Modulation of occluding junctions alters the hematopoietic niche to trigger immune activation
Figures
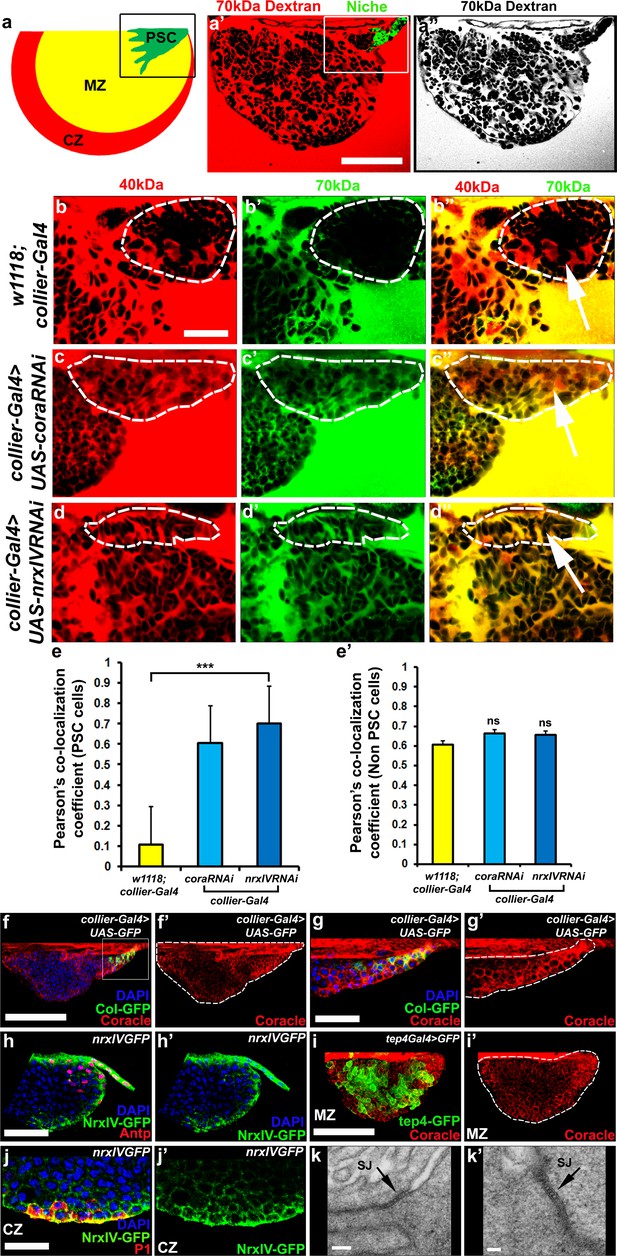
Septate junctions form a permeability barrier at the PSC in the lymph gland.
(a) Schematic representation of the lymph gland (PSC-Posterior Signalling Centre; MZ-Medullary Zone; CZ-Cortical Zone). (a’,a’’) 70kDa-dextran (Red) is excluded from the PSC (Green). (b–d’’) Dual Dye Assay with 70kDa-dextran (Green) and 40kDa-dextran (Red) dextran in wild-type (b–b’’) and following PSC specific knockdown of cora or NrxIV (c–d’’). (e,e’) Pearson’s co-localization co-efficient quantification of data in b-d in PSC and non-PSC cells. (f–f’’) Coracle expression (red) in PSC cells (collier-Gal4 >UAS GFP; green). (g–g’) Enlarged view of boxed region in (f). (h–h’). NrxIV expression (green) in PSC cells (Antp antibody; Red). (i–i’) Coracle expression (red) in MZ cells (tep4-Gal4 > UAS GFP; green). (j–j’) NrxIV expression (green) in CZ cells (P1 antibody; red). (k–k’) Electron micrographs showing septate junctions in between PSC cells. Nuclei labeled with DAPI (Blue). (a’–a’’,f,g) ***=P < 0.001; ns = non significant. Error bars represent s.d. Scale Bars:(a’,a’’,f–f’, i–i’) 50 μm, (b–d’’,g–h’, j–j’) 40 μm, (k) 100 nm (k’) 50 nm.
-
Figure 1—source data 1
Contains numerical quantitation represented in Figure 1e.
- https://doi.org/10.7554/eLife.28081.005
-
Figure 1—source data 2
Contains numerical quantitation represented in Figure 1e’.
- https://doi.org/10.7554/eLife.28081.006
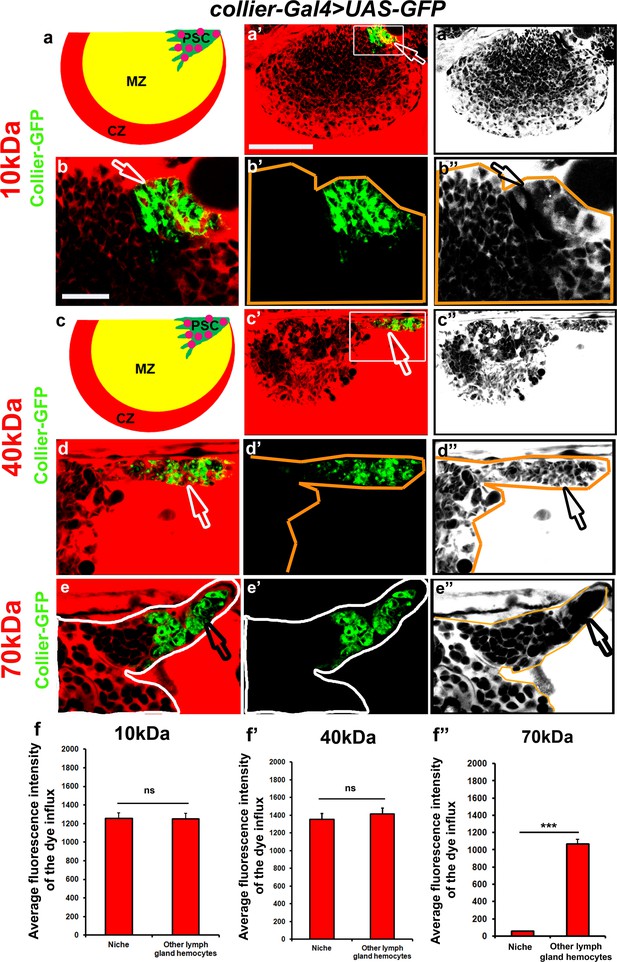
Low molecular weight dyes are not excluded from the PSC.
(a’,a’’) 10 and (c’,c’’) 40 kDa dextran (Red) are not excluded from the PSC also shown in the (a,c) schematic representation of lymph glands. (b–b’’ and d–d’’) High-magnification images of boxed region in (a’ and c’). (e–e’’) 70 kDa dextran (Red) is excluded from the PSC. Pink circles represent the 10 and 40 kDa dextran entering the PSC. (f–f’’) Quantitation of 10, 40 and 70 kDa dye influx in the PSC. (a’,b–b’,c’,d–d’ and e–e’) PSC is labeled with Collier-GFP (green; UAS-GFP driven by collier-Gal4). Scale Bar: (a’,a’’ and c’,c’’) 50 μm and (b–b’’,d–d’’ and e–e’’) 40 μm.
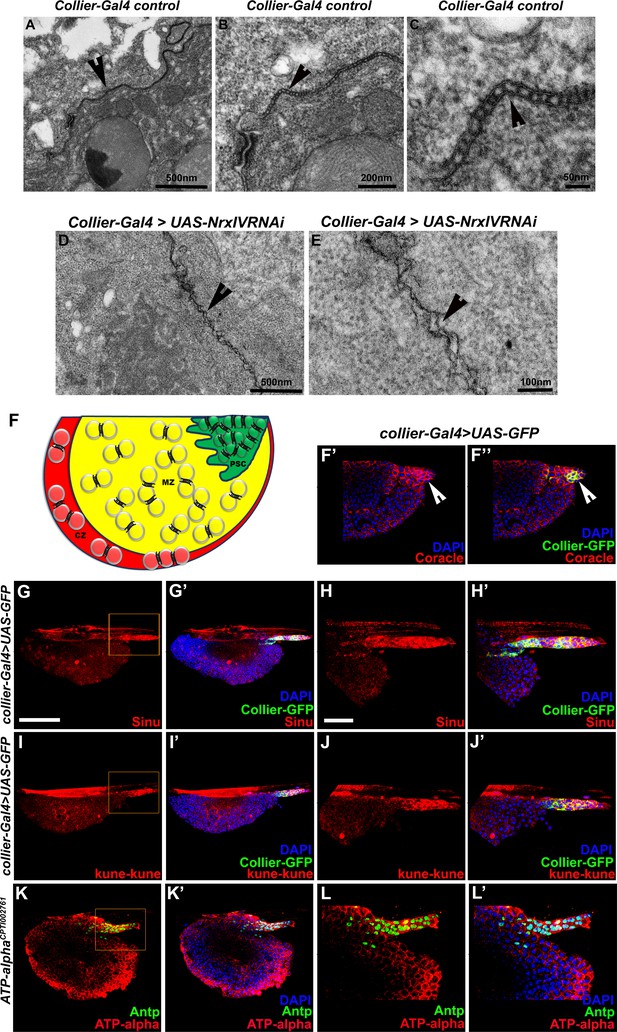
Septate junctions in the PSC are absent upon PSC- specific depletion of septate junctions.
