Asymmetry of movements in CFTR's two ATP sites during pore opening serves their distinct functions
Figures
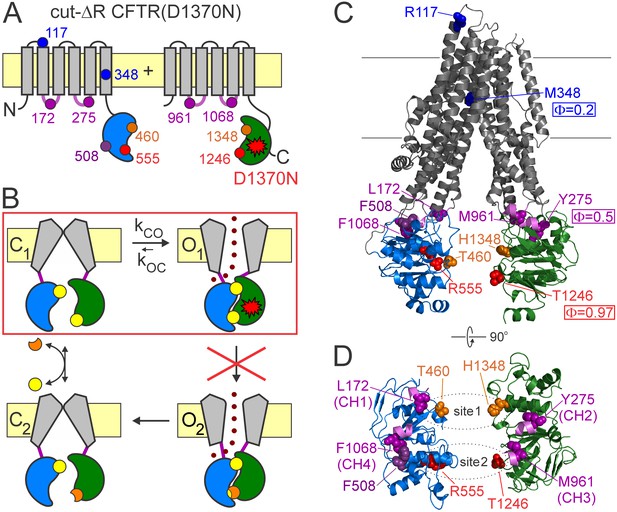
CFTR domain topology, gating mechanism, and localization of target positions.
(A) Domain topology of the CFTR cut-ΔR(D1370N) background construct. TMDs, gray; NBD1, blue; NBD2, green; intracellular loops containing coupling helices (CH1-4, from left to right), violet; target positions, colored circles. Red star denotes mutation D1370N. (B) Cartoon gating cycle of WT CFTR; and two-state equilibrium gating (red box) in saturating ATP of the background construct in which the D1370N mutation (red star) disrupts ATP hydrolysis (red cross). ATP, yellow circles; ADP, orange crescents, upper ATP binding site represents non-catalytic site 1, lower site represents catalytic site 2. (C–D) Target positions highlighted in colored spacefill on the cryoEM structure of dephosphorylated closed human CFTR (PDBID: 5UAK). (C) Full-length structure with all target positions shown, color coding as in A. Φ values for positions 1246, 348, and 275 are taken from (Sorum et al., 2015). (D) Only NBDs and coupling helices (CH1-4) shown, viewed from an angle orthogonal to the membrane. Dotted ellipses identify ATP sites 1 and 2.
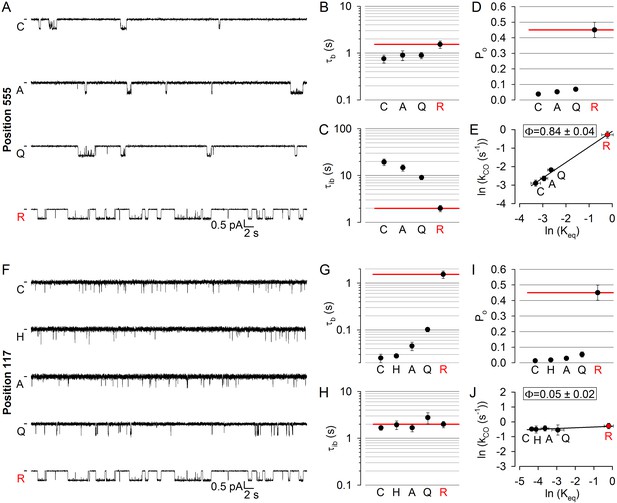
Longitudinal Φ value gradient extends from catalytic site 2 to the extracellular surface.
(A, F) Inward unitary currents of the cut-ΔR(D1370N) CFTR background construct, and of channels bearing mutations at either position 555 (A) or position 117 (F), in the same background. Letters to the left of the traces indicate the amino acid present in the target position; the native residue is marked by red. Currents were recorded in symmetrical 140 mM Cl-, at −80 mV in (A) but at −100 mV in (F); dashes on the left mark zero-current level. (B–D) and (G–I), Mean burst (B, G, τb) and interburst (C, H, τib) durations and open probabilities (D, I, Po) of the constructs in A and F, respectively. Red horizontal lines highlight the respective control values of the background construct which is identified by the red letter representing the native target residue. All data are shown as mean ± SEM (n = 6 for data in B–D, n = 5–7 for data in G–I). (E, J) Brønsted plots for position 555 (E) and 117 (J). Red symbol and letter identifies the background construct. Solid lines are linear regression fits with slope Φ indicated.
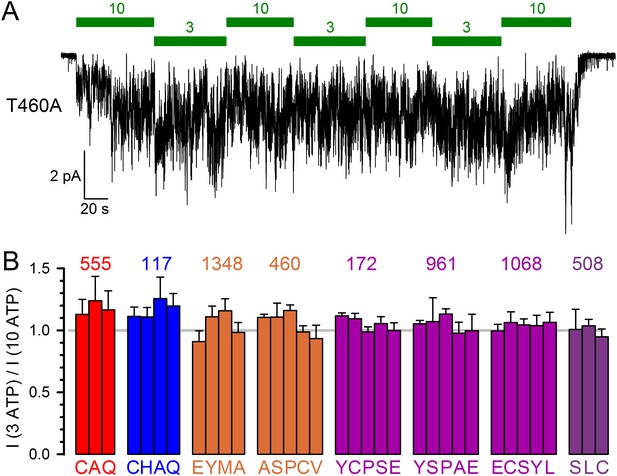
All tested constructs are saturated by 10 mM ATP.
(A) Representative macroscopic inward current of CFTR channels bearing mutation T460A in the cut-ΔR(D1370N) background, recorded in an inside-out patch at −80 mV in response to alternating exposures to 3 and 10 mM MgATP (green bars). (B) Fractional macroscopic currents in 3 mM ATP, normalized to the mean of the currents in the presence of bracketing applications of 10 mM ATP, for cut-ΔR(D1370N) channels harbouring the indicated amino-acid substitutions (letters) at the respective target positions (numbers above groups of bars). Bars are grouped by target position, color coding as in Figures 1 and 6. All data are shown as mean ± SEM (n = 3–11). For the background construct I3 ATP / I10 ATP=1.00 ± 0.07 (n = 6) (cf., Sorum et al., 2015).
