Automated cell-type classification in intact tissues by single-cell molecular profiling
Figures
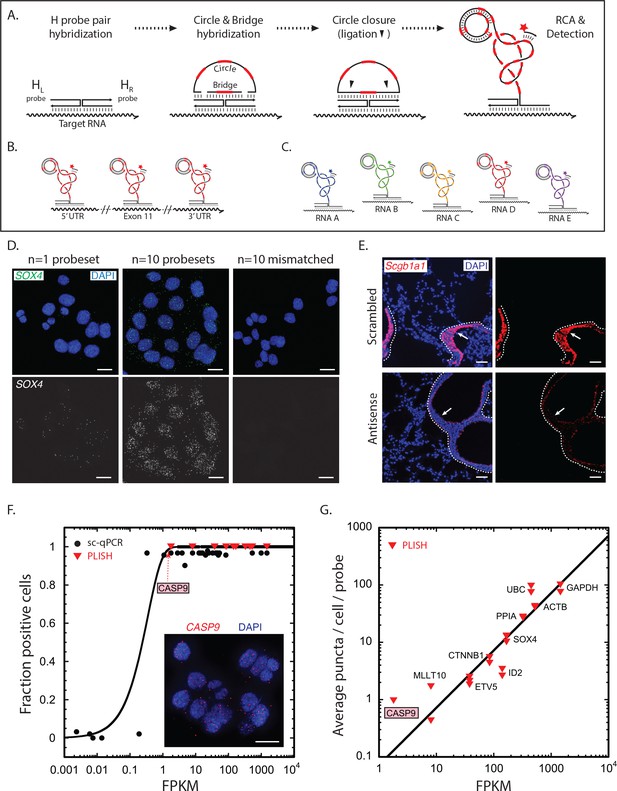
RNA detection by proximity ligation at RNA-DNA Holliday junctions.
(A) Mechanism of RNA detection by PLISH. Left (HL) and right (HR) DNA 'H' probes targeting adjacent sites hybridize to a target RNA. Subsequent addition of circle and bridge oligonucleotides harboring a specific 'barcode' sequence (red dash) results in an RNA-DNA Holliday junction. The nicks in the junction are then sealed by ligation to create a covalently closed circle, and rolling-circle amplification (RCA) generates complementary tandem repeats. The single-stranded amplicons are detected with a fluorescently-labeled 'imager' oligonucleotide (red star) that is complementary to the specific 'barcode' sequence. See note in 1C. (B) To increase efficiency of detection for low abundance transcripts, H probe pairs embedded with the same barcode sequence can be 'tiled' along the length of the target mRNA. (C) Up to five distinct transcripts can be simultaneously detected using five different barcode sequences (one unique sequence for each RNA), and five complementary imager oligonucleotides that are conjugated to spectrally-distinct fluorophores. (D) PLISH RNA detection requires coincident hybridization of both left and right H probes, and redundant probesets increase detection efficiency. Cultured HCT116 cells were stained with one (left panel) or ten (middle panel) pairs of H probes targeting the SOX4 transcript. Many more RNA molecules are detected with ten probesets. No puncta are observed when the RNA-recognition sequences of the left H probes are scrambled (right panel). SOX4, SRY-box 4; Scale bar, 10 μm. (E) PLISH RNA detection in tissues is highly sequence-specific. Mouse lung was hybridized with a single pair of H probes targeting nucleotides 228–268 of the Scgb1a1 transcript. The section in the bottom row was pre-incubated with a 60-base antisense blocking oligonucleotide complementary to nucleotides 219–278, whereas the section in the top row was pre-incubated with a scrambled 60-base blocking oligonucleotide. The antisense blocking oligonucleotide dramatically reduces the Scgb1a1 signal (bottom), whereas the scrambled blocking oligonucleotide has no effect (top). Note that the PLISH signal is tightly restricted to the bronchial Club cells (arrow). The dashed lines indicate the basal surface of bronchial epithelium. Scgb1a1, secretoglobin family 1A1 member 1; Scale bar, 40 μm. (F) PLISH RNA detection sensitivity in cultured cells matches single-cell qPCR sensitivity. FPKM values for 36 mRNAs are plotted against the fraction of HCT116 cells in which they were detected by single-cell qPCR (Wu et al., 2014) (filled black circles) or by PLISH (red inverted triangles). The black line is the prediction of a Poisson sampling model for the fraction of cells with at least one transcript, assuming that the transcript abundance increases proportionately with FPKM, and that one FPKM unit corresponds to 2.5 copies per cell. The inset shows PLISH staining for CASP9, which has an FPKM value of 2. CASP9, caspase 9; FPKM, Fragments Per Kilobase of transcript per Million mapped reads; qPCR, quantitative reverse transcription polymerase chain reaction; Scale bar, 20 μm. (G) RNA abundances measured by PLISH and by single-cell RNA sequencing are highly correlated. A log-log plot showing the average single-cell FPKM value for 10 mRNAs in HCT116 cells (Wu et al., 2014) plotted against the number of puncta per cell per probe measured by PLISH. Multiple points at each FPKM value are independent experiments. The data fit to a line of slope 1 with R2 = 0.8.
