Incomplete inhibition of HIV infection results in more HIV infected lymph node cells by reducing cell death
Figures
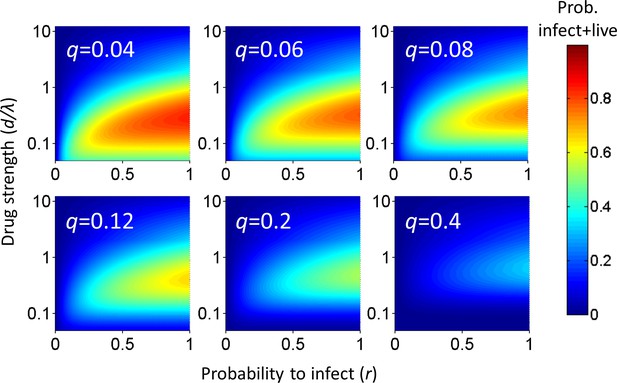
Probability for a cell to be infected and live as a function of inhibitor.
Probability for a cell to be infected and live was calculated for 20 infection attempts (λ) and represented as a heat map. Drug strength (d/λ) is on the y-axis, and the probability per infection attempt to infect (r) is on the x-axis. Each plot is the calculation for one value of the probability per infection attempt to die (q) denoted in white in the top left corner.
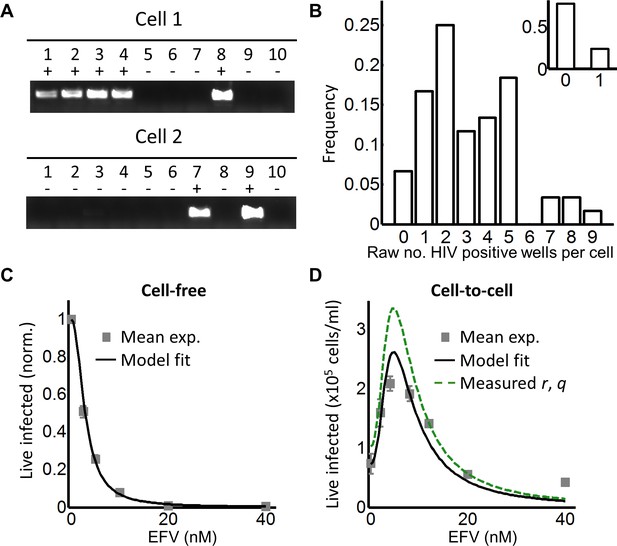
Partial inhibition increases the number of live infected cells.
(A) To quantify HIV DNA copy number per cell, GFP-positive cells were sorted into individual wells and lysed. Each lysate was subdivided into 10 wells and PCR performed to detect HIV DNA, with the sum of positive wells being the raw HIV copy number for that cell. (B) Histogram of raw HIV DNA copies per cell in coculture infection (n = 60 cells, four independent experiments). Inset shows raw HIV DNA copies per cell in cell-free infection (n = 30, two independent experiments) (C) Number of live infected cells normalized by maximum number of live infected cells in cell-free infection with EFV. Black line is best-fit for EFV suppression of cell-free infection (IC50 = 2.9 nM, h = 2.1). Means and standard errors for three independent experiments. (D) Number of live infected cells/ml 2 days post-infection resulting from coculture infection of 106 cells/ml in the presence of EFV. Means and standard errors for three independent experiments. Black line is best-fit of Equation (3) with r = 0.22 and q = 0.17. Dashed green line is the result of Equation (3) with experimentally measured r = 0.28, q = 0.15.
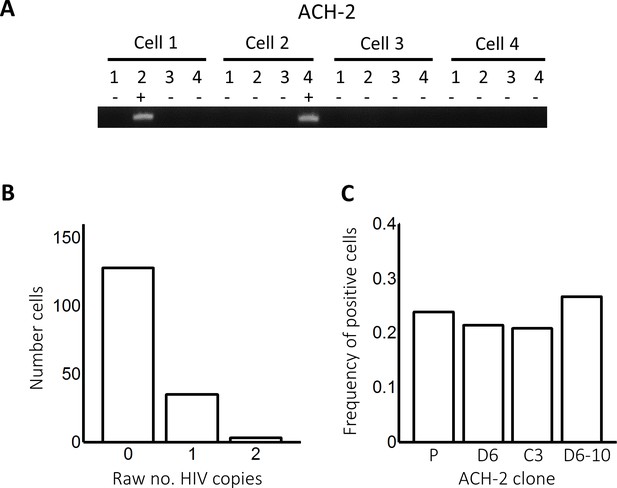
Detected integrations in ACH-2 cells.