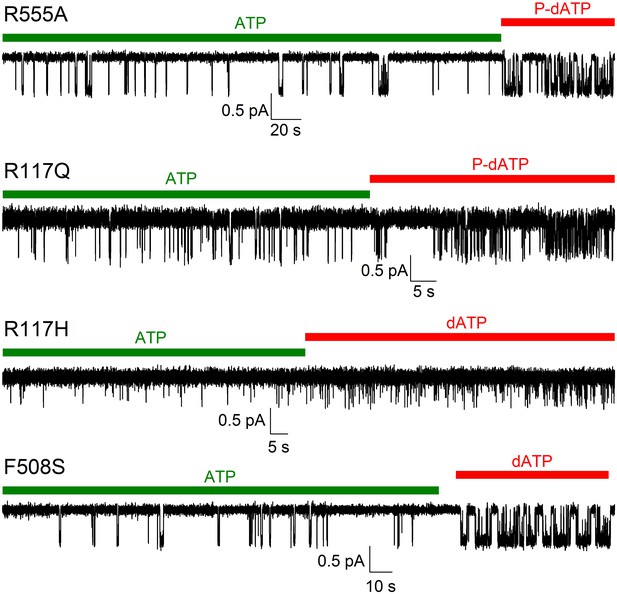
Stimulation of open probability by 2'-deoxy-ATP (dATP) or N6-(2-phenylethyl)-dATP (P-dATP) facilitates counting channels for low-Po mutants.
Inward currents of single cut-ΔR(D1370N) channels harbouring the indicated mutations. After several minutes of low-Po activity in 10 mM MgATP (green bars), exposure to 5 mM Mg-dATP or 25–50 μM Mg-P-dATP (red bars) robustly stimulated Po. A statistical test based on comparison of the observed opening rate and cumulative open time (Csanády et al., 2000) allowed high-confidence exclusion of the presence of a second active channel for each of the four patches shown (p<10−9, p<10−11, p=0.015, and p<10−14, respectively).
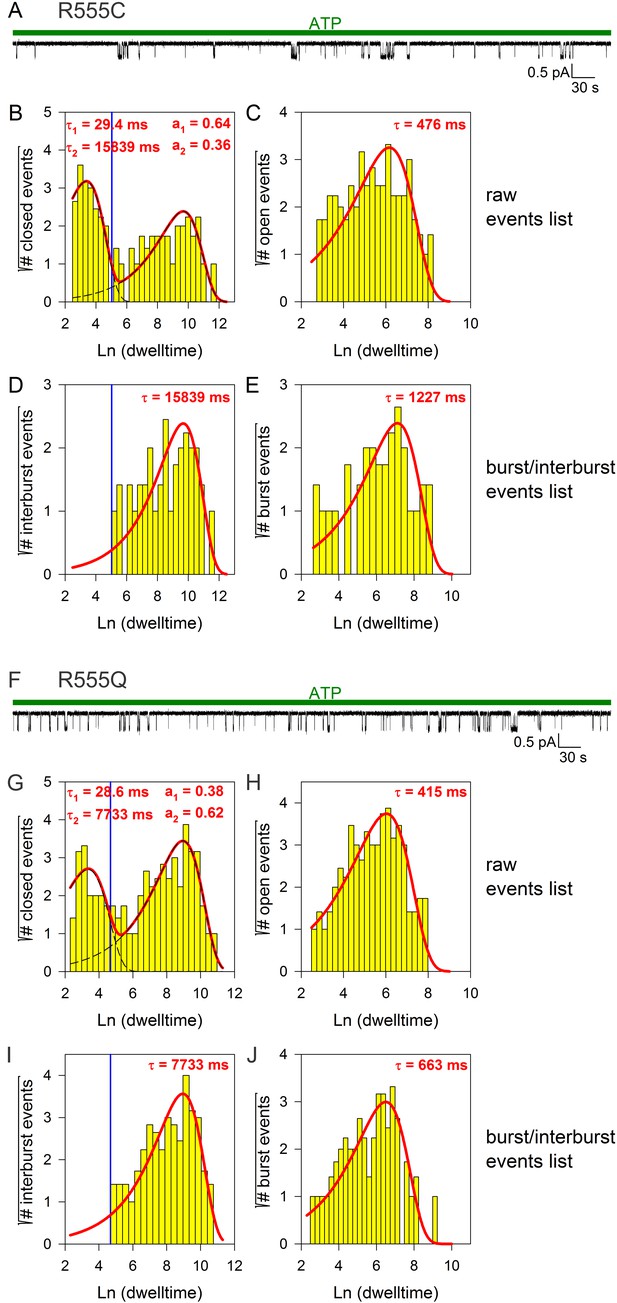
Burst analysis of single-channel recordings for position 555 mutants.
(A, F) Inward currents of single cut-ΔR(D1370N) channels bearing mutation R555C (A) or R555Q (F) in 10 mM MgATP (green bars). (B–C, G–H), Closed (B, G) and open (C, H) dwell-time histograms (Sigworth and Sine, 1987) constructed from the events list of the records in (A, F); lower binning limit is 12 ms. Solid red lines are maximum likelihood fits of the dwell-times by a double- and a single-exponential distribution, respectively, with time constants (τ(1, 2)) and fractional amplitudes (a1, a2) printed in the panels. Dotted black lines in panels (B, G) depict the two components of the fit; vertical blue line marks the burst delimiter (tcrit) calculated using the method of (Magleby and Pallotta, 1983): this method ensures unbiased estimation of mean burst and interburst durations by equalizing the probabilities of misassigning interburst events as flickery closures and vice versa. (D–E), (I-J) Dwell-time histograms of reconstructed interburst (D, I) and burst (E, J) events following suppression of closed events shorter than tcrit. Solid red lines in D, I replot the longer component of the double-exponential fits in (B, G): note, some flickery closures remain uncancelled (events above red fit line, right of blue line), but an equal number of interburst closures is cancelled (lack of events below red fit line, left of blue line). Solid red lines in (E, J) are maximum likelihood fits of the burst durations by single-exponential distributions (τ, time constant).
