A novel SH2 recognition mechanism recruits Spt6 to the doubly phosphorylated RNA polymerase II linker at sites of transcription
Figures
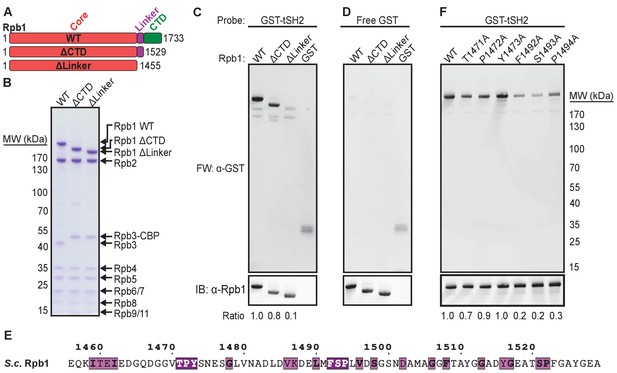
Conserved residues in the Rpb1 linker are necessary for binding Spt6 tSH2.
(A) Full-length Rpb1 (WT) and deletion constructs (ΔCTD and ΔLinker) tested for binding. (B) Coomassie-stained SDS-PAGE of purified RNAPII with the Rpb1 variants depicted in panel A. Rpb3 from ΔCTD and ΔLinker strains retains a TAP calmodulin-binding peptide (CBP) tag not found in wild type. (C) Proteins in (B) were probed with GST-Spt6 tSH2 (Far Western; FW) or an antibody against the Rbp1 N-terminus (y-80; Santa Cruz Biotechnology). The ratio of signal for the far western over α-Rpb1 normalized to WT is indicated. (D) As in (C), except probed with free GST. (E) Sequence of the Saccharomyces cerevisiae (S.c.) Rpb1 linker colored by conservation across Saccharomyces cerevisiae, Schizosaccharomyces pombe, Caenorhabditis elegans, Drosophila melanogaster, Danio rerio, and Homo sapiens aligned using T-Coffee (RRID:SCR_011818; Notredame et al., 2000) and visualized using ESPript (RRID:SCR_006587; Robert and Gouet, 2014). White on purple represents strictly conserved residues, black on magenta is a global similarity score >0.7. (F) Far western blot (FW) measuring binding of Spt6 tSH2 to point mutants in the Rpb1 linker. Ratios were determined as in (C) except that the antibody 8WG16 against the Rpb1 CTD was used (Covance).
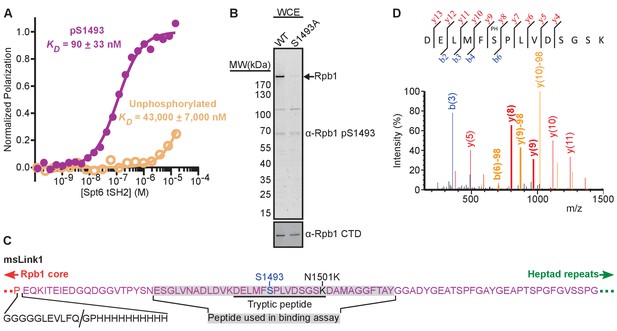
Phosphorylation of Rpb1 linker S1493 enhances binding and occurs in vivo.
(A) Spt6 tSH2 was titrated into Fl-Rpb11476-1511 peptides with or without S1493 phosphorylated, yielding the indicated changes in polarization. Average binding affinities were calculated from at least 3 independent experiments, with the estimated standard deviations indicated. (B) Whole cell extracts from strains with normal (WT) or mutant (S1493A) Rpb1 were tested by western blotting using antibodies against an Rpb1 pS1493 peptide or the Rpb1 CTD. (C) Sequence of the msLink1 linker. A five-glycine linker, PreScission protease site, and 10x-histidine tag were inserted between P1455 (the last ordered residue in most RNAPII crystal structures; red) and E1456 (beginning of the Rpb1 linker; magenta). The PreScission cut site is indicated with a slash. N1501 was mutated to a lysine (black) to allow generation of a tryptic peptide (underlined) containing S1493 (blue). The peptide used in (A) is indicated with a gray background. (D) Mass spectrum of the peptide underlined in (C), indicating phosphorylation on S1493, after purification of msLink1 from yeast. Peaks representing b ions, blue; y ions, red; neutral loss of 98 Da (phosphate), orange. Peaks above 30% intensity are labeled. Peaks flanking the phosphorylation site are labeled and bold. See also Figure 2—figure supplement 1 and Figure 2—source data 1.
