CDK9-dependent RNA polymerase II pausing controls transcription initiation
Figures
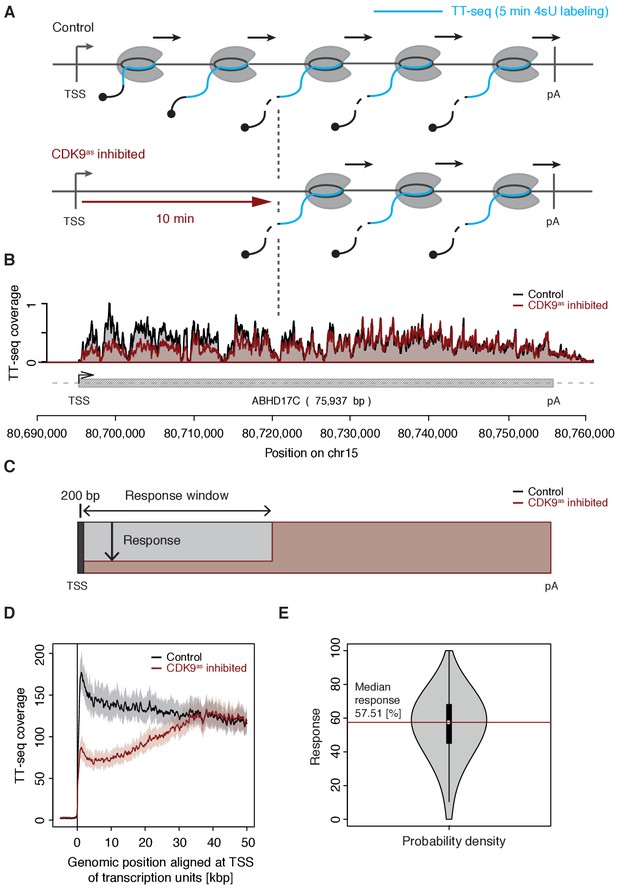
CDK9 inhibition decreases RNA synthesis in the 5’-region of genes.
(A) Experimental design. TT-seq was carried out with CDK9as cells after treatment with solvent DMSO (control) or 1-NA-PP1 (CDK9as inhibited). (B) TT-seq signal before (black) and after (red) CDK9 inhibition at the ABHD17C gene locus (75,937 [bp]) on chromosome 15. Two biological replicates were averaged. The grey box depicts the transcript body from the transcription start site (TSS, black arrow) to the polyA site (pA). (C) Schematic representation of changes in TT-seq signal showing the definition of the response window. Colors are as in (B). (D) Metagene analysis comparing the average TT-seq signal before and after CDK9 inhibition. The TT-seq coverage was averaged for 954 out of 2538 investigated TUs that exceed 50 [kbp] in length (Materials and methods). TUs were aligned with their TSS. Shaded areas around the average signal (solid lines) indicate confidential intervals (Materials and methods). (E) Violin plot showing the relative response to CDK9 inhibition for 2538 investigated TUs defined as 1 - (CDK9as inhibited/Control)100 for a window from the TSS to 10 [kbp] downstream, excluding the first 200 [bp] (C). A red line indicates the median response (58%).
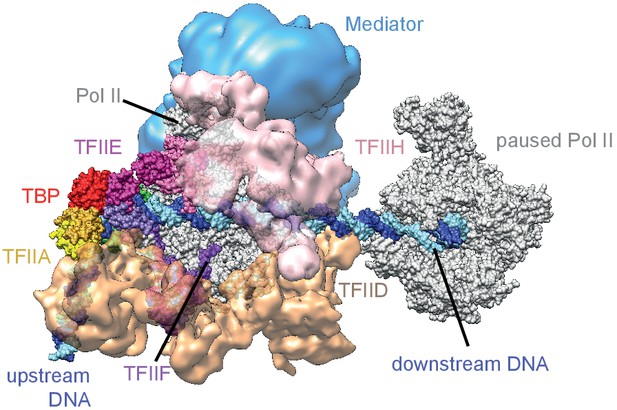
Model of a paused polymerase positioned up to around 50 bp downstream of the TSS.
Modeling shows that paused Pol II (silver, right) positioned 50 bp downstream of the transcription start site (TSS) allows for formation of the Pol II initiation complex (different colors, left). Shorter distances between the active sites of paused and initiating Pol II are predicted to lead to steric clashes. Modeling is based on the latest structural information (Mediator EMD-8307 [Robinson et al., 2016], TFIID EMD-3305 [Louder et al., 2016], TFIIH EMD EMD-3307 [He et al., 2016], closed complex PDB-code 5FZ5 [Plaschka et al., 2016], EC PDB-code 1WCM [Kettenberger et al., 2004]).
