SPIN1 promotes tumorigenesis by blocking the uL18 (universal large ribosomal subunit protein 18)-MDM2-p53 pathway in human cancer
Figures
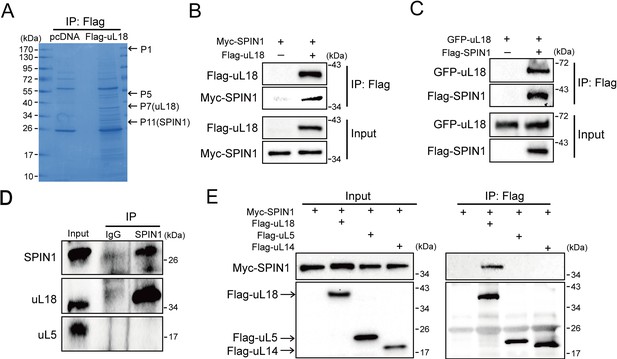
SPIN1 binds to uL18, but not uL5, or uL14.
(A) Identification of SPIN1 as a candidate of uL18 binding protein by immunopurification and mass spectrometric analysis. Lysates from HEK 293 cells were immunoprecipitated with the anti-Flag antibody. Bound proteins were visualized on a coomassie staining SDS-PAGE gel. Several bands were excised and subjected to mass spectrometry. One of them was identified as SPIN1 (Spindlin 1). The polypeptides identified from these bands are listed in Table 1. (B) and (C) SPIN1 interacts with uL18. (B) HCT116p53-/- cells were transfected with plasmids encoding Myc-SPIN1 and Flag-uL18, and 48 hr later cell lysates were collected for immunoprecipitation (IP) analysis using the anti-Flag antibody. (C) HCT116p53-/- cells were transfected with plasmids encoding Flag-SPIN1 and GFP-uL18 for 48 hr and harvested for IP/WB analysis with indicated antibodies. (D) The interaction between endogenous SPIN1 and uL18. The HEK 293 cell lysates were immunoprecipitated with anti-SPIN1 or control immunoglobulin G (IgG), followed by WB analysis with anti-SPIN1, anti-uL18 and anti-uL5. (E) SPIN1 was specifically co-immunoprecipitated by uL18, but not uL5 or uL14. H1299 cells were co-transfected with Myc-SPIN1 and Flag-uL18, Flag-uL5 or Flag-uL14 as indicated and subjected to IP with the anti-Flag antibody, followed by WB analysis with indicated antibodies.
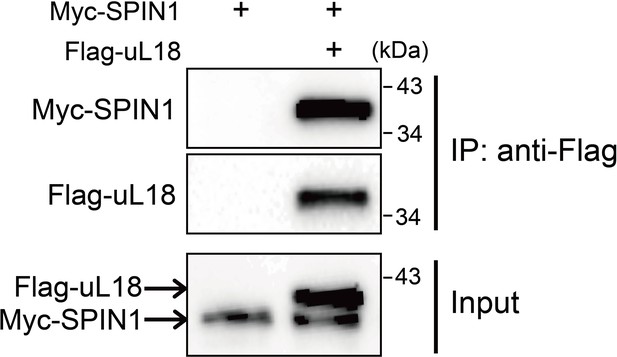
SPIN1 interacts with uL18 in HEK293 cells.
HEK293 cells were transfected with plasmids encoding Myc-SPIN1 and Flag-uL18, and 48 hr later cell lysates were collected for IP-WB analysis using anti-Flag antibody.
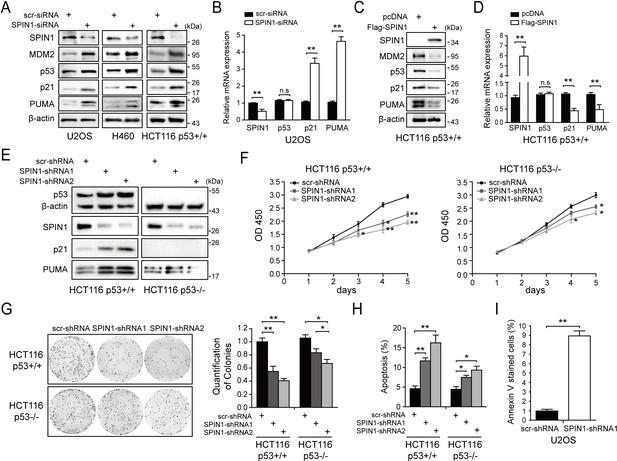
SPIN1 knockdown inhibits cell proliferation and induces apoptosis.