(A–C) Low (A), middle (B) and high (C) magnification electron micrographs of septate junctions in between the cells in the PSC. (D–E) Electron micrographs of the PSC cells showing that septate junctions fail to form and are disorganized upon PSC- specific depletion of NrxIV (collier-Gal4 >UAS NrxIVRNAi). (F) Septate junction localization in the PSC and the primary lymph gland lobe of the LG. High expression of Coracle (Red) is also found in the PSC cells that are close to the MZ region in the inner z-confocal sections of the lymph gland lobe (F’–F’’). (H–L’) are high magnification images of the boxed regions in (G–K’) showing high levels of expression of Sinuous (Red, G–H’), Kune-kune (Red, I–J’) and ATPα tagged with YFP (Green, K–L’). PSC is labeled with Collier-GFP (green; UAS-GFP driven by collier-Gal4, F’–J’) or Antennapedia (Red, K–L’). White arrows indicate high expression of Coracle in the PSC cells that are closer and further inside the MZ. Nuclei are labeled with DAPI (Blue). Scale Bar: (A,D) 500 nm, (B) 200 nm, (C) 50 nm, (E) 100 nm, (G–K’) 50 μm and (F–F’’, H–L’) 40 μm.
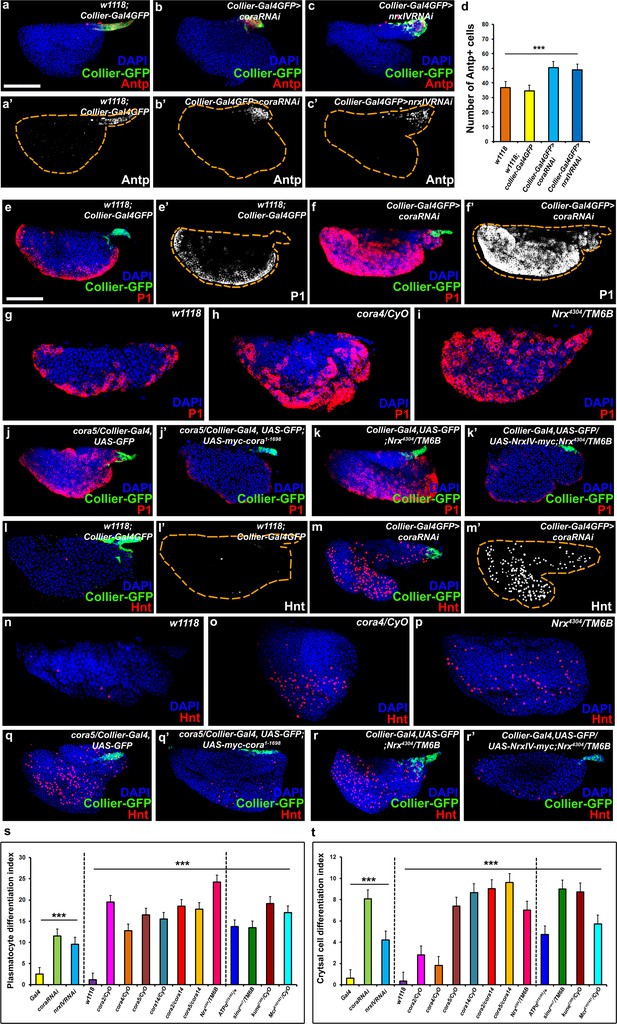
The septate junction-mediated permeability barrier is essential for PSC integrity and function.
(a–d) PSC (red and green) in wild-type, Cora or NrxIV knockdown LGs, (d) quantification of PSC cell numbers. (e–i) Plasmatocyte differentiation (red, white in e’,f’) in LG of control (e,e’,g), Cora knockdown (f,f’), cora (h;cora4/CyO), Nrx (i;Nrx4304/TM6B) heterozygous mutant alleles. PSC specific rescue of Cora (j’) or NrxIV (k’) in cora or NrxIV mutants compared to their respective controls (j,k). (l–p) Crystal Cell differentiation (red; white in l’,m’) in LG of control (l,l’,n), Cora knockdown (m,m’), cora (o;cora4/CyO), Nrx (p;Nrx4304/TM6B) heterozygous mutant alleles. PSC specific rescue of Cora (q’) or NrxIV (r’) in cora or NrxIV mutants compared to their respective controls (q,r). Quantitation of plasmatocyte (s) and crystal cell (t) differentiation for LGs with the phenotype shown in panels e-p as well as for a collection of hypomorphic mutations in SJ (septate junction) components as compared to the w1118 control. (a–c,e,f,j–k’,l,m,q–r’) PSC labeled with GFP (green; collier-Gal4 >UAS GFP) and/or Antennapedia (Red:a-c; white a’-c'). Plasmatocytes labeled with P1 (Red:e,f,g-k’;white:e’,f’), crystal cells labeled with Hnt (Red:l,m,n-r’; white:l’,m’). ***=P < 0.001. Error bars = s.e.m. PSC- specific Cora or NrxIV knockdown and mutant genotypes were compared to respective parental or wild type controls for the statistical analysis with the t-test. Scale Bar: (a–c’,e–r’) 50 μm.
-
Figure 2—source data 1
Contains numerical data for plasmatocyte differentiation indices presented in Figure 2—figure supplement 2s.
- https://doi.org/10.7554/eLife.28081.013
-
Figure 2—source data 2
Contains numerical data for crystal cell differentiation indices presented in Figure 2t.
- https://doi.org/10.7554/eLife.28081.014
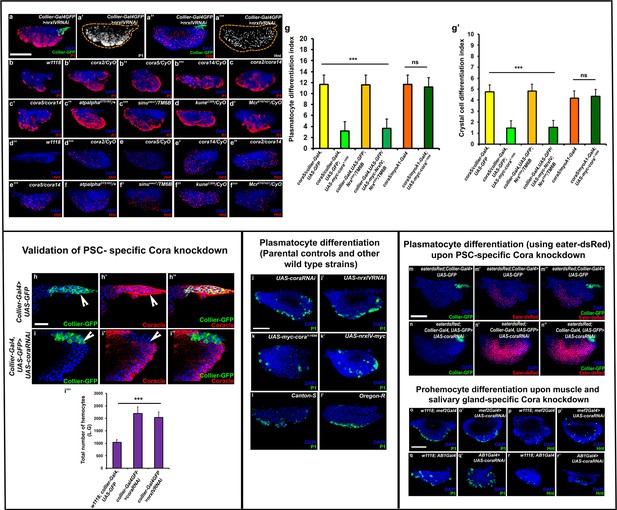
Reduced expression of septate junction components affects blood cell homeostasis in the lymph gland.
(a–l’) P1 and Hnt positive plasmatocyte and crystal cell numbers increase in PSC specific (a–a’’’) Nrx knockdown and in septate junction related mutant alleles and hypomorphs namely (b’,d’’’) cora2/CyO, (b’’,e) cora5/CyO, (b’’’,e’) cora14/CyO, (c,e’’) cora2/cora14, (c’,e’’’) cora5/cora14, (c’’,f) atpalphaDTS1R2/+, (c’’’,f’) sinunwu7/TM6B, (d,f’’) kuneC309/CyO, (d’,f’’’) McrEY07421/CyO larvae as compared to the (b,d’’) wild-type control. Mutant alleles of cora were recombined to obtain viable allelic combinations of cora2/cora14 and cora5/cora14. (g–g’) Quantitation of plasmatocyte and crystal cell differentiation indices upon over-expression of Cora in the PSC or gut in the cora mutant genetic background. (h–i’’) Validation of Coracle knockdown using anti-Coracle antibody upon collier-Gal4 mediated PSC- specific Coracle knockdown as compared to control. (i’’’) Total number of DAPI positive hemocytes are increased in the collier-Gal4 mediated cora or nrxIV depleted L.G’s as compared to the control. (j–l’) P1 positive plasmatocyte differentiation in the parental UAS controls and wild type strains namely (j) UAS-coraRNAi, (j’) UAS-NrxIVRNAi, (k) UAS-myc-cora1-1698, (k’) UAS-NrxIV-myc, (l) Canton-S, (l’) Oregon-R. (m–n’’) Eater-dsRed positive plasmatocyte differentiation upon collier-Gal4 mediated PSC- specific depletion of Coracle as compared to its control. (o–r’) P1 or Hnt positive plasmatocyte or crystal cell differentiation upon mef2-Gal4 mediated muscle- specific or AB1-Gal4 mediated salivary gland- specific depletion of Coracle as compared to their control. (a–a’’’, h–i’’ and m–n’’) PSC is labeled in green with Collier-GFP (UAS-GFP driven by collier-Gal4). Plasmatocytes are labeled with P1 (Red:a,b-d’; white:a’; green:j-l’, o–o’ and q–q’) or Eater-dsRed (Red:m-n’’), crystal cells with Hnt (Red:a’’,d’’-f’’’; white:a’’’; green:p-p’ and r-r’). Coracle expression marked by anti-Coracle antibody (Red:h-i’’). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001, ns indicates non-significant. Error bars represent s.e.m. Scale Bar: (a–f’’’ and j–r’) 50 μm, (h–i’’) 40 μm.
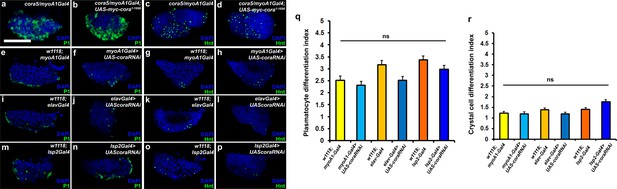
Depletion of septate junctions in other Drosophila organs does not affect lymph gland hematopoiesis.