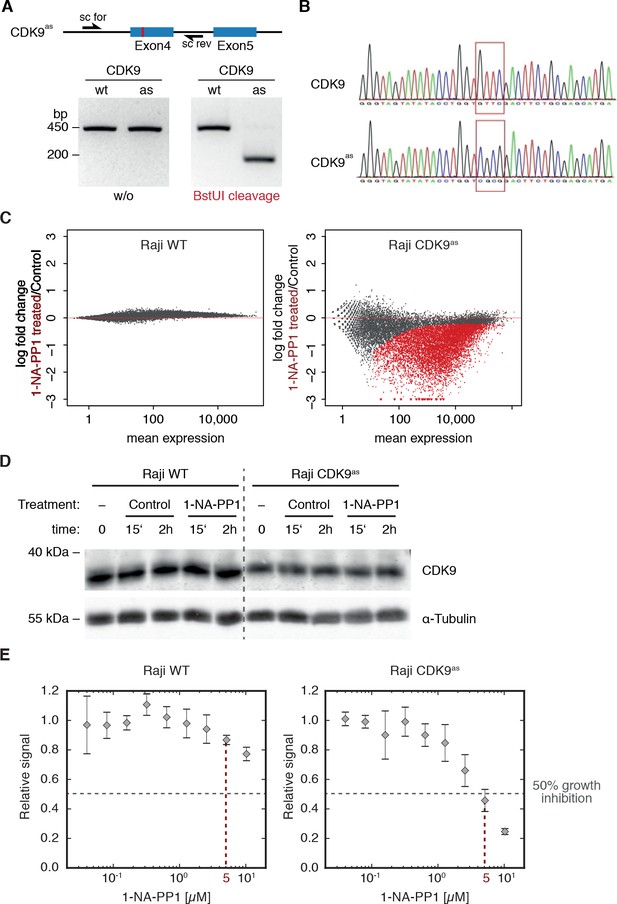
CRISPR-Cas9 directed engineering, cellular and biochemical characterization of CDK9as Raji B cell line.
(A) BstUI restriction enzyme recognition site used for screening is indicated in the HDR template sequence (highlighted in red). Agarose gels of screening PCRs followed by restriction digest with BstUI of wild type (wt) and CDK9as (as) Raji B cell line. (B) Validation of CDK9as Raji B cell line by sequencing. (C) Log fold change upon 1-NA-PP1 treatment (5 µM for 15 min) versus the normalized mean read count across replicates and conditions for wild type Raji B cells (left panel) and CDK9as Raji B cells (right panel). Significantly up- or downregulated TUs (adjusted p-value<0.01) are marked in red. (D) Wild type and CDK9as Raji B cells were treated with 10 µM of 1-NA-PP1 for 15 min or 2 hr. DMSO was used as control. Stable CDK9 protein levels were detected by Western blotting (Materials and methods). α-Tubulin was used as loading control. (E) Cell proliferation at increasing 1-NA-PP1 inhibitor concentrations (log scale) was determined using a colorimetric assay based on MTS metabolization (Materials and methods). Cell proliferation of CDK9as Raji B cells was dramatically reduced by >50% when 1-NA-PP1 concentrations of 5 µM (indicated with dashed red line) or higher were used, whereas wild type Raji B cells were largely unaffected. Error bars indicate the standard deviation (n = 4).
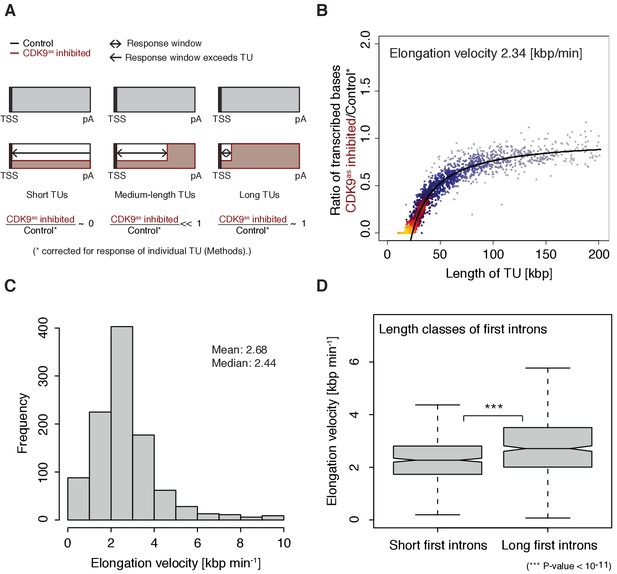
Pol II elongation velocity.