-
Figure 2—source data 1
Binding of fluorescently labeled Rpb1 linker peptides to Spt6 tSH2 wild type protein, related to Figure 2A.
- https://doi.org/10.7554/eLife.28723.005
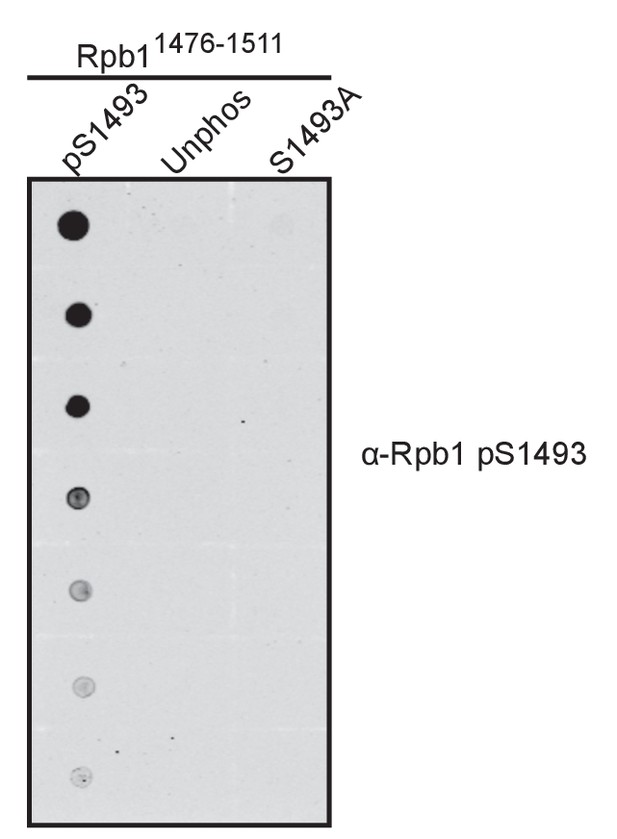

The antibody recognizing S1493 phosphorylated Rpb1 does not recognize unphosphorylated Rpb1 peptides.
A nitrocellulose membrane was spotted with the indicated Rpb11476-1511 peptides and probed with the antibody raised against a pS1493 peptide derived from the Rpb1 linker.
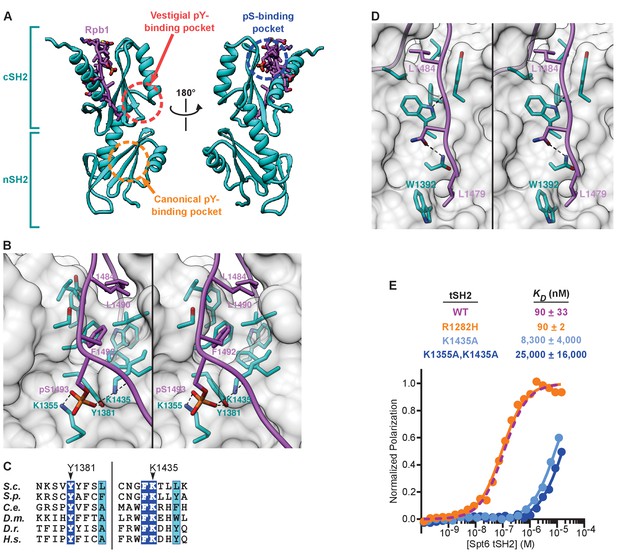
Structure of the Spt6 tSH2-Rpb1 linker complex.
(A) Front and back views. The Spt6 tSH2 nSH2 and cSH2 domains are indicated. (B) Stereo view of the pS1493 binding pocket and hydrophobic groove. (C) Conservation of Spt6 residues that contact pS1493, as described in Figure 1E. Residues contacting the phosphate are indicated (S. cerevisiae numbering). Blue background, strictly conserved. Cyan background, highly similar (global similarity score >0.7). (D) Stereo view of part of the Rpb1 binding groove on the front of Spt6 cSH2. (E) Spt6 tSH2 mutant proteins were titrated into the Fl-Rpb11476-1511 pS1493 peptide and polarization was detected. Average binding affinities calculated from at least 3 independent experiments and estimated standard deviations are shown. The wild type curve from Figure 2 is reproduced here for reference. See also Figure 3—source data 1.
