Cdc48 regulates a deubiquitylase cascade critical for mitochondrial fusion
Figures
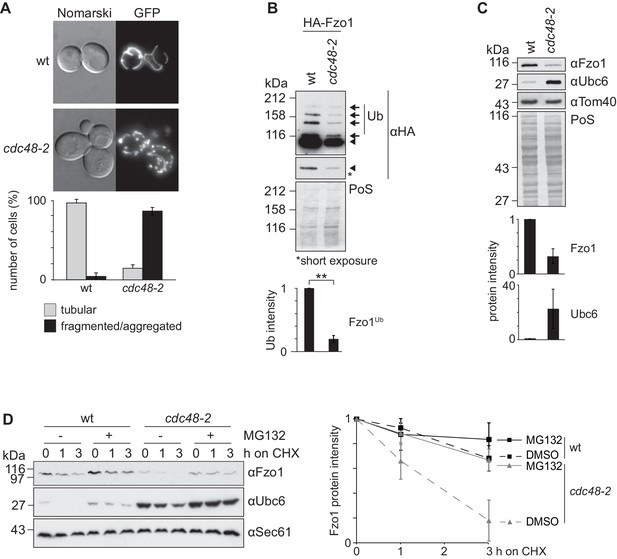
Cdc48 regulates Fzo1 and mitochondrial fusion.
(A) Mitochondrial morphology of CDC48 mutant cells. Wild-type (wt) or cdc48-2 mutant cells were analyzed for mitochondrial tubulation after expressing a mitochondrial-targeted GFP plasmid. Cellular (Nomarski) and mitochondrial (GFP) morphology were visualized by fluorescence microscopy. Bottom panel, quantification of four independent experiments (with more than 200 cells each) including mean and standard deviation (SD), as described (Cumming et al., 2007). (B) Ubiquitylation of Fzo1 upon mutation of CDC48. Crude mitochondrial extracts from wt or cdc48-2 mutant cells expressing HA-Fzo1, or the corresponding empty vector, were solubilized and analyzed by SDS-PAGE and immunoblotting using HA-specific antibodies. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated by a black arrowhead or black arrows, respectively. Ubiquitylated forms of Fzo1 are labeled with Ub. Bottom panel, quantification of three independent experiments, normalized to PoS and including SD. **, p≤0.01 (paired t-test). (C) Steady state levels of Fzo1 upon mutation of CDC48. Total cellular extracts of wt or cdc48-2 mutant cells were analyzed by SDS-PAGE and immunoblotting using Fzo1- or Ubc6-specific and, as a loading control, Tom40-specific antibodies. Bottom panels, quantification of three independent experiments, including SD. (D) Proteasome dependence of Fzo1 degradation in cdc48-2 mutant cells. The turnover of endogenous Fzo1 expressed in Δpdr5 Δsnq2 and Δpdr5 Δsnq2 cdc48-2 cells was assessed after inhibition of cytosolic protein synthesis with cycloheximide (CHX), for the indicated time points in exponentially growing cultures in absence or presence of the proteasomal inhibitor MG132. Samples were analyzed by SDS-PAGE and immunoblotting using Fzo1-specific, Ubc6-specific (as an unstable protein control) and Sec61-specific (as a loading control) antibodies. Right panel, quantification of five independent experiments, including SD. PoS, PonceauS staining.
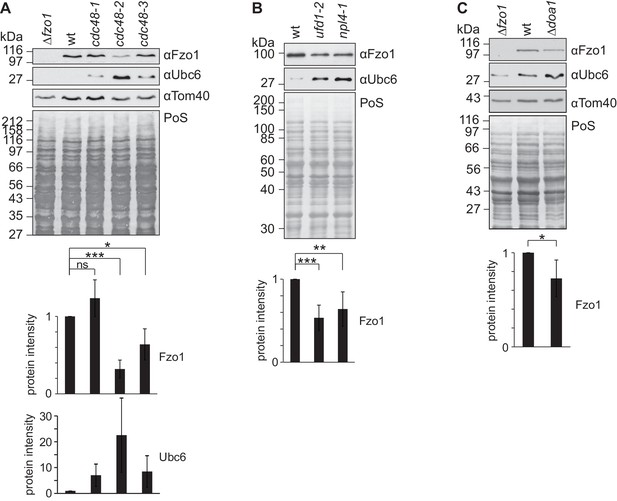
Cdc48 regulates Fzo1 and mitochondrial fusion.
(A) Steady state levels of Fzo1 upon mutation of CDC48. Total cellular extracts of Δfzo1 or wt cells or different CDC48 mutant cells were analyzed by SDS-PAGE and immunoblotting using Fzo1-, Ubc6- and Tom40-specific antibodies. Bottom panels, quantification of five independent experiments, including SD. ns, p>0.05; *, p≤0.05; ***, p≤0.001 (One-way ANOVA, Tukey’s multiple comparison test). (Β) Role of Cdc48 cofactors in the steady state levels of Fzo1. Total cellular extracts of wt cells or ufd1-2 and npl4-1 mutant cells were analyzed by SDS-PAGE and immunoblotting using Fzo1- or Ubc6-specific antibodies. Bottom panels, quantification of seven (ufd1-2) or nine (npl4-1) independent experiments, including SD. **p≤0.01; ***p≤0.001 (paired t-test). (C) Steady state levels of Fzo1 upon deletion of DOA1. Total cellular extracts of Δfzo1, wt or Δdoa1 cells were analyzed by SDS-PAGE and immunoblotting using Fzo1-, Ubc6- and Tom40-specific antibodies. Bottom panel, quantification of five independent experiments, including SD. *p≤0.05 (paired t-test). PoS, PonceauS staining.

Cdc48 regulates Fzo1 and mitochondrial fusion.
(A) Rescue analysis of Fzo1 steady state levels in cdc48-2 cells. Total cellular extracts of wt or cdc48-2 mutant cells expressing Cdc48, Cdc48A547T or the corresponding empty vector were analyzed by SDS-PAGE and immunoblotting using an HA-specific antibody. (B) Rescue analysis of Fzo1 ubiquitylation in cdc48-2 cells. Crude mitochondrial extracts from wt or cdc48-2 mutant cells, additionally expressing HA-Fzo1 and Cdc48, Cdc48A547T or the corresponding empty vector, as indicated, were lysed and HA-tagged Fzo1 was precipitated using HA-coupled beads. Samples were analyzed by SDS-PAGE and immunoblotting using an HA-specific antibody. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated as in Figure 1B. (C) Rescue analysis of mitochondrial morphology in cdc48-2 cells. Wt or cdc48-2 mutant cells expressing Cdc48 or Cdc48A547T or the corresponding empty vector as indicated were analyzed for mitochondrial tubulation after expressing a mitochondrial-targeted GFP plasmid, as in Figure 1A. Quantification from three different experiments (with more than 200 cells each), including SD, as described (Cumming et al., 2007). IP, immunoprecipitation. PoS, PonceauS staining.
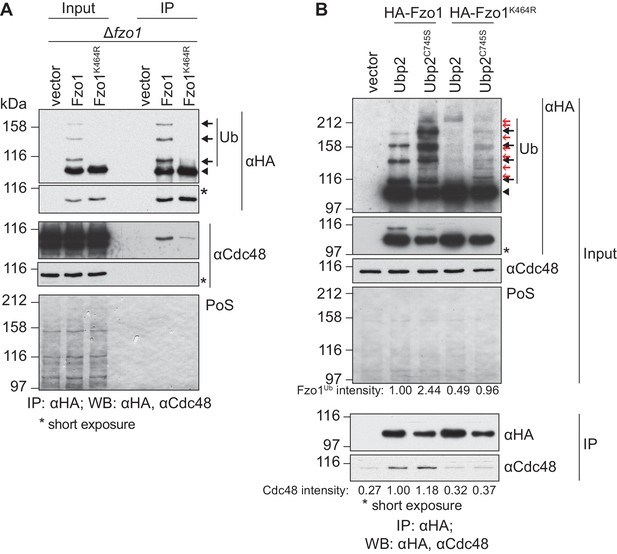
Cdc48 specifically affects ubiquitylated Fzo1.