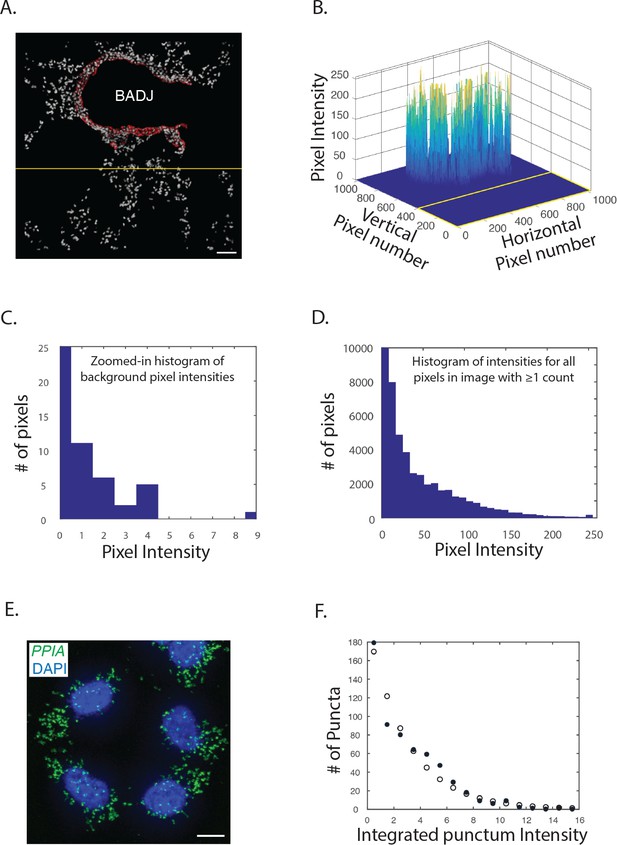
Signal-to-noise in PLISH images.
(A) An unprocessed micrograph of mouse lung interrogated with H probes targeting Scgb1a1 (red) and DAPI (white). The yellow line demarcates a region of the micrograph used to measure the background signal. Scale bar, 50 μm. (B) A plot of pixel intensities in the red channel from the image in panel A. The intensities ranged from 0 to 255. (C) A zoomed-in histogram of the red pixel intensities within the field demarcated by the yellow line, which was used to measure the background signal. 435,175 of these 435,200 background pixels had 0 counts (the vertical axis is truncated), and the mean intensity was 1.3*10^−4 counts. (D) A histogram of the red pixel intensities for all the non-zero pixels in panel A. The mean intensity of the non-zero pixels was 42 counts. (E) A micrograph of PLISH puncta for PP1A (green) and DAPI (blue) in cultured HCT116 cells. PP1A, protein phosphatase 1A; Scale bar, 10 μm. (F) A histogram of the integrated intensities for the PLISH puncta in panel E (filled circles). The histogram fitted well to a negative binomial distribution with a single 'fail' event (open circles). Mean-events-to-failure parameters between 1000 and 60,000 gave similar agreement with the data.
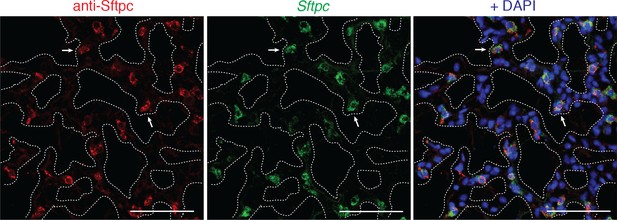
Benchmarking PLISH specificity against a validated antibody by co-staining in tissue.