-
Figure 3—source data 1
Binding of fluorescently labeled Rpb1 linker peptides to Spt6 tSH2 mutant proteins, related to Figure 3E.
- https://doi.org/10.7554/eLife.28723.007
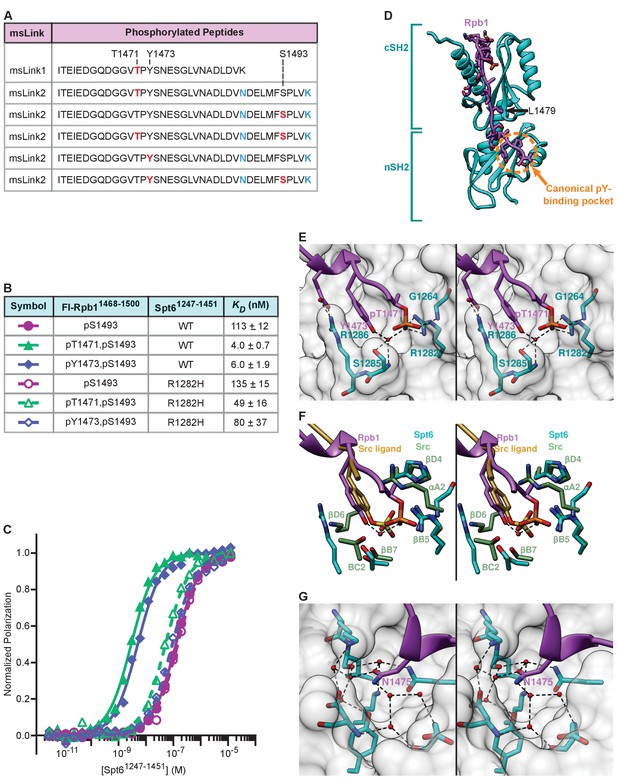
Binding of Rpb1 pT1471 and pY1473 to Spt6 tSH2, and mimicry of canonical SH2 interactions.
(A) Tryptic peptides from msLink1 and msLink2 observed by mass spectrometry, with phosphorylated sites indicated in red, and mutated residues in blue. (B) Spt6 tSH2 (wild type or R1282H) was titrated into Fl-Rpb11468-1500 pS1493, Fl-Rpb11468-1500 pT1471,pS1493, or Fl-Rpb11468-1500 pY1473,pS1493 peptides and polarization was detected. Average binding affinities calculated from at least 3 independent experiments and estimated standard deviations are shown. (C) Fluorescence polarization-derived binding curves from B. (D) Spt6 tSH2-Rpb11468-1500 pT1471,pS1493 linker structure. L1479 is the first ordered residue in the pS1493 structure of Figure 3, and is indicated with an arrow. (E) Stereo view of the Spt6 tSH2 pT1471-binding pocket. An ordered water molecule is shown as a red sphere. (F) Stereo view of the Spt6 tSH2 pT-binding pocket superimposed with the pY-binding pocket from the Src SH2 domain (PDP: 1SPS). (G) Stereo view of the Spt6 tSH2 specificity pocket. Ordered water molecules are shown as red spheres. See also Figure 4—figure supplement 1, Videos 1–5, and Figure 4—source data 1.
-
Figure 4—source data 1
Binding of fluorescently labeled, doubly phosphorylated Rpb1 linker peptides to Spt6 tSH2 proteins, related to Figure 4B and C.