(A) Schematic representation of observed response window of TT-seq signal with CDK9as inhibitor (red) or control (black) for TUs of three different length classes (short TUs < 25 [kbp], medium-length TUs 25–50 [kbp] and long TUs > 100 [kbp]). (B) Scatter plot of the ratio of transcribed bases (CDK9as inhibited/control) (Materials and methods) against the length of the TUs in nucleotides [kbp] revealed that the schematic representation in (A) holds true for 2443 investigated TUs (Materials and methods). Modeling of the observed relation allows estimation of a robust average elongation velocity of 2.3 [kbp/min] (solid black line, Materials and methods). (C) Distribution of gene-wise elongation velocity depicted as a histogram (mean 2.7 [kbp/min], median 2.4 [kbp/min]). (D) Distributions of elongation velocity [kbp/min] depicted for 513 TUs with short first intron (<50% quantile, left) and 514 TUs with long first intron (>50% quantile, right).
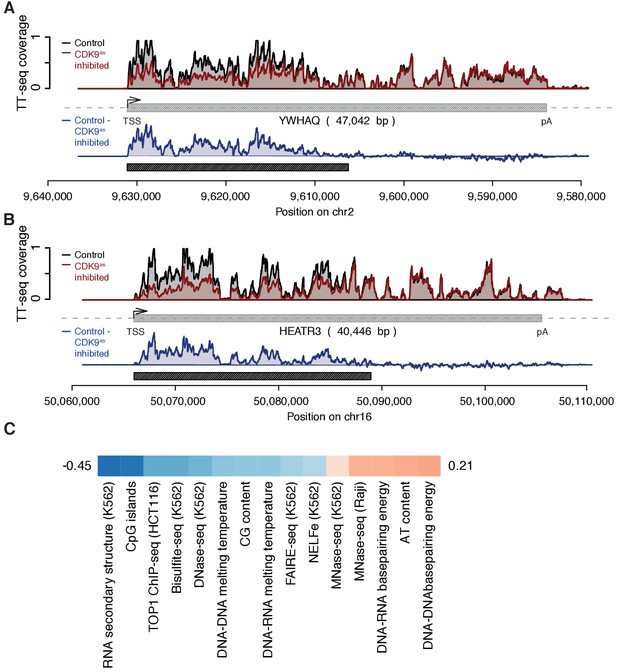
Example genome browser views of TT-seq signals in CDK9as cells with estimated response window and genomic features correlating with elongation velocity.
(A) YWHAQ gene locus (47,042 [bp]) on chromosome 2. The upper panel shows TT-seq signal with CDK9as inhibitor (red) and control (black). Grey box depicts transcript body from transcription start site (TSS, black arrow) to polyA site (pA). Lower panel shows the difference of TT-seq signal (control – CDK9as inhibited in blue). Black rectangle depicts the estimated response window according to elongation velocity estimate (Materials and methods). (B) HEATR3 gene locus (40,446 [bp]) on chromosome 16 depicted as in (A). (C) Color encoded Spearman correlation coefficients (color encoded, −0.45 in blue to 0.21 in red) of elongation velocity [kbp/min] against genomic features and measures of transcriptional context (Materials and methods, Supplementary file 1).
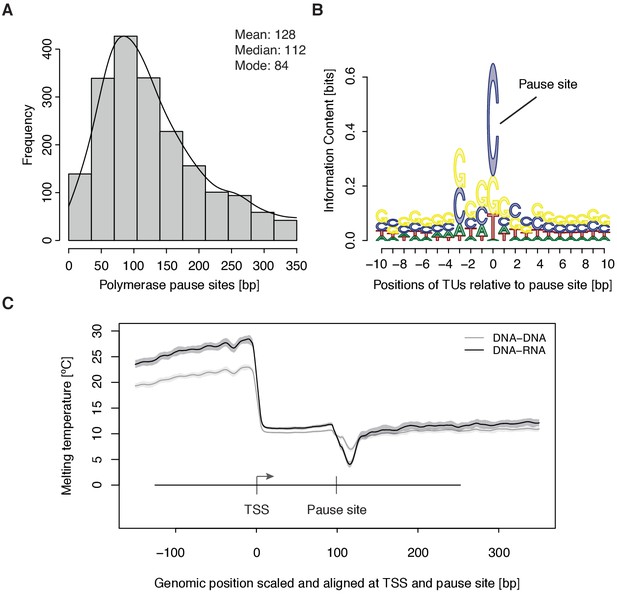
Distribution and sequence of promoter-proximal pause sites.
