Haploinsufficiency of Trp53 dramatically extends the lifespan of Sirt6-deficient mice
Figures
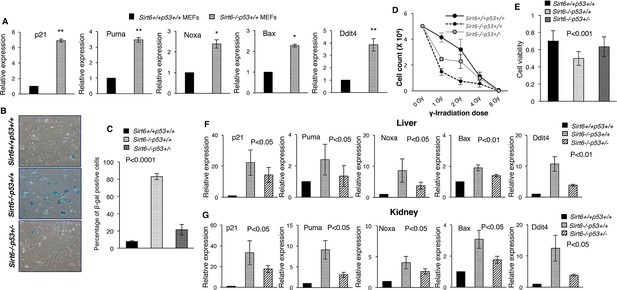
Heterozygosity of Trp53 rescues premature senescence in Sirt6-deficient scenario, (Trp53 has been denoted as p53 in the figures).
(A) Quantification for qPCR analyses for gene expression of p53 targets (with respect to Gapdh controls) in Sirt6+/+Trp53+/+ (WT) and Sirt6-/-Trp53+/+ (Sirt6 KO) MEFs, respectively. Data represent mean ± SEM, n = 3. *p<0.05, and **p<0.01 calculated using Student’s t-test. (B) Representative images of senescence-associated β-galactosidase staining in Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- MEFs at P6. Scale bar, 100 µm. (C) Quantification of data presented in (B). Data represent mean ± SEM, approximately 100 cells were counted from each genotype in three replicates. P value calculated using one-way ANOVA. (D) Graph showing survival of Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- MEFs of passage 3 (P3) one week after exposure to different doses of γ-irradiation. Data represent mean ± SEM, n=3. (E) Graphical representation of cell viability of Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- MEFs at P3 as measured by MTT assay. Data represent mean ± SEM, n=3. P value calculated using one-way ANOVA. (F) Quantification for qPCR analyses of gene expression of p53 targets (with respect to Gapdh controls) in liver from Sirt6+/+Trp53+/+(WT), Sirt6-/-Trp53+/+(Sirt6 KO) and Sirt6-/-Trp53+/- (compound mutant) mice, respectively. Data represent mean ± SEM, n = 3. P value calculated using one-way ANOVA. (G) Quantification for qPCR analyses of gene expression of p53 targets (with respect to Gapdh controls) in kidneys of Sirt6+/+Trp53+/+(WT), Sirt6-/-Trp53+/+(Sirt6 KO) and Sirt6-/-Trp53+/- (compound mutant) mice, respectively. Data represent mean ± SEM, n = 3. P value calculated using one-way ANOVA.
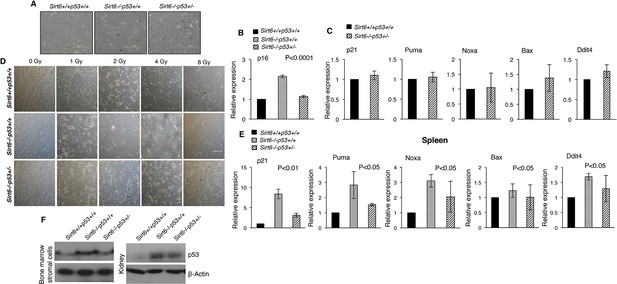
Attenuation of p53 downstream targets and rescue of premature senescence-associated phenotypes in Sirt6-deficient scenario by haploinsufficiency of Trp53, (Trp53 has been denoted as p53 in the figures).
(A) Representative images of MEFs derived from Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- embryos at P6. Scale bar, 100 µm. (B) qPCR analyses of p53 mRNA levels in Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- MEFs at P6 (with respect to Gapdh controls). Data represent mean ± SEM, n=3. P value has been calculated using one-way ANOVA. (C) Quantification for qPCR analyses for gene expression of p53 targets (with respect to Gapdh controls) in Sirt6+/+Trp53+/+ and Sirt6-/-Trp53+/- MEFs. Data represent mean ± SEM, n=3. (D) Representative images of Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- MEFs at P3 one week after exposure to different doses of γ-irradiation-induced DNA damage. Scale bar, 100 µm. (E) qPCR analyses of the gene expression of p53 downstream targets in the spleen of Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice (with respect to Gapdh controls). Data represent mean ± SEM, n=3. P values calculated using one-way ANOVA. (F) Western blotting analysis of p53 expression in the bone marrow stromal cells and kidney of around 24 days old Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- littermate mice.