- https://doi.org/10.7554/eLife.28723.011
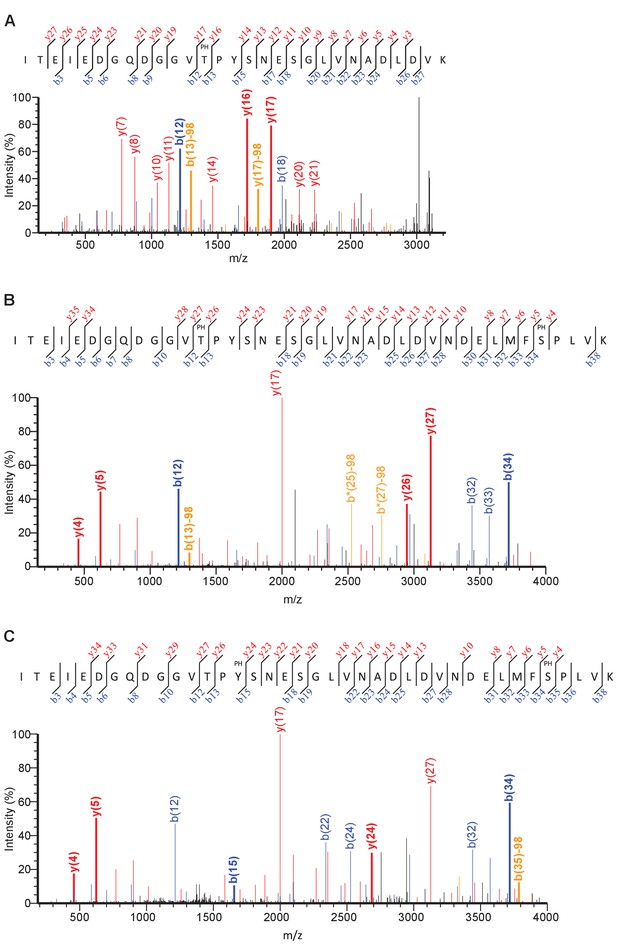
Mass spectra of phosphorylated Rpb1 linker peptides identified by mass spectrometry.
(A) A mass spectrum of a peptide obtained from msLink1 that indicates phosphorylation on T1471. Peaks representing b ions are colored blue. Peaks representing y ions are colored red. Orange peaks represent ions that have a neutral loss of 98 Da, which corresponds to a phosphate (H3PO4). Peaks above 30% intensity are labeled. Peaks representing ions flanking the phosphorylation site are labeled and bold. (B) A mass spectrum of a peptide obtained from msLink2 that indicates phosphorylation on T1471 and S1493 in the same peptide. Colored as in A. (C) A mass spectrum of a peptide from msLink2 that indicates phosphorylation on Y1473 and S1493 in the same peptide. Colored as in A.
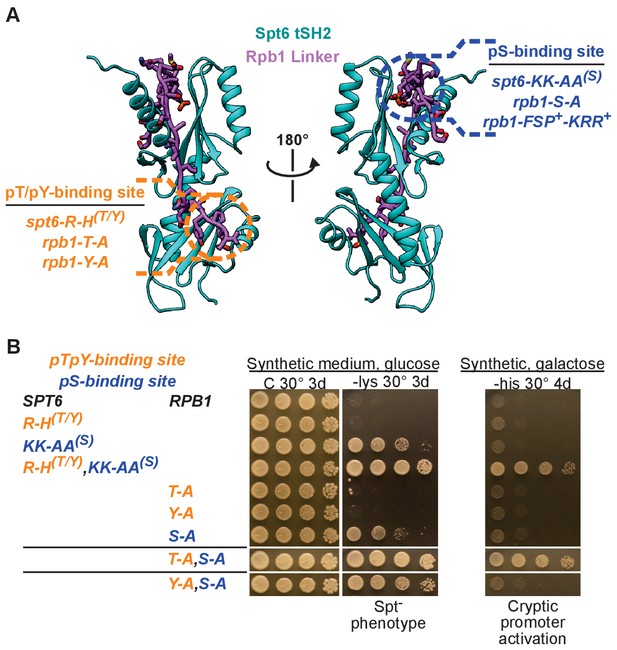
The Spt6-Rpb1 interaction is important for maintaining repressive chromatin.
(A) Structural map of Spt6 and Rpb1 mutations that were integrated at their native loci. For a description of the rpb1-FSP+-KRR+ allele and its phenotypes, refer to Figure 5—figure supplement 1. (B) Growth conditions/phenotype (above/below) for the mutations shown in A. Growth on media lacking lysine indicates the Spt⁻ phenotype and growth on galactose media lacking histidine indicates activation of the cryptic promoter within FLO8 with high levels of transcription. See also Figure 5—figure supplement 1 and Figure 5—figure supplement 2.