(A) Physical interaction between Cdc48 and ubiquitylated Fzo1. HA-Fzo1, HA-Fzo1K464R or the corresponding vector were expressed in ∆fzo1 cells. Crude mitochondrial extracts were lysed and HA-tagged Fzo1 was precipitated using HA-coupled beads and analyzed by SDS-PAGE and immunoblotting using HA- and Cdc48-specific antibodies. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated as in 1B. (B) Effect of the anti-fusion ubiquitylation of Fzo1 on its interaction with Cdc48. HA-Fzo1 or HA-Fzo1K464R, expressed in the presence of Ubp2 (∆fzo1 cells plus empty vector) or Ubp2C745S (∆ubp2 ∆fzo1 cells plus Ubp2C745S-Flag), or the corresponding vector control (the empty vectors corresponding to HA-Fzo1 and Ubp2C745S-Flag, expressed in ∆ubp2 ∆fzo1 cells), were analyzed for Cdc48 interaction, as in 2A. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated by a black arrowhead or black arrows, respectively. Red arrows with no fill indicate Fzo1 ubiquitylated species specifically accumulating upon expression of Ubp2C745S. PoS, PonceauS staining; IP, immunoprecipitation; WB, western blot.
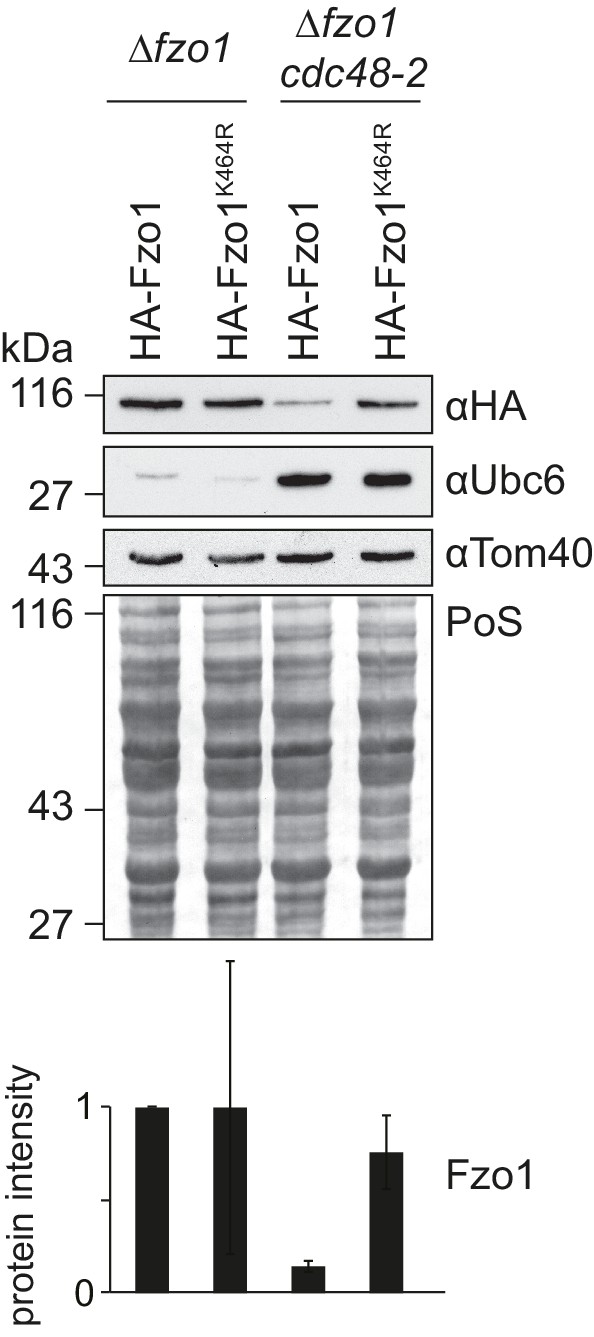
Cdc48 specifically affects ubiquitylated Fzo1.
Steady state levels of HA-Fzo1K464R upon mutation of CDC48. Total cellular extracts of ∆fzo1 or ∆fzo1 cdc48-2 mutant cells expressing HA-Fzo1 or HA-Fzo1K464R were analyzed by SDS-PAGE and immunoblotting using Fzo1-,Ubc6- and Tom40-specific antibodies. Bottom panel, quantification of four independent experiments, including SD. PoS, Ponceau S staining.
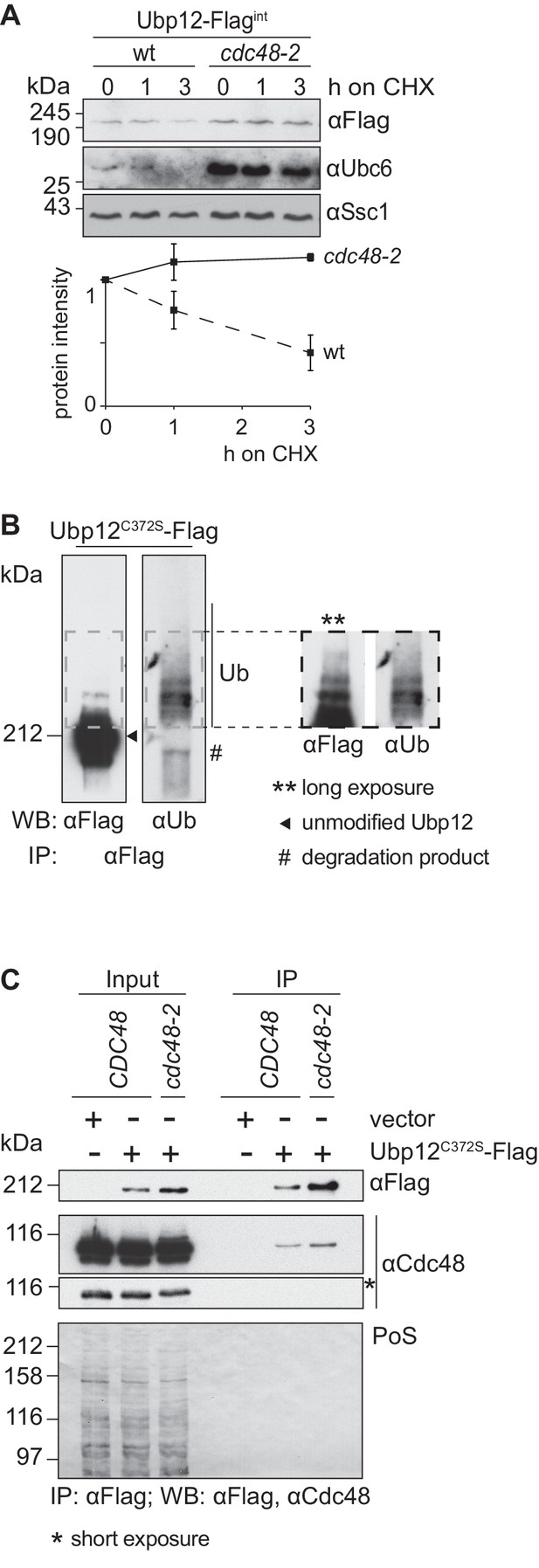
Cdc48 supports ubiquitin-dependent turnover of Ubp12.
(A) Stability of the Ubp12 protein. The turnover of Ubp12 endogenously Flag tagged (Ubp12-Flagint), in wt or cdc48-2 cells, was assessed with CHX chase, as in 1D. Samples were analyzed by SDS-PAGE and immunoblotting using a Flag-, Tom40- and, as an unstable protein control, a Ubc6-specific antibody. Bottom panel, quantification of three independent experiments, including SD. (B) Ubiquitylation of Ubp12. The Ubp12C372S-Flag inactive variant, expressed from an episomal plasmid, was immunoprecipitated from total soluble extracts using Flag-coupled beads. After elution, Ubp12 was analyzed by western blot using Flag- or ubiquitin (Ub - P4D1)-specific antibodies. Ubiquitylated forms of Ubp12C372S-Flag are labeled with Ub. (C) Physical interaction between Cdc48 and Ubp12. The catalytically inactive Ubp12C372S-Flag variant, expressed from an episomal plasmid, or the corresponding empty vector, were expressed in Δubp12 (CDC48) or Δubp12 cdc48-2 (cdc48-2) mutant cells and analyzed for Cdc48 interaction. Crude mitochondrial extracts were lysed, Flag-tagged Ubp12 was precipitated using Flag-coupled beads, and the eluate analyzed by SDS-PAGE and immunoblotting using Flag- and Cdc48-specific antibodies. PoS, Ponceau S staining; IP, immunoprecipitation; WB, western blot.

Cdc48 supports ubiquitin-dependent turnover of Ubp12.