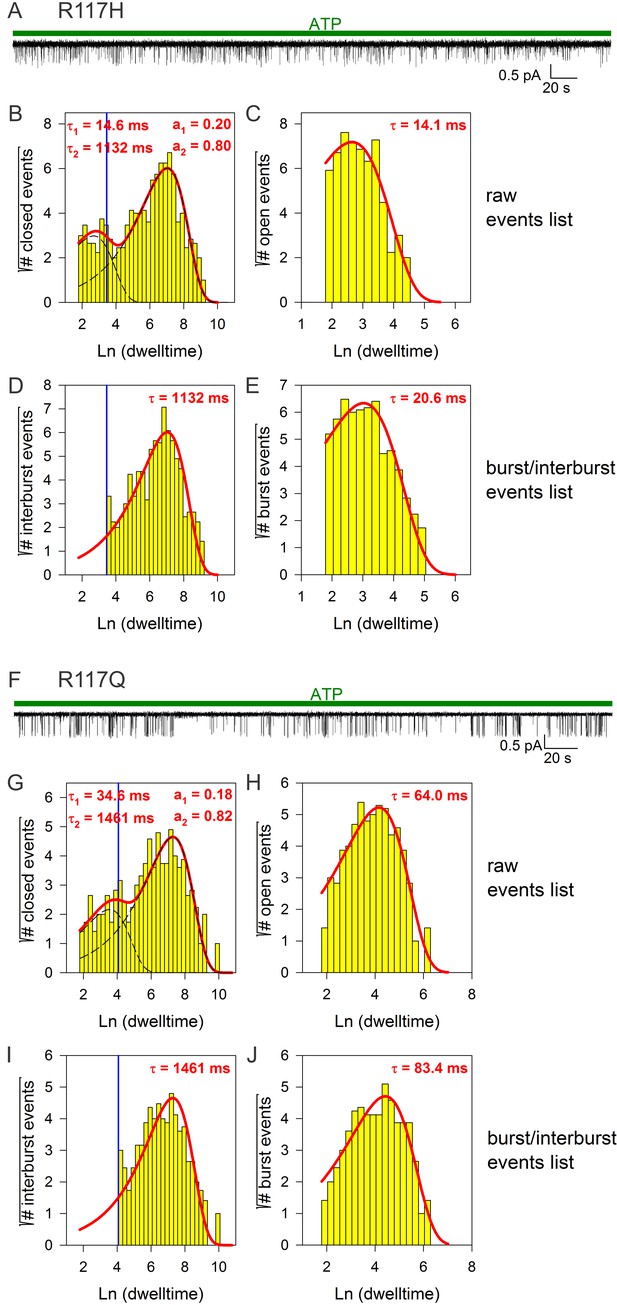
Burst analysis of single-channel recordings for position 117 mutants.
(A, F) Inward currents of single cut-ΔR(D1370N) channels bearing mutation R117H (A) or R117Q (F) in 10 mM MgATP (green bars). (B–C, G–H) Closed (B, G) and open (C, H) dwell-time histograms (Sigworth and Sine, 1987) constructed from the events list of the records in (A, F); lower binning limit is 6 ms. Solid red lines are maximum likelihood fits of the dwell-times by a double- and a single-exponential distribution, respectively, with time constants (τ(1, 2)) and fractional amplitudes (a1, a2) printed in the panels. Dotted black lines in panels (B, G) depict the two components of the fit; vertical blue line marks the burst delimiter (tcrit) calculated using the method of (Magleby and Pallotta, 1983). (D–E, I–J) Dwell-time histograms of reconstructed interburst (D, I) and burst (E, J) events following suppression of closed events shorter than tcrit. Solid red lines in (D, I) replot the longer component of the double-exponential fits in (B, G). Solid red lines in (E, J) are maximum likelihood fits of the burst durations by single-exponential distributions (τ, time constant).
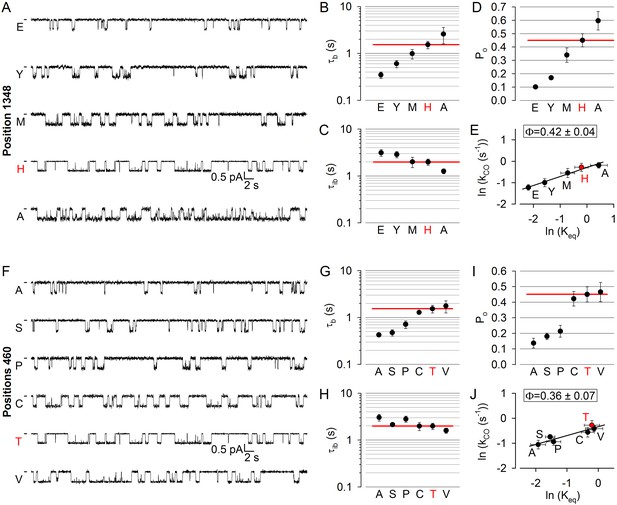
Delayed movement in non-catalytic site 1 during pore opening.
(A, F) Inward unitary currents of the cut-ΔR(D1370N) CFTR background construct, and of channels bearing mutations at either position 1348 (A) or position 460 (F), in the same background. Letters to the left of the traces indicate the amino acid present in the target position; the native residue is marked by red. Currents were recorded at −80 mV, in symmetrical 140 mM Cl-; dashes on the left mark zero-current level. (B–D) and (G–I), Mean burst (B, G, τb) and interburst (C, H, τib) durations and open probabilities (D, I, Po) of the constructs in A and F, respectively. Red horizontal lines highlight the respective control values of the background construct which is identified by the red letter representing the native target residue. All data are shown as mean ± SEM (n = 5–10 for data in B-D, n = 5–10 for data in G–I). (E, J) Brønsted plots for position 1348 (E) and 460 (J). Red symbol and letter identifies the background construct. Solid lines are linear regression fits with slope Φ indicated.