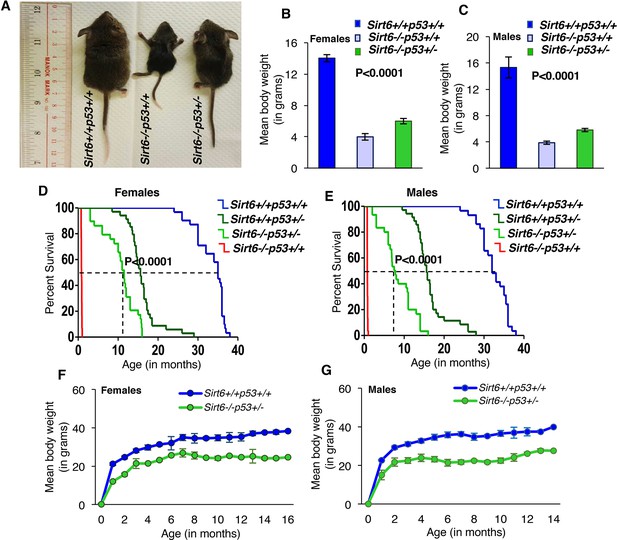
Haploinsufficiency of Trp53 significantly extends the lifespan of Sirt6-deficient mice, (Trp53 has been denoted as p53 in the figures).
(A) Representative images of Sirt6+/+Trp53+/+(WT), Sirt6-/-Trp53+/+ (Sirt6 KO), and -/-Trp53+/- (compound mutant) littermate mice at the age of 24 days. (B) Mean body weights of WT, Sirt6 KO and compound mutant female mice at the age of ~24 days. Data represent mean ± SEM; n = 5. P value calculated using one-way ANOVA. (C) Mean body weights of WT, Sirt6 KO and compound mutant male mice at the age of ~24 days. Data represent mean ± SEM; n = 5. P value calculated using one-way ANOVA. (D) Kaplan-Meier survival curve of Sirt6+/+Trp53+/+(n = 30), Sirt6-/-Trp53+/+ (n = 30), Sirt6-/-Trp53+/-(n = 31) and Sirt6+/+Trp53+/- (n = 35) female mice. P value calculated using log-rank (Mantel-Cox) test. (E) Kaplan-Meier survival curve of Sirt6+/+Trp53+/+ (n = 30), Sirt6-/-Trp53+/+ (n = 30), Sirt6-/-Trp53+/- (n = 31) and Sirt6+/+Trp53+/- (n = 35) male mice. P value calculated using log-rank (Mantel-Cox) test. (F) Graphical representation of increasing body weights in female Sirt6-/-Trp53+/- and WT littermate mice. Data represent mean ± SEM. (G) Graphical representation of increasing body weights in male Sirt6-/-Trp53+/- and WT littermate mice. Data represent mean ± SEM.
-
Figure 2—source data 1
Serum Igf-1 levels in 24-day old mice.
- https://doi.org/10.7554/eLife.32127.007
-
Figure 2—source data 2
Serum Igf-1 levels in 10-16 months old mice.
- https://doi.org/10.7554/eLife.32127.008
-
Figure 2—source data 3
Serum glucose levels in 24-day old mice.
- https://doi.org/10.7554/eLife.32127.009
-
Figure 2—source data 4
Serum glucose levels in 10-16 months old mice.
- https://doi.org/10.7554/eLife.32127.010
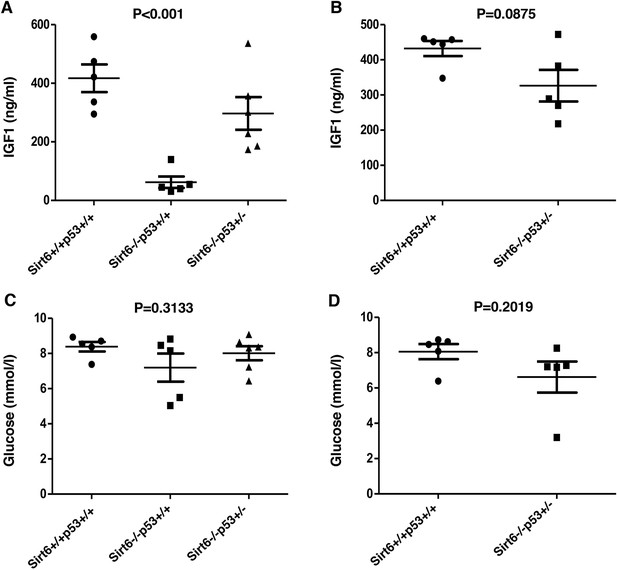
Analysis of serum IGF-1 and glucose levels in Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice, (Trp53 has been denoted as p53 in the figures).