(A) To quantify HIV DNA copy number per cell in ACH-2 cells which contain single inactive integrations of HIV, cells were sorted into individual wells of a multi-well plate at 1 cell per well and lysed. Each lysed cell was further subdivided into four wells and nested PCR performed to a region of the reverse transcriptase gene to detect whether the well contained a copy of HIV DNA. A total of 166 cells were tested. (B) Histogram of the raw number of HIV DNA copies per cell. (C) Histogram of the frequency of ACH-2 cells positive for HIV DNA copies on the parental ACH-2 population, two subclones of the parental line, and one of the subclones split over 10 wells. P, parental (n = 71 cells); D6, subclone D6 (n = 56); C3, subclone C3 (n = 24), and D6-10, subclone D6 with each lysed cell split over 10 wells (n = 15).
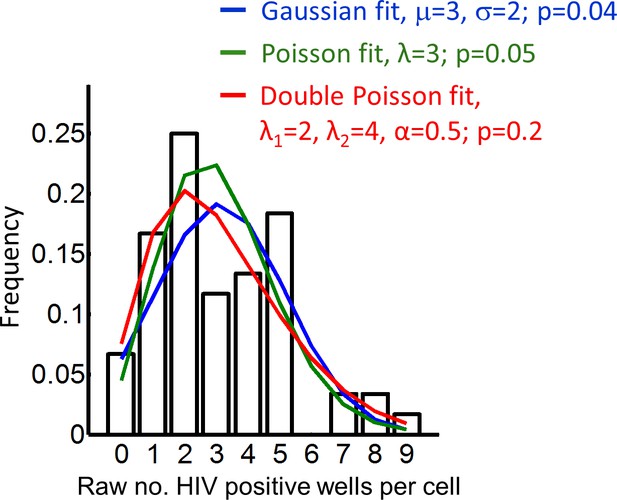
Fitting of different distributions to the frequency of HIV DNA copies per cell.
The experimentally determined distribution of the raw number of HIV-positive wells per cell was fitted by a Gaussian distribution (blue line), a Poisson distribution (green line), or a two-peaked Poisson distribution (red line) described by p(m; α,λ1,λ2)=(1-α)λ1me-λ1/m!+αλ2me-λ2/m!, where λ1 and λ2 are the means of the first and second peak, respectively, and α is the fraction of transmissions which fall within the second peak. Only the double Poisson distribution could not be rejected using a chi-square goodness-of-fit test (p=0.2).
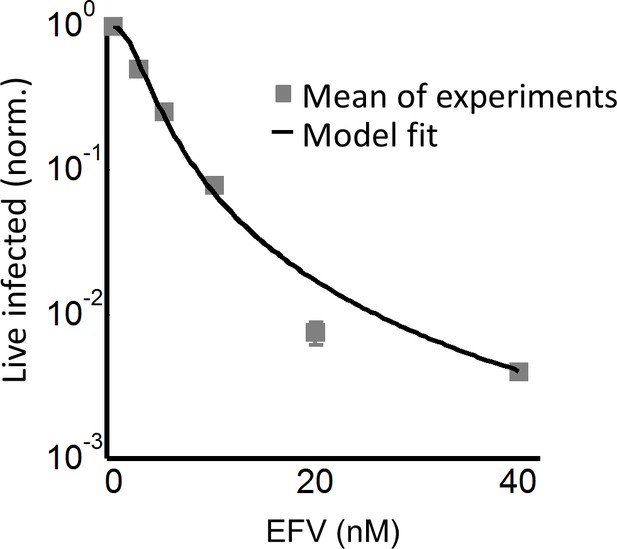
The number of live infected cells in cell-free infection with wild-type HIV.
Live infected cells are normalized by the maximum number of live infected cells in cell-free infection as a function of EFV. Black line represents best-fit for the effect of EFV on infection with single infection attempts, with IC50 = 2.4 nM and Hill coefficient of 1.8. Shown are means and standard errors for three independent experiments. Y-axis shown on a log scale to better view fits at low live infected cell numbers.