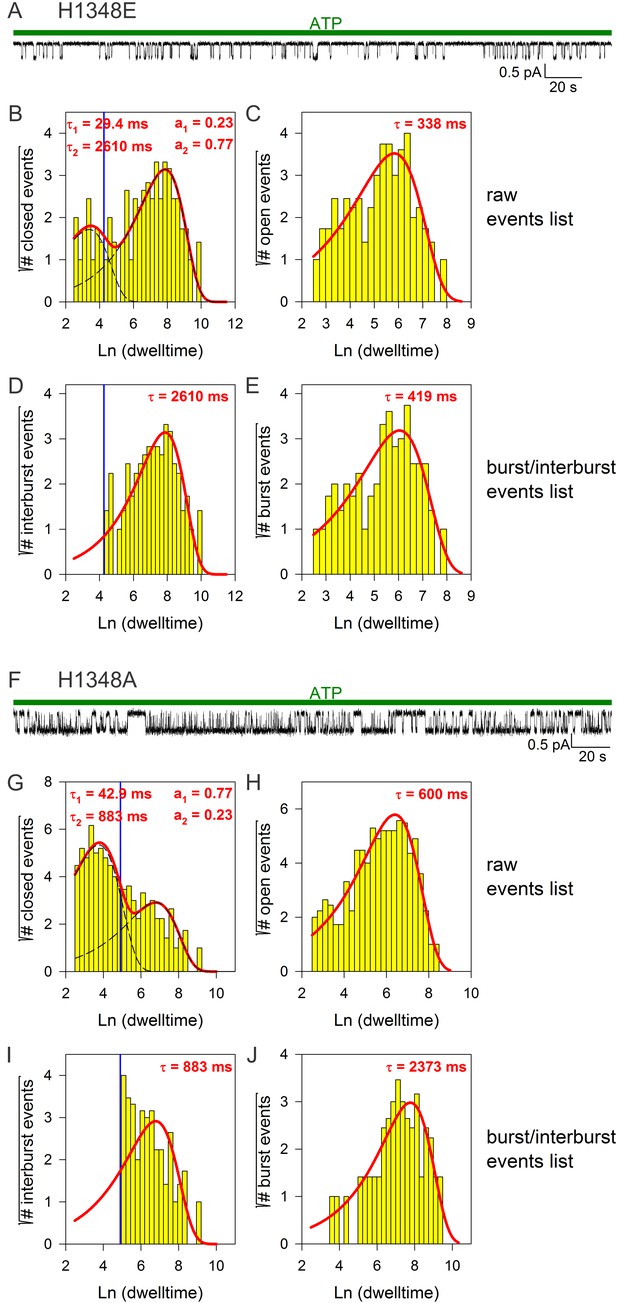
Burst analysis of single-channel recordings for position 1348 mutants.
(A, F) Inward currents of single cut-ΔR(D1370N) channels bearing mutation H1348E (A) or H1348A (F) in 10 mM MgATP (green bars). (B–C, G–H) Closed (B, G) and open (C, H) dwell-time histograms (Sigworth and Sine, 1987) constructed from the events list of the records in (A, F); lower binning limit is 12 ms. Solid red lines are maximum likelihood fits of the dwell-times by a double- and a single-exponential distribution, respectively, with time constants (τ(1, 2)) and fractional amplitudes (a1, a2) printed in the panels. Dotted black lines in panels (B, G) depict the two components of the fit; vertical blue line marks the burst delimiter (tcrit) calculated using the method of (Magleby and Pallotta, 1983). (D–E, I–J), Dwell-time histograms of reconstructed interburst (D, I) and burst (E, J) events following suppression of closed events shorter than tcrit. Solid red lines in (D, I) replot the longer component of the double-exponential fits in (B, G). Solid red lines in (E, J) are maximum likelihood fits of the burst durations by single-exponential distributions (τ, time constant).
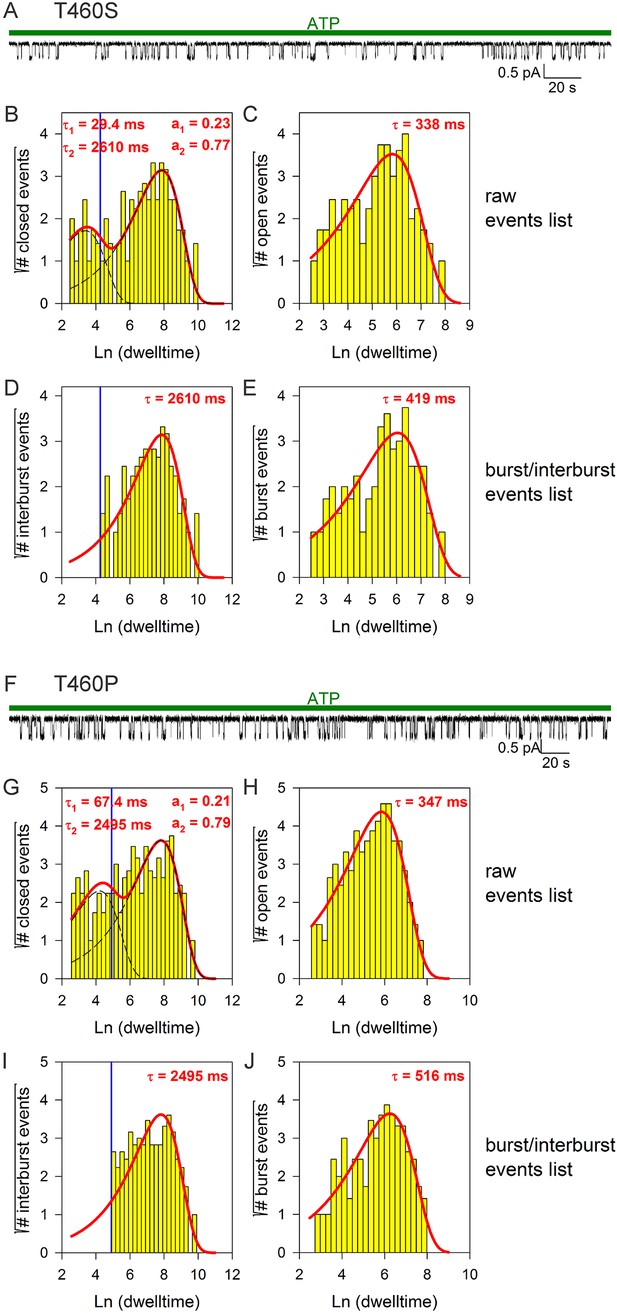
Burst analysis of single-channel recordings for position 460 mutants.