(A) Serum IGF-1 levels in Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice at around 24 days. P value calculated using one-way ANOVA. (B) Serum IGF-1 levels in Sirt6+/+Trp53+/+ and Sirt6-/-Trp53+/- mice at around 10-16 months. P value calculated using one-way ANOVA. (C) Serum glucose levels in Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice at around 24 days. P value calculated using one-way ANOVA. (D) Serum glucose levels in Sirt6+/+Trp53+/+ and Sirt6-/-Trp53+/- mice at around 10-16 months. P value calculated using one-way ANOVA.
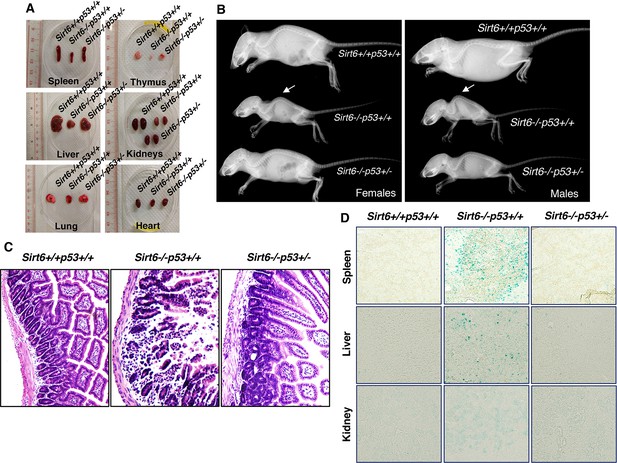
Amelioration of premature-aging associated phenotypes in Sirt6 KO mice upon haploinsufficiency of p53, (Trp53 has been denoted as p53 in the figures).
(A) Representative images of spleens, thymus, livers, kidneys, lungs and hearts from 24 days old Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+, and Sirt6-/-Trp53+/- mice. (B) Whole body X-ray images of female and male Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice (note the lordokyphosis/curved spine in Sirt6-/-p53+/+ mice) at the age of 24 days. (C) Histology of small intestinal cross-sections of WT, Sirt6 KO and compound mutant mice to detect colitis, at the age of ~24 days. (D) Representative images of senescence-associated β-galactosidase staining in spleen, liver and kidneys of 3 weeks old Sirt6+/+Trp53+/+ (WT), Sirt6-/-Trp53+/+ (Sirt6 KO) and Sirt6-/-Trp53+/- (compound mutant) mice.
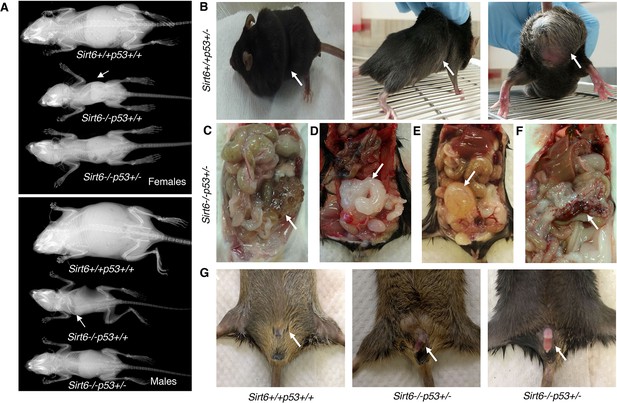
Incidence of cancer and other age-associated disorders in Sirt6-/-Trp53+/- mice, (Trp53 has been denoted as p53 in the figures).
(A) Whole body X-ray images of female and male Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice (note the lordokyphosis/curved spine in Sirt6-/-Trp53+/+ mice) at the age of 24 days. (B) Representative images of sarcoma in Sirt6+/+Trp53+/- mice at the age of 17–20 months. (C) Representative image of kidney tumor in Sirt6-/-Trp53+/- mice at around 10-16 months. (D) Representative image of prostate tumor in Sirt6-/-Trp53+/- mice at around 10-16 months. (E) Representative image of bladder tumor in Sirt6-/-Trp53+/- mice at around 10-16 months. (F) Representative image of pancreatic tumor in Sirt6-/-Trp53+/- mice at around 10-16 months. (G) Representative images of penile protrusion in Sirt6-/-Trp53+/- mice at around 9-11 months.