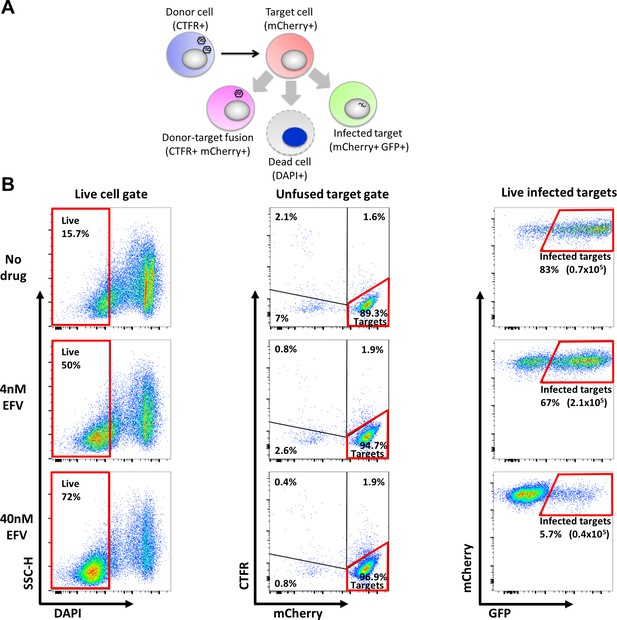
Gating strategy for coculture infection with wild type HIV.
(A) Possible outcomes of donor cell-target cell interactions are target cell infection, fusion, and target cell death. Each outcome is traced by fluorescent markers for donor cells (Cell Trace Far Red, CTFR), target cells (mCherry), infected cells (GFP) and death (DAPI). Live infected target cells were DAPI negative and mCherry and GFP positive. Donor-target fusions were excluded by excluding all CTFR-positive cells. (B) Representative flow cytometry scatter plots used in gating strategy. First, live cells were selected using the absence of DAPI fluorescence (left column). Next, target cells were separated from donor cells and donor-target fusions using CTFR and mCherry (middle column). Last, the number of infected targets was determined by GFP fluorescence (right column). Percentages represent fraction of the population in the gate and numbers in parentheses represent the number of infected cells/ml. Areas marked in red represent selection gates for live infected targets.
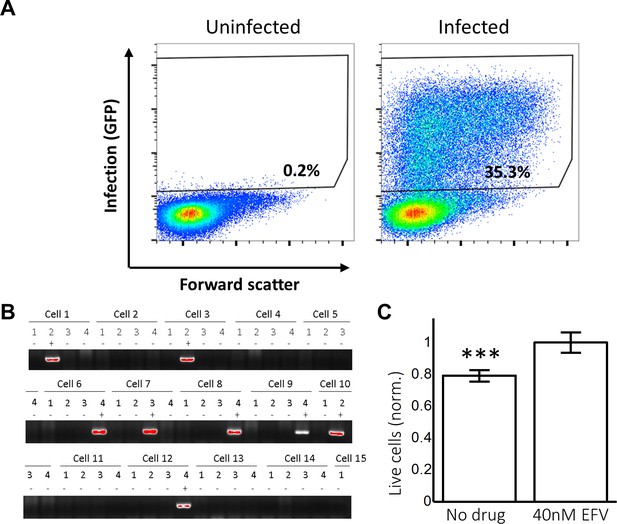
Experimental measurement of r and q.
(A) The probability of a cell to be infected and live (Pλ) as measured by the fraction of live cells which express GFP 2 days post-infection. Left plot shows uninfected control and right plot shows infected cells. Results from one of two independent experiments. Maximum GFP expression in this reporter system with highly concentrated virus was 70%. (B) The number of HIV DNA copies per cell 1 day post cell-free HIV infection for experiments in (A). First 14 cells from one of two independent experiments are shown. (C) To determine the fraction of live cells Lλ remaining after infection, cells were serum starved to prevent cell division. The concentration of live cells 2 days post cell-free infection was measured using trypan blue exclusion and divided by the concentration of live reporter cells in a separate cell culture infected with the identical amount of virus, but with infection was inhibited with 40 nM EFV. Results are of two independent experiments, with 10 fields of view counted per condition. ***p=7×10−3 by two tailed t-test.