(A, F) Inward currents of single cut-ΔR(D1370N) channels bearing mutation T460S (A) or T460P (F) in 10 mM MgATP (green bars). (B–C, G–H) Closed (B, G) and open (C, H) dwell-time histograms (Sigworth and Sine, 1987) constructed from the events list of the records in (A, F); lower binning limit is 12 ms. Solid red lines are maximum likelihood fits of the dwell-times by a double- and a single-exponential distribution, respectively, with time constants (τ(1, 2)) and fractional amplitudes (a1, a2) printed in the panels. Dotted black lines in panels (B, G) depict the two components of the fit; vertical blue line marks the burst delimiter (tcrit) calculated using the method of (Magleby and Pallotta, 1983). (D–E, I–J) Dwell-time histograms of reconstructed interburst (D, I) and burst (E, J) events following suppression of closed events shorter than tcrit. Solid red lines in (D, I) replot the longer component of the double-exponential fits in (B, G). Solid red lines in (E, J) are maximum likelihood fits of the burst durations by single-exponential distributions (τ, time constant).
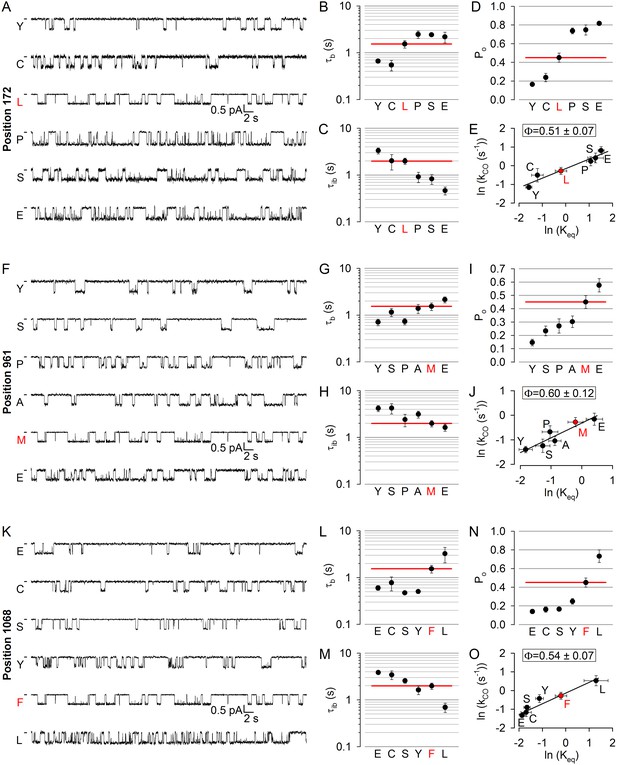
No asymmetry in the timing of motions can be detected at the level of the coupling helices.
(A, F, K) Inward unitary currents of the cut-ΔR(D1370N) CFTR background construct, and of channels bearing mutations at position 172 (A), 961 (F), or 1068 (K), in the same background. Letters to the left of the traces indicate the amino acid present in the target position; the native residue is marked by red. Currents were recorded at −80 mV, in symmetrical 140 mM Cl-; dashes on the left mark zero-current level. (B–D, G-I) and (L–N), Mean burst (B, G, L, τb) and interburst (C, H, M, τib) durations and open probabilities (D, I, N, Po) of the constructs in A, F, and K, respectively. Red horizontal lines highlight the respective control values of the background construct which is identified by the red letter representing the native target residue. All data are shown as mean ± SEM (n = 3–8 for data in (B–D), n = 4–15 for data in G-I, n = 4–7 for data in L-N). (E, J, O) Brønsted plots for position 172 (E), 961 (J), and 1068 (O). Red symbol and letter identifies the background construct. Solid lines are linear regression fits with slope Φ indicated.
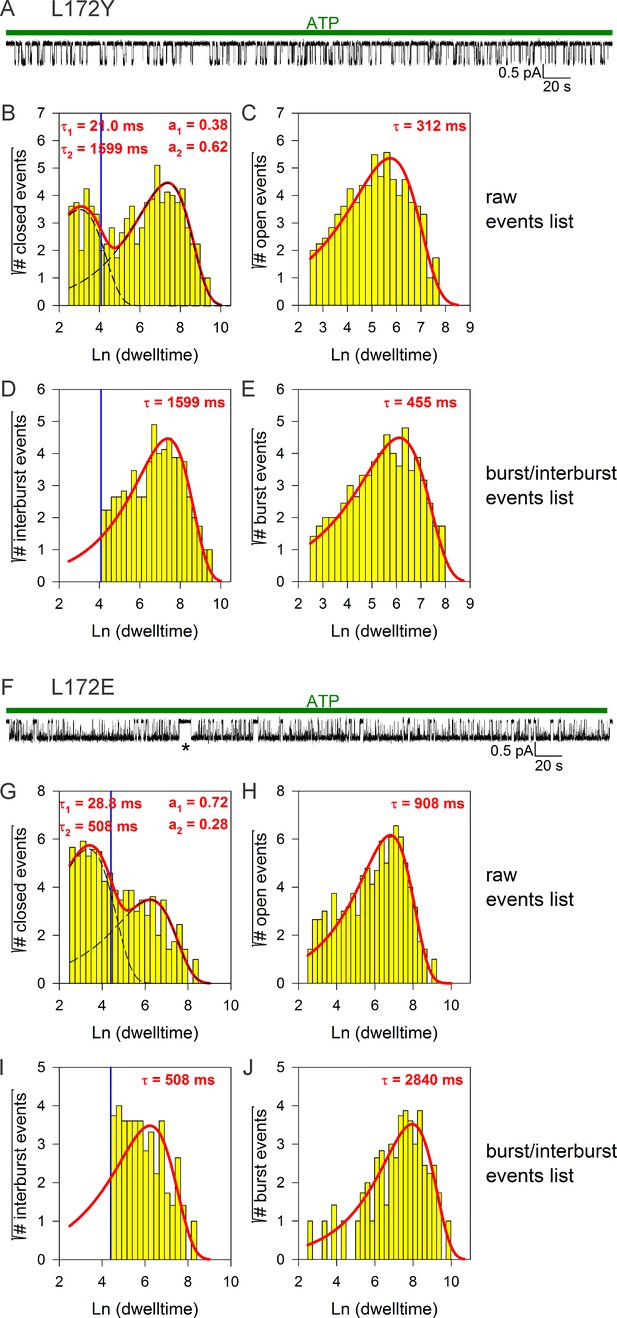
Burst analysis of single-channel recordings for position 172 mutants.