(a–d) Over-expression of Cora in the gut using myo1A-Gal4 in the cora mutant does not rescue the plasmatocyte and crystal cell differentiation defect. (e–p) Systemic knockdown of Cora using gut specific myo1A-Gal4, pan- neuronal elav-Gal4 or fat body specific lsp2-Gal4 does not affect the number of P1 positive plasmatocytes or Hnt positive crystal cells. (q–r) Quantitation of plasmatocyte and crystal cell differentiation indices upon systemic knockdown of Cora using gut specific myo1A-Gal4, pan- neuronal elav-Gal4 or fat body specific lsp2-Gal4. Plasmatocytes are labeled with P1 (Green:a-b,e-f,i-j and m-n), crystal cells with Hnt (Green:c-d,g-h,k-l and o-p). Nuclei are labeled with DAPI (Blue). ns indicates non-significant. Error bars represent s.e.m. Scale Bar: (a–p) 50 μm.
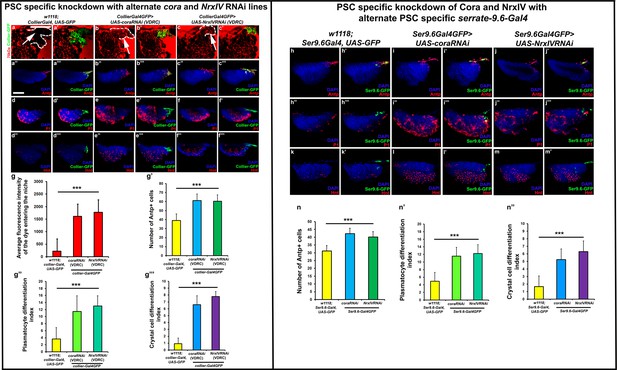
Depletion of septate junctions using alternate RNAi lines and additional PSC specific Ser9.6-Gal4 driver results in breakdown of permeability barrier and loss of blood cell homeostasis.
(a–c’) 70 kDa Dextran (Red) enters the PSC upon collier-Gal4 mediated knockdown of SJ’s using alternate VDRC RNAi lines as compared to the control. (g) Quantitation of dye influx in PSC upon SJ knockdown using collier-Gal4. Antp positive PSC cells (b’’–c’’’), P1 positive plasmatocytes (e–f’) and Hnt positive crystal cell numbers (e’’–f’’’) increase upon depletion of SJ’s using alternate VDRC RNAi lines driven by collier-Gal4 as compared to the control (a’’–a’’’,d–d’,d’’–d’’’). (g’–g’’’) Quantitation of PSC, plasmatocyte and crystal cell numbers upon collier-Gal4 mediated knockdown of SJ’s using alternate VDRC RNAi lines. Antp positive PSC cells (i–j’), P1 positive plasmatocytes (i’’–j’’’) and Hnt positive crystal cell counts (l–m’) also increase upon depletion of SJ’s using additional PSC specific – Ser9.6 driver as compared to their respective controls (h–h’’’,k and k’). (n–n’’) Quantitation of PSC, plasmatocyte and crystal cell numbers upon collier-Gal4 mediated knockdown of SJ’s using additional Ser9.6-Gal4. PSC is co-labeled in green with collier-Gal4 or Ser9.6-Gal4 driven GFP or Antp (Red). Plasmatocytes are labeled with P1 (Red), crystal cells with Hnt (Red). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001. Error bars represent s.e.m. Scale Bar: (a–c’) 40 μm, (a’’–f’’’, h–m’) 50 μm.
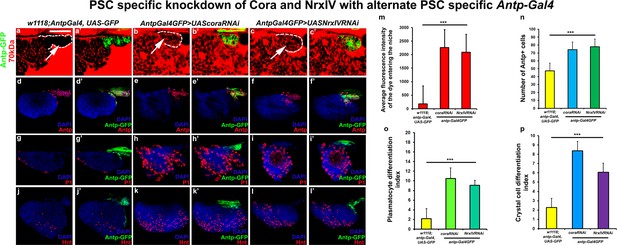
Depletion of septate junctions using additional PSC specific Antp-Gal4 results in breakdown of permeability barrier and loss of blood cell homeostasis.
(a–c’) 70 kDa Dextran (Red) enters the PSC upon Antp-Gal4 mediated knockdown of SJ’s as compared to control. (m) Quantitation of 70 kDa dye influx upon SJ knockdown with Antp-Gal4. Antp positive PSC cells (e–f’), P1 positive plasmatocytes (h–i’) and Hnt positive crystal cell counts (k–l’) also increase upon depletion of SJ’s using additional PSC specific –Antp-Gal4 driver as compared to respective controls (d–d’, g–g’ and j–j’). (n–p) Quantitation of PSC, plasmatocyte and crystal cell numbers upon PSC mediated knockdown of SJ’s using additional Antp-Gal4. PSC is co-labeled in green with Antp-Gal4 driven GFP or Antp (Red). Plasmatocytes are labeled with P1 (Red), crystal cells with Hnt (Red). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001. Error bars represent s.e.m. Scale Bar: (a–c’) 40 μm, (d–l’) 50 μm.
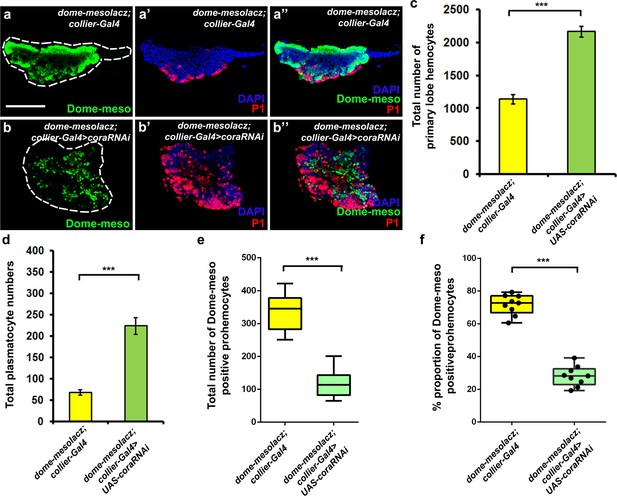
Depletion of septate junctions in the PSC results in a decrease in the proportion of Dome-MESO positive prohemocytes.
(a–b’’) PSC- specific depletion of septate junctions leads to a decrease in the dome-MESO lacZ positive (Green) prohemocyte population and an increase in plasmatocyte differentiation (b–b’’) as compared to controls (a–a’’). (c–e) Quantitation of total number of hemocytes, plasmatocytes and Dome-MESO lacZ positive prohemocytes in the primary lobe upon depletion of septate junctions in the PSC. Quantitation of percent (%) proportion of Dome-MESO lacZ positive prohemocytes upon PSC- specific depletion of septate junctions (f). Plasmatocytes are labeled with P1 (Red, a’–b’’), Dome-MESO lacZ positive prohemocytes (Green, a–b’’). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001. Error bars represent s.e.m. Scale Bar: (a–b’’) 50 μm.
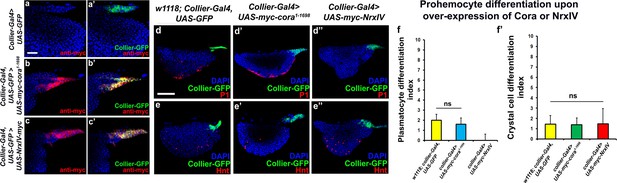
Effects of septate junction components overexpression on blood cell homeostasis.
(a–c’) Validation of myc-tagged Cora or NrxIV over-expression using collier-Gal4 mediated PSC- specific over-expression using anti-myc tag antibody as compared to its control. P1 positive plasmatocyte counts (d’) remain unaltered upon collier-Gal4 mediated over-expression of Cora whereas they are completely suppressed upon collier-Gal4 mediated over-expression of NrxIV (d’’) as compared to their respective controls (d). Hnt positive crystal cell counts are not affected upon collier-Gal4 mediated over-expression of Cora (e’) or Nrx (e’’) as compared to control (e). (f–f’) Quantitation of plasmatocyte and crystal cell counts upon collier-Gal4 mediated over-expression of cora or nrxIV. PSC is co-labeled in green with collier-Gal4 driven GFP. Plasmatocytes are labeled with P1 (Red), crystal cells with Hnt (Red). Nuclei are labeled with DAPI (Blue). ns indicates non-significant. Error bars represent s.d. Scale Bar: (a–c’) 40 μm, (d–e’’) 50 μm.
-
Figure 3—source data 1
Contains numerical data plotted in Figure 3f.
- https://doi.org/10.7554/eLife.28081.017
-
Figure 3—source data 2
Contains numerical data plotted in Figure 3f’.
- https://doi.org/10.7554/eLife.28081.018
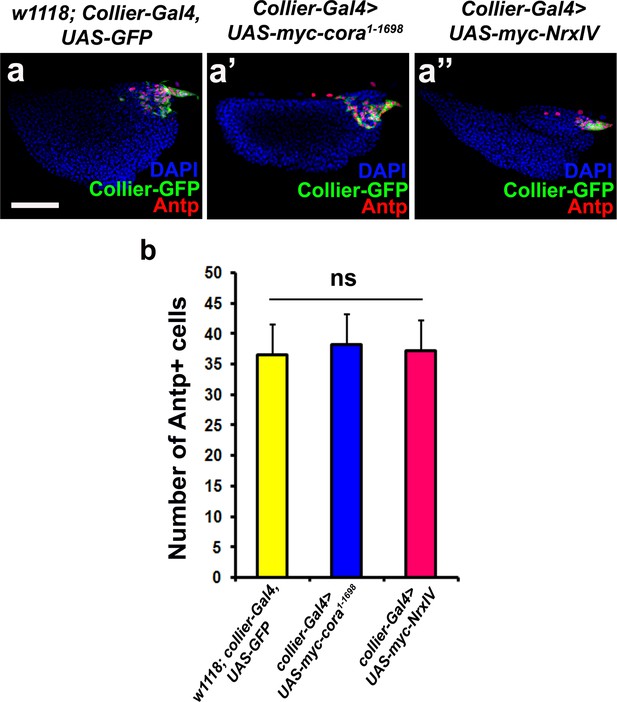
PSC- specific over-expression of Cora or NrxIV does not alter the PSC size.