Mouse lung co-stained for Sftpc by indirect immunohistochemistry (red) and PLISH (green) with DAPI (blue). 181 of 184 PLISH+ cells were also antibody+, and 181 of 195 antibody+ cells were PLISH+. White arrows indicate representative co-labeled AT2 cells. Dotted lines demarcate alveolar septae. AT2, alveolar epithelial type II; Scale bar, 40 μm.
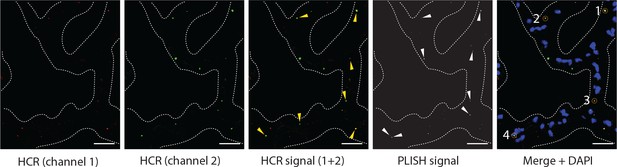
Estimating efficiency of PLISH probes in tissue.
Double in situ hybridization for Axin2 using HCR and PLISH was performed in mouse lung. HCR signals (yellow arrowheads, third panel) were identified based on overlap of puncta from two different HCR channels, and PLISH puncta were imaged in the same field (white arrowheads, fourth panel). Colocalized HCR and PLISH puncta are enumerated (dashed orange circles) in the fifth panel. Over three fields of view, we observed 92 HCR puncta and 140 PLISH puncta, with 29 cases of co-localized HCR and PLISH signal. Thus, the four PLISH probe pairs gave a combined detection efficiency of 32%, with a per-site efficiency of 9%. Dashed line demarcates alveolar septae. Scale bar, 40 μm.
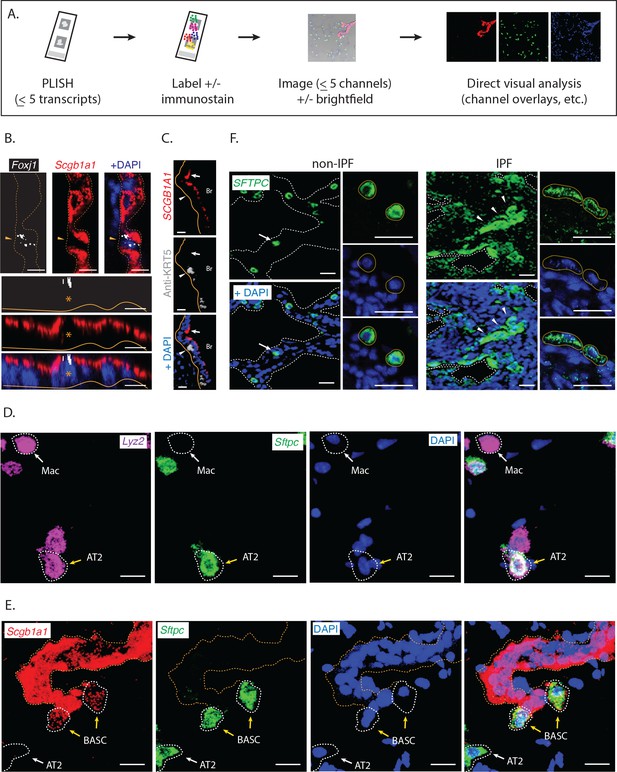
Direct visual analysis of single-molecule and single-cell gene expression in diverse specimens.
(A) PLISH experimental workflow. After an initial probe hybridization and enzymatic amplification step, up to five distinct channels can be simultaneously detected and imaged by conventional fluorescence microscopy, enabling direct visualization of RNA abundance. (B) PLISH detects single RNA molecules with single-cell resolution in tissues. PLISH staining for Foxj1 (white) and Scgb1a1 (red) in the bronchial epithelium of mouse lung shows a single ciliated cell (Foxj1+, arrowhead and asterisk) between Club cells (Scgb1a1+) in a planar view (top) and with orthogonal reconstruction (bottom). Note the discrete white puncta in the ciliated cell, which correspond to single Foxj1 transcripts. Dashed lines indicate the lateral and solid lines indicate the basal surface of airways. Foxj1, forkhead box J1; Scgb1a1, secretoglobin family 1A member 1; Scale bars, 10 μm. (C) Simultaneous RNA and protein detection in FFPE sections. FFPE human lung co-stained by PLISH (SCGB1A1, red) and indirect immunohistochemistry (anti-KRT5, grey) shows the expected localization