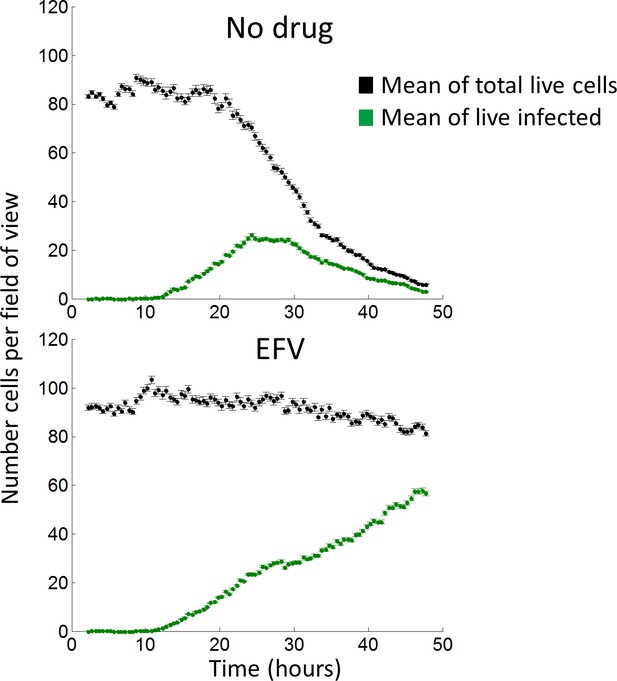
Time-lapse microscopy of HIV infection in the absence and presence of EFV.
Infection was performed by an input of 20% infected mCherry-negative donor cells. Black points represent the number of total live cells, as detected by the presence of cellular mCherry fluorescence. Green points represent the number of live infected target cells, as detected by co-expression of mCherry and HIV-dependent GFP. Top graph shows infection in the absence of drug and bottom graph in the presence of 30 nM EFV. Time-lapse interval was 20 min between frames, and three independent experiments were performed with 15 fields of view per condition in each experiment. The 45 fields of view from three independent experiments were averaged, and each point is the mean and standard error of the number of cells per field of view. For clarity, every second measurement is shown.
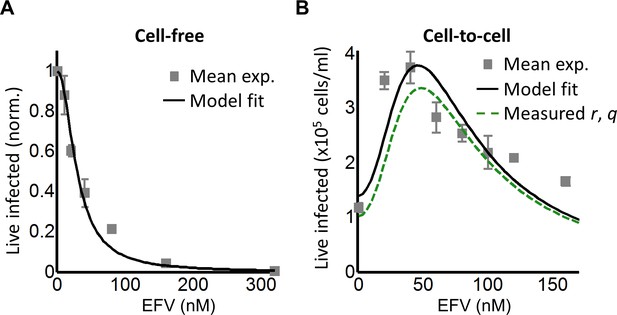
Partial inhibition of the EFV-resistant L100I mutant shifts the peak of live infected cells to higher EFV concentrations.
(A) The number of live infected cells normalized by the maximum number of live infected cells in cell-free infection as a function of EFV for the L100I mutant. Black line is best-fit for EFV suppression of cell-free infection (IC50 = 29 nM, h = 2.0). Shown are means and standard errors for three independent experiments. (B) Number of live infected cells/ml 2 days post-infection resulting from coculture infection of 106 cells/ml in the presence of EFV. Means and standard errors for three independent experiments. Black line is best-fit of Equation (3) with r = 0.29 and q = 0.13. Dashed green line is the result of Equation (3) with the experimentally measured r = 0.28 and q = 0.15 for wild-type HIV infection.
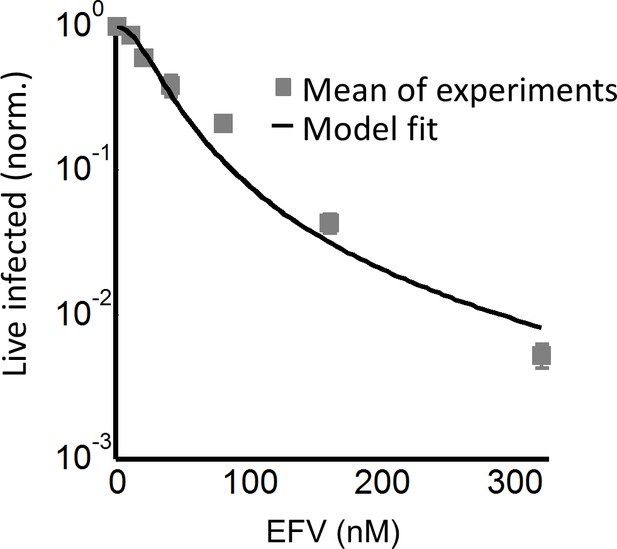
The number of live infected cells in cell-free infection with the L100I mutant.