Antp positive PSC cell counts (a–a’’) do not show a significant change upon collier-Gal4 mediated over-expression of Cora or NrxIV. (b) Quantitation of PSC cell counts upon collier-Gal4 mediated over-expression of cora or NrxIV. PSC is co-labeled in green with collier-Gal4 driven GFP and Antp (Red). Nuclei are labeled with DAPI (Blue). ns indicates non-significant. Error bars represent SD. Scale Bar: (a–a’’) 50 μm.
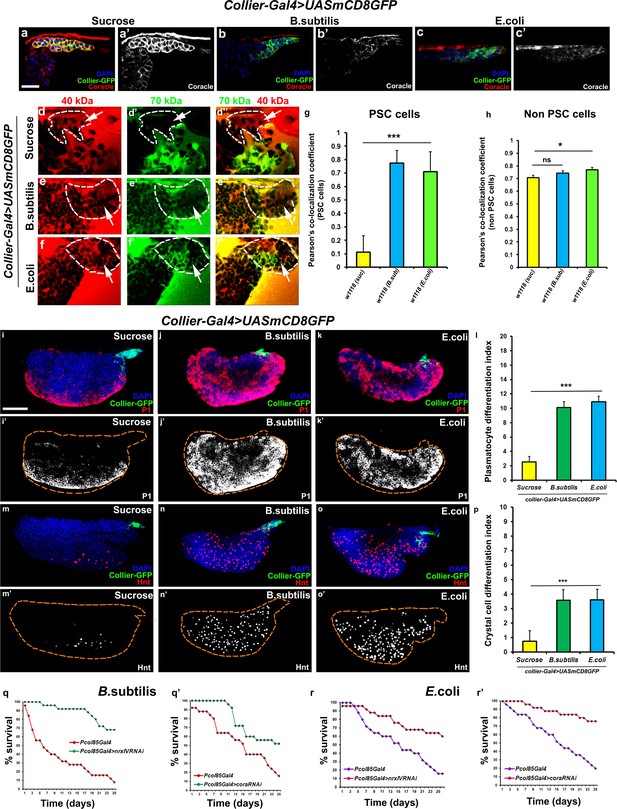
Bacterial infection results in breakdown of the permeability barrier leading to prohemocyte differentiation.
(a–c’) Coracle (Red:a-c; white:a’-c’) expression in the PSC (Green) decreased upon infection with (b,b’) B. subtilis and (c,c’) E. coli as compared to (a,a’) sucrose treated control. (d–h) Dual Dye Assay with 70kDa-dextran (Green) and 40kDa-Dextran (Red) in control, (d–d’’) and B. subtilis (e–e’’) E. coli (f–f’’) infected larva. (g,h) Pearson’s co-localization co-efficient quantification of data in d-h in PSC and non-PSC cells. (i–k) Plasmatocyte differentiation (red; white in i’,j’,k’) in (i) control, and (j) B. subtilis or (k) E. coli infected larva. Quantitation of plasmatocyte (t) differentiation data. (m–o) Crystal cell differentiation (red; white in i’,j’,k’) in (m) control, and (n) B. subtilis or (o) E. coli infected larva. Quantitation of crystal cell (p) differentiation data. (q–r’) Survival plots for flies infected with B. subtilis (q,q’) or E. coli (r,r’) over time in control versus PSC specific Nrx or Cora knockdown flies. (a–c,d–I,k–m,o–q) PSC labelled in green (collier-Gal4 >UAS GFP). Plasmatocytes labeled with P1 (Red:i-k; white:i’-k’), crystal cells with Hnt(Red:m-o; white:m’-o’). Nuclei labeled with DAPI (Blue). ***=P < 0.001, *=P < 0.1 and ns = non significant. Error bars represent s.e.m. Scale Bar: (i–k’,m–o’) 50 μm, (a–f’’) 40 μm.
-
Figure 4—source data 1
Contains numerical data plotted in Figure 4g.
- https://doi.org/10.7554/eLife.28081.023
-
Figure 4—source data 2
Contains numerical data plotted in Figure 4h.
- https://doi.org/10.7554/eLife.28081.024
-
Figure 4—source data 3
Contains numerical data plotted in Figure 4l.
- https://doi.org/10.7554/eLife.28081.025
-
Figure 4—source data 4
Contains numerical data plotted in Figure 4p.
- https://doi.org/10.7554/eLife.28081.026
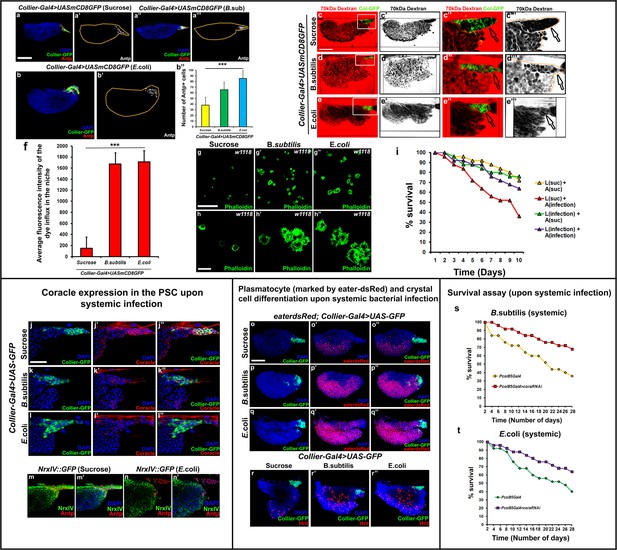
Bacterial infection induces changes in the prohemocyte microenvironment.
(a–b’) PSC cell numbers increase upon infection with (a’’,a’’’) B. subtilis and (b,b’) E. coli as compared to (a,a’) sucrose treated control. (b’’) Quantitation of PSC cell numbers upon infection. 70 kDa dextran (Red) can access the PSC cells upon infection with either B. subtilis (d–d’’’) or E. coli (e–e’’’) as compared to the sucrose control (c–c’’’). (f) Quantitation of 70 kDa dye influx upon bacterial infection. Circulating hemocytes are activated and show increased phalloidin (Green) positive filopodial extensions upon infection with B. subtilis (g’,h’) or E. coli (g’’,h’’) as compared to the sucrose control (g,h). (i) Flies having prior exposure to E. coli in larval stages (L) have better survival ability upon subsequent infection in adult stages (A) as compared to the respective controls. (j–l’’) Coracle (Red) expression in the PSC is down-regulated upon systemic infection with B. subtilis (k–k’’) or E. coli (l–l’’) as compared to sucrose control (j–j’’). (m–n’) NrxIV::GFP expression (Green) in the PSC marked with Antp (Red) is down-regulated upon systemic infection with E. coli (n–n’) as compared to sucrose control (m–m’). (o–r’’) Eater-dsRed (Red) positive plasmatocytes or Hnt (Red) positive crystal cells are increased upon systemic infection with B. subtilis (p–p’’,r’) or E. coli (q–q’’,r’’) as compared to sucrose control (o–o’’,r). (s–t) Survival plots for flies systemically infected with B. subtilis or E. coli over time in control versus PSC specific Cora knockdown flies. (a,a’’,b,c–e’’, j–l’’ and o–r’’) PSC is labeled in green with Collier-GFP (UAS-GFP driven by collier-Gal4) and Antennapedia (Red:a,a’’,b, m-n’; white:a’,a’’’,b’). (i) Suc indicates sucrose control while infection indicates bacterial infection for the survival analysis. Plasmatocytes are labeled with Eater-dsRed (Red:o-q’’), crystal cells with Hnt (Red: r-r’’). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001. Error bars represent s.e.m. Scale Bar: (a–e’ and o–r’’) 50 μm, (c’’–e’’’, j–n’) 40 μm and (g–h’’) 100 μm.
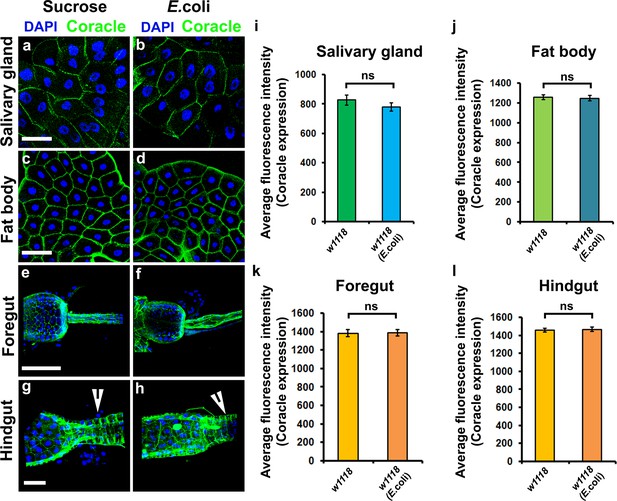
Bacterial infection does not affect Coracle expression in other Drosophila organs.
(a–l) Coracle (Green) expression in the salivary gland (a,b); fat body cells (c,d); foregut (e,f); hindgut (g,h) is not affected upon systemic infection of Drosophila larvae with E. coli as compared to sucrose controls. Quantitation of Coracle expression in the different organs upon infection with E. coli (i–l). Nuclei are labeled with DAPI (Blue). ns represents non-significant. Scale Bar: (a–d) 40 μm, (e–f) 80 μm and (g–h) 50 μm.