of Club cells (SCGB1A1+, arrow) and basal cells (KRT5+, arrowhead) along the bronchial (Br) epithelium. Solid lines indicate the basal surface of airways. FFPE, formalin-fixed, paraffin-embedded; KRT5, keratin 5; SCGB1A1, secretoglobin family 1A member 1; Scale bar, 5 μm. (D) Discrimination of AT2 cells from macrophages by visual inspection of RNA abundance. PLISH staining in mouse lung for Lyz2 (purple) and Sftpc (green) allows clear discrimination of alveolar macrophages (Lyz2+ Sftpc-, white arrow) from AT2 cells (Sftpc+ Lyz2+, yellow arrow). AT2, Alveolar epithelial type II; Lyz2, lysozyme 2; Mac, macrophage; Sftpc, surfactant protein C; Scale bar, 20 μm. (E) Discrimination of AT2 cells from BASC cells by visual inspection of RNA abundance. PLISH staining for the Club cell marker (Scgb1a1, red) and AT2 cell marker (Sftpc, green) shows AT2 (Sftpc+), Club (Scgb1a1+) and BASC (Sftpc+ Scgb1a1Lo) cells. Note the discrete red puncta in the 'BASC' cells, which correspond to single Scgb1a1 transcripts. The cell types localize appropriately, with the AT2 cell in an alveolus (white arrow), the Club cells in the terminal bronchiole, and the double-positive BASCs at the bronchioalveolar junction (yellow arrows). Dashed lines demarcate the airway. AT2, alveolar epithelial type II; BASC, bronchioalveolar stem cell; Scgb1a1, secretoglobin family 1A member 1; Sftpc, surfactant protein C; Scale bar, 20 μm. (F) PLISH in patient tissue samples for molecular analysis of human disease. PLISH staining for SFTPC (green) in non-IPF human lung (left) marks AT2 cells (white arrow) distributed within alveolar septae (dashed lines). The adjacent panels show a magnified image of healthy cuboidal AT2 cells (dashed circles). PLISH staining in IPF human lung (right) shows densely cellular regions with architectural distortion of alveolar septae (dashed lines). SFTPCHi AT2 cells are inappropriately clustered (white arrowheads) and have abnormal flattened morphologies, as seen at higher magnification in right panels (dashed ellipses). AT2, alveolar epithelial type II; IPF, Idiopathic Pulmonary Fibrosis; SFTPC, surfactant protein C; Scale bar, 100 μm.
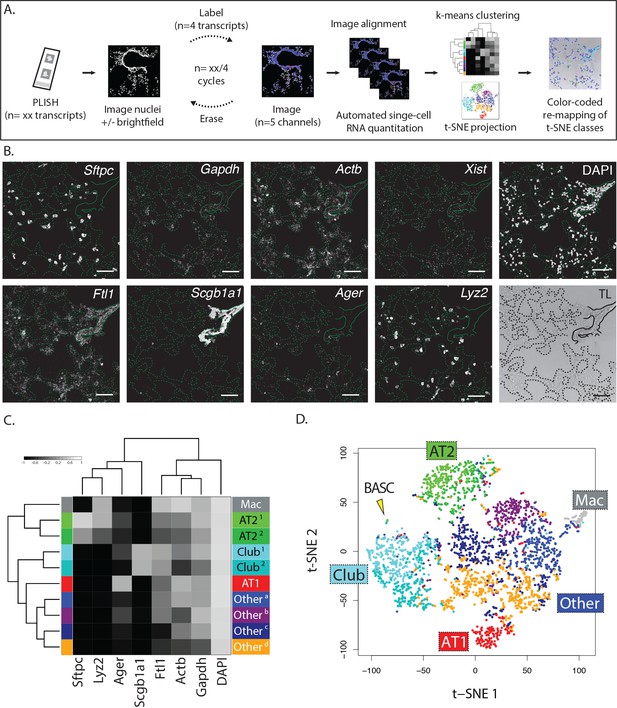
Multiplexed PLISH: rapid label-image-erase cycles, automated data analysis, and unsupervised cell classification.