Live infected cells are normalized by the maximum number of live infected cells in cell-free infection as a function of EFV. Black line represents best-fit for the effect of EFV on infection with single HIV infection attempts, with IC50 = 26.5 nM and Hill coefficient of 2.0. Shown are means and standard errors for three independent experiments.
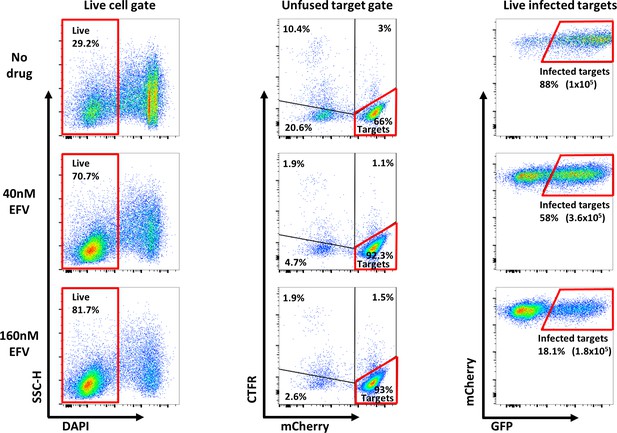
Gating strategy for coculture infection with mutant HIV.
Representative flow cytometry scatter plots used in gating strategy. First, live cells were selected using the absence of DAPI fluorescence (left column). Next, target cells were separated from donor cells and donor-target fusions using CTFR and mCherry (middle column). Last, the number of infected targets was determined by GFP fluorescence (right column). Percentages represent fraction of the population in the gate and numbers in parentheses shown in the right column represent the number of infected cells/ml. Areas marked in red represent selection gates for live infected targets.
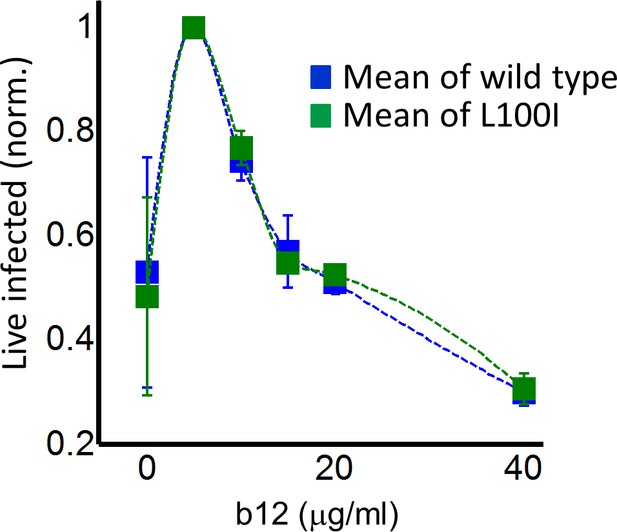
Partial inhibition of coculture infection with neutralizing antibody results in higher numbers of live infected cells.
Shown are the numbers of live infected cells normalized by the maximum number of live infected cells in coculture infection as a function of b12 antibody concentration. Infection was by either EFV-sensitive HIV (blue) or the L100I EFV-resistant mutant (green). Dashed lines are a guide to the eye. Shown are means and standard errors for three independent experiments.
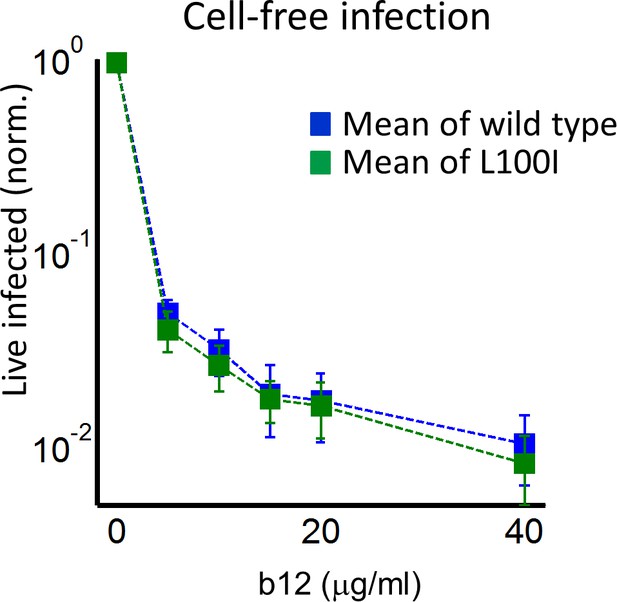
The number of live infected cells with cell-free infection in the face of neutralizing antibody b12.