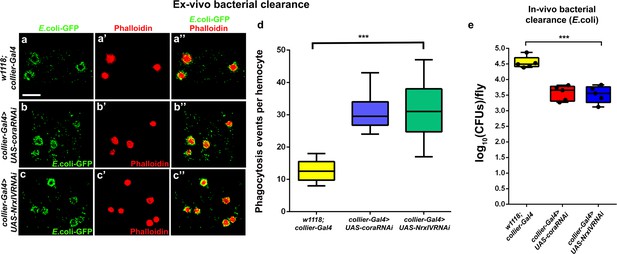
Flies bearing PSC- specific depletion of septate junctions have a better ex-vivo and in-vivo bacterial clearance ability.
(a–d) Hemocytes derived from flies bearing PSC- specific depletion of cora (b-b’’, collier-Gal4 >UAS coraRNAi) or NrxIV (c-c’’, collier-Gal4 >UAS NrxIVRNAi) have a better phagocytic ability as compared to circulating hemocytes derived from controls (a-a’’, w1118; collier-Gal4) in the ex-vivo bacterial clearance phagocytosis assay. (d) Quantitation of phagocytic events of the GFP tagged E. coli strain per hemocyte. (e) Flies bearing PSC- specific depletion of septate junctions have a lower bacterial load 24 hr post infection with Ampicillin resistant E. coli as compared to controls in the in-vivo bacterial clearance assay. Cell boundaries are marked with Phalloidin (Red, (a’–c’’), GFP tagged E. coli marked in green. *** indicates p<0.001. Scale Bar: (a–c’’) 50 μm.
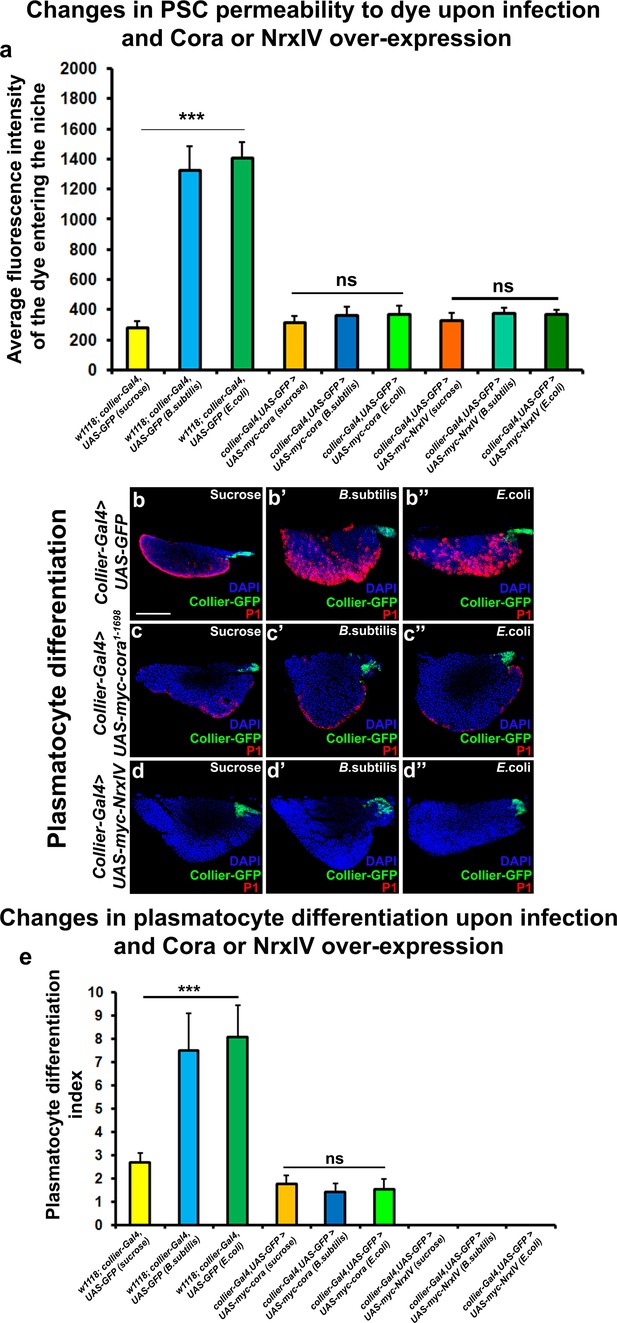
Over-expression of septate junction components in the PSC blocks infection-induced permeability barrier breakdown and prohemocyte differentiation.
(a) Quantitation of 70 kDa dye influx in the PSC upon infection of larvae over-expressing SJ’s in the PSC using collier-Gal4. (b–e) In contrast to controls (b–b’’), P1-positive plasmatocytes counts are not increased upon B. subtilis or E. coli mediated infection of larvae over-expressing Cora (c–c’’) or NrxIV (d–d’’) in the PSC using collier-Gal4. (e) Quantitation of plasmatocyte numbers upon infection of larvae over-expressing SJ’s in the PSC using collier-Gal4. PSC is co-labeled in green with collier-Gal4 driven GFP. Plasmatocytes are labeled with P1 (Red). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001 and ns indicates non-significant. Error bars represent s.d. Scale Bar: (b–d’’) 50 μm.
-
Figure 5—source data 1
Contains numerical data for quantitation in Figure 5a.
- https://doi.org/10.7554/eLife.28081.030
-
Figure 5—source data 2
Contains numerical data for quantitation in Figure 5e.
- https://doi.org/10.7554/eLife.28081.031
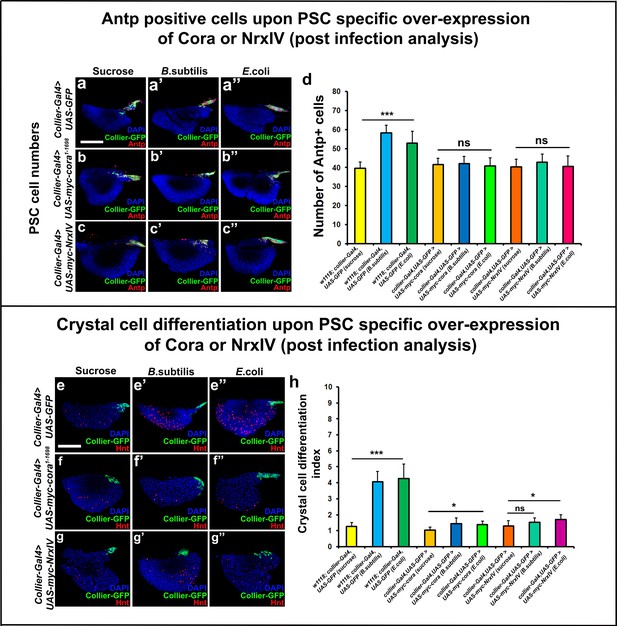
Over-expression of septate junctions in the PSC blocks infection induced prohemocyte differentiation.
Antp positive PSC cell counts are unaltered (a–c’’) and Hnt positive crystal cell counts remain unaltered (e–g’’) upon infection with B. subtilis or E. coli of larvae over-pressing Cora or NrxIV using collier-Gal4. (d,h) Quantitation of PSC and crystal cell numbers upon infection of larvae over-expressing SJ’s in the PSC using collier-Gal4. PSC is co-labeled in green with collier-Gal4 driven GFP and Antp (Red), crystal cells with Hnt (Red). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001, * indicates p<0.1 and ns indicates non-significant. Error bars represent SD. Scale Bar: (a–c’’, e–g’’) 50 μm.
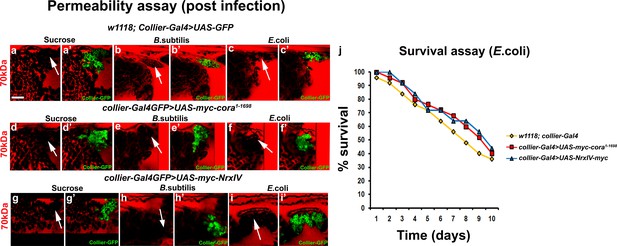
Over-expression of septate junctions in the PSC blocks infection induced permeability barrier breakdown.
(a–i’) 70 kDa Dextran (Red) cannot access the PSC upon B. subtilis (e,e’,h,h’) or E. coli (c,c’,f,f’) mediated infection of larvae having collier-Gal4 driven over-expression of Cora or Nrx as compared to their respective sucrose controls (a–c’,d,d’,g,g’). (j) Survival plot for flies systemically infected with E. coli over time in control versus PSC specific Cora or NrxIV over-expression conditions. PSC is co-labeled in green with collier-Gal4 driven GFP. Scale Bar: (a–i’) 40 μm.
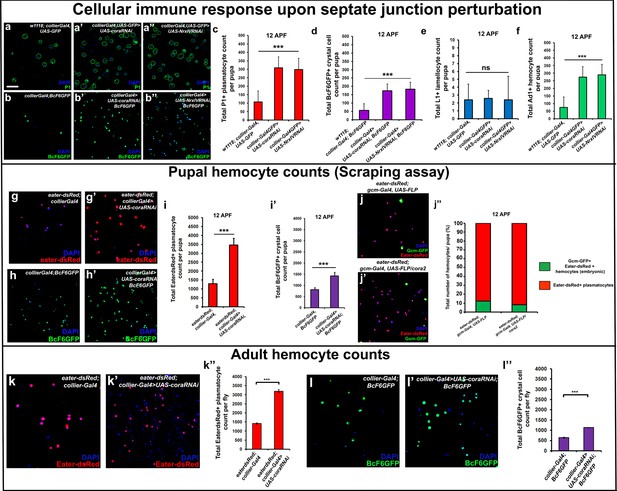
Depletion of septate junctions in the PSC triggers the hemocyte-mediated cellular immune response.