(A, F) Inward currents of single cut-ΔR(D1370N) channels bearing mutation L172Y (A) or L172E (F) in 10 mM MgATP (green bars). (B–C, G–H), Closed (B, G) and open (C, H) dwell-time histograms (Sigworth and Sine, 1987) constructed from the events list of the records in (A, F); lower binning limit is 12 ms. Solid red lines are maximum likelihood fits of the dwell-times by a double- and a single-exponential distribution, respectively, with time constants (τ(1, 2)) and fractional amplitudes (a1, a2) printed in the panels. Dotted black lines in panels (B, G) depict the two components of the fit; vertical blue line marks the burst delimiter (tcrit) calculated using the method of (Magleby and Pallotta, 1983). (D–E, I–J) Dwell-time histograms of reconstructed interburst (D, I) and burst (E, J) events following suppression of closed events shorter than tcrit. Solid red lines in (D, I) replot the longer component of the double-exponential fits in (B, G). Solid red lines in (E, J) are maximum likelihood fits of the burst durations by single-exponential distributions (τ, time constant). A single long interburst event in F marked by an asterisk was omitted from the analysis.
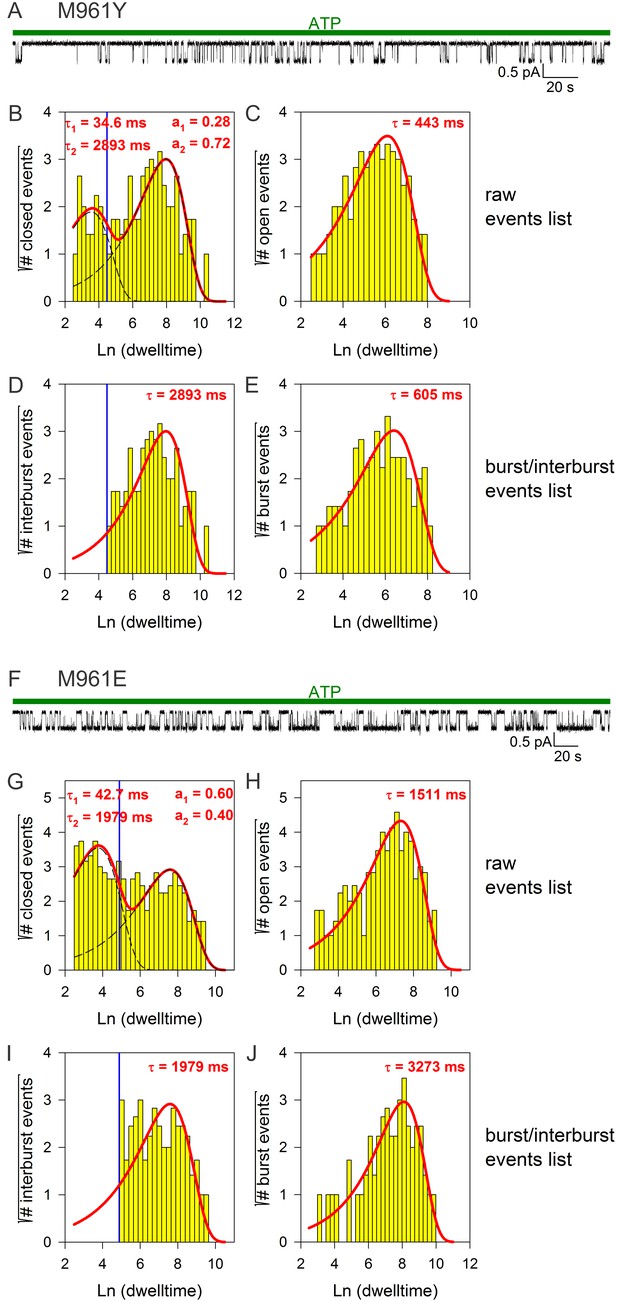
Burst analysis of single-channel recordings for position 961 mutants.
(A, F) Inward currents of single cut-ΔR(D1370N) channels bearing mutation M961Y (A) or M961E (F) in 10 mM MgATP (green bars). (B–C, G–H) Closed (B, G) and open (C, H) dwell-time histograms (Sigworth and Sine, 1987) constructed from the events list of the records in (A, F); lower binning limit is 12 ms. Solid red lines are maximum likelihood fits of the dwell-times by a double- and a single-exponential distribution, respectively, with time constants (τ(1, 2)) and fractional amplitudes (a1, a2) printed in the panels. Dotted black lines in panels (B, G) depict the two components of the fit; vertical blue line marks the burst delimiter (tcrit) calculated using the method of (Magleby and Pallotta, 1983). (D–E, I–J), Dwell-time histograms of reconstructed interburst (D, I) and burst (E, J) events following suppression of closed events shorter than tcrit. Solid red lines in (D, I) replot the longer component of the double-exponential fits in (B, G). Solid red lines in (E, J) are maximum likelihood fits of the burst durations by single-exponential distributions (τ, time constant).