(A) SPIN1 knockdown induces protein levels of p53 and its target genes. U2OS, H460 and HCT116p53+/+ cells were transfected with scramble siRNA (scr-siRNA) or SPIN1 siRNA and harvested 48 hr post-transfection for WB analysis with indicated antibodies. (B) SPIN1 knockdown induces mRNA levels of p53 target genes without effect on TP53 mRNA level. U2OS cells were transfected with scramble siRNA (scr-siRNA) or SPIN1 siRNA, and harvested 72 hr post-transfection for RT-qPCR (mean ± SEM, n = 2). (C) SPIN1 overexpression reduces protein levels of p53 and its target genes. HCT116p53+/+ cells were transfected with pcDNA or Flag-SPIN1 and harvested 48 hr post-transfection for WB analysis with indicated antibodies. (D) SPIN1 overexpression reduces mRNA levels of p53 target genes without effect on TP53 mRNA levels. HCT116p53+/+ cells were transfected with pcDNA or Flag-SPIN1 and harvested 72 hr post-transfection for RT-qPCR (mean ± SEM, n = 2). (E) Knockdown of SPIN1 causes p53-dependent induction of p21 and PUMA. The protein levels of p53 and its targets in HCT116p53+/+ cells and HCT116p53-/- cells that stably express scramble shRNA (scr-shRNA) or SPIN1 shRNAs were detected by WB analysis with indicated antibodies. (F) SPIN1 knockdown suppresses cell survival. HCT116p53+/+ and HCT116p53-/- cells that stably expressed scramble or SPIN1 shRNAs were seeded in 96-well plate and cell viability was evaluated every 24 hr by CCK-8 assays (mean ± SEM, n = 2). (G) Knockdown of SPIN1 inhibits clonogenic ability of colorectal cancer cells, more significantly when the cells harbor wild-type p53. HCT116p53+/+ cells and HCT116p53-/-cells that stably expressed scramble or SPIN1 shRNAs were seeded on 60 mm plates. Puromycin selection was performed for 14 days. Colonies were fixed with methanol, and visualized by staining with crystal violet (mean ± SEM, n = 3). (H) The effect of SPIN1 knockdown on apoptosis of HCT116p53+/+ cells and HCT116p53-/-cells that stably expressed scramble or SPIN1 shRNAs (mean ± SEM, n = 3). (I) U2OS cells were transfected with scramble or SPIN1 shRNA and incubated in IncuCyte S3 chamber in the presence of IncuCyte Annexin V Green Reagent for apoptosis. Positively stained cells were determined using IncuCyte analysis software. *p<0.05, **p<0.01 by two-tailed t-test (C, D, G, H,I).
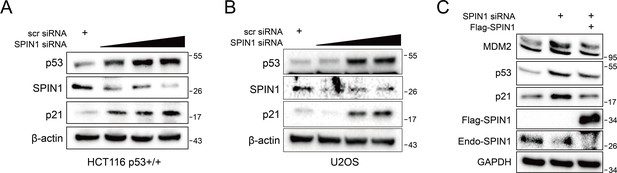
SPIN1 knockdown increase p53 and its targets protein levels in a dose-dependent manner.
HCT116 p53+/+ (A) and U2OS (B) cells were transfected with a titrated concentration of SPIN1 siRNA (20 nM, 40 nM and 60 nM). And cells were harvested 48 hr after transfection and followed by WB analysis with indicated antibodies. (C) U2OS cells were co-transfected with scramble siRNA (40 nM), SPIN1-siRNA (40 nM), with or without FLAG-SPIN1 as indicated. Cells were harvested 48 hr after transfection for WB analysis.