(a–b’’, c, d, f) Total number of P1 (Green) positive circulating plasmatocytes, BcF6GFP (Green) positive circulating crystal cells and Ad1 positive circulating adult hemocytes is highly increased in 12 hr APF (after puparium formation) early pupal circulation upon collier-Gal4 mediated depletion of Cora or NrxIV as compared to the controls. (e) L1 positive circulating lamellocyte counts are not affected. (g–i’) Eater-dsRed (Red) positive plasmatocyte or BcF6GFP (Green) positive crystal cell counts in 12 hr APF pupae using the scraping method for hemocyte isolation upon collier-Gal4 mediated PSC- specific depletion of Coracle as compared to its control. (j–j’’) Total number of hemocytes (represented as %) in 12 hr APF pupae using hemocyte scraping assay in pupae where the Act5C > FRT > CD2>FRT > Gal4 and UAS-FLP and UAS-GFP (Green) transgenes were used to permanently label the hemocytes that express the embryonic plasmatocyte driver, gcm-Gal4 in wild type and cora mutant genetic background with Eater-dsRed (Red) as the plasmatocyte marker. (k–k’’) EaterdsRed (Red) positive adult plasmatocytes isolated from adult flies are increased upon PSC- specific depletion of Coracle using collier-Gal4 as compared to the control. (l–l’’) BcF6GFP (Green) positive adult crystal cells isolated from adult flies are also increased upon PSC- specific depletion of Coracle using collier-Gal4 as compared to its control. Nuclei are labeled with DAPI (Blue). *** indicates p<0.001, * indicates p<0.1 and ns indicates non-significant. Error bars represent s.e.m. Scale Bar: (a–b’’, g–h’,j–j’, k–k’ and l–l’) 50 μm.
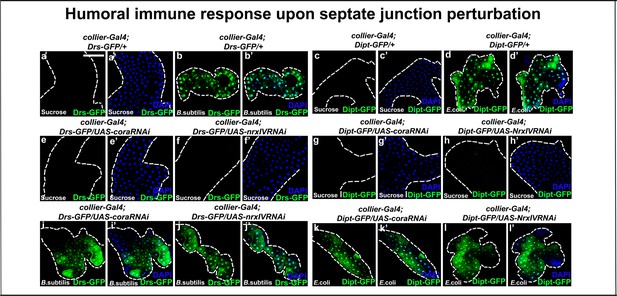
Depletion of septate junctions in the PSC does not trigger endogenous activation of the humoral immune response in the absence of bacterial infection.
(a–l’) Toll or Imd pathway reporters- Drosomycin-GFP (Green) or Diptericin-GFP (Green) are not expressed in the fat bodies of larvae having collier-Gal4 mediated depletion of Cora or NrxIV. GFP expression (Green) in fat bodies of larvae having collier-Gal4 mediated SJ depletion and their respective controls is activated only upon infection with either B. subtilis or E. coli as compared to their sucrose controls respectively. Nuclei are labeled with DAPI (Blue). Scale Bar: (a–l’) 100 μm.
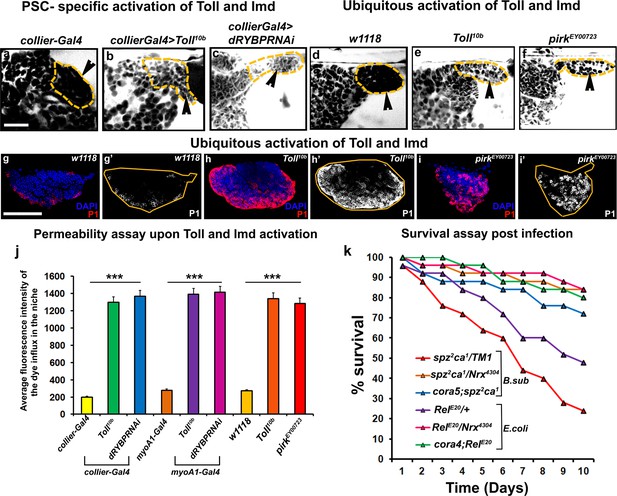
Barrier breakdown contributes to immune activation by triggering prohemocyte differentiation.
(a–f) 70kDa-dextran (white) is excluded from the PSC in controls (a,d) but labels the PSC in genetic backgrounds that activate the Toll (b,e) or Imd (c,f) pathways either in the PSC specifically (b,c) or ubiquitously (e,f); quantitation of this data (j). (g–i) Ubiquitous activation of the Toll (h) and Imd (i) pathways results in plasmatocyte differentiation compared to wild-type controls (g). (k) Survival Plots post infection with B. subtilis or E. coli of Toll (spz2ca1) and Imd (RelE20) pathway mutants combined with hypomorphic NrxIV and cora alleles. (b,c) (l–q’) PSC labelled with GFP (green; collier-Gal4 >UAS GFP). Plasmatocytes labeled with P1 (Red:g-i,l-q; white:g’-i’,l’-q’). Nuclei labeled with DAPI (Blue). ***=P < 0.001. Error bars represent s.e.m. Scale Bar:(g–i’, l–q’) 50 μm,(a–f) 40 μm.
-
Figure 7—source data 1
Contains numerical data for quantitation in Figure 7j.
- https://doi.org/10.7554/eLife.28081.037
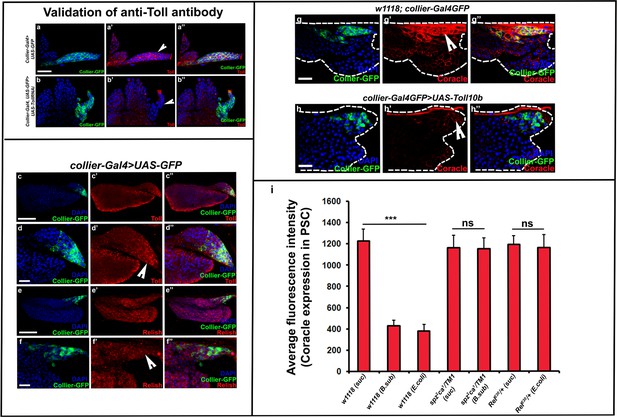
Toll and Imd pathway components are expressed in the lymph gland and regulate lymph gland mediated cellular immune response to infection.
(a–b’’) Validation of PSC- specific Toll (Red) knockdown using anti-Toll antibody as compared to its control in order to validate the specificity of the anti-Toll antibody. (c–f’’) Toll pathway component, Toll (Red) and Imd pathway component, Relish (Red) are uniformly expressed in collier-Gal4 driven GFP positive PSC cells and other hemocytes of the L.G. (g–h’’) Coracle (Red) expression in the PSC is down-regulated upon Toll pathway activation using collier-Gal4 as compared to the control. (i) Coracle expression in the PSC remains unaffected upon infection with B. subtilis or E. coli in spz2ca1 or RelE20 immune mutants as compared to the wild type control. PSC is co-labeled in green with collier-Gal4 driven GFP. Nuclei are labeled with DAPI (Blue). *** indicates p<0.001, * indicates p<0.1 and ns indicates non-significant. Error bars represent s.e.m. Scale Bar: (c–c’’,e–e’’,j–m’’’ and o–r’’’) 50 μm and (a–b’’,d–d’’,f–f’’ and g–h’’) 40 μm.
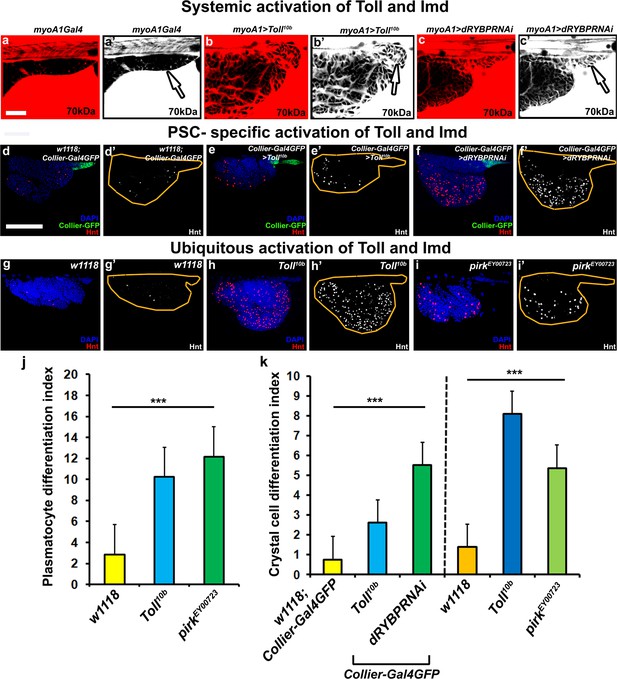
Toll and Imd pathway activation alter the homeostatic balance in the lymph gland.
(a–c) 70 kDa dye (Red:a–c; white:a’–c’) is not excluded from the PSC upon gut specific systemic (b,b’) Toll or (c,c’) Imd activation using myo1AGal4 as compared to the (a,a’) control. Hnt positive crystal cell numbers increase upon localized activation of (e,e’) Toll or (f,f’) Imd in the PSC in contrast to the (d,d’) control. (k) Quantitation of prohemocyte differentiation into crystal cells upon PSC specific activation of Toll or Imd pathways. Hnt positive crystal cell numbers increase upon ubiquitous activation of (h,h’) Toll or (i,i’) Imd as compared to the (g,g’) wild type control. (k) Quantitation of crystal cell differentiation upon ubiquitous activation of Toll and Imd pathways. (j) Quantitation of prohemocyte differentiation into plasmatocytes upon ubiquitous activation of Toll or Imd pathways. (b–c’,e–f’ and k) PSC and gut specific Toll and Imd pathway activation is achieved by expression of UAS- Toll10b and UAS-dRYBPRNAi, respectively. (h–i’,j and k) Ubiquitous activation of the Toll and Imd pathway is achieved using the Toll10b and pirkEY00723 mutant alleles respectively. (d–f) PSC is labeled in green with Collier-GFP (UAS-GFP driven by collier-Gal4). Crystal cells with Hnt (Red:d-i; white:d’-i’). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001. Error bars represent s.e.m. Scale Bar: (a–c’) 40 μm, (d–i’) 50 μm.