(A) Turnover of episomal Ubp12 in wt or cdc48-2 cells. Ubp12-Flag stability was assessed after inhibition of cytosolic protein synthesis with cycloheximide (CHX), for the indicated time points in exponentially growing cultures. Samples were analyzed by SDS-PAGE and immunoblotting using Flag-, Ubc6- and Tom40-specific antibodies. Bottom panel, quantification of three independent experiments, including SD. (B) Proteasome dependence of Ubp12-Flag degradation. The turnover of Ubp2-Flag, expressed from an episomal plasmid, was assessed as in 1D. Samples were analyzed by SDS-PAGE and immunoblotting using Flag-, Ubc6- and Ssc1-specific antibodies. (C) Ubp12 expression levels. Expression levels of endogenously Flag-tagged Ubp12 (Ubp12-Flagint), Ubp12-Flag expressed from an episomal plasmid and endogenously Flag-tagged Ubp12 under the control of a pGAL promoter (pGAL-Ubp12-Flagint) (grown in glucose or galactose as indicated) were analyzed by SDS-PAGE and immunoblotting using Flag- and Ssc1-specific antibodies. Pos, PonceauS staining.
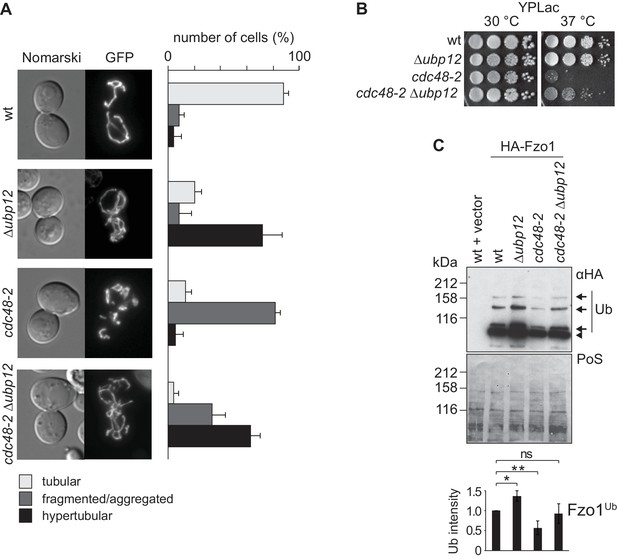
Interdependence of Cdc48 and Ubp12 for Fzo1 regulation.
(A) Mitochondrial morphology upon deletion of UBP12 and/or mutation of CDC48. The indicated mutant cells were analyzed for mitochondrial tubulation after expressing a mitochondrial-targeted GFP plasmid, as in Figure 1A. Right panel, quantification from three different experiments (with more than 200 cells each), including SD, as described (Cumming et al., 2007) (B) Respiratory capacity of cells upon deletion of UBP12 and/or mutation of CDC48. Fivefold serial dilutions of exponentially growing cells of wt or the mutant strains Δubp12, cdc48-2, and Δubp12 cdc48-2 were spotted on YP media supplemented with lactate (YPLac) and incubated at 30°C for two days or 37°C for five days. (C) Ubiquitylation levels of Fzo1 upon deletion of UBP12 and/or mutation of CDC48. Crude mitochondrial extracts from the indicated strains additionally expressing HA-Fzo1, or the corresponding empty vector, were analyzed by SDS-PAGE and immunoblotting using an HA-specific antibody. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated as in Figure 1B. Bottom panel, quantification of four independent experiments, normalized to PoS and including SD. ns, p>0.05. *, p≤0.05, **, p≤0.01 (One-way ANOVA, Tukey’s multiple comparison test). PoS, PonceauS staining.
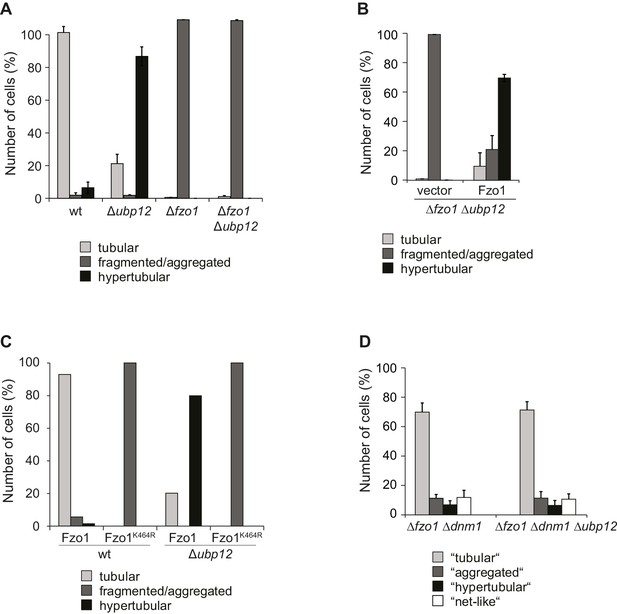
Interdependence of Cdc48 and Ubp12 for Fzo1 regulation.
(A) Mitochondrial morphology upon deletion of UBP12 in Δfzo1 cells. The indicated mutant cells were analyzed for mitochondrial tubulation after expressing a mitochondrial-targeted GFP plasmid, as in Figure 1A. Quantification from three different experiments (with more than 200 cells each), including SD, as described (Cumming et al., 2007). (B) Mitochondrial morphology upon expression of HA-Fzo1 in Δfzo1 Δubp12 cells. The indicated mutant cells were analyzed for mitochondrial tubulation after expressing a mitochondrial-targeted GFP plasmid, as in Figure 1A. Quantification from three different experiments (with more than 200 cells each), including SD, as described (Cumming et al., 2007). (C) Mitochondrial morphology upon endogenous expression of HA-Fzo1 or HA-Fzo1K464R in Δubp12 cells. The indicated mutant cells were analyzed for mitochondrial tubulation after expressing a mitochondrial-targeted GFP plasmid, as in Figure 1A. Quantification from one experiment (with more than 200 cells each). (D) Mitochondrial morphology upon deletion of UBP12 in Δfzo1 Δdnm1 cells. The indicated mutant cells were analyzed for mitochondrial tubulation after expressing a mitochondrial-targeted GFP plasmid, as in Figure 1A. Quantification from three different experiments (with more than 200 cells each), including SD, as described (Cumming et al., 2007).
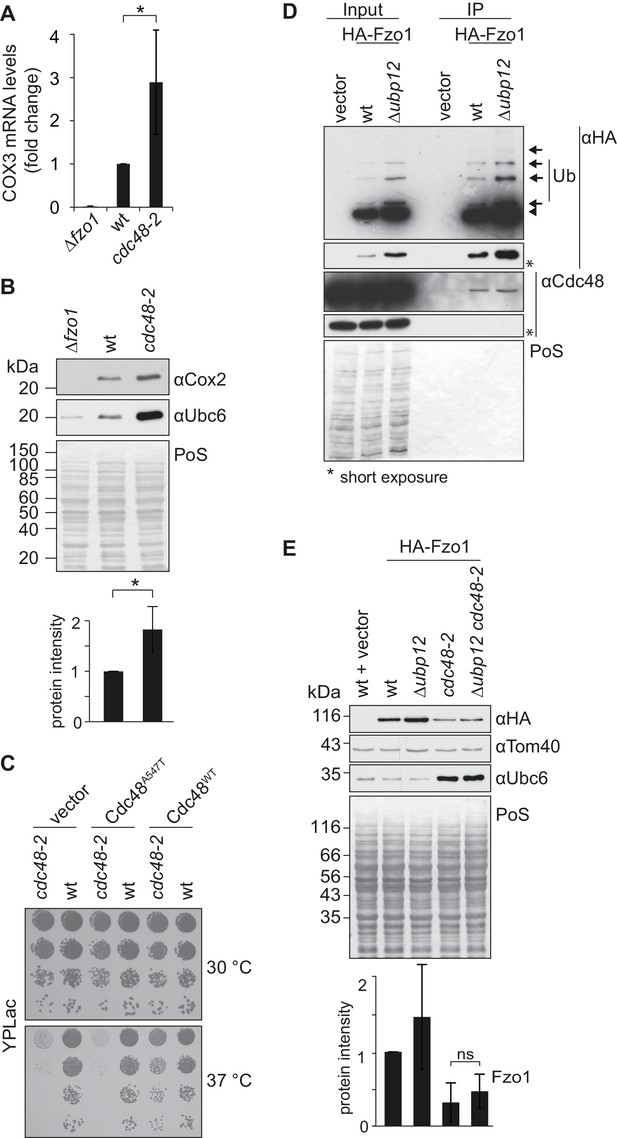
Interdependence of Cdc48 und Ubp12 for Fzo1 regulation.