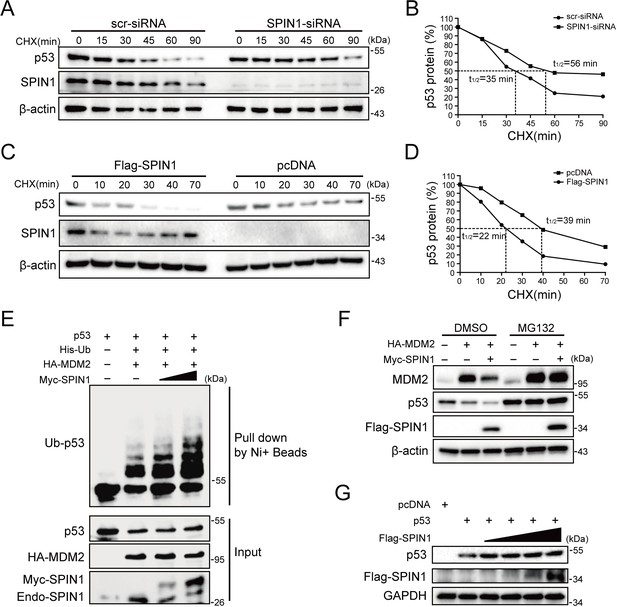
SPIN1 reduces p53 stability by enhancing MDM2-mediated ubiquitination.
(A) and (B) p53-half-life is increased by SPIN1 knockdown. (A) HCT116p53+/+ cells transfected with scramble or SPIN1 siRNA for 48 hr, were treated with 100 μg/ml of cycloheximide (CHX), and harvested at different time points as indicated. The p53 protein was detected by WB analysis, quantified by densitometry and plotted against time to determine p53-half-lives (B). (C) and (D) SPIN1 overexpression shortens the half-life of p53. HCT116p53+/+ cells transfected with pcDNA or Flag-SPIN1 for 48 hr were treated with 100 μg/ml of cycloheximide and harvested at indicated time points for WB analysis with indicated antibodies (C). The intensity of each band was quantified, and normalized with β-actin and plotted (D). (E) SPIN1 promotes MDM2-induced p53 ubiquitination. HCT116p53-/- cells were transfected with combinations of plasmids encoding HA-MDM2, p53, His-Ub or Myc-SPIN1, and treated with MG132 for 6 hr before being harvested for in vivo ubiquitination assay. Bound and input proteins were detected by WB analysis with indicated antibodies. (F) SPIN1 enhances MDM2-mediated p53 proteasomal degradation. HCT116p53+/+ cells were transfected with plasmids encoding HA-MDM2 and Flag-SPIN1, and treated with MG132 for 6 hr before harvested, followed by WB analysis with antibodies as indicated. (G) Ectopic SPIN1 does not change p53 protein level without MDM2. MEFp53-/-; Mdm2-/- cells were transfected with combinations of plasmids encoding p53 with or without Flag-SPIN1, followed by WB analysis using antibodies as indicated.
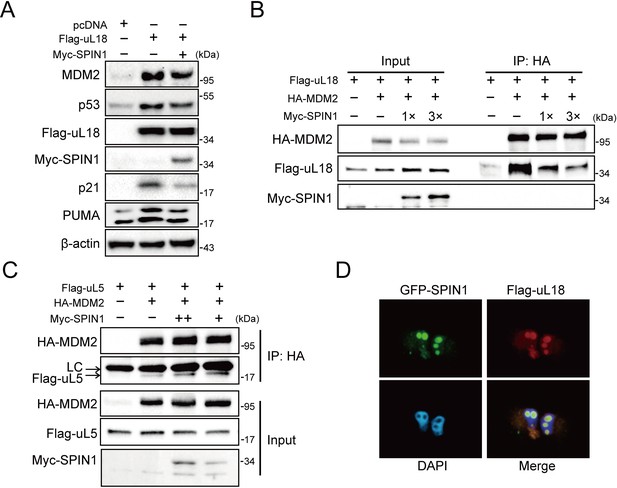
SPIN1 blocks uL18-MDM2 interaction by sequestering uL18 in the nucleolus.
(A) SPIN1 overexpression attenuates p53 activation induced by ectopic uL18. U2OS cells were co-transfected with plasmids encoding Flag-uL18 or Myc-SPIN1 for 36 hr and harvested for WB analysis with indicated antibodies. (B) Overexpression of SPIN1 disrupts the uL18-MDM2 binding. Lysates were prepared from HCT116p53-/- cells co-transfected with HA-MDM2, Flag-uL18, Myc-SPIN1 or the corresponding empty vectors for 48 hr and analyzed by immunoprecipitated with the anti-HA antibody. Immunoprecipitates and 5% of inputs were immunoblotted with the indicated antibodies. (C) Overexpression of SPIN1 fails to disrupt the uL5-MDM2 interaction. Lysates were prepared from HCT116p53-/- cells co-transfected with HA-MDM2, Flag-uL5 and Myc-SPIN1 for 48 hr and analyzed by immunoprecipitated with the anti-HA antibody. Immunoprecipitates and 5% of inputs were immunoblotted with the indicated antibodies. (LC: light chain). (D) SPIN1 and uL18 co-localize in the nucleolus. H1299 cells were transfected with GFP-SPIN1 and Flag-uL18 for 36 hr and then immunostained with the anti-Flag antibody (red), and counterstained with DAPI.