(A) Multiplexed PLISH experimental workflow. Probes for many different RNAs are hybridized and amplified in a single reaction. The PLISH amplicons marking four RNA species are then labeled with four fluorescent imager oligonucleotides, imaged on a microscope, and 'erased' by elimination of the imager oligonucleotides. Amplicons marking a different subset of four RNAs are then labeled with four new imager oligonucleotides, imaged, and erased. This cycle is repeated until all of the RNA species have been visualized and photo-documented. The images are automatically aligned and processed, and the signal for each RNA species in each cell is summed to produce single-cell expression profiles. Unsupervised k-means clustering of the expression profiles (or other computational tools) empirically identifies distinct cell classes. t-SNE plots (or other data visualization tools) show the differences in gene expression and the classification for all of the cells in a tissue. The location of individual cells or cell classes is spatially remapped onto images of the tissue in order to integrate molecular, histological, and spatial features. (B) A multiplexed PLISH data set. Eight different transcripts in mouse lung were visualized with two label-image-erase cycles. A micrograph for each channel in one field of view is shown. Solid lines indicate the basal surface of airways and dashed lines indicate alveolar septae. Actb, beta actin; Ager, advanced glycosylation end product-specific receptor; Ftl1, ferritin light polypeptide 1; Gapdh, glyceraldehyde-3-phosphate dehydrogenase; Lyz2, lysozyme 2; Scgb1a1, secretoglobin family 1A member 1; Sftpc, surfactant protein C; Xist, inactive X specific transcripts. Scale bar, 80 μm. (C) Automated cell classification. K-means clustering partitions ~2900 single cells into one of ten molecularly distinct classes, with the expression profile of each cluster centroid displayed in a heat map. Based on marker gene expression, individual clusters were inferred to be macrophages (mac, white), two classes of AT2 cells (light and dark green), two classes of Club cells (light and dark cyan), AT1 cells (red), and four categories of unassigned cell types (shades of blue, purple, and orange). (D) An overview of the cells in a murine lung. Differences in gene expression for ~2900 cells are displayed as a two-dimensional t-SNE plot. Each cell is represented by a single dot, colored according to its cluster assignment. Labels mark the location of each cell class. The yellow arrowhead indicates a small island of cells that exhibit the profile of BASCs. AT1, alveolar epithelial type I cell; AT2, alveolar epithelial type II cell; BASC, bronchioalveolar stem cell; Mac, macrophage.
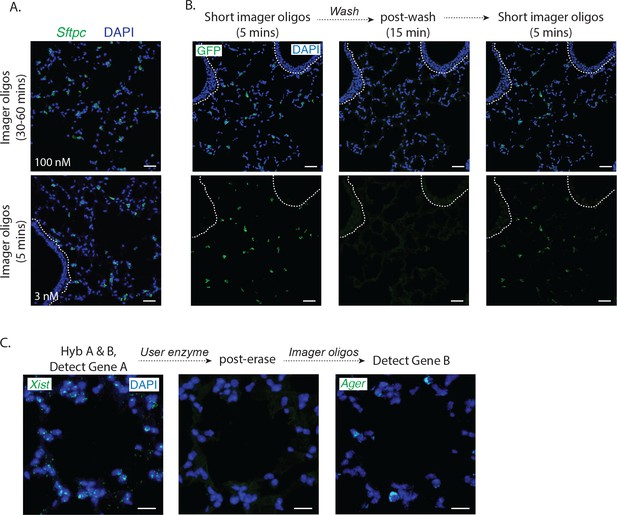
Rapid label-image-erase strategies without tissue degradation.
(A) Imaging with and without washout of fluorophore-labeled 'imager' oligonucleotides gives identical signal-to-noise ratios. Mouse lung tissue sections were interrogated with a single PLISH probe pair targeting Sftpc (green). In the top panel, the tissue section was hybridized with 100 nM imager oligonucleotides then washed (to remove excess imager oligonucleotides) prior to imaging, which took 30–60 min. In the bottom panel, the tissue section was hybridized with 3 nM imager oligonucleotides then imaged without a wash step, taking only 5 min. Bronchioles indicated by dashed lines. Sftpc, surfactant protein C; Scale bar, 40 μm. (B) Signal erasure by dissociation of short imager oligonucleotides. PLISH puncta for Sftpc (green) were visualized with short (11 base pair) imager oligonucleotides (first column). The signal was then erased by washing at 37°C for 15 min and re-imaged (middle column), showing no residual fluorescence. The sample was then re-stained with short imager oligonucleotides and re-imaged (third column), showing re-emergence of the fluorescent signal. The cycle time was 25 min. Bronchioles indicated by dashed lines. Sftpc, surfactant protein C; Scale bar, 40 μm. (C) Signal erasure by enzymatic digestion of imager oligonucleotides. PLISH puncta for Xist (green) were visualized in mouse lung with uracil-containing imager oligonucleotides (left panel). The tissue section was then incubated with the New England Biolabs USER enzyme cocktail at 37°C for 20 min to digest the imager oligonucleotides and re-imaged (middle panel), then re-stained with a new set of imager oligonucleotides for Ager (green, right panel). The cycle time was 25 min. Ager, advanced glycosylation end product-specific receptor; Xist, inactive X specific transcripts; Scale bar, 20 μm.