(A) Analysis of mtDNA content in cdc48-2 cells using RT-PCR. mtDNA content in Δfzo1, wt and cdc48-2 cells was analyzed by measuring COX3 and ACT1 (as housekeeping gene) RNA levels using RT-PCR. Quantification of six independent experiments, including SD. *p≤0.05 (paired t-test). (B) Analysis of mtDNA content in cdc48-2 cells using the Cox2 protein amount. Total cellular extracts of Δfzo1, wt and cdc48-2 cells were analyzed by SDS-PAGE and immunoblotting using Cox2- (as mtDNA marker) or Ubc6-specific antibodies. Bottom panel, quantification of five independent experiments, including SD. *p≤0.05 (paired t-test). (C) Respiratory capacity of cdc48-2 cells upon expression of wt or mutant Cdc48. A spot assay was performed as described in Figure 4B with the indicated cells but using YPLac, grown at 30°C for 1 day and at 37°C for 3 days. (D) Physical interaction between Cdc48 and Fzo1 in Δubp12 cells. HA-Fzo1 or the corresponding empty vector was expressed in wt or Δubp12 cells and analyzed for Cdc48 interaction, as in 2A. Crude mitochondrial extracts were lysed, HA tagged Fzo1 was precipitated using HA-coupled beads, and the eluate was analyzed by SDS-PAGE and immunoblotting using HA- and Cdc48-specific antibodies. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated as in Figure 1B. (E) Steady state levels of Fzo1 upon deletion of UBP2 and/or mutation of CDC48. Total cellular extracts of wt cells or Δubp12, cdc48-2 and Δubp12 cdc48-2 mutant cells were analyzed by SDS-PAGE and immunoblotting using HA-, Ubc6- and Tom40-specific antibodies. Bottom panel, quantification of six independent experiments, including SD. ns, p>0.05 (One-way ANOVA; Tukey’s multiple comparison test). PoS, Ponceau S staining; IP, immunoprecipitation; WB, western blot.
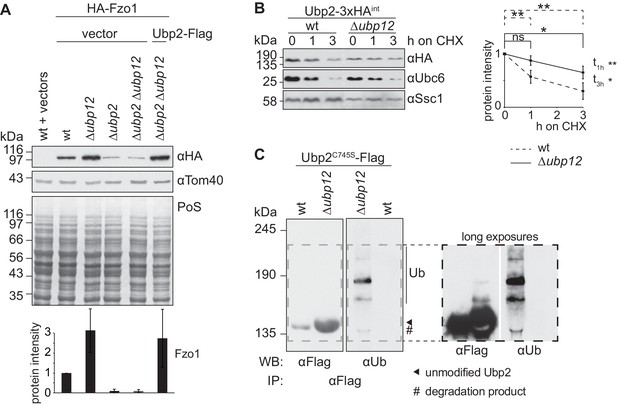
Ubp12 modulates Ubp2 ubiquitylation and turnover.
(A) Interdependent role of Ubp2 and Ubp12 for the steady state levels of Fzo1. Total cellular extracts of wt or Δubp2, Δubp12, and Δubp2 Δubp12 mutant cells expressing HA-Fzo1 and also expressing either Ubp2-Flag or the corresponding empty vector, as indicated, were analyzed by SDS-PAGE and immunoblotting using HA- and Tom40-specific antibodies. Bottom panel, quantification of four independent experiments, including SD. (B) Turnover of endogenous Ubp2 in wt or Δubp12 cells. The turnover of endogenously 3xHA-tagged Ubp2 (Ubp2-3xHAint) was assessed as in 3A. Samples were analyzed by SDS-PAGE and immunoblotting using antibodies against HA, Ubc6 and Ssc1. Right panel, quantification of four independent experiments, including SD. For the statistical analysis of the degradation kinetics of each strain, a paired t-test was used; for the statistical analysis of the difference in steady state levels of both strains at the indicated time points (t1h, t3h) an unpaired t-test was used. ns, p>0.05; *, p≤0.05; **, p≤0.01. (C) Ubiquitylation of Ubp2. The Ubp2C745S-Flag inactive variant, expressed in wt or Δubp12 cells, was immunoprecipitated from total soluble extracts using Flag-coupled beads. Eluted Ubp2 was analyzed by western blot using Flag- or ubiquitin (Ub - P4D1)-specific antibodies. Ubiquitylated forms of Ubp2C745S-Flag are labeled with Ub. PoS, Ponceau S staining; IP, immunoprecipitation; WB, western blot.
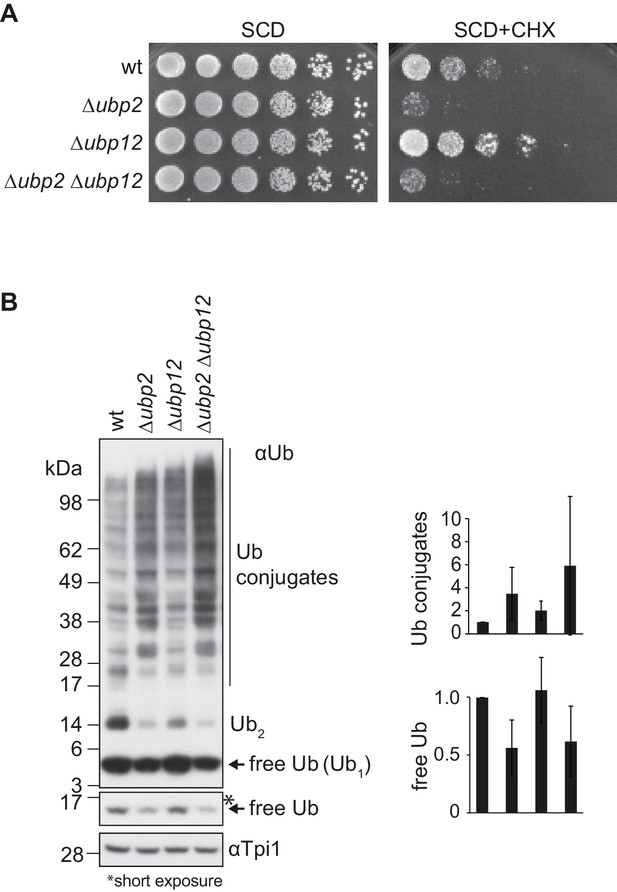
Ubp12 modulates Ubp2 ubiquitylation and turnover.
(A) Opposing roles of Ubp2 and Ubp12 for CHX resistance. A spot assay was performed, as described in Figure 4B, but on synthetic media supplemented with glucose (SCD) in the absence or presence of 0.5 µg/ml CHX and incubated at 30°C for one or five days, respectively. (B) Distinct roles of Ubp2 and Ubp12 for cellular ubiquitylation. Total cellular extracts of the indicated strains were analyzed by SDS-PAGE and immunoblotting using ubiquitin (Ub; αP4D1) and Tpi1-specific antibodies, used as loading control. Free ubiquitin or ubiquitylated conjugates are labeled with Ub. Right panels, quantification of three independent experiments showing the levels of free Ub or Ub conjugates, including SD.

Ubp12 modulates Ubp2 ubiquitylation and turnover.
(A) Proteasome dependence of Ubp2-Flag degradation in Δpdr5 Δsnq2 mutant cells. The turnover of ectopically expressed Ubp2-Flag was assessed as in Figure 1D. Samples were analyzed by SDS-PAGE and immunoblotting using Flag- and Ubc6-specific antibodies. (B) Physical interaction between Ubp2 and Ubp12. Catalytically inactive variants ectopically expressed Ubp2C745S-Flag and non-tagged Ubp12C372S, or their corresponding empty vectors, were expressed in Δubp2 Δubp12 cells. Total soluble extracts were prepared and Ubp12C372S was precipitated using Sepharose beads in the presence or absence of a Ubp12-specific antibody, as indicated. The eluates were analyzed by SDS-PAGE and immunoblotting using Flag- and Ubp12-specific antibodies. (C) Ubiquitylation of Ubp2. The Ubp2C745S-Flag inactive variant, expressed in wt, Δubp12 and Δubp12Δmdm30 cells, was immunoprecipitated from total soluble extracts using Flag-coupled beads. Eluted Ubp2 was analyzed by western blot using antibodies specific for Flag or ubiquitin (Ub; αP4D1). Ubiquitylated forms of Ubp2C745S-Flag are labeled with Ub. PoS, PonceauS staining; IP, immunoprecipitation; WB, western blot.