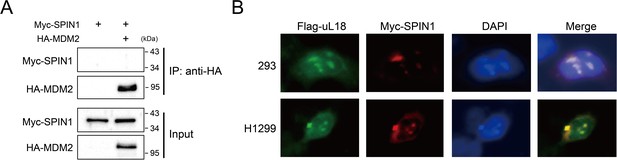
SPIN1 does not bind to MDM2, and SPIN1 and uL18 co-localize in the nucleolus.
(A) SPIN1 does not bind to MDM2. HCT116p53-/- cells were transfected with combination of plasmids encoding Myc-SPIN1 and HA-MDM2, followed by IP-WB analysis with indicated antibodies. (B) SPIN1 and uL18 co-localize in the nucleolus. HEK293 and H1299 SPIN1 stable cells were transfected with Flag-uL18 for 36 hr and then immunostained with anti-Myc (red) and anti-Flag antibody (green), and counterstained with DAPI.
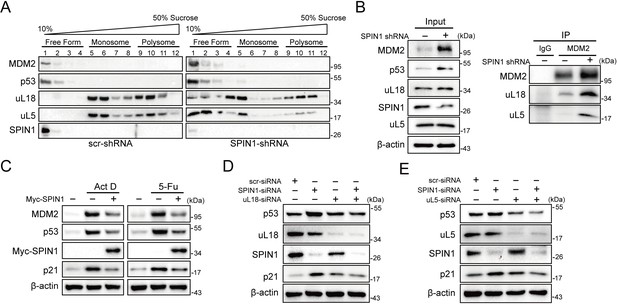
SPIN1 depletion increases ribosome-free uL18 and uL5.
(A) Knockdown of SPIN1 releases free forms of uL18 and uL5. HCT116p53+/+ were transfected with scramble or SPIN1 shRNA for 36 hr and subjected to sucrose gradient fractionation assay followed by WB analysis with indicated antibodies. (B) SPIN1 knockdown increases the endogenous uL18/uL5-MDM2 interaction. Cell lysates of HCT116p53+/+ cells transfected with scramble or SPIN1 shRNA were immunoprecipitated with MDM2 or control IgG, and analyzed by WB analysis with indicated antibodies. (C) SPIN1 overexpression counteracts p53 activation induced by ActD or 5-Fu. U2OS cells were transfected with pcDNA or Flag-SPIN1 for 48 hr, and treated with ActD or 5-Fu for 12 hr before harvested for WB analysis with indicated antibodies. (D) and (E) Knockdown of uL18 or uL5 compromises the induction of p53 by SPIN1 depletion. U2OS cells were transfected with scramble siRNA, SPIN1 siRNA, uL18 siRNA (D) or uL5 siRNA (E) as indicated for 48 hr. Cell lysates were subjected to WB analysis with indicated antibodies.
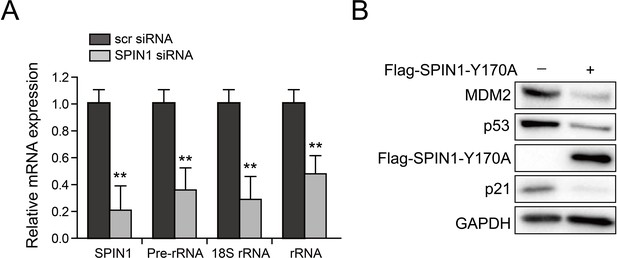
SPIN1 knockdown reduces rRNA expression and SPIN1-Y170A mutant retains activity to repress p53.
(A) Scramble siRNA or SPIN1 siRNA was introduced into U2OS cells. RNA levels were analyzed using Q-PCR (*p value < 0.05; **p value < 0.01 by tailed t-test; n = 6). (B). SPIN1-Y170A inhibits p53. U2OS cells were transfected with control plasmids or FLAG-SPIN1-Y170A and the cells were collected for western blot analysis 48 hr after transfection.