(A) Distribution of pause site distance from the TSS for 2135 investigated TUs depicted as a histogram (mean 128 [bp], median 112 [bp], mode 84 [bp]). Two biological replicates were averaged. (B) Position weight matrix (PWM) logo representation of bases at positions –10 to +10 [bp] around the pause site (position 0). (C) Mean melting temperature of the DNA-RNA and DNA-DNA hybrid aligned at the TSS and the pause site (signal between the TSS and the pause site is scaled to common length of 100 [bp]). Shaded areas around the average signal (solid lines) indicate confidence intervals.
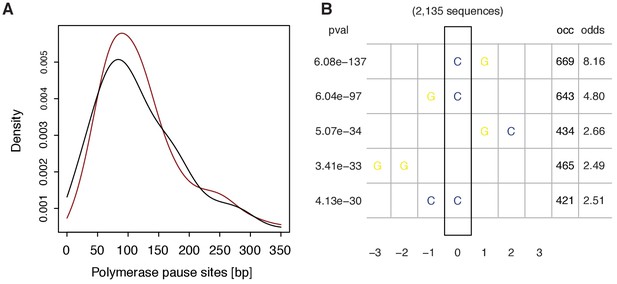
Features of underlying DNA sequence around promoter-proximal pause sites.
(A) Distributions of pause site depicted as densities for TUs with a response ratio >75% quantile (574 TUs, red) and TUs with a response ratio <25% quantile (469 TUs, black). (B) Plot showing the top 5 enriched 2-mers found by comparing the frequency of all possible 2-mers in a window of ±10 bp around the estimated pause site for fixed positions. Testing was done via Fisher’s exact test against the (background) frequency of the respective 2-mer obtained from a window of the same size shifted 500 bp downstream. The respective p-values and odd-ratios are given in the left and right panel.
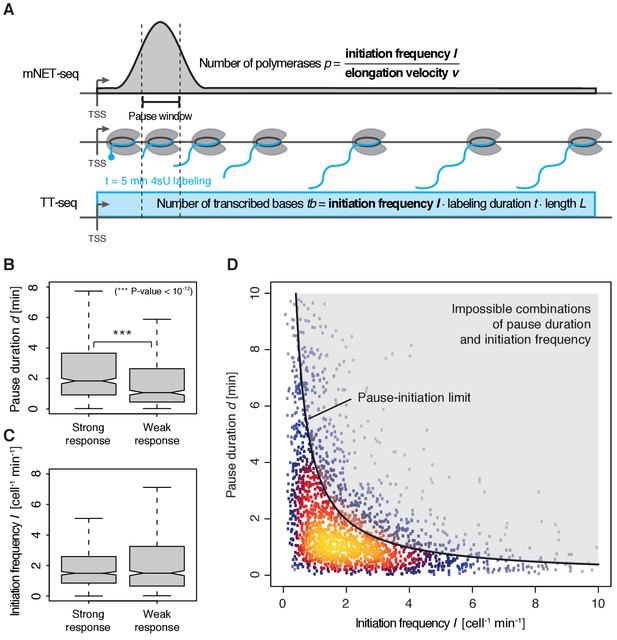
Pol II pausing generally limits transcription initiation (‘pause-initiation limit’).
(A) Schematic representation of polymerase flow in the promoter-proximal region. The mNET-seq signal (top) is the ratio of the initiation frequency I over the elongation velocity v. The TT-seq signal (bottom) corresponds to initiation frequency I. Thus, v can be derived from the ratio of the TT-seq over the mNET-seq signal, and the reciprocal of v in the pause window corresponds to the pause duration d. (B) Distributions of gene-wise pause duration d [min] for TUs with a CDK9 response ratio >75% quantile (574 TUs) and TUs with a response ratio <25% quantile (469 TUs). (C) Distributions of gene-wise initiation frequency I [cell−1min−1] for TUs with a CDK9 response ratio >75% quantile (635 TUs) and TUs with a response ratio <25% quantile (635 TUs). (D) Scatter plot between the initiation frequency I [cell−1min−1] and the pause duration d [min] for 2135 common TUs with color-coded density estimation. The grey shaded area depicts impossible combinations of I and d according to published kinetic theory (Ehrensberger et al., 2013) and assuming that steric hindrance occurs below a distance of 50 [bp] between the active sites of the initiating Pol II and the paused Pol II.