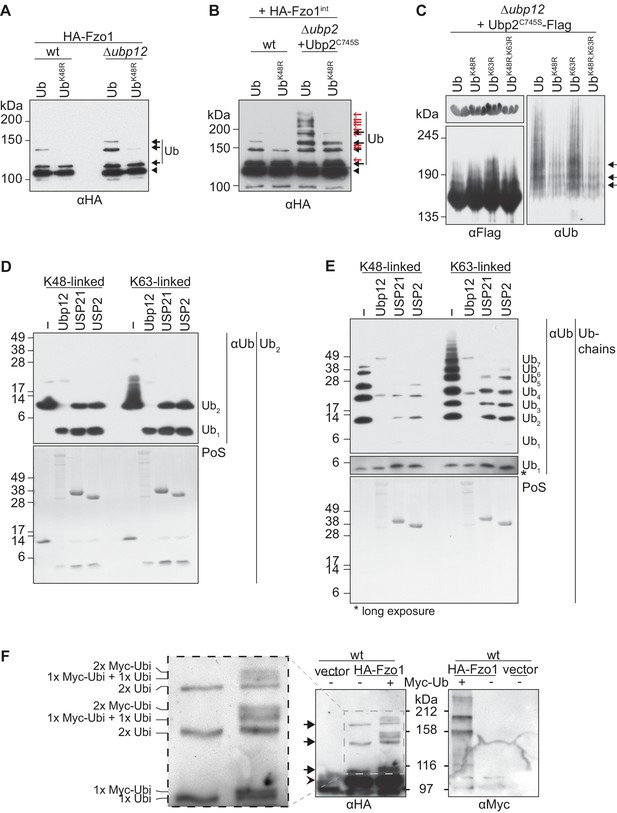
Characterization of the deubiquitylation reaction by Ubp12.
(A) Analysis of ubiquitin chain-type composition of Fzo1. Crude mitochondrial extracts from wt or Δubp12 mutant cells expressing HA-Fzo1, and over-expressing either wt ubiquitin (Ub) or ubiquitin with a K48R mutation (UbK48R), were solubilized, subjected to HA-immunoprecipitation and analyzed by SDS-PAGE and immunoblotting using an HA-specific antibody. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated as in 1B. (B) Ubiquitin chain-type analysis of Fzo1 upon Ubp2C745S expression. Crude mitochondrial extracts from wt or Δubp2 (expressing Ubp2C745S) cells expressing HA-Fzo1 endogenously, and overexpressing either wt ubiquitin (Ub) or UbK48R, were analyzed as in A. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated as in 2B (C) Analysis of Ubp2 ubiquitin chain composition in Δubp12 cells. Soluble extracts from Δubp12 cells expressing Ubp2C745S-Flag and different ubiquitin variants (as indicated) were prepared and Flag-tagged Ubp2C745S was precipitated using Flag-coupled beads. The eluate was analyzed by SDS-PAGE and immunoblotting using antibodies against Flag and ubiquitin (Ub; αP4D1). (D) Deubiquitylation (DUB) assay using Ub2 chains. Purified di-ubiquitin chains (Ub2) composed of either only K48- or K63-linkages were treated with the purified DUBs Ubp12, USP21 and USP2. Treated chains were analyzed by SDS-PAGE and immunoblotting using a ubiquitin-specific antibody (Ub; αP4D1). Mono-ubiquitin or di-ubiquitin chains are labeled with Ub1 or Ub2, respectively. (E) DUB assay using Ub-chains. Purified poly-ubiquitin chains (Ub-chains) composed of either only K48- or K63-linkages were treated with the purified DUBs Ubp12, USP21 or USP2. Treated chains were analyzed by SDS-PAGE and immunoblotting as in C. Ubiquitin chains were labeled as in D with the subscript value indicating the amount of ubiquitin moieties in the respective chain. (F) Ubiquitylation pattern of Fzo1. Wt cells expressing HA-Fzo1 were analyzed for Fzo1 ubiquitylation upon the expression of Myc-ubiquitin, or the respective empty vector. HA-Fzo1 was immunoprecipitated from mitochondrial extracts using HA-coupled beads. Eluted Fzo1 was split into two and samples were analyzed by SDS-PAGE and immunoblotting using HA- or Myc-specific antibodies. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated as in 1B. The composition of the additional species apparent upon co-expression of Myc-tagged ubiquitin is explained in the inset. PoS, PonceauS staining.
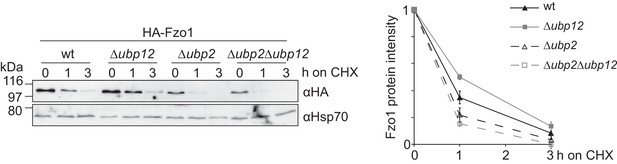
Characterization of the deubiquitylation reaction by Ubp12.
Opposite effects of Ubp12 and Ubp2 in Fzo1 stability. The turnover of HA-Fzo1 in wt, Δubp12, Δubp2 or Δubp12 Δubp2 cells was assessed after inhibition of cytosolic protein synthesis with cycloheximide (CHX), for the indicated time points in exponentially growing cultures. Samples were analyzed by SDS-PAGE and immunoblotting using a HA- and Hsp70-specific antibodies. Left panel, quantification of three independent experiments, including SD.
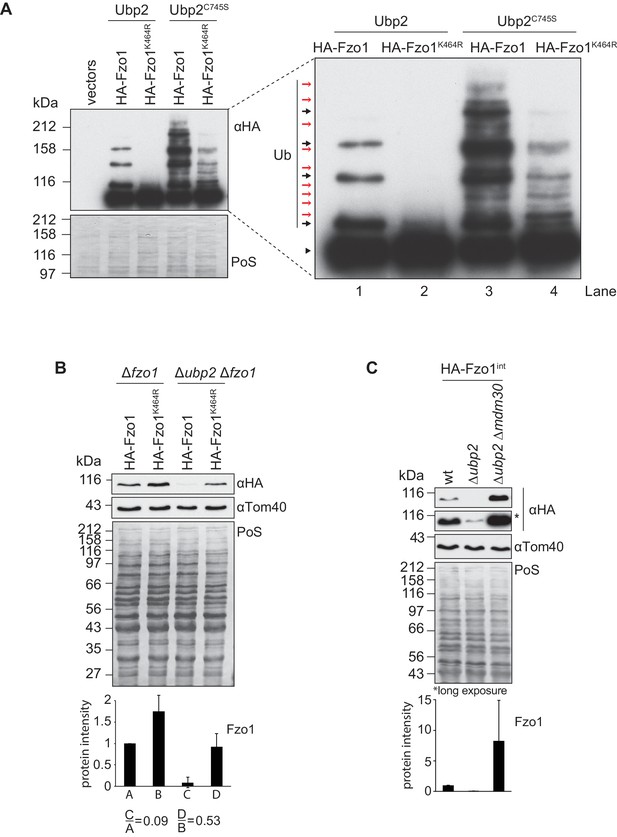
Interdependent roles of Ubp2 and Ubp12.
(A) Effect of Ubp2C745S on Fzo1K464R ubiquitylation. HA-Fzo1 or HA-Fzo1K464R were expressed in the presence of Ubp2 (∆fzo1 cells plus empty vector) or instead in the presence of Ubp2C745S (∆ubp2 ∆fzo1 plus Ubp2C745S-Flag), as indicated. Crude mitochondrial extracts were solubilized and HA-tagged Fzo1 was analyzed by SDS-PAGE and immunoblotting using an HA-specific antibody. Unmodified and ubiquitylated forms of HA-Fzo1 are indicated as in 2B. (B) Effect of UBP2 deletion on the steady state levels of Fzo1K464R. Total cellular extracts of indicated strains expressing HA-Fzo1 or HA-Fzo1K464R as indicated were analyzed by SDS-PAGE and immunoblotting using HA- and Tom40-specific antibodies. Bottom panel, quantification of five independent experiments, including SD. (C) Effect of Ubp2 and Mdm30 on the steady state levels of Fzo1. Total cellular extracts of wt, Δubp2 and Δubp2 Δmdm30 cells expressing HA-tagged Fzo1 endogenously (HA-Fzo1int) were analyzed by SDS-PAGE and immunoblotting using HA- and Tom40-specific antibodies. Bottom panel, quantification of three independent experiments, including SD. PoS, Ponceau S staining.