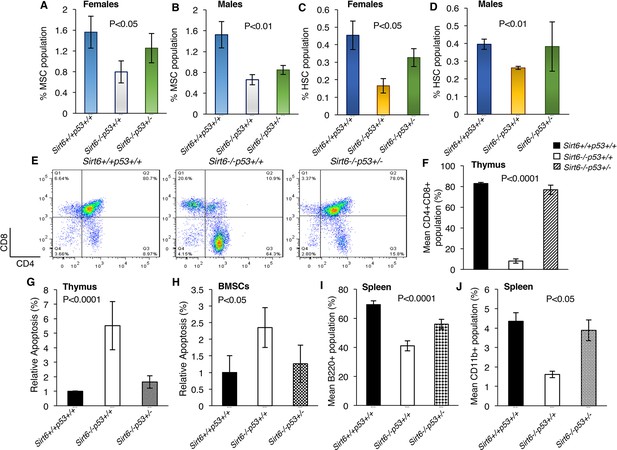
Haploinsufficiency of Trp53 prevents bone marrow stem cell decline, suppresses apoptosis, and restores immune cell populations in Sirt6-deficient mice, (Trp53 has been denoted as p53 in the figures).
(A) Percentage of mesenchymal stem cell (MSC) population in the female Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice at the age of ~24 days. Data represent mean ± SEM; n=5. (B) Percentage of mesenchymal stem cell (MSC) population in the male Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice at the age of ~24 days. Data represent mean ± SEM; n=5. (C) Percentage of hematopoietic stem cell (HSC) population in the female Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice at the age of ~24 days. Data represent mean ± SEM; n=5. (D) Percentage of hematopoietic stem cell (HSC) population in the male Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice at the age of ~24 days. Data represent mean ± SEM; n=5. (E) Representative flow cytometry analysis of CD4+CD8+ cells in the thymus of Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice at the age of ~24 days. Y-axis and X-axis represent the numbers of CD8+ cells and CD4+ cells respectively. (F) Graphical representation of data presented in (E). Data represent mean ± SEM; n = 5. (G) Quantification of apoptotic cells in the thymus of Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice by Annexin V staining. Data represent mean ± SEM; n=4. (H) Quantification of apoptotic cells in the thymus and bone marrow stroma of Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice by Annexin V staining. Data represent mean ± SEM; n=4. (I) Graphical representation of peripheral B cell population (B220+ cells) in the spleens from Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice. Data represent mean ± SEM; n=5. (J) Graphical representation of peripheral monocyte counts (CD11b+ cells) in the spleens from Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- mice. Data represent mean ± SEM; n=5. All P values have been calculated using one-way ANOVA.
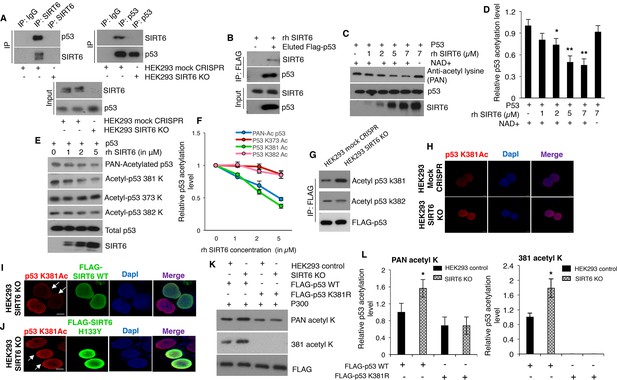
SIRT6 directly deacetylates p53 at lysine 381, but not K382.