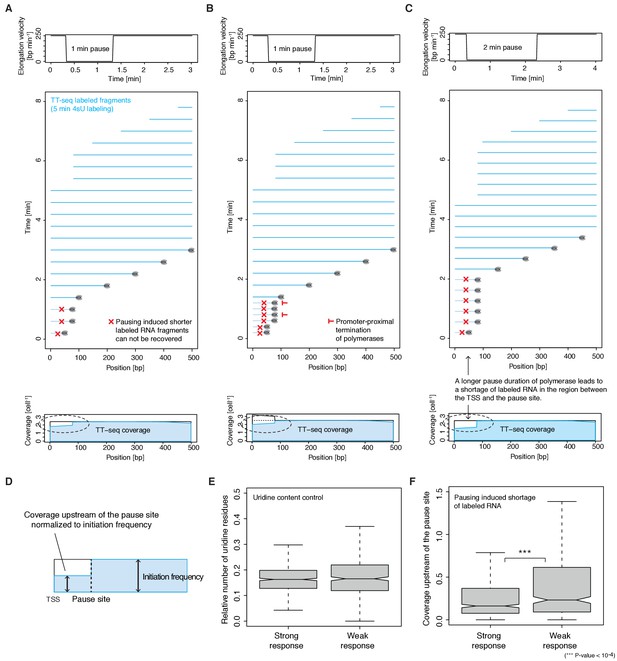
A longer pause duration but not promoter-proximal termination of polymerase leads to shortage of labeled RNA in the region between TSS and pause site.
(A) Simulation of labeled RNA fragments synthesized in 5 min labeling duration (TT-seq fragments depicted for polymerases with a distance corresponding to 40 s of elongation, middle panel) for a pause site 80 bp downstream of the TSS with a given elongation velocity profile [bp min−1] comprising a pause duration of 1 min (upper panel) and a initiation frequency of 0.5 [cell−1min−1]. Lower panel shows the resulting TT-seq coverage. Shorter fragments have a higher probability to escape labeled RNA purification and can not be recovered fully. (B) Simulation of labeled RNA fragments as in (A) with two times the initiation frequency (1 [cell−1min−1]) and a promoter-proximal termination of every second polymerase. The resulting TT-seq coverage (lower panel) shows less effect (higher coverage) upstream of the pause site. For reasons of simplicity the promoter-proximal termination of every second polymerase is modeled by overlaying two simulation instances with an initiation frequency of 0.5 [cell−1min−1]. One as in (A) and one with a constantly terminating polymerase. Note that polymerases that terminate in the pause window do not contribute signal to the region downstream of the pause site. (C) Simulation of labeled RNA fragments as in (A) with a pause duration of 2 min (upper panel) leading to a greater shortage of labeled RNA in the region between the TSS and the pause site. (D) Schematic representation of coverage ratio calculation for real TT-seq coverage. (E) Distributions of gene-wise uridine content in the region between the TSS and the pause site for TUs with a response ratio >75% quantile (603 TUs) and TUs with a response ratio <25% quantile (527 TUs). (F) Distributions of gene-wise mean real TT-seq signal in the region between the TSS and the pause site normalized to initiation frequency for subsets as in (E).