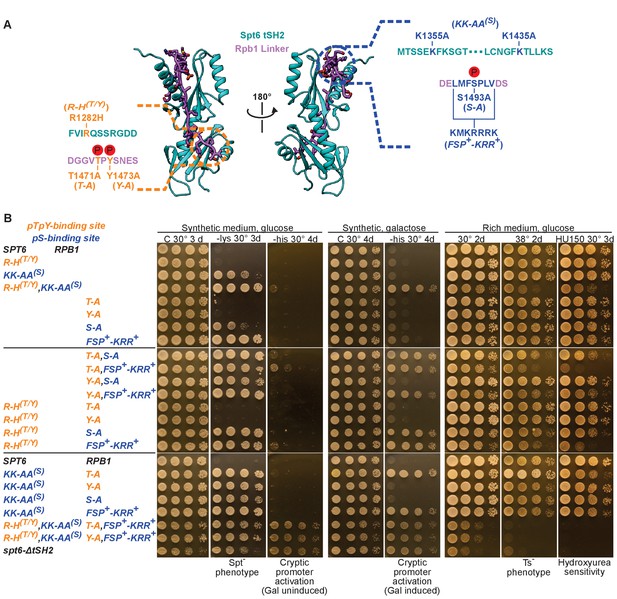
Complete genetic analysis of mutations that disrupt the Spt6-Rpb1 interaction.
(A) Structural map of Spt6 and Rpb1 mutations that were integrated at their native loci. The abbreviated strain names used in panel B are shown in parentheses. (B) Growth conditions/phenotype (above/below) for the mutations shown in A. Growth on media lacking lysine indicates the Spt⁻ phenotype and growth on media lacking histidine indicates activation of the cryptic promoter within FLO8. Galactose drives elevated transcription of the reporter, so growth on galactose -his indicates a weaker defect than activation on glucose -his. HU150 indicates addition of 150 mM hydroxyurea. Discussion of phenotypes. Mutating the pS1493 interface caused the Spt− phenotype with the severity of the phenotype correlating with the observed effect of the mutation on binding affinity in vitro. The spt6-KK-AA(S), rpb1-S-A double mutant had a stronger phenotype than either single mutant, suggesting that each mutation caused only a partial loss of chromatin maintenance. The additive effects of these alleles suggest that these residues may have additional functions outside of this specific interaction. A more severe Rpb1 mutation that replaces residues LMFSPLV1490-1496 with KMKRRRK1490-1496 (alias = rpb1-FSP+-KRR+) saturated the Spt− phenotype. Because greater phenotypic suppression is not possible, combining rpb1-FSP+-KRR+ with spt6-KK-AA(S) showed no further defect. The rpb1-FSP+-KRR+ strain also activated a cryptic promoter located within the FLO8 gene to a low level (growth on galactose). This defect was enhanced by adding the spt6-KK-AA(S) mutation, indicating that even rpb1-FSP+-KRR+ retains partial Spt6 binding activity or, as mentioned above, that these residues have additional functions that contribute to chromatin maintenance during transcription. Combining the ‘dual site’ spt6 and rpb1 alleles slightly enhanced the phenotypes caused by individual mutations, including activation of the cryptic promoter under low levels of transcription (growth on glucose), suggesting that the individual mutations do not completely disrupt the interaction. This is consistent with the large interface that extends beyond the specific residues mutated in these alleles. We also note the possibility of additional functions of these regions such as interactions of other factors with the Spt6 pockets or Rpb1 residues. These mutants displayed additional defects similar to those caused by a full tSH2 domain deletion, including slow growth at normal and elevated temperatures and hydroxyurea (HU) sensitivity, suggesting that these two interfaces represent essentially all of the biological functions of this domain.