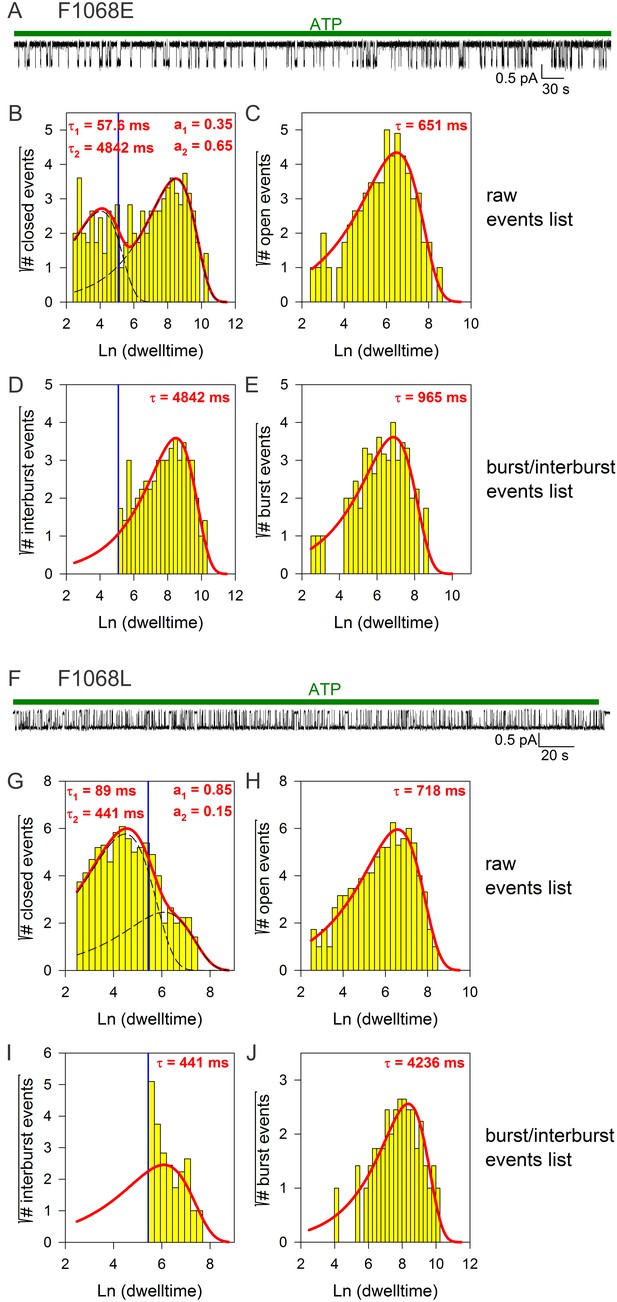
Burst analysis of single-channel recordings for position 1068 mutants.
(A, F) Inward currents of single cut-ΔR(D1370N) channels bearing mutation F1068E (A) or F1068L (F) in 10 mM MgATP (green bars). (B–C, G–H), Closed (B, G) and open (C, H) dwell-time histograms (Sigworth and Sine, 1987) constructed from the events list of the records in (A, F); lower binning limit is 12 ms. Solid red lines are maximum likelihood fits of the dwell-times by a double- and a single-exponential distribution, respectively, with time constants (τ(1, 2)) and fractional amplitudes (a1, a2) printed in the panels. Dotted black lines in panels (B, G) depict the two components of the fit; vertical blue line marks the burst delimiter (tcrit) calculated using the method of (Magleby and Pallotta, 1983). (D–E, I–J) Dwell-time histograms of reconstructed interburst (D, I) and burst (E, J) events following suppression of closed events shorter than tcrit. Solid red lines in (D, I) replot the longer component of the double-exponential fits in (B, G). Solid red lines in (E, J) are maximum likelihood fits of the burst durations by single-exponential distributions (τ, time constant).
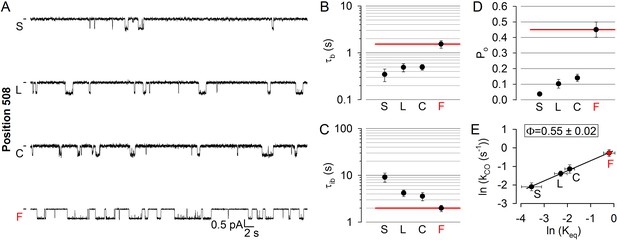
Disease hotspot position 508 is on the move in the transition state for opening.
(A), Inward unitary currents of the cut-ΔR(D1370N) CFTR background construct, and of channels bearing mutations at position 508, in the same background. Letters to the left of the traces indicate the amino acid present in the target position; the native residue is marked by red. Currents were recorded at −80 mV, in symmetrical 140 mM Cl-; dashes on the left mark zero-current level. (B–D) Mean burst (B, τb) and interburst (C, τib) durations and open probabilities (D, Po) of the constructs in (A). Red horizontal lines highlight the respective control values of the background construct which is identified by the red letter representing the native target residue. All data are shown as mean ± SEM (n = 5–7). E, Brønsted plot for position 508. Red symbol and letter identifies the background construct. Solid line is a linear regression fit with slope Φ indicated.