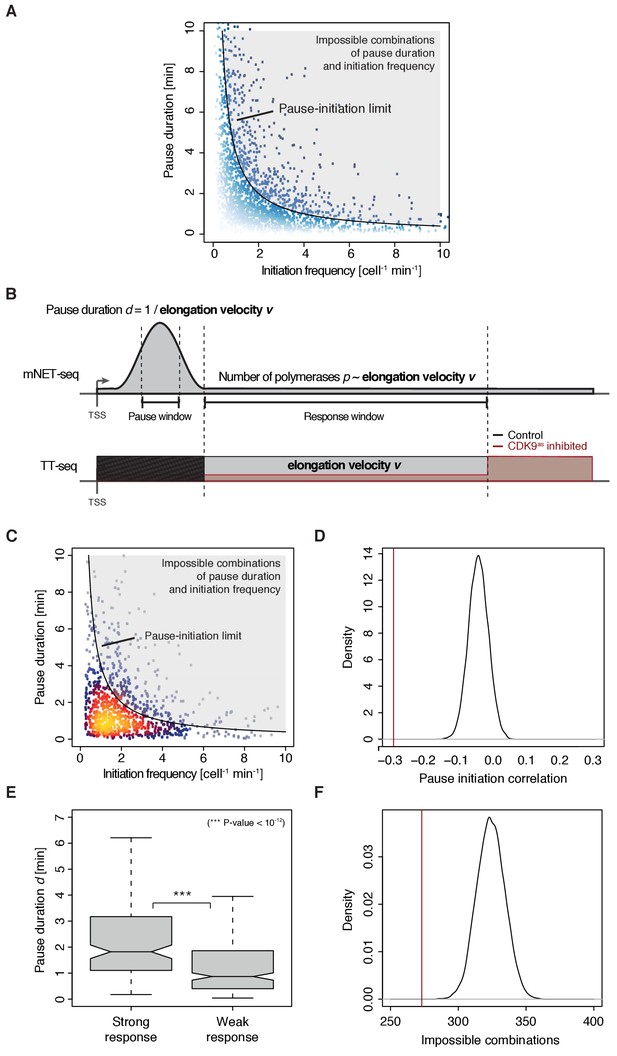
Verification of anti-correlation between initiation frequency I and pause duration d including ‘pause-initiation limit’.
(A) Scatter plot comparing the initiation frequency [cell−1min−1] against the pause duration [min] for 2135 common TUs with color encoding according to mNETseq signal strength (weak in white to strong in blue). The grey shaded area depicts impossible combinations of I and d according to published kinetic theory (Ehrensberger et al., 2013) and assuming that steric hindrance occurs below a distance of 50 bp between the active sites of the initiating Pol II and the paused Pol II. (B) Schematic representation of polymerase flow in the promoter-proximal region. The number of polymerases in a region of interest (mNET-seq signal, top) corresponds to the average elongation velocity v in that region. The width of the response window (TT-seq signal, bottom) informs on Pol II elongation velocity v. The pause duration can be derived (without the initiation frequency I) as the reciprocal of v in the pause window. The elongation velocity v in the pause window relates directly to v in the response window which can be adjusted to the elongation velocity obtained from CDK9 inhibition (Materials and methods). (C) Scatter plot revealing an anti-correlation between the initiation frequency I [cell−1min−1] and the pause duration [min] for 974 common TUs with color-coded density estimation (Spearman correlation coefficient −0.3). The grey shaded area depicts impossible combinations of I and according to published kinetic theory (Ehrensberger et al., 2013). (D) Density showing the spearman correlation coefficient of pause duration and initiation frequency I for repeated randomly shuffled mNET-seq signal assignment to TUs. Original spearman correlation coefficient is indicated with a red line. (E) Distributions of gene-wise pause duration [min] for TUs with a CDK9 response ratio >75% quantile (155 TUs) and TUs with a response ratio <25% quantile (271 TUs). (F) Density showing the number of impossible combinations of pause duration and initiation frequency I (above pause-initiation limit) for repeated randomly shuffled mNET-seq signal assignment to TUs. Original observation is indicated with a red line.
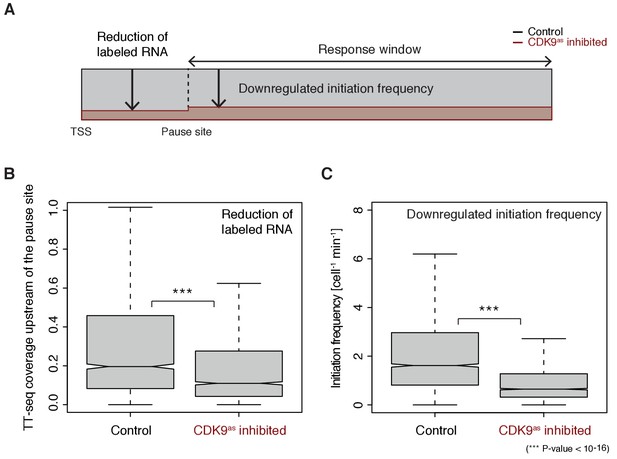
Increasing Pol II pause duration decreases the frequency of transcription initiation.
(A) Schematic representation of observed decrease in TT-seq signal upon CDK9 inhibition, upstream and downstream of the pause site. (B) Distributions of gene-wise mean TT-seq signals in the region between the TSS and the pause site, before (control) and after CDK9 inhibition, normalized to the initiation frequency before CDK9 inhibition. (C) Distributions of gene-wise initiation frequencies before (control) and after CDK9 inhibition.