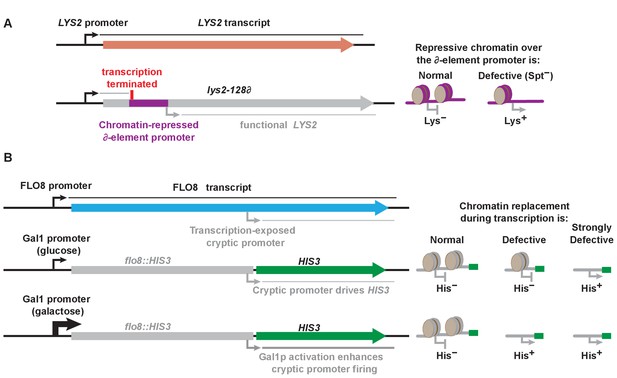
Schematic of Spt− and cryptic promoter activation reporters used to assay yeast phenotypes.
(A) The suppressor of Ty1 (Spt−) phenotype was scored by growth of strains carrying the lys2-128∂ allele on media lacking lysine. This allele arose by insertion of the transposon Ty1 into the 5’ end of the LYS2 gene, followed by resolution to a single ∂ element (Simchen et al., 1984). This terminates transcription from the normal LYS2 promoter and provides a weak promoter that makes a truncated but functional transcript. This promoter is repressed in normal cells but can be activated when chromatin formation is impaired. (B) The FLO8 gene contains a cryptic promoter (Kaplan et al., 2003). This promoter is inactive in normal chromatin but can be activated if nucleosomes are not rapidly redeposited after passage of an RNAPII complex (Cheung et al., 2008). Integration of the HIS3 gene downstream of the cryptic promoter allows its activation to be detected as growth of cells on media lacking histidine. Substitution of the GAL1 promoter for the FLO8 promoter allows activation to be detected under more stringent conditions (cultures grown on glucose, leading to minimal transcription of the reporter and therefore low levels of exposure of the cryptic promoter) or less stringent conditions (cultures grown without glucose in the presence of galactose, driving high levels of transcription, greater disturbance of chromatin, and higher levels of exposure of the cryptic promoter), allowing greater dynamic range for determining the extent of a defect in restoring chromatin after transcription.
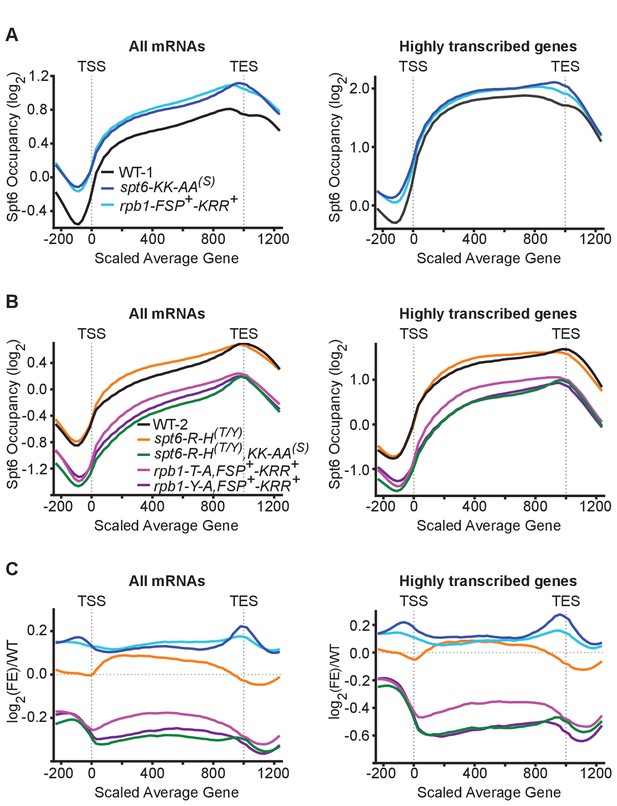
The Spt6-Rpb1 interaction is important for maintaining Spt6 occupancy at transcribed genes.
(A) The log2 enrichment ratios of Spt6 across all protein coding genes (left) and the top 20% of most frequently transcribed genes (right) in wild type yeast or yeast harboring spt6-KK-AA(S) or rpb1-FSP+-KRR+ mutant alleles (see Figure 5 and Figure 5—figure supplement 1 for allele designations). −200 to 0 and 1000–1200 are in base pairs, 0–1000 is normalized to the average transcript size. (B) As in A except mutant alleles include spt6-R-H(T/Y), spt6-R-H(T/Y),KK-AA(S), rpb1-T-A,FSP+-KRR+, or rpb1-Y-A,FSP+-KRR+. (C) The log2 enrichment ratios of Spt6 in panels A and B normalized to the respective wild type occupancy. See also Figure 6—figure supplement 1 and Figure 6—figure supplement 2.