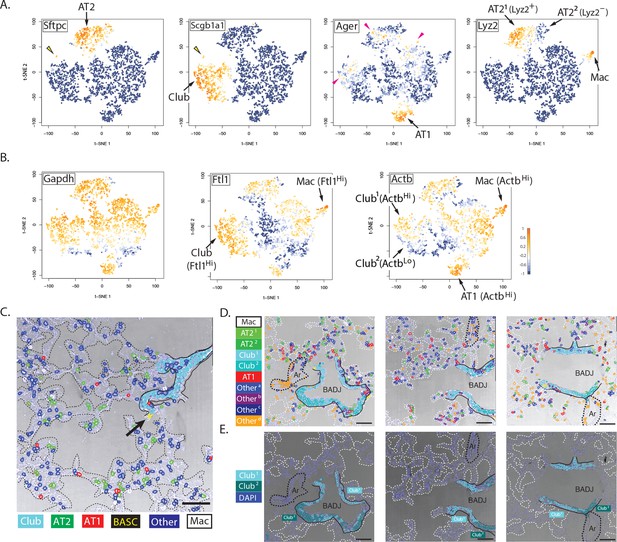
Biological insights from integrated molecular and spatial information.
(A) Specificity and promiscuity in marker gene expression. t-SNE plots are colored according to the expression of four cell-type marker genes. High expression (yellow) in the first two panels highlights AT2 and Club cells, as indicated, while the arrowhead indicates rare double-positive BASCs. The third panel shows high levels of Ager in AT1 cells (arrow), but promiscuous expression in a subset of AT2, Club1 and Otherb cell classes (red arrowheads). Lyz2 expression in the fourth panel is restricted to the macrophage and AT21 classes. Note that differential Lyz2 expression splits the canonical AT2 cell type into two sub-classes (AT21 and the AT22). (B) Differential expression of 'housekeeping' genes in canonical cell types. t-SNE plots are colored according to the expression of three ubiquitous 'housekeeping' genes. Gapdh (left) is the most evenly and broadly distributed, while Ftl1 (middle) is highest in the macrophage and Club cell classes. Actb is the highest in the macrophage and AT1 cell classes. Unexpectedly, differential Actb expression splits the canonical Club cell type into two sub-classes (Club1 and Club2). (C) Spatial organization of lung cell classes. The nuclei of cells in a transmitted light image of a bronchioalveolar duct junction (BADJ) are pseudocolored according to their basic cluster assignment. The Club class (cyan) localizes to the bronchial epithelium, while the AT1 (red) and AT2 (green) cell classes are distributed throughout the alveolar compartment. The macrophage class (white) is primarily found in the alveolar lumen. Rare BASCs (yellow) localize precisely to the bronchioalveolar junctions (arrow), where they have been shown to reside by immunostaining (Kim et al., 2005) (Figure 4C and Figure 4—figure supplement 1E). This image demonstrates how PLISH can be used to localize specific cells of interest within their anatomical context. Solid lines indicate the basal surface of airways and dashed lines demarcate alveolar septae. Scale bar, 80 μm. (D) Spatial organization in the terminal airway. The nuclei in three terminal airway fields of view are pseudocolored according to their cluster assignment. Note the presence of both Club1 and Club2 cell classes (light and dark cyan), and all four Other cell classes. The Otherd class (orange) is enriched in pulmonary arteries, indicated by black dashed lines, and therefore might represent endothelial or perivascular cells. Solid lines indicate the basal surface of airways, black dashed lines indicate pulmonary arteries, and dashed white lines demarcate alveolar septae. Ar, artery; Scale bar, 80 μm. (E) Two sub-classes of Club cells, defined by a difference in Actb expression, segregate anatomically. Club cells in the three fields of view from panel D are pseudocolored with enhanced contrast, revealing a striking spatial pattern. The Club1 sub-class (cyan, ActbHi AgerLo) localizes to the BADJ, whereas the Club2 sub-class (blue, ActbLo Ager–) segregates more proximally in the terminal airways. Differential Actb expression might reflect region-specific differences in mechanical stress. The integration of molecular and spatial information in this image reveals biology that would be missed with either piece of information alone. Solid lines indicate the basal surface of airways, black dashed lines indicate pulmonary arteries, and dashed white lines demarcate alveolar septae. Ar, artery; BADJ, bronchioalveolar duct junction; Scale bar, 80 μm.