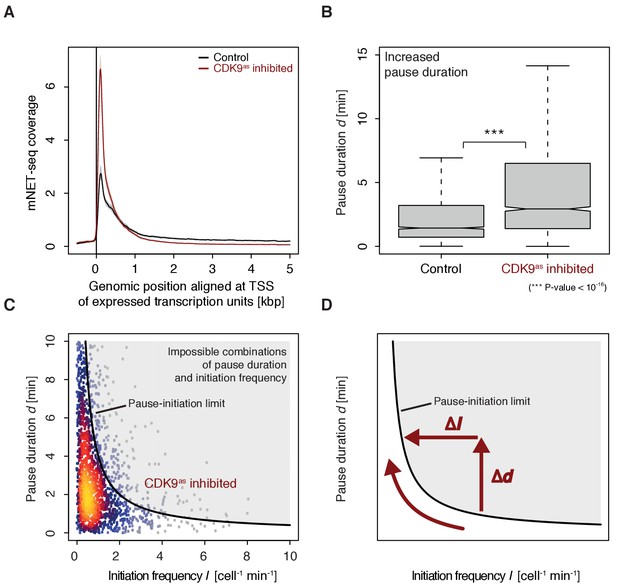
CDK9 inhibition leads to increased pause duration.
(A) Metagene analysis comparing the average mNET-seq signal before and after CDK9 inhibition. Two biological replicates were averaged. The mNET-seq coverage was averaged for 2538 investigated TUs (Materials and methods). TUs were aligned with their TSS. Shaded areas around the average signal (solid lines) indicate confidentiality intervals (Materials and methods). (B) Distributions of gene-wise pause duration d [min] before (control) and after CDK9 inhibition. (C) Scatter plot between the initiation frequency I [cell−1min−1] and the pause duration d [min] after CDK9 inhibition for 2135 common TUs with color-coded density estimation. The grey shaded area depicts impossible combinations of I and d (Ehrensberger et al., 2013) assuming that steric hindrance occurs below a distance of 50 [bp] between the active sites of the initiating Pol II and the paused Pol II. (D) Schematic of changes in pause duration (Δd) and initiation frequency (ΔI) upon CDK9 inhibition. As a consequence, data points in panel (D) are moved to the left and upwards.
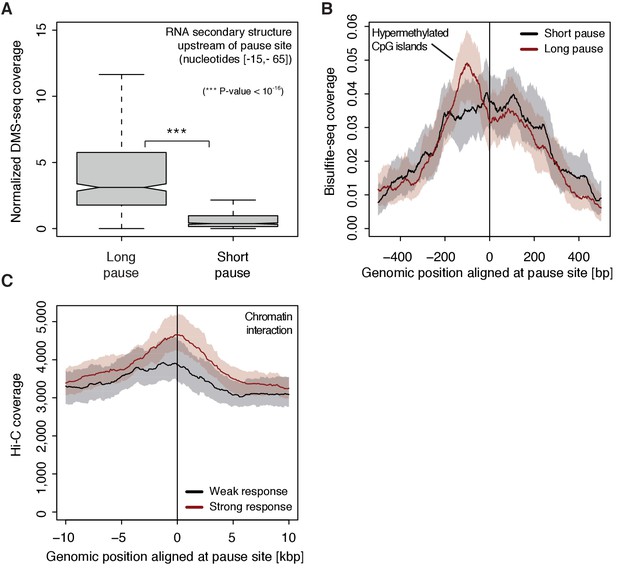
Determinants of CDK9-dependent promoter-proximal pausing.
(A) Distribution of gene-wise mean in vivo DMS-seq signals (detecting RNA secondary structure) for a window between −65 and −15 [bp] upstream of the pause site for TUs with long pause durations (pause duration >75% quantile, 534 TUs) and with short pause durations (pause duration <25% quantile, 534 TUs) normalized to denatured DMS-seq coverage (Materials and methods). (B) Metagene analysis comparing the average Bisulfite-seq signal (detecting methylated DNA) for subsets as in (A) aligned at the pause site (red, long pause duration, and black, short pause duration). Shaded areas around the average signal (solid lines) indicate confidence intervals. (C) Metagene analysis comparing the average Hi-C signal (detecting long-range chromatin interactions) for strongly CDK9-responding TUs (red, response ratio >75% quantile, 552 TUs) and weakly CDK9-responding TUs (black, response ratio <25% quantile, 440 TUs) aligned at the pause site. Shaded areas around the average signal (solid lines) indicate confidence intervals (Materials and methods, Supplementary file 1).