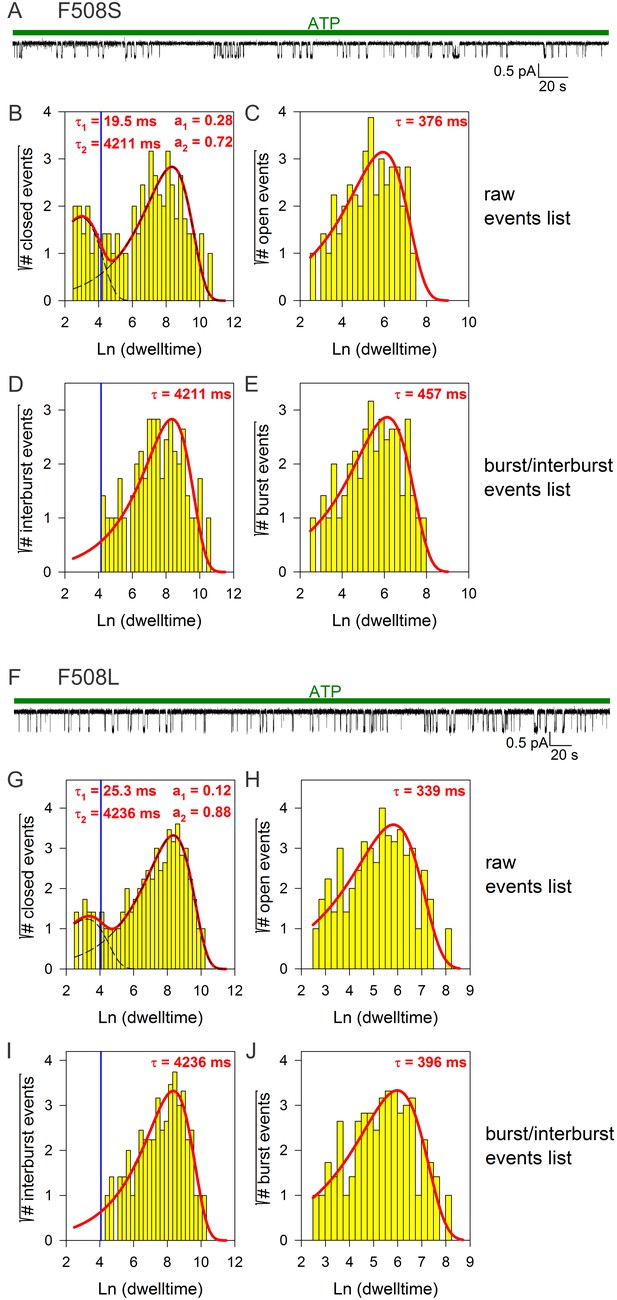
Burst analysis of single-channel recordings for position 508 mutants.
(A, F) Inward currents of single cut-ΔR(D1370N) channels bearing mutation F508S (A) or F508L (F) in 10 mM MgATP (green bars). (B–C, G–H) Closed (B, G) and open (C, H) dwell-time histograms (Sigworth and Sine, 1987) constructed from the events list of the records in (A, F); lower binning limit is 12 ms. Solid red lines are maximum likelihood fits of the dwell-times by a double- and a single-exponential distribution, respectively, with time constants (τ(1, 2)) and fractional amplitudes (a1, a2) printed in the panels. Dotted black lines in panels (B, G) depict the two components of the fit; vertical blue line marks the burst delimiter (tcrit) calculated using the method of (Magleby and Pallotta, 1983). (D–E, I–J) Dwell-time histograms of reconstructed interburst (D, I) and burst (E, J) events following suppression of closed events shorter than tcrit. Solid red lines in (D, I) replot the longer component of the double-exponential fits in (B, G). Solid red lines in (E, J) are maximum likelihood fits of the burst durations by single-exponential distributions (τ, time constant).
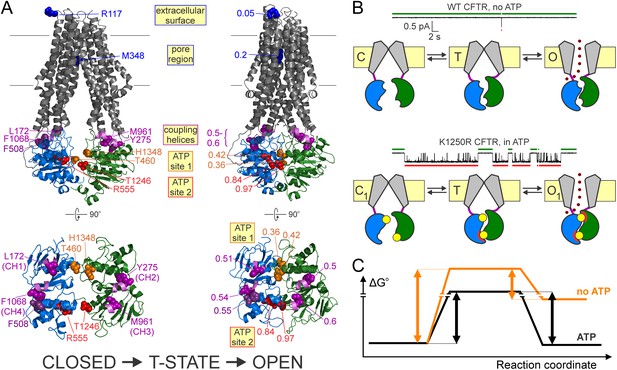
Distinct roles of the two ATP sites in promoting CFTR channel gating.
(A) Homology models (Corradi et al., 2015) of closed (left) and open (right) conformations of phosphorylated CFTR gating in ATP, based on the structures of inward-facing Tm287-288 (left) and outward-occluded McjD (right); target positions are highlighted in spacefill. Labels identify the native residues (left) and the Φ values of each position (right). For both conformations full structures (top) and NBDs with coupling helices only (bottom) are shown. (B) Current traces showing a single pore opening event in the absence of ATP in a patch containing hundreds of WT CFTR channels (top trace), and gating of a single K1250R CFTR channel in 5 mM ATP (bottom trace). Green and red bars identify closed interburst and open burst events, respectively. Cartoons below the traces illustrate the mechanism of gating of WT CFTR in the absence (top) and of K1250R CFTR in the presence (bottom) of ATP. Color coding as in Figure 1B, the upper ATP binding site represents non-catalytic site 1, the lower site represents catalytic site 2. Red semi-circles around ATP (yellow circles) represent tight bonding of the nucleotide with the opposing NBD surface. (C) Standard Gibbs energy profiles of a CFTR channel in the absence (orange profile) and presence (black profile) of bound ATP during progress from the closed state (left) through the transition state (center) to the open state (right). Vertical arrows illustrate the heights of the energetic barriers for the forward (opening) and backward (closing) steps under both conditions.
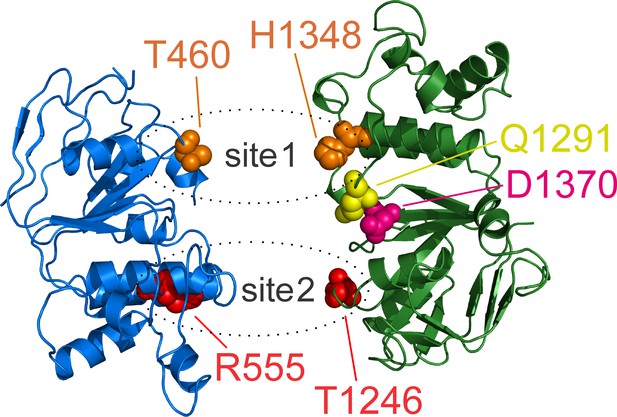
Location of aspartate 1370, mutated in our background construct.
Structures of NBD1 (blue) and NBD2 (green) of human dephosphorylated CFTR in a closed ATP-free state (PDBID: 5UAK) viewed top-down from the direction of the membrane. ATP sites 1 and 2 are highlighted by dotted ellipses, target positions in both sites are shown in spacefill. Aspartate 1370 (salmon spacefill) is located between the two ATP sites, at the boundary between the core and α-helical subdomains of NBD2, and interacts with Q-loop glutamine 1291 (yellow spacefill).
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.29013.019