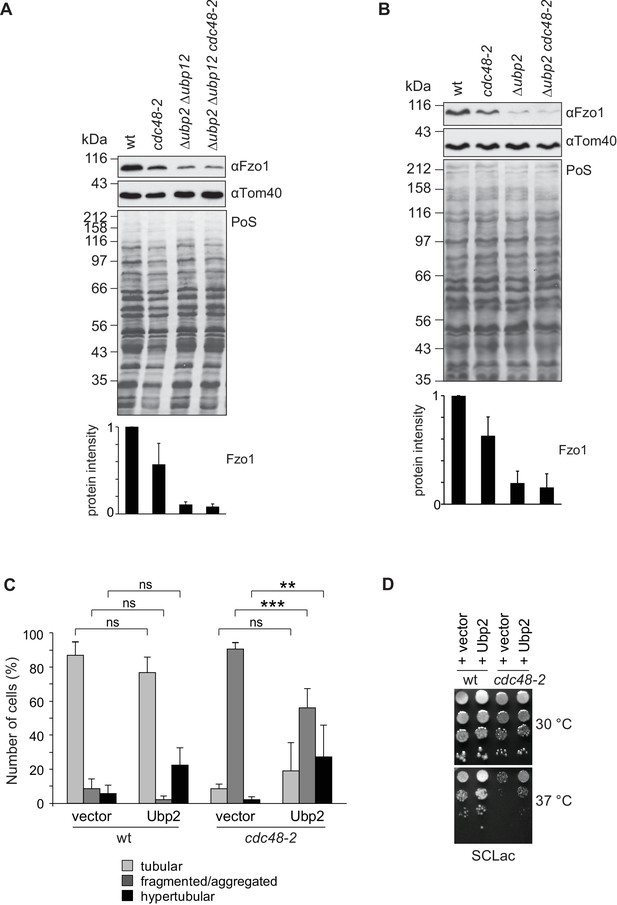
Cdc48 regulates mitochondrial fusion via Ubp12 and Ubp2.
(A) Steady state levels of Fzo1 in Δubp2 Δubp12 upon mutation of CDC48. Total cellular extracts of wt, cdc48-2, Δubp2 Δubp12 and Δubp2 Δubp12 cdc48-2 cells were analyzed by SDS-PAGE and immunoblotting using Fzo1- and Tom40-specific antibodies. Bottom panel, quantification of five independent experiments, including SD. (B) Steady state levels of Fzo1 in Δubp2 cells upon deletion of CDC48. Total cellular extracts of wt, cdc48-2, Δubp2 and Δubp2 cdc48-2 cells were analyzed by SDS-PAGE and immunoblotting using Fzo1- and Tom40-specific antibodies. Bottom panel, quantification of five independent experiments, including SD. (C) Mitochondrial morphology of cdc48-2 cells upon overexpression of Ubp2. Wt or cdc48-2 mutant cells expressing Ubp2 or the corresponding empty vector were analyzed for mitochondrial tubulation after expressing a mitochondrial-targeted GFP plasmid, as in Figure 1A. Quantification from three different experiments (with more than 200 cells each), including SE, as described (Cumming et al., 2007). ns, p>0.05. **p≤0.01, ***p≤0.001 (One-way ANOVA, Tukey’s multiple comparison test). (D) Role of Ubp2 overexpression on the respiratory capacity of CDC48-deficient cells. A spot assay was performed as described in Figure 4B with the indicated cells but using synthetic media supplemented with lactate (SCLac) and incubated for 4 days. PoS, Ponceau S staining.

Cdc48 regulates mitochondrial fusion via Ubp12 and Ubp2.
Physical interaction between Ubp2 and Cdc48. The catalytically inactive variant Ubp2C745S-Flag or the corresponding empty vector were expressed in Δubp12 cells and analyzed for Cdc48 interaction, as in 2A. Crude mitochondrial extracts were lysed and Flag-tagged Ubp2C745S was precipitated using Flag-coupled beads. The eluate was analyzed by SDS-PAGE and immunoblotting using Flag- and Cdc48-specific antibodies. PoS, Ponceau S staining; IP, immunoprecipitation; WB, western blot.
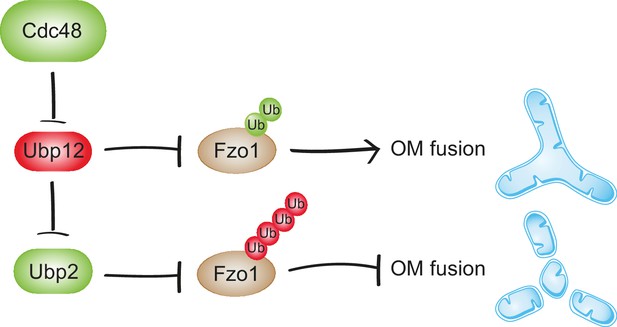
Synergistic regulation of mitochondrial fusion by the Cdc48 cascade.
Cdc48 supports turnover of Ubp12, stabilizing ubiquitylation on Fzo1 that promotes mitochondrial fusion (green ubiquitins). Moreover, degradation of Ubp12 stabilizes Ubp2, facilitating the removal of ubiquitin chains on Fzo1 inhibiting mitochondrial fusion (red ubiquitins). Thereby, Cdc48 activates mitochondrial fusion via Ubp12 and Ubp2. In contrast, Cdc48 impairment blocks progression of mitochondrial fusion by actively preventing Ubp12 turnover. Ubp12 then leads to a cascade of events inhibiting mitochondrial fusion: A) removal of the pro-fusion ubiquitylated forms and B) inhibition of Ubp2, consequently leading to the accumulation of the anti-fusion ubiquitylated forms. This cascade allows a synergistic effect of Cdc48, via a DUB regulatory cascade, to effectively promote or inhibit mitochondrial fusion.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| strain (Saccharomyces cerevisiae) | ∆fzo1 | PMID: 9483801 | Escobar_lab_stock_number: FA2 | |
| strain (S. cerevisiae) | cdc48-1 | PMID: 21441928 | Escobar_lab_stock_number: FA230 | |
| strain (S. cerevisiae) | cdc48-2 | PMID: 21441928 | Escobar_lab_stock_number: FA231 | |
| strain (S. cerevisiae) | cdc48-3 | PMID: 21441928 | Escobar_lab_stock_number: FA232 | |
| strain (S. cerevisiae) | ∆ubp2 | PMID: 9483801 | Escobar_lab_stock_number: FA260 | |
| strain (S. cerevisiae) | ∆ubp12 | PMID: 9483801 | Escobar_lab_stock_number: FA269 | |
| strain (S. cerevisiae) | ∆fzo1 ∆ubp2 | PMID: 23317502 | Escobar_lab_stock_number: FA362 | |
| strain (S. cerevisiae) | ∆ubp2 ∆ubp12 | PMID: 23317502 | Escobar_lab_stock_number: FA382 | |
| strain (S. cerevisiae) | ∆ubp12 ∆mdm30 | this study | Escobar_lab_stock_number: FA390 | UBP12::kanMX4; MDM30::kanMX4;obtained by crossing |
| strain (S. cerevisiae) | HA-Fzo1int in wt | PMID: 23317502 | Escobar_lab_stock_number: FA407 | |
| strain (S. cerevisiae) | HA-Fzo1int in ∆ubp2 | PMID: 23317502 | Escobar_lab_stock_number: FA415 | |
| strain (S. cerevisiae) | HA-Fzo1int in ∆ubp2 ∆mdm30 | PMID: 23317502 | Escobar_lab_stock_number: FA427 | |
| strain (S. cerevisiae) | ∆fzo1 ∆ubp12 | this study | Escobar_lab_stock_number: FA432 | FZO1::kanMX4; UBP12::kanMX4;obtained by crossing |
| strain (S. cerevisiae) | HA-Fzo1-K464Rint in wt | this study | Escobar_lab_stock_number: FA451 | HA-Fzo1K464R genomically integrated with NatNT2 into RS140 |
| strain (S. cerevisiae) | wt (BY4741) | PMID: 9483801 | Escobar_lab_stock_number: RS140 | |
| strain (S. cerevisiae) | cdc48-2 ∆fzo1 | this study | Escobar_lab_stock_number: RS430 | FZO1::natNT2 in FA231 |
| strain (S. cerevisiae) | cdc48-2 ∆ubp12 | this study | Escobar_lab_stock_number: RS466 | FZO1::hphNT1 in FA231 |
| strain (S. cerevisiae) | cdc48-2 ∆ubp2 ∆ubp12 | this study | Escobar_lab_stock_number: RS499 | UBP12::natNT2; UBP2::hphNT1 in FA231 |
| strain (S. cerevisiae) | ∆doa1 | PMID: 9483801 | Escobar_lab_stock_number: RS518 | |
| strain (S. cerevisiae) | ∆pdr5 ∆snq2 | other | Escobar_lab_stock_number: RS527 | gift by J. Dohmen (YGA58): MATa, ADE2 his3-D200 leu2-3,112 lys2-801, trp1D63 ura3-52 PDR5::hphNT1 SNQ2::kanMX4 |
| strain (S. cerevisiae) | Ubp12-Flagint in cdc48-2 | this study | Escobar_lab_stock_number: RS546 | Ubp12-Flag genomically integrated with NatNT2 into FA231 |