Live infected cells are normalized by the maximum number of live infected cells in cell-free infection as a function of b12. Infection was by either EFV-sensitive HIV (blue) or the L100I EFV-resistant mutant (green). Dashed lines are a guide to the eye. Shown are means and standard errors for three independent experiments. Log scale on y-axis used for better resolution.
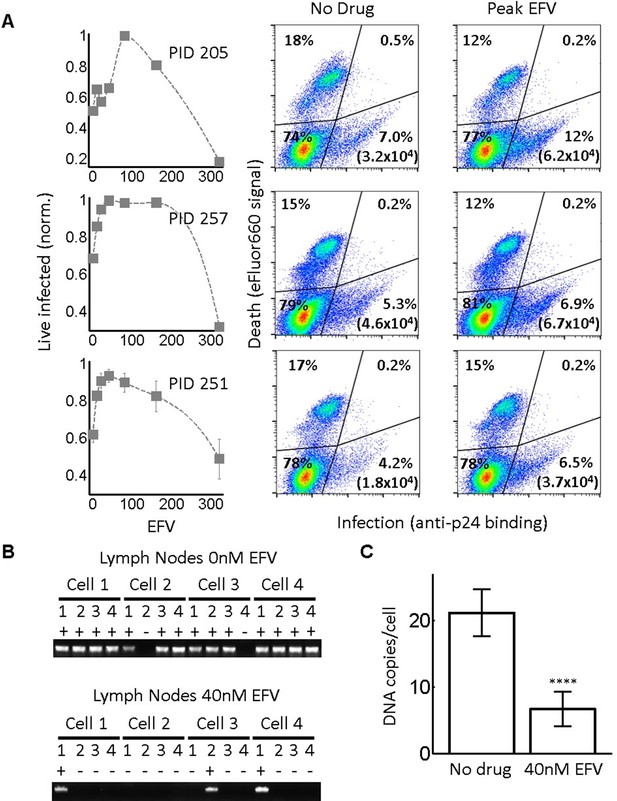
Infection optimum with EFV in lymph node cells.
(A) Number of live infected cells as a function of EFV. Each row shows in vitro infected lymph node cells from one participant. Left column is the number of live infected cells normalized by maximum number of live infected cells in coculture infection. Middle and right columns are flow cytometry dot plots of infection without drug and at the infection optimum, with HIV p24 on x-axis and death detection by eFluor660 on y-axis. Infected live cells are bottom right. Number in brackets represents live infected cell density per ml. PID, participant identification number. For PID205 and 257, cells were sufficient for one experiment. For PID251, means and standard errors for three independent experiments are shown. Dashed lines are a guide to the eye. (B) HIV DNA copy number per cell was quantified by sorting fixed p24-positive cells from PID251 into individual wells. Cells were lysed and de-crosslinking performed. Each lysate was divided into four wells and PCR performed to detect HIV DNA. First row is representative of cells with no drug, second row is representative of 40 nM EFV. (C) Mean and standard error of the number of HIV DNA copies without drug and with 40 nM EFV after assay sensitivity correction. N = 56 cells from three independent experiments for each condition. ****p=4×10−9, two tailed t-test.
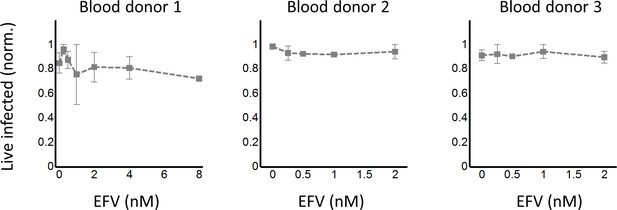
Cell-to-cell infection of peripheral blood mononuclear cells in the presence of EFV does not lead to a discernable peak in live infected cells.