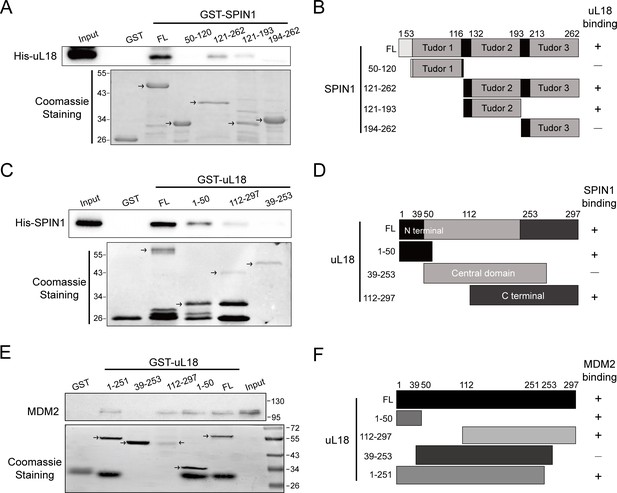
Mapping of domains responsible for uL18-SPIN1 and uL18-MDM2 binding.
(A) uL18 interacts with the second Tudor like domain of SPIN1. Purified GST-tagged SPIN1 fragments, including aa 1-262(FL), aa 50–120, aa 121–262, aa 121–193, aa 194–262 and GST protein alone were incubated with purified His-uL18 protein for 3 hr at 4°C. Bound proteins were detected by WB analysis using anti-uL18 or coomassie staining. (B) A schematic diagram of uL18-binding regions on SPIN1 based on the result from (A). (C) SPIN1 interacts with both the N- and C-termini of uL18. Purified GST-tagged uL18 fragments, including aa1-297(FL), aa 1–50, aa 112–297, aa 39–253 or GST protein alone were rotated with purified His-SPIN1 protein for 1 hr at 4°C. Bound proteins were detected by WB analysis using anti-SPIN1 or coomassie staining. (D) A schematic diagram of SPIN1-binding regions on uL18 derived from the result from (C). (E) MDM2 interacts with both the N- and C-termini of uL18. Purified GST-tagged uL18 fragments, including aa1-297(FL), aa 1–50, aa 112–297, aa 39–253, aa 1–251 or GST protein alone were rotated with purified His-MDM2 protein for 4 hr at 4°C. Bound proteins were detected by WB analysis using anti-MDM2 (2A10) or coomassie staining. (F) A schematic diagram of uL18 binding regions on MDM2 based on the result from (E).
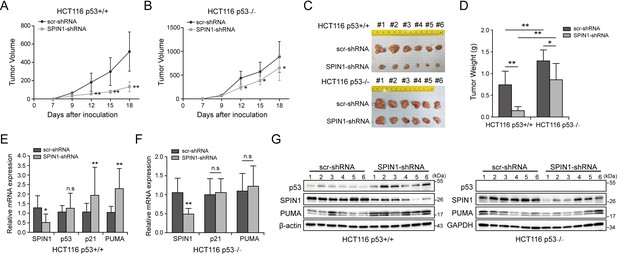
SPIN1 knockdown retards tumor growth more dramatically by inducing p53 activity.
(A) and (B) Growth curves of xenograft tumors derived from HCT116p53+/+ cells and HCT116p53-/- cells that expressed scramble or SPIN1 shRNA. Data are represented as mean ± SEM, n = 6. (C) The images of xenograft tumors that were harvested at the end of experiment. (D) Quantification of the average weights of collected tumors from the above experiments. (E) and (F) The mRNA levels of SPIN1, p53 and p53 target genes were detected in six tumors by RT-qPCR (mean ± SEM, n = 6). (G) The protein levels of SPIN1, p53 and p53 targets were detected in six tumors samples by WB analysis with indicated antibodies. *p<0.05, **p<0.01 by two-tailed t-test (D, E, F, G).