| strain (S. cerevisiae) | Ubp12-Flagint in wt | this study | Escobar_lab_stock_number: RS547 | Ubp12-Flag genomically integrated with NatNT2 into BY4741 |
| strain (S. cerevisiae) | ∆pdr5 ∆snq2 | this study | Escobar_lab_stock_number: RS554 | PDR5::NatNT2; SNQ2::hphNT1 in RS140 |
| strain (S. cerevisiae) | ∆fzo1 ∆dnm1 ∆ubp12 | this study | Escobar_lab_stock_number: RS556 | UBP12::NatNT2 in TS1028 |
| strain (S. cerevisiae) | ∆pdr5 ∆snq2 cdc48-2 | this study | Escobar_lab_stock_number: RS559 | PDR5::NatNT2; SNQ2::hphNT1 in FA231 |
| strain (S. cerevisiae) | cdc48-2 ∆ubp2 | this study | Escobar_lab_stock_number: TS686 | UBP2::hphNT1 in FA231 |
| strain (S. cerevisiae) | ∆fzo1 ∆dnm1 | other | Escobar_lab_stock_number: TS1028 | gift by B. Westermann (SB95): FZO1::kanMX4; DNM1::kanMX4; obtained by crossing |
| strain (S. cerevisiae) | wt (DF5) | PMID: 11007476 | Escobar_lab_stock_number: TS1124 | |
| strain (S. cerevisiae) | ufd1-2 | PMID: 11847109 | Escobar_lab_stock_number: TS1125 | |
| strain (S. cerevisiae) | npl4-1 | PMID: 8930904 | Escobar_lab_stock_number: TS1126 | |
| strain (S. cerevisiae) | Ubp2-9Mycint in wt | this study | Escobar_lab_stock_number: TS1134 | Ubp2-9Myc genomically integrated with NatNT2 into RS140 |
| strain (S. cerevisiae) | Ubp2-3HAint in wt | this study | Escobar_lab_stock_number: TS1144 | Ubp2-3HA genomically integrated with hphNT1 in RS140 |
| strain (S. cerevisiae) | Ubp2-3HAint in ∆ubp12 | this study | Escobar_lab_stock_number: TS1147 | Ubp2-3HA genomically integrated with hphNT1 in FA269 |
| strain (S. cerevisiae) | pGAL-Ubp12-Flagint in wt | this study | Escobar_lab_stock_number: TS1153 | pGAL-Ubp12-Flag genomically integratedwith kanMX4 into RS544 |
| recombinant DNA reagent | pRS316 (plasmid) | PMID: 2659436 | Escobar_lab_stock_number: p8 | |
| recombinant DNA reagent | HA-Fzo1 on pRS316 (plasmid) | PMID: 23317502 | Escobar_lab_stock_number: p10 | |
| recombinant DNA reagent | HA-Fzo1-K464R on pRS316 (plasmid) | PMID: 23317502 | Escobar_lab_stock_number: p14 | |
| recombinant DNA reagent | YEplac181 (plasmid) | PMID: 3073106 | Escobar_lab_stock_number: p58 | |
| recombinant DNA reagent | Ubp2-Flag on YEplac181(plasmid) | PMID: 23317502 | Escobar_lab_stock_number: p59 | |
| recombinant DNA reagent | Ubp2-C745S-Flag on YEplac181(plasmid) | PMID: 23317502 | Escobar_lab_stock_number: p60 | |
| recombinant DNA reagent | Ubp12-Flag on YEplac181(plasmid) | PMID: 23317502 | Escobar_lab_stock_number: p61 | |
| recombinant DNA reagent | Ubp12-C372S-Flag on YEplac181(plasmid) | PMID: 23317502 | Escobar_lab_stock_number: p62 | |
| recombinant DNA reagent | YEplac195 (plasmid) | PMID: 3073106 | Escobar_lab_stock_number: p63 | |
| recombinant DNA reagent | Ubp12C372S on YEplac195 (plasmid) | this study | Escobar_lab_stock_number: p65 | Ubp12C372S (non-tagged) on YEplac195, 2µ, Ura3 |
| recombinant DNA reagent | mt-GFP on pYX142 (plasmid) | PMID: 11054823 | Escobar_lab_stock_number: p70 | |
| recombinant DNA reagent | Cdc48 wt on pRS313 (plasmid) | PMID: 22580068 | Escobar_lab_stock_number: p75 | |
| recombinant DNA reagent | pRS313 (plasmid) | PMID: 2659436 | Escobar_lab_stock_number: p79 | |
| recombinant DNA reagent | Cdc48-A547T on pRS313 (plasmid) | this study | Escobar_lab_stock_number: p150 | Cdc48A547T on pRS313, cen, His3 |
| recombinant DNA reagent | Ub on pKT10 (plasmid) | PMID: 2164637 | Escobar_lab_stock_number: p341 | |
| recombinant DNA reagent | Ub-K48R on pKT10 (plasmid) | PMID: 2164637 | Escobar_lab_stock_number: p342 | |
| recombinant DNA reagent | Ub-K63R on pKT10 (plasmid) | PMID: 2164637 | Escobar_lab_stock_number: p343 | |
| recombinant DNA reagent | Ub-K48R,K63R on pKT10 (plasmid) | PMID: 2164637 | Escobar_lab_stock_number: p344 | |
| recombinant DNA reagent | Myc-Ub on pRS426 (plasmid) | PMID: 25620559 | Escobar_lab_stock_number: p356 | |
| recombinant DNA reagent | pRS426 (plasmid) | PMID: 25620559 | Escobar_lab_stock_number: p375 | |
| Antibody | anti-Cdc48 | other | gift by T. Sommer; (1:1,000/1:10,000) | |
| Antibody | anti-Cox2 | other | gift by W. Neupert; (1:5,000) | |
| Antibody | anti-Flag M2 | Sigma | Sigma: F1804 | (1:1,000) |
| Antibody | anti-Fzo1 | this study | Produced by GenScript using the peptide CHGDRKPDDDPYSSS; (1:1,000) | |
| Antibody | anti-HA | Roche | Roche: 11867423001 | (1:1,000) |
| Antibody | anti-Myc | Cell Signaling | Cell_Signaling: #2276 | (1:1,000) |
| Antibody | anti-Sec61 | other | gift by T. Sommer; (1:10,000) | |
| Antibody | anti-Ssc1 | Fölsch et al., 1998 | (1:40,000) | |
| Antibody | anti-Tom40 | other | gift by W. Neupert; (1:40,000) | |
| Antibody | anti-Tpi1 | other | gift by J. Dohmen; (1:5,000) | |
| Antibody | anti-Ub (P4D1) | Cell Signaling | Cell_Signaling: #3936 | (1:1,000) |
| Antibody | anti-Ubc6 | other | gift by T. Sommer; (1:10,000) | |
| Antibody | anti-Ubp12 | this study | (1:200) | |
| software | Microsoft Office 2010 | Micosoft Corporation | ||
| software | Adobe Photoshop CS6 | Adobe | ||
| software | Adobe Illustrator CS6 | Adobe | ||
| software | Clone Manager | Sci-Ed Software | ||
| software | Image Quant | GE Healthcare Life Sciences | ||
| software | Axiovision | Zeiss | ||
| software | StepOne System | Thermo Fisher Scientific | ||
| kit | NucleoSpin RNA | Machery Nagel | REF:740955 | |
| kit | SuperScript III First-Strand Synthesis System | Invitrogen | Catalogue_number:18080051 |
Yeast strains used in this study.