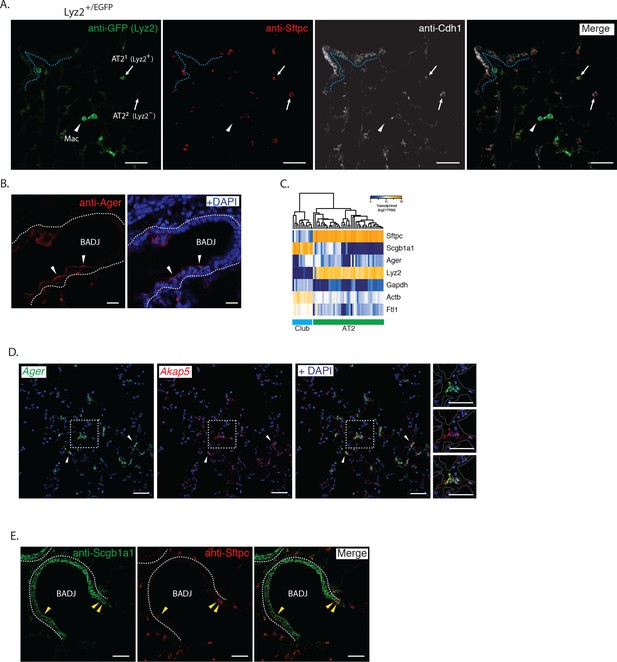
Immunohistochemical and single cell RNA-sequencing correlation of PLISH results.
(A) Lyz2+/EGFP mouse lung stained with anti-GFP (green) to mark Lyz2+ cells, anti-Sftpc (red, AT2), and anti-Cdh1 (white, epithelial) discriminates three cell populations. Macrophages (arrowhead, Lyz2+ Sftpc- Cdh1-), AT21 (Sftpc+ Lyz2+ Cdh1+) and AT22 (Sftpc+ Lyz2- Cdh1+). AT2 cells indicated by arrows. Dashed line indicates basal surface of bronchiole. AT2, alveolar epithelial type II; Mac, macrophage; Cdh1, cadherin 1; Lyz2, lysozyme 2; Sftpc, surfactant protein C; Scale bar, 40 μm. (B) Anti-Ager (red) labels the apical surface of cells in the bronchial epithelium (arrowheads). Dashed lines indicate the basal surface of airways. Ager, advanced glycosylation end product-specific receptor; BADJ, bronchioalveolar duct junction; Scale bar, 20 μm. (C) Heat-map of single cell RNA-sequencing (Treutlein et al., 2014) of Club (cyan bar) and AT2 (green bar) cells shows low expression of Ager in a subset of cells from both populations, supporting demonstration by PLISH of promiscuous expression of this AT1 cell marker. Note also broad expression of Lyz2 by AT2 cells (see t-SNE of Ager and Lyz2 in Figure 4A). AT1, alveolar epithelial type I; AT2, alveolar epithelial type II; Ager, advanced glycosylation end product-specific receptor; Lyz2, lysozyme 2. (D) Mouse lung co-stained for known (Ager, green) and novel (Treutlein et al., 2014) (Akap5, red) AT1 markers by PLISH. Arrowheads indicate representative AgerHi AT1 cells also expressing Akap5. Area in dashed box shown enlarged in right panels. Dashed lines demarcate alveolar septae. AT1, alveolar epithelial type I; Ager, advanced glycosylation end product-specific receptor; Akap5, A-Kinase Anchoring Protein 5; Scale bars, 50 μm. (E) Mouse lung stained with anti-Scgb1a1 (green) and anti-Sftpc (red) shows double positive BASCs (yellow arrowheads) localized to the BADJ. Dashed lines indicate the basal surface of airway epithelium; BADJ, bronchioalveolar duct junction; Scgb1a1, secretoglobin family 1a member 1; Sftpc, surfactant protein C; Scale bar, 40 μm.
Additional files
-
Source code 1
Custom scripts.
- https://doi.org/10.7554/eLife.30510.012
-
Supplementary file 1
Oligonucleotide sequences.
- https://doi.org/10.7554/eLife.30510.013
-
Transparent reporting form
- https://doi.org/10.7554/eLife.30510.014