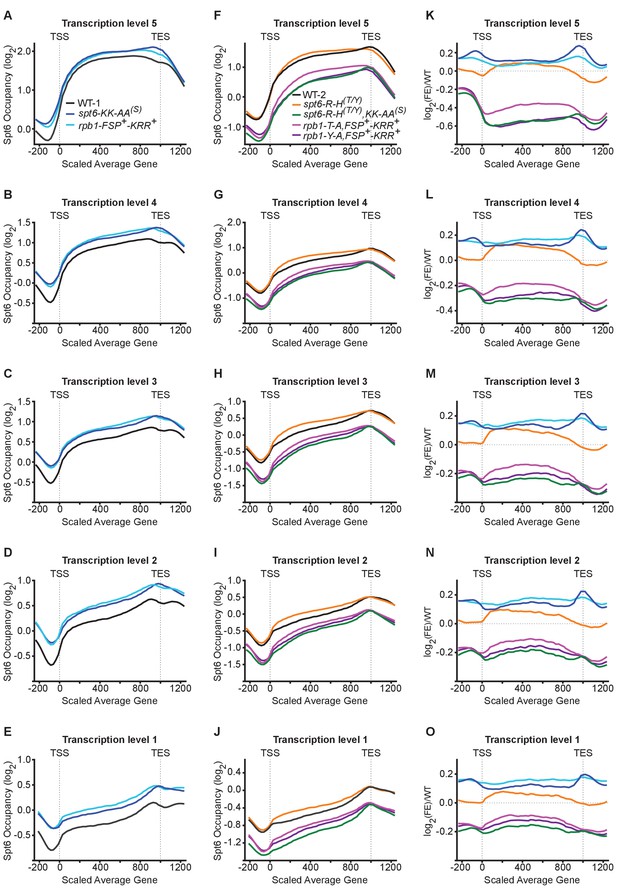
Mutations that impair the Spt6-Rpb1 linker interface reduce Spt6 occupancy at highly transcribed genes.
(A–E) The log2 enrichment ratios of Spt6 across genes split into quintiles based on transcription frequency in wild type yeast or yeast harboring spt6-KK-AA(S) or rpb1-FSP+-KRR+ mutant alleles. Transcription level 1 refers to the least frequently transcribed genes and Transcription level 5 refers to the most frequently transcribed genes. Panel A is taken from Figure 6A and is included here for clarity. −200 to 0 and 1000–1200 are in base pairs, 0–1000 is normalized to the size of each transcript. (F–J) As in A except mutant alleles include spt6-R-H(T/Y), spt6-R-H(T/Y),KK-AA(S), rpb1-T-A,FSP+-KRR+, or rpb1-Y-A,FSP+-KRR+. Panel F is taken from Figure 6B and is included here for clarity. (K–O) The log2 enrichment ratios of Spt6 in panels A-J normalized to the respective wild type occupancy at genes in the indicated quintile. Panel K is taken from Figure 6C and is included here for clarity.
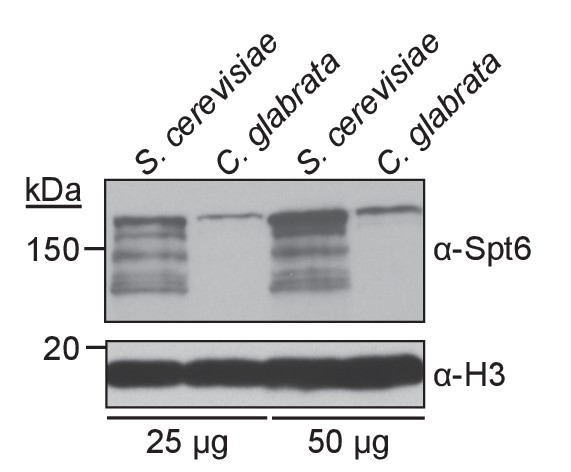
Antibody raised against S. cerevisiae Spt6 cross-reacts with C. glabrata Spt6.