https://doi.org/10.7554/eLife.30015.022| Strain # | Strain name | Genotype | Reference |
|---|---|---|---|
| FA2 | ∆fzo1 | FZO1::kanMX4 in BY4741 | Brachmann et al., 1998 |
| FA230 | cdc48-1 | cdc48-1::KanMX4 in BY4741 | Li et al. (2011) |
| FA231 | cdc48-2 | cdc48-2::KanMX4 in BY4741 | Li et al. (2011) |
| FA232 | cdc48-3 | cdc48-3::KanMX4 in BY4741 | Li et al. (2011) |
| FA260 | ∆ubp2 | UBP2::kanMX4 in BY4741 | Brachmann et al., 1998 |
| FA269 | ∆ubp12 | UBP12::kanMX4 in BY4741 | Brachmann et al., 1998 |
| FA362 | ∆fzo1 ∆ubp2 | FZO1::kanMX4; UBP2::kanMX4; obtained by crossing | Anton et al. (2013) |
| FA382 | ∆ubp2 ∆ubp12 | UBP12::kanMX4; UBP2::kanMX4; obtained by crossing | Anton et al. (2013) |
| FA390 | ∆ubp12 ∆mdm30 | UBP12::kanMX4; MDM30::kanMX4; obtained by crossing | this study |
| FA407 | HA-Fzo1int in wt | HA-Fzo1 genomically integrated with NatNT2 into RS140 | Anton et al. (2013) |
| FA415 | HA-Fzo1int in ∆ubp2 | HA-Fzo1 genomically integrated with NatNT2 into FA260 | Anton et al. (2013) |
| FA427 | HA-Fzo1int in ∆ubp2 ∆mdm30 | HA-Fzo1 genomically integrated with NatNT2 into ∆ubp2 ∆mdm30 | Anton et al. (2013) |
| FA432 | ∆fzo1 ∆ubp12 | FZO1::kanMX4; UBP12::kanMX4; obtained by crossing | this study |
| FA451 | HA-Fzo1-K464Rint in wt | HA-Fzo1K464R genomically integrated with NatNT2 into RS140 | this study |
| RS140 | wt | BY4741; S288C isogenic yeast strain; MATa, his3Δ1, leu2Δ0, met15Δ0, ura3Δ0 | Brachmann et al., 1998 |
| RS430 | cdc48-2 ∆fzo1 | FZO1::natNT2 in FA231 | this study |
| RS466 | cdc48-2 ∆ubp12 | FZO1::hphNT1 in FA231 | this study |
| RS499 | cdc48-2 ∆ubp2 ∆ubp12 | UBP12::natNT2; UBP2::hphNT1 in FA231 | this study |
| RS518 | ∆doa1 | DOA1::kanMX4 in BY4741 | Brachmann et al., 1998 |
| RS527 | ∆pdr5 ∆snq2 | MATa, ADE2 his3-D200 leu2-3,112 lys2-801, trp1D63 ura3-52 PDR5::hphNT1 SNQ2::kanMX4 | J. Dohmen (YGA58) |
| RS546 | Ubp12-Flagint in cdc48-2 | Ubp12-Flag genomically integrated with NatNT2 into FA231 | this study |
| RS547 | Ubp12-Flagint in wt | Ubp12-Flag genomically integrated with NatNT2 into BY4741 | this study |
| RS554 | ∆pdr5 ∆snq2 | PDR5::NatNT2; SNQ2::hphNT1 in RS140 | this study |
| RS556 | ∆fzo1 ∆dnm1 ∆ubp12 | UBP12::NatNT2 in TS1029 | this study |
| RS559 | ∆pdr5 ∆snq2 cdc48-2 | PDR5::NatNT2; SNQ2::hphNT1 in FA231 | this study |
| TS686 | cdc48-2 ∆ubp2 | UBP2::hphNT1 in FA231 | this study |
| TS1029 | ∆fzo1 ∆dnm1 | FZO1::kanMX4; DNM1::kanMX4; Mat α, BY background, obtained by crossing | B. Westermann (#94) |
| TS1124 | wt (DF5) | MATα, trp1-1(am), ura3-52, his3∆200, leu2-3, lys2-801 | Hoppe et al. (2000) |
| TS1125 | ufd1-2 | ufd1-2ts in TS1124 | Braun et al. (2002) |
| TS1126 | npl4-1 | npl4-1ts in TS1124 | DeHoratius and Silver (1996) |
| TS1134 | Ubp2-9Mycint in wt | Ubp2-9Myc genomically integrated with NatNT2 into RS140 | this study |
| TS1144 | Ubp2-3HAint in wt | Ubp2-3HA genomically integrated with hphNT1 in RS140 | this study |
| TS1147 | Ubp2-3HAint in ∆ubp12 | Ubp2-3HA genomically integrated with hphNT1 in FA269 | this study |
| TS1153 | pGAL-Ubp12-Flagint in wt | pGAL-Ubp12-Flag genomically integrated with kanMX4 into RS544 | this study |
Plasmids used in this study.
https://doi.org/10.7554/eLife.30015.023| Plasmid # | Plasmid name | Description | Bacterial selection | Reference |
|---|---|---|---|---|
| 8 | pRS316 | pRS316, cen, Ura3 | Amp | Sikorski and Hieter, 1989 |
| 10 | HA-Fzo1 on pRS316 | HA-Fzo1 on pRS316, Fzo1 prom, cen, Ura3 | Amp | Anton et al. (2013) |
| 14 | HA-Fzo1-K464R on pRS316 | HA-Fzo1K464R on pRS316, Fzo1 prom, cen, Ura3 | Amp | Anton et al. (2013) |
| 58 | YEplac181 | YEplac181, 2µ, Leu2 | Amp | Gietz and Sugino, 1988 |
| 59 | Ubp2-Flag on YEplac181 | Ubp2-Flag on YEplac181, Adh1 prom, 2µ, Leu2 | Amp | Anton et al. (2013) |
| 60 | Ubp2-C745S-Flag on YEplac181 | Ubp2C745S-Flag on YEplac181, Adh1 prom, 2µ, Leu2 | Amp | Anton et al. (2013) |
| 61 | Ubp12-Flag on YEplac181 | Ubp2-Flag on YEplac181, Adh1 prom, 2µ, Leu2 | Amp | Anton et al. (2013) |
| 62 | Ubp12-C372S-Flag on YEplac181 | Ubp2C372S-Flag on YEplac181, Adh1 prom, 2µ, Leu2 | Amp | Anton et al. (2013) |
| 63 | YEplac195 | YEplac195, 2µ, Ura3 | Amp | Gietz and Sugino, 1988 |
| 65 | Ubp12C372S on YEplac195 | Ubp12C372S (non-tagged) on YEplac195, 2µ, Ura3 | Amp | this study |
| 70 | mt-GFP on pYX142 | mt-GFP on pYX142, cen, Leu2 | Amp | Westermann and Neupert, 2000 |
| 75 | Cdc48 wt on pRS313 | Cdc48 wt on pRS313, cen, His3 | Amp | Esaki and Ogura (2012) |
| 79 | pRS313 | pRS313, cen, His3 | Amp | Sikorski and Hieter, 1989 |
| 150 | Cdc48-A547T on pRS313 | Cdc48A547T on pRS313, cen, His3 | Amp | this study |
| 341 | Ub on pKT10 | Ub on pK10, 2µ, Ura3 | Amp | Tanaka et al., 1990 |
| 342 | Ub-K48R on pKT10 | UbK48R on pK10, 2µ, Ura3 | Amp | Tanaka et al., 1990 |
| 343 | Ub-K63R on pKT10 | UbK63R on pK10, 2µ, Ura3 | Amp | Tanaka et al., 1990 |
| 344 | Ub-K48R,K63R on pKT10 | UbK48R,K63R on pK10, 2µ, Ura3 | Amp | Tanaka et al., 1990 |
| 356 | Myc-Ub on pRS426 | pCup1-Myc-Ub on pRS426, 2µ, Ura3 | Amp | Li et al., 2015 |
| 375 | pRS426 | pRS426, 2µ, Ura3 | Amp | Li et al., 2015 |
Antibodies used in this study.
https://doi.org/10.7554/eLife.30015.024| Name | Dilution | Reference |
|---|---|---|
| Cdc48 | 1:1000/1:10,000 | T. Sommer |
| Cox2 | 1:5000 | W. Neupert |
| Flag M2 | 1:1000 | Sigma (F1804) |
| Fzo1 | 1:1000 | this study |
| HA | 1:1000 | Roche (11867423001) |
| Myc | 1:1000 | Cell Signaling (#2276) |
| Sec61 | 1:10,000 | T. Sommer |
| Ssc1 | 1:40,000 | Fölsch et al., 1998 |
| Tom40 | 1:40,000 | W. Neupert |
| Tpi1 | 1:5000 | J. Dohmen |
| Ub (P4D1) | 1:1000 | Cell Signaling (#3936) |
| Ubc6 | 1:10,000 | T. Sommer |
| Ubp12 | 1:200 | this study |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.30015.025





