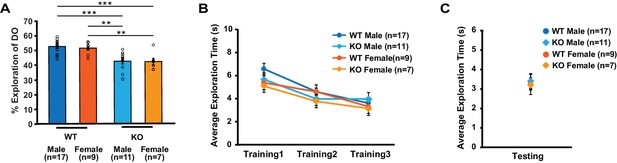
Male and female mice show similar exploratory behavior during object-location memory task.
Data from Figures 1D and 7F2 (non-injected mice and mice injected with eGFP virus) are combined to test gender difference in behavior. (A) Within each genotype, male and female mice explored the DO at the similar level (two-way ANOVA: genotype, F(1,40) = 36.2, p<0.0001; gender, F(1,40) = 0.2, p=0.6; genotype X treatment, F(1,40) = 0.07, p=0.8; Tukey’s post-hoc, WTmale vs. KOmale: p<0.0001, WTmale vs. KOfemale: p=0.0002, WTfemale vs. KOmale: p=0.002, WTfemale vs. KOfemale: p=0.004). (B) Total object exploration time across training sessions was similar between groups (two-way repeated measures ANOVA, F(3,36) = 2, p=0.1). (C) Total object exploration time during testing was similar across groups (one-way ANOVA, F(3,40) = 0.1, p=0.9). ** indicates p<0.005, *** indicates p<0.0005. n, number of mice. Error bars reflect S.E.M.
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.27872.016