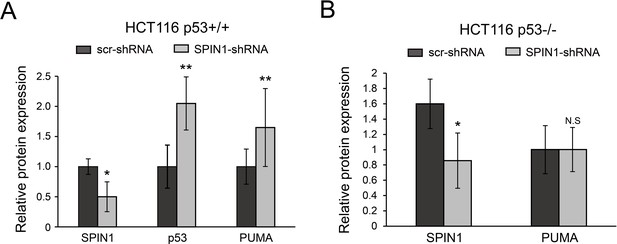
Quantification of protein expression analyzed from xenograft tumors by Image J software.
All data were presented as mean ± SEM, n = 6, *p<0.05, **p<0.01 by two-tailed t-test.
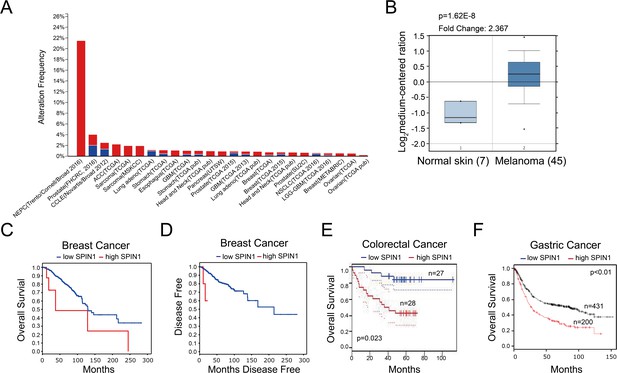
High expression of SPIN1 is detected in multiple cancers and associated with poor prognosis in cancer patients.
(A) TCGA database was utilized, and the data were modified from the cBioPortal for Cancer Genomics (http://www.cbioportal.org/). (B) The expression profile of SPIN1 in cancers and normal tissues was searched in Oncomine Gene Browser (http://www.oncomine.org/). The results were from Talantov Melanoma database. Seven cases of normal skin and 45 cases of melanoma were analyzed in this figure. Correlation between SPIN1 upregulation and tumor stage, poorer prognosis or treatment resistance is not clear. (C and D) Overexpression of SPIN1 is correlated with overall survival and disease-free survival in breast cancer (http://www.cbioportal.org/), although the sample number of high SPIN1 patients is small and more samples are desired. (E) SPIN1 overexpression was associated with poor prognosis in colorectal cancer patients in an expression profile study from GSE17537 (http://www.PrognoScan.org/). (F) High expression of SPIN1 was correlated with poor overall survival in gastric cancer patients (http://www.kmplot.com).
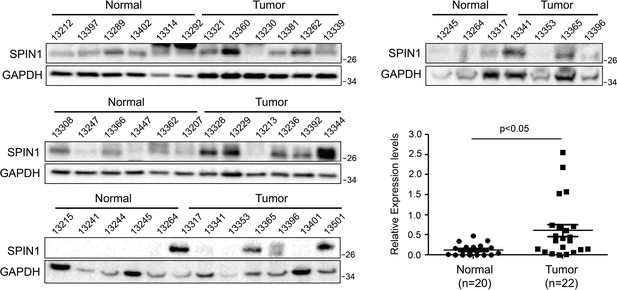
Western blotting analyses of human colon cancer tissues (n = 22) and normal colon tissue (n = 20) and quantification of SPIN1 expression (Mean ± SEM, p<0.05).
https://doi.org/10.7554/eLife.31275.016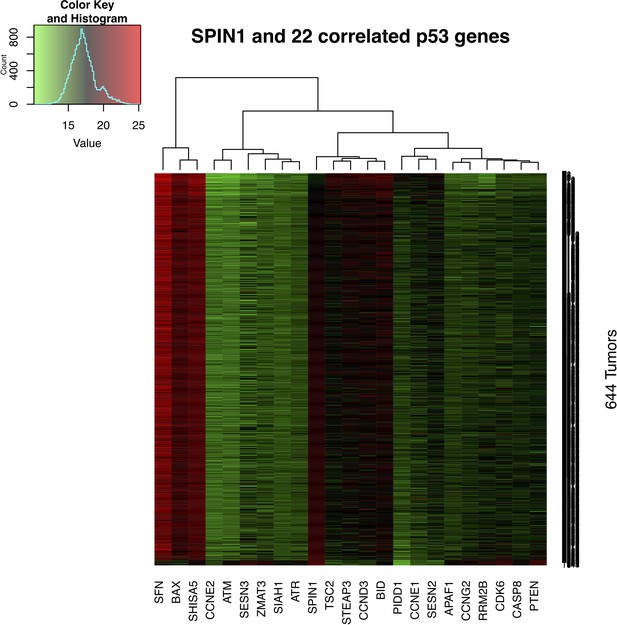
Expression of genes involved in p53 pathway is correlated with SPIN1 expression.