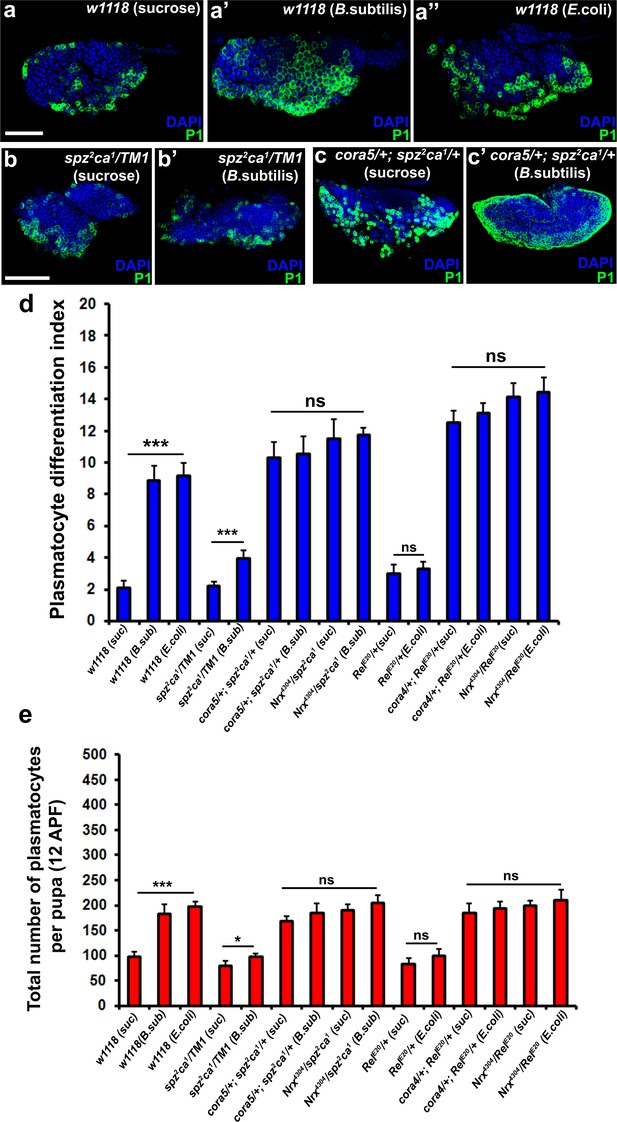
Modulation of permeability barrier at the PSC is a key target of both Toll and Imd pathway in order to mount a robust immune response.
Number of P1 (Green) positive plasmatocytes (b–b’) in spz2ca1 immune mutants is lower in uninfected or infected conditions as compared to respective wild type controls (a–a’’). P1 (Green) positive differentiated plasmatocyte (c–c’) numbers are increased in double mutants of spz2ca1 with cora mutant allele in both uninfected and infected conditions as compared to respective wild type controls. (d) Quantitation of plasmatocyte differentiation index in the immune deficient mutants and immune deficient mutants combined with mutations in SJ components in uninfected and infected conditions. (e) Total number of circulating plasmatocytes in spz2ca1 or RelE20 immune mutants is lesser in uninfected or infected conditions as compared to respective wild type controls. Total number of circulating plasmatocytes is highly increased in double mutants of spz2ca1 with cora or nrxIV mutant allele or RelE20 with cora or nrxIV mutant allele in both uninfected and infected conditions as compared to respective wild type controls. Nuclei are labeled with DAPI (Blue). *** indicates p<0.001, * indicates p<0.1 and ns indicates non-significant. Error bars represent s.e.m. Scale Bar: (a–a’’) 50 μm (b–c’) 60 μm.
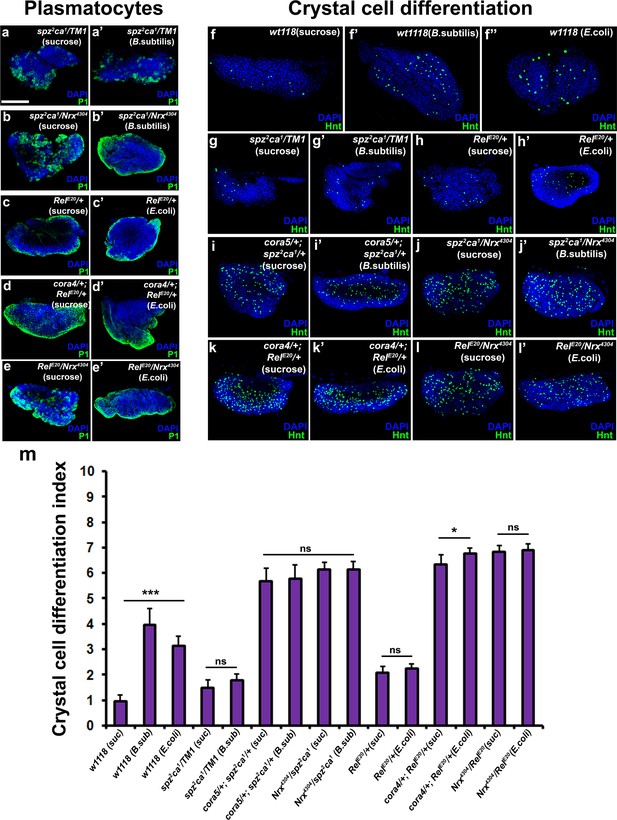
Cellular immune response is upregulated upon partial loss of septate junctions in the immune-deficient mutants.
(a–e) P1 (Green) labeled plasmatocytes in immune deficient spz2ca1 or RelE20 mutants (a–a’ and c–c’) in control larva and larva infected with bacteria. (b–b’ and d–e’) Plasmatocyte numbers are increased by introducing a mutant allele of cora or NrxIV to an immune deficient spz2ca1 or RelE20 mutant background in both uninfected and infected conditions as compared to controls. (f–l) Hnt (Green) labeled crystal cells in wild type controls (f) and in spz2ca1 or RelE20 immune deficient mutants in control larva and larva infected with bacteria. (i–l) Number of Hnt (Green) labeled crystal cells is increased by introducing a mutant allele of cora or NrxIV to an immune deficient spz2ca1 or RelE20 mutant background in both uninfected and infected conditions as compared to controls. (m) Quantitation of crystal cell differentiation indices in the immune deficient mutants and the double mutants with SJ mutant alleles in uninfected and infected conditions. Nuclei are labeled with DAPI (Blue). *** indicates p<0.001, * indicates p<0.1 and ns indicates non-significant. Error bars represent s.e.m. Scale Bar: (a–l’) 50 μm.
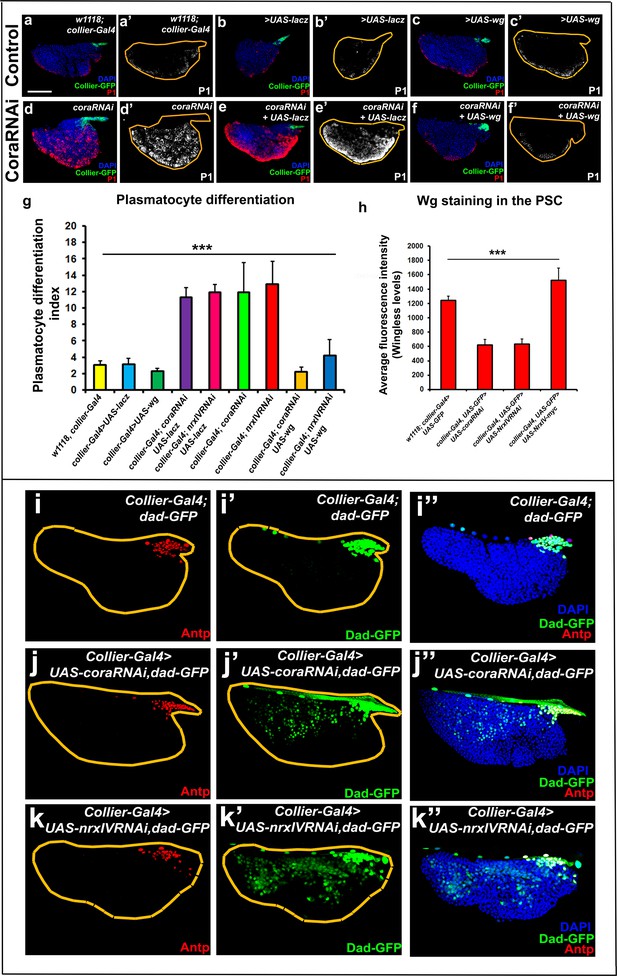
Signalling microenvironment at the PSC is altered upon PSC-specific depletion of septate junction components.