Whole cell lysates (25 or 50 μg total protein) prepared from S. cerevisiae and C. glabrata cells (experimental and reference genomes, respectively) were subjected to western blotting and detected with the antiserum used for the ChIP-Seq experiments. Histone H3 serves as an additional loading control.
Videos
Overview of the Spt6 tSH2-Rpb1 linker complex.
Related to Figure 4.
Unbiased Fo-Fc omit map around Rpb1.
Related to Figure 4. The electron density map was phased on a tSH2 model with random 0–0.1 Å shifts applied to every atom and was refined in the absence of the Rpb1 peptide. The base contour level for the Fo-Fc map is 2.5 x RMSD with areas of weaker electron density contoured to 1.8 x RMSD. Density within 2 Å of an atom in the Rpb1 peptide is shown.
Interaction network of the Spt6 tSH2-Rpb1 linker interface.
Related to Figure 4. Water molecules that bridge the Spt6 tSH2-Rpb1 linker interaction are shown as red spheres. Hydrogen bonds identified using UCSF Chimera FindHBond (Pettersen et al., 2004) without relaxed constraints are depicted as dashed yellow lines.
Comparison of the Spt6 tSH2-Rpb1 pT1471/Y1473 interaction with other SH2-pY interactions.
Related to Figure 4. Structures of SH2 domains and their pY ligands (Src, PDB: 1SPS; BRDG1, PDB: 3MAZ; CBL-N, PDB: 2CBL; Gads, PDB: 5GJH; Syp, PDB: 1AYA) were aligned on the Spt6 tSH2-Rpb1 linker structure. Alignments were performed on the Cα atoms of conserved residues in the pY-binding pocket (αA2, βB5, βB7, and βD4; the CBL-N alignment also included the Cα atoms of Spt6 R1286 and CBL R299).
Structural model of the Spt6 tSH2 interaction with Rpb1 pY1473.
Related to Figure 4. A phosphate with appropriate geometry was modeled onto the hydroxyl group of Rpb1 Y1473 without any other structural adjustment. Hydrogen bonds are displayed as dashed lines (precise geometric criteria (Pettersen et al., 2004), yellow; within 0.4 Å and 20° of precise geometric criteria, purple).
Tables
Crystallographic Data Collection and Refinement
https://doi.org/10.7554/eLife.28723.008| Data collection | tSH2-Rpb1pS1493 | tSH2-Rpb1pT1471,pS1493 |
|---|---|---|
| Beamline | Home source | Home source |
| Wavelength | 1.54178 | 1.54178 |
| Space group | C2221 | C2221 |
| Unit cell dimensions: | ||
| a, b, c (Å) | 43.01, 103.54, 115.49 | 42.45, 102.10, 115.74 |
| Resolution (Å) | 40.00–2.20 (2.28–2.20) | 40.00–1.80 (1.86–1.80) |
| I/σI | 25 (2) | 8 (1) |
| CC1/2 | 0.977 | 0.641 |
| Completeness (%) | 89.81 (60) | 99.67 (96) |
| Redundancy | 24.7 (17.0) | 7.9 (5.2) |
| Rmeas | 0.056 (0.458) | 0.099 (1.238) |
| Rpim | 0.011 (0.106) | 0.035 (0.518) |
| Refinement | ||
| Resolution (Å) | 27.64–2.20 (2.28–2.20) | 32.45–1.80 (1.85–1.80) |
| Number of reflections | 12,178 | 23,722 |
| Rwork/Rfree (%) | 17.7/24.3 | 16.2/19.8 |
| Number of protein atoms | 1859 | 1954 |
| Number of solvent atoms | 83 | 171 |
| RMSD bond lengths (Å)/angles (°) | 0.007/0.878 | 0.009/1.013 |
| ϕ/ψ most favored/allowed (%) | 96.33/3.67 | 97.75/1.80 |
-
Values in parentheses refer to the high-resolution shell. Refinement statistics were determined by PHENIX and MolProbity.
Additional files
-
Supplementary file 1
Plasmids.
- https://doi.org/10.7554/eLife.28723.023
-
Supplementary file 2
Yeast strains.
- https://doi.org/10.7554/eLife.28723.024
-
Transparent reporting form
- https://doi.org/10.7554/eLife.28723.025