mRNA expression levels of 664 colorectal tumors were retrieved from Genomic Data Commons (https://portal.gdc.cancer.gov/). In the data set, gene expression levels were measured with FPKM (Fragments Per Kilobase of transcript per Million mapped reads) and normalized using the Upper Quantile method. All 644 tumor samples were sorted based on the expression level of SPIN1 from low to high.
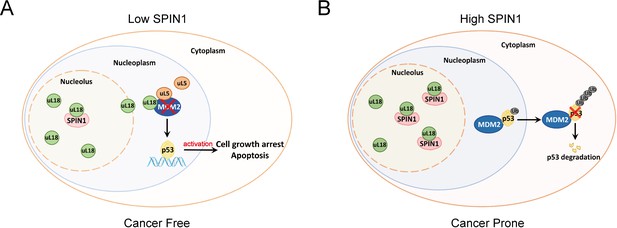
A model for SPIN1 regulation of the uL18-MDM2-p53 pathway in cancer.(see text in the Discussion for details).
https://doi.org/10.7554/eLife.31275.018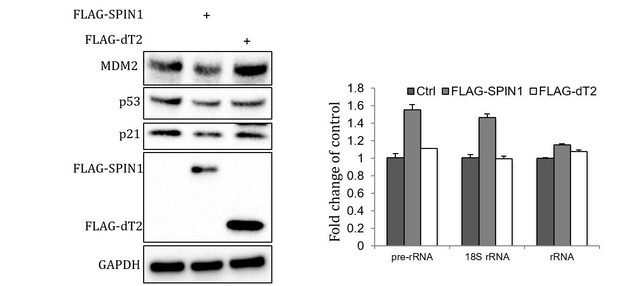
SPIN1-dT2 fails to inhibit p53.
U2OS cells were transfected with wild type and Tudor 2 domain deletion mutant (FLAG-dT2) SPIN1 and cells were harvested 48 hrs after transfection for Western Blot (Left panel) and qRT-PCR (Right panel) analyses.
Tables
uL18-associated polypeptides identified from mass spectrometry analysis of proteins as shown in Figure 1A.
https://doi.org/10.7554/eLife.31275.004| Gel slice number | Protein | Accession | Molecular weight | Score |
|---|---|---|---|---|
| P1 | Myb-binding protein 1A (MYBBP1A) | gi|6959304 | 149727 | 149 |
| P5 | Protein arginine N-methyltransferase 5 (PRMT5) | gi|2323410 | 72685 | 121 |
| P7 | 60S ribosomal protein L5 uL18(RPL5) | gi|14591909 | 34569 | 1014 |
| P11 | Serine/arginine-rich splicing factor 1 (SRSF1) | gi|5902076 | 27746 | 75 |
| P11 | Spindlin 1 (SPIN1) | gi|5410330 | 29602 | 95 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (human) | SPIN1 | National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/gene/10927) | Gene ID: 10927; Accession number: NM_006717; UniPro ID: Q9Y657 | |
| Gene (human) | RPL5/uL18 | National Center for Biotechnology Information https://www.ncbi.nlm.nih.gov/gene/6125 | gene ID: 6125; Accession number: NM_000969; UniPro ID: P46777 | |
| Gene (human) | RPL11/uL5 | National Center for Biotechnology Information https://www.ncbi.nlm.nih.gov/gene/6135 | gene ID: 6135; Acctssion number: NM_000975; UniPro ID: P62913 | |
| Gene (human) | RPL23/uL14 | National Center for Biotechnology Information https://www.ncbi.nlm.nih.gov/gene/9349 | gene ID: 9349; Accession number: NM_000978; UniPro ID: P62829 | |
| Gene (human) | TP53 | National Center for Biotechnology Information https://www.ncbi.nlm.nih.gov/gene/7157 | gene ID: 7157; Accession number: NM_000546; UniPro ID: P04637 | |
| Gene (human) | p21/CDKN1A | National Center for Biotechnology Information https://www.ncbi.nlm.nih.gov/gene/1026 | gene ID: 1026; Accession number: NM_000389; UniPro ID: Q42580 | |
| Gene (human) | PUMA/BBC3 | National Center for Biotechnology Information https://www.ncbi.nlm.nih.gov/gene/27113 | gene ID: 27113; Accession number: NM_001127240; UniPro ID: Q9BXH1 | |
| Gene (human) | MDM2 | National Center for Biotechnology Information https://www.ncbi.nlm.nih.gov/gene/4193 | gene ID: 4193; Accession number: NM_001145337; UniPro ID: Q00987 | |