The number of live infected cells was normalized by the maximum number of live infected cells after a 2-day coculture infection with wild-type HIV. For each donor, the means and standards errors are shown for three independent experiments. Dashed lines are a guide to the eye.
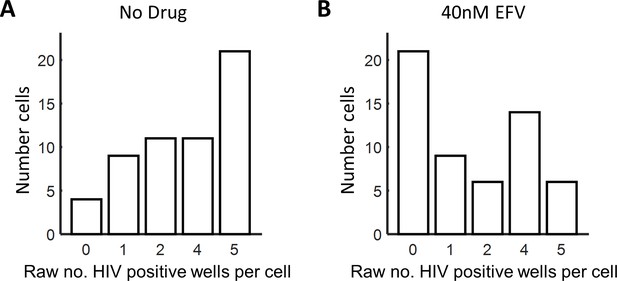
Raw HIV DNA copy numbers per lymph node cell.
Fixed, p24-positive cells from PID251 were sorted into individual wells of a multi-well plate at 1 cell per well and lysed. Each lysed cell was further divided into four wells and nested PCR performed on a region of the reverse transcriptase gene. Shown are histograms of the raw numbers of positive wells per cell. (A) No drug. (B) 40 nM EFV. Results are from three independent experiments.
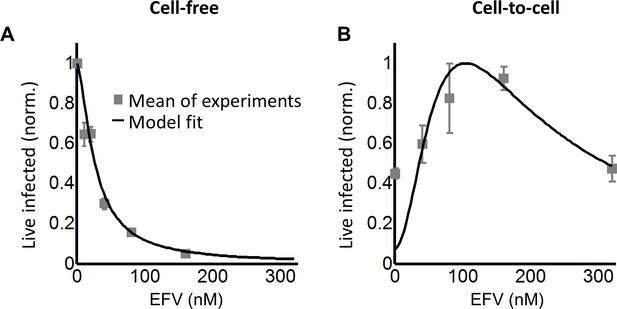
Infection with the K103N mutant shows an infection optimum at clinically observed lymph node EFV concentrations.
(A) The number of live infected cells normalized by the maximum number of live infected cells in cell-free infection as a function of EFV for the K103N mutant. Black line is best-fit for EFV suppression of cell-free infection (IC50 = 26 nM, h = 1.5). Shown are means and standard errors for two independent experiments using cells from PID251. (B) The number of live infected cells normalized by the maximum number of live infected cells in coculture infection as a function of EFV for the K103N mutant. Black line represents best-fit model for the effect of EFV on coculture infection according to Equation (3), with d values calculated based on the cell-free infection data for the K103N mutant, and the mean number of HIV DNA copies in the absence of drug determined for L100I. The fits recapitulated the experimental results when r = 0.91 and q = 0.15. Shown are means and standard errors for three independent experiments. There were sufficient lymph node cells from PID251 for two of the three experiments, and the third experiment was performed with lymph node cells from PID274.
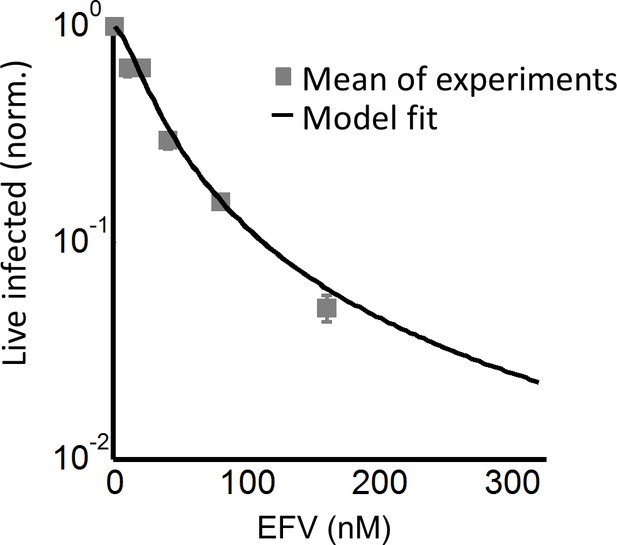
The number of live infected cells in cell-free infection with the K103N mutant.
Live infected cells are normalized by the maximum number of live infected cells in cell-free infection as a function of EFV. Black line represents best-fit model for the effect of EFV on infection with single infection attempts, with IC50 = 26.0 nM and Hill coefficient of 1.5. Shown are means and standard errors for three independent experiments.