(a–c’) Plasmatocyte differentiation in controls: collier-Gal4, collier-Gal4 >UAS LacZ, collier-Gal4 >UAS wg. (d,d’) Cora depletion (collier-Gal4 >UAS coraRNAi) increases plasmatocyte differentiation but this defect is rescued by co-expression of wg (f,f’, collier-Gal4 >UAS coraRNAi, UAS-wg) but not LacZ (e,e’, collier-Gal4 >UAS coraRNAi, UAS-LacZ). (g) Quantitation of prohemocyte differentiation into plasmatocytes upon wg over-expression in Nrx or Cora depleted PSC. (h) Quantitation of Wg levels in the PSC analyzed by immuno-staining with anti-Wg antibody. Compared to control, lower staining is observed following collier-Gal4 mediated cora or NrxIV depletion in the PSC whereas higher levels are observed after over-expression of UAS-NrxIV-myc in the PSC. Range of BMP signals marked by Dad-GFP (Green) is extended further from the PSC upon collier-Gal4 mediated depletion of cora (j–j’’) or NrxIV (k–k’’) as compared to the control (i–i’’). (a–f’ and i–k’’) PSC labelled with GFP (green; collier-Gal4 >UAS GFP) or with Antennapedia (Red:i-k). Plasmatocytes labeled with P1 (Red:a–f; white:a’–f’). Nuclei labeled with DAPI (Blue). ***=P < 0.001. Plasmatocyte differentiation indices of lymph glands bearing PSC- specific lacZ or Wg over-expression in the PSC following cora or NrxIV knockdown were compared to respective parental controls or cora/NrxIV knockdown genotypes for the statistical analysis with the t-test. Error bars for plasmatocyte differentiation data represent s.e.m. Error bars for the wg fluorescence intensity data represent s.d. Scale Bar:(a–f’, i–k’’) 50 μm.
-
Figure 9—source data 1
Contains numerical data for quantitation in Figure 9g.
- https://doi.org/10.7554/eLife.28081.044
-
Figure 9—source data 2
Contains numerical data for quantitation in Figure 9h.
- https://doi.org/10.7554/eLife.28081.045
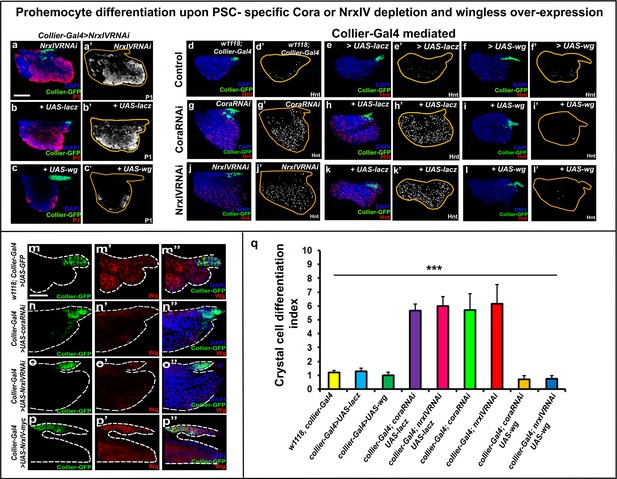
Wg over-expression suppresses prohemocyte differentiation induced by depletion of septate junction components in the PSC.
(a–c’) Nrx depletion (a-a’, collier-Gal4 >UAS NrxIVRNAi) increases plasmatocyte differentiation, this defect is rescued by expression of wg (c-c’, collier-Gal4 >UAS NrxIVRNAi, UAS-wg) but not LacZ (b-b’, collier-Gal4 >UAS NrxIVRNAi, UAS-LacZ). (d–l’) Crystal cell differentiation is normal in (d–f’) collier-Gal4, collier-Gal4 >UAS LacZ, collier-Gal4 >UAS wg controls. (g,g’) Cora or (j,j’) Nrx depletion (collier-Gal4 >UAS coraRNAi or collier-Gal4 >UAS NrxIVRNAi) increases crystal cell differentiation, this defect is rescued by expression of wg (i-i’ and l-l’, collier-Gal4 >UAS coraRNAi, UAS-wg or collier-Gal4 >UAS NrxIVRNAi, UAS-wg) but not LacZ (h-h’ and k-k’, collier-Gal4 >UAS coraRNAi, UAS-LacZ or collier-Gal4 >UAS NrxIVRNAi, UAS-LacZ). (m–p’’) Wg (Red) levels are less in the collier-Gal4 mediated cora or NrxIV depleted PSC whereas the levels are higher upon over-expression of UAS-NrxIV-myc in the PSC as compared to the control. (q) Quantitation of prohemocyte differentiation into crystal cells upon wg over-expression in Nrx or Cora depleted PSC. (a–p’’) PSC is labeled in green with Collier-GFP (UAS-GFP driven by collier-Gal4). Plasmatocytes are labeled with P1 (Red:a-c;white:a’-c’), crystal cells with Hnt (Red:d-l;white:d’-l’). Nuclei are labeled with DAPI (Blue). Crystal cell differentiation indices of lymph glands bearing PSC- specific lacZ or Wg over-expression in the PSC following cora or NrxIV knockdown were compared to respective parental controls or cora/NrxIV knockdown genotypes for the statistical analysis with the t-test. *** indicates p<0.001. Error bars represent s.e.m. Scale Bar: (a–l’) 50 μm, (m–p’’) 40 μm.
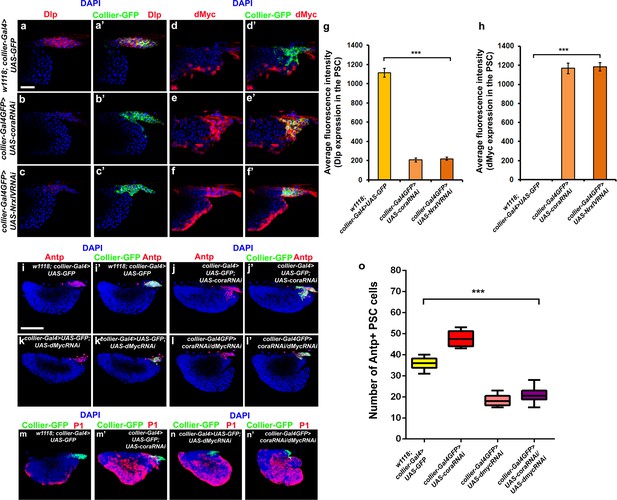
Depletion of septate junction in the PSC changes the signalling micro-environment of the PSC affecting PSC cell numbers and blood cell homeostasis in the LG.
(a–c’, g) Dally- like protein (Dlp) expression is down-regulated upon PSC- specific cora (b-b’, collier-Gal4 >UAS coraRNAi) or NrxIV (c-c’,collier-Gal4 >UAS NrxIVRNAi) knockdown as compared to controls (a-a’, w1118; collier-Gal4 >UAS GFP) quantitated as average fluorescence intensity of Dlp expression in the PSC (g). (d–f’, h) dMyc is expressed upon PSC- specific cora (e-e’, collier-Gal4 >UAS coraRNAi) or NrxIV (f-f’, collier-Gal4 >UAS NrxIVRNAi) knockdown as compared to controls (d-d’, w1118; collier-Gal4 >UAS GFP) quantitated as average fluorescence intensity of dMyc expression in the PSC (h). (i–n’) PSC cell numbers are lesser upon dMyc knockdown (k-k’, collier-Gal4 >UAS dMycRNAi) as compared to wild type controls (i-i’, w1118; collier-Gal4 >UAS GFP) or upon cora knockdown (j-j’, collier-Gal4 >UAS coraRNAi). dMyc knockdown in the cora knockdown genetic background (l-l’, collier-gal4 >UAS coraRNAi/UAS-dMycRNAi) leads to a decrease in the PSC cell numbers as compared to cora knockdown lymph glands (j–j’). There is no significant effect on the plasmatocyte differentiation in the lymph glands upon PSC- specific depletion of dMyc in wild type (n) or cora knockdown (n’) genetic background as compared to the cora knockdown (m’) or wild type controls (m). PSC is labeled in green with Collier-GFP (UAS-GFP driven by collier-Gal4, (a–n’). Plasmatocytes are labeled with P1 (Red:m-n’), PSC cells are labeled with Antp (Red:i-l’). Dlp and dMyc are marked in red (a–f). Nuclei are labeled with DAPI (Blue). *** indicates p<0.001. Error bars represent s.e.m. Scale Bar: (a–f’) 40 μm, (i–n’) 50 μm.
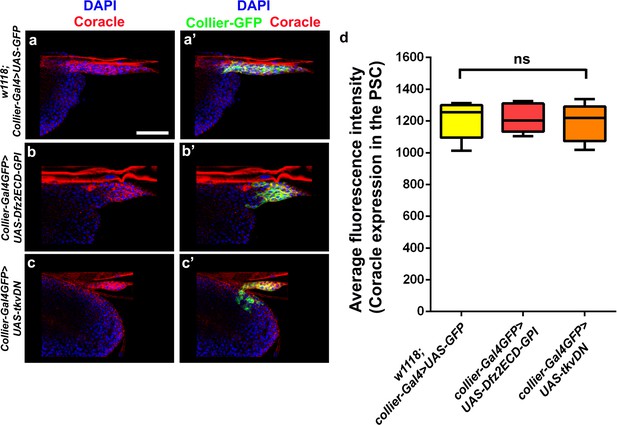
Abrogation of Wg or BMP signalling in the PSC does not affect Coracle expression in the PSC.
(a–d) PSC- specific abrogation of Wg (b-b’, collier-Gal4 >UAS-Dfz2ECD-GPI) or BMP (c-c’, collier-Gal4 >UAS tkvDN) signalling does not affect Coracle (Red) expression as compared to controls (a-a’, w1118; collier-Gal4 >UAS GFP). Quantitation of Coracle expression in the PSC upon abrogation of Wg or BMP signalling (d). Nuclei are labeled with DAPI (Blue). PSC is labeled in green with Collier-GFP (UAS-GFP driven by collier-Gal4, (a–c’). ns represents non-significant. Scale Bar: (a–c’) 40 μm.
Additional files
-
Source code 1
Hemocyte counter.
MATLAB source code for counting prohemocytes, differentiated cells and circulating hemocytes.
- https://doi.org/10.7554/eLife.28081.046
-
Source code 2
Supporting accessory MATLAB file for the hemocyte counter code file.
- https://doi.org/10.7554/eLife.28081.047
-
Transparent reporting form
- https://doi.org/10.7554/eLife.28081.048