| Strain, strain background (mouse) | NOD-SCID | Jackson Laboratories https://www.jax.org/strain/001303 | Stock No: 001303 | |
| Cell line (human) | 293 | ATCC https://www.atcc.org/Products/All/CRL-1573.aspx | Catalog number: ATCC CRL-1573; RRID:CVCL_0045 | |
| Cell line (human) | H1299 | ATCC https://www.atcc.org/Products/All/CRL-5803.aspx | Catalog number: ATCC CRL-5803; RRID: CVCL_0060 | |
| Cell line (human) | U2OS | ATCC https://www.atcc.org/Products/All/HTB-96.aspx | Catalog number: ATCC HTB-96; RRID:RRID:CVCL_0042 | |
| Cell line (human) | H460 | ATCC https://www.atcc.org/Products/All/HTB-177.aspx | Catalog number: ATCC HTB-177; RRID:CVCL_0459 | |
| Cell line (human) | HCT116 p53+/+ | from Dr. Bert Vogelstein at the John Hopkins Medical institutes | ||
| Cell line (human) | HCT116 p53-/- | from Dr. Bert Vogelstein at the John Hopkins Medical institutes | ||
| Cell line (human) | MEF (Mdm2-/-; p53-/-) | from Dr. Guillermina Lozano from MD Anderson Cancer Center, the University of Texas. | ||
| Antibody | Mouse anti-human Flag monoclonal antibody | Sigma-Aldrich | Catalog number: F1804; RRID: AB_262044 | Applications: WB; Immunofluoresce |
| Antibody | Mouse anti-human Myc monoclonal antibody | Santa Cruz Technology | Catalogue number: sc-40 | Applications: WB; Immunofluoresce |
| Antibody | Mouse anti-human GFP monoclonal antibody | Santa Cruz Technology | Catalogue number: sc-9996; RRID: AB_627695 | Applications: WB; Immunofluoresce |
| Antibody | Mouse anti-human GST monoclonal | ProteinTech | Catalogue number: HRP-66001; RRID: AB_10951482 | Applications: WB |
| Antibody | Rabbit anti-bacterial His polyclonal antibody | ProteinTech | Catalogue number:10560–1-lg; RRID: AB_1607770 | Applications: WB |
| Antibody | Rabbit anti-human SPIN1 polyclonal antibody | ProteinTech | Catalogue number:12105–1-AP; RRID: AB_2196111 | Applications: WB |
| Antibody | Mouse anti-human p53 monoclonal antibody | Santa Cruz Technology | Catalogue number: sc-126; RRID: AB_628082 | Applications: WB |
| Antibody | Mouse anti-human p21 monoclonal antibody | Neomarkers, Fremont, | Catalogue number: MS-891-P0; RRID:AB_143907 | Applications: WB |
| Antibody | Rabbit anti-human PUMA polyclonal antibody | ProteinTech | Catalogue number:55120–1-AP; RRID:AB_10859944 | Applications: WB |
| Antibody | Mouse anti-human β-actin monoclonal antibody | Santa Cruz Technology | Catalogue number: sc-47778; RRID:AB_2714189 | Applications: WB |
| Antibody | Rabbit anti-human GAPDH polyclonal antibody | Proteintech | Catalogue number:10494–1-AP; RRID:AB_2263076 | Applications: WB |
| Chemical compound, drug | Cycloheximide | Sigma-Aldrich | Catalogue number: 66-81-9 | |
| Chemical compound, drug | MG-132 | Sigma-Aldrich | Catalogue number:474787 | |
| Chemical compound, drug | 5-FU | Sigma-Aldrich | Catalogue number:51218 | |
| Chemical compound, drug | Actinomycin D (Act D) | Sigma-Aldrich | Catalogue number: 50-76-0 |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.31275.019