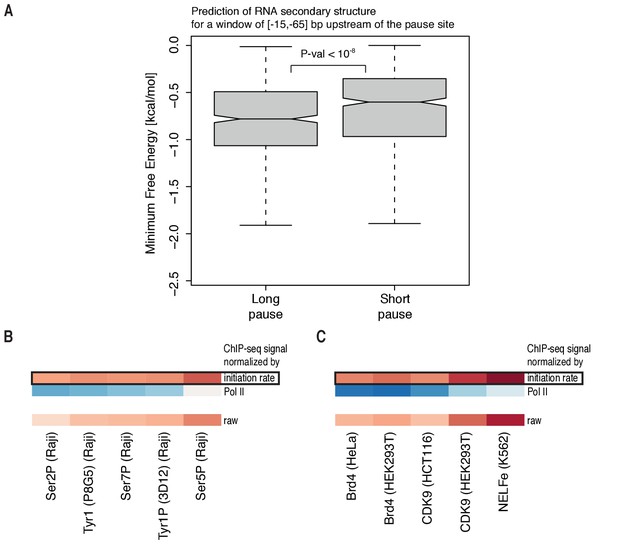
Features of promoter-proximal pausing.
(A) Distributions of gene-wise mean minimum free energy (Materials and methods) for a window of [−15,–65] bp upstream of the pause site for TUs with long pause durations (pause duration >75% quantile, 534 TUs) and TUs with short pause durations (pause duration <25% quantile, 534 TUs). (B) Heatmap showing the pairwise Spearman correlation (color encoded, −0.12 in blue to 0.15 in red) using ChIP measurements of Pol II phospho-isoforms S2P, S5P, S7P and T1P in the pause window against the pause duration in three different variants: ChIP measurements normalized to productive initiation rate (initiation rate), normalized to total Pol II (Pol II), raw signal (raw). (C) Heatmap as in (B) using ChIP measurements of CDK9, NELFe and Brd4 (color encoded, −0.19 in blue to 0.22 in red) (Supplementary file 1).
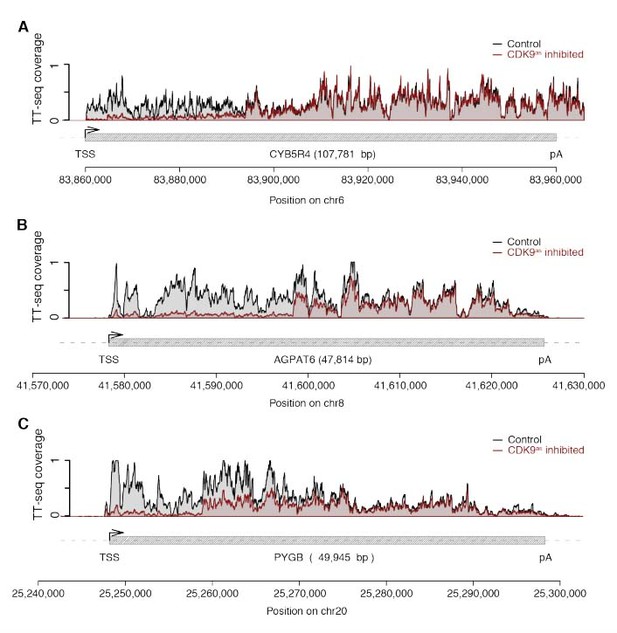
Example genome browser views of TT-seq signals in CDK9as cells with high responsiveness (~ 90%).
(A) CYB5R4 gene locus (107,781 [bp]) on chromosome 6. The upper panel shows TT-seq signal with CDK9as inhibitor (red) and control (black). Grey box depicts transcript body from transcription start site (TSS, black arrow) to polyA site (pA). (B) AGPAT6 gene locus (47,814 [bp]) on chromosome 8 depicted as in (A). (C) PYGB gene locus (49,945 [bp]) on chromosome 20 depicted as in (A).
Additional files
-
Supplementary file 1
Published datasets used for analysis.
Note that the conclusions we draw across different cell-lines are all based on metagene analysis, involving from 500 up to more than 2000 genes. Thus, we assume cell-line specific differences to have an insignificant influence and that the tendencies we observe rather suggest strong conservation.
- https://doi.org/10.7554/eLife.29736.018
-
Transparent reporting form
- https://doi.org/10.7554/eLife.29736.019