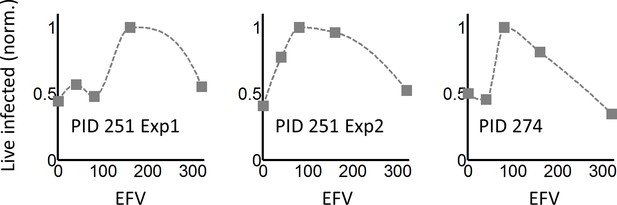
Response of K103N coculture infection to EFV in individual experiments.
Shown is the number of live infected cells normalized by the maximum number of live infected cells in coculture infection as a function of EFV for each experiment and study participant used. Two experiments per performed for lymph nodes resected from PID251, and one for PID274. Dashed lines are a guide to the eye.
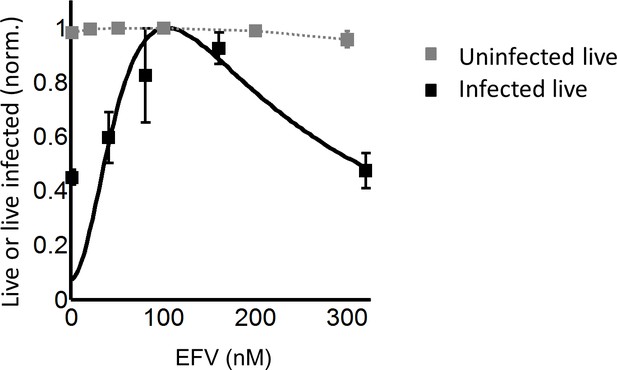
Effect of EFV on viability of uninfected lymph node cells from PID274.
The fraction of live cells at different EFV concentrations was normalized by the maximum fraction of live cells in the experiment. Shown are the means (grey squares) and standard errors for three independent experiments. Black squares are the normalized live K103N-infected cells as a function of EFV. Grey dotted line is a guide to the eye. Black line represents best-fit model for the effect of EFV on coculture infection according to Equation (3), with d values calculated based on the cell-free infection data for the K103N mutant, and the mean number of HIV DNA copies in the absence of drug determined for L100I. Best-fit parameters were r = 0.91 and q = 0.15.
Additional files
-
Supplementary file 1
S Table 1: Parameters and definitions.
- https://doi.org/10.7554/eLife.30134.023
-
Supplementary file 2
S Table 2: Measurement of r and q
- https://doi.org/10.7554/eLife.30134.024
-
Supplementary file 3
S Table 3: Participant information.
- https://doi.org/10.7554/eLife.30134.025
-
Source code 1
script1r2.m: Matlab source code for Figure 1.
- https://doi.org/10.7554/eLife.30134.026
-
Source code 2
script2r2: Matlab source code for copy number correction for RevCEM clones based on HIV copy detection efficiency in ACH-2 cells.
- https://doi.org/10.7554/eLife.30134.027
-
Source code 3
script3r2.m: Matlab source code for cell-free and coculture infection model fits for RevCEM cells infected with wild-type HIV in Figure 2.
- https://doi.org/10.7554/eLife.30134.028
-
Source code 4
Script4.m: Matlab source code for histograms of raw HIV copy numbers for cell-free and coculture infection and fits of different distributions to coculture copy number frequencies.
- https://doi.org/10.7554/eLife.30134.029
-
Source code 5
Script5.m: Matlab source code for image analysis of time-lapse data for Figure 2—figure supplement 5.
- https://doi.org/10.7554/eLife.30134.030
-
Source code 6
script6r2.m: Matlab source code for cell-free and coculture infection model fits for RevCEM cells infected with L100I mutant HIV in Figure 3.
- https://doi.org/10.7554/eLife.30134.031
-
Source code 7
Script7.m: Matlab source code for copy number correction for lymph node cells based on HIV copy detection efficiency is ACH-2 cells.
- https://doi.org/10.7554/eLife.30134.032
-
Source code 8
script8r2.m: Matlab source code for cell-free and coculture infection model fits for lymph node cells infected with K103N mutant HIV in Figure 6.
- https://doi.org/10.7554/eLife.30134.033
-
Transparent reporting form
- https://doi.org/10.7554/eLife.30134.034