(A) Western blotting analysis of endogenous interaction between SIRT6 and p53 in mock CRISPR control cells and SIRT6 KO cells using specific antibodies for SIRT6 and p53, keeping respective IgG controls. (B) Western blotting analysis to examine direct interaction between SIRT6 and p53 in vitro using recombinant human (rh) SIRT6 and eluted FLAG-tagged p53 from HEK293 cells. (C) Western blotting analysis of deacetylation of p53 in vitro with increasing concentration of recombinant (rh) SIRT6 and purified p53, in the presence and absence of NAD+. (D) Quantification of data presented in (C). Data represent mean ± SEM, n = 3. calculated using Student’s t-test. (E) Analyses of SIRT6-mediated deacetylation of p53 at lysine (K) 381, 373 or 382 in vitro by Western blotting using specific antibodies and PAN-acetyl lysine antibodies. (F) Quantification of data presented in (E). Data represent mean ± SEM, n = 3. (G) Western blotting data showing acetylation at lysine 381 and 382 in immunoprecipitated FLAG-tagged p53 in SIRT6 KO cells and mock CRISPR control HEK293 cells. (H) Immunofluorescence staining of p53 acetylation at K381 in HEK293 mock CRISPR and SIRT6 KO cells. Scale bar, 10 µm. (I) Immunofluorescence staining of p53 acetylation at K381 (red fluorescence) in SIRT6 KO cells with ectopically expressed wild-type (WT) SIRT6 (green fluorescence). Scale bar, 10 µm. (J) Immunofluorescence staining of p53 acetylation at K381 (red fluorescence) in SIRT6 KO cells with ectopically expressed catalytically inactive mutant SIRT6 (H133Y) (green fluorescence). Scale bar, 10 µm. (K) Western blotting analysis of SIRT6-mediated deacetylation of WT and lysine 381 to arginine (K381R) mutant p53 by antibodies against pan-acetyl lysine and acetylation at lysine 381 of p53. (L) Quantification of data presented in (K). Data represent mean ± SEM, n = 3. *p<0.05 calculated using Student’s t-test.
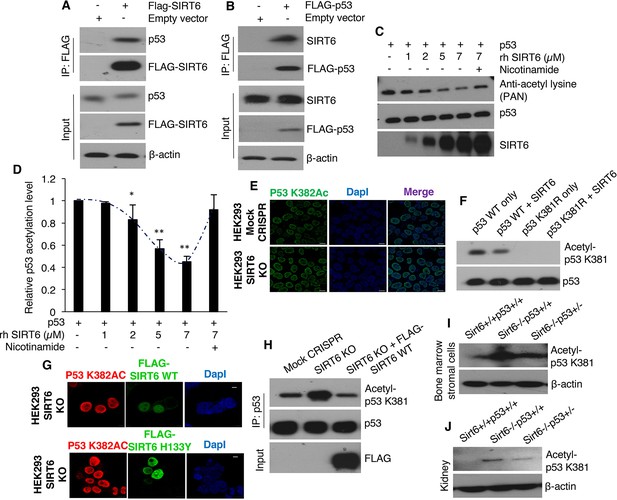
Analyses of physical and functional interaction between SIRT6 and p53, (Trp53 has been denoted as p53 in the figures).
(A) Western blotting analysis of endogenous p53 pulled down by FLAG-tagged SIRT6 in HEK293 cells. (B) Western blotting analysis of endogenous SIRT6 pulled down by FLAG-tagged p53 in HEK293 cells. (C) SIRT6-mediated deacetylation of p53 in vitro using recombinant (rh) SIRT6 and eluted FLAG-tagged p53 from HEK293 cells in the presence and absence of sirtuin inhibitor nicotinamide, as analyzed by Western blotting. (D) Quantification of data presented in (C). Data represent mean ± SEM, n = 3 for. *p<0.05 and **p<0.01 calculated using Student’s t-test. (E) Immunofluorescence staining of p53 acetylation at lysine 382 in HEK293 mock CRISPR and SIRT6 KO cells. Scale bar, 10 µm. (F) Western blotting analysis of p53 deacetylation in vitro using p53 WT and p53 K381R mutant as substrates for recombinant (rh) SIRT6. (G) Immunofluorescence staining of p53 acetylation at lysine 382 (red fluorescence) in SIRT6 WT or catalytically inactive mutant H133Y (green fluorescence) reconstituted SIRT6 KO cells. Scale bar, 10 µm. (H) Endogenous p53 immunoprecipitated from mock CRISPR, SIRT6 KO and SIRT6 KO cells with ectopic expression of FLAG-tagged wild-type (WT) SIRT6 followed by western blotting analysis of p53 acetylation at K381. (I) Western blotting analysis of p53 acetylation at K381 in the bone marrow stromal cells of around 24 days old Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- littermate mice. (J) Western blotting analysis of p53 acetylation at K381 in the kidneys of around 24 days old Sirt6+/+Trp53+/+, Sirt6-/-Trp53+/+ and Sirt6-/-Trp53+/- littermate mice.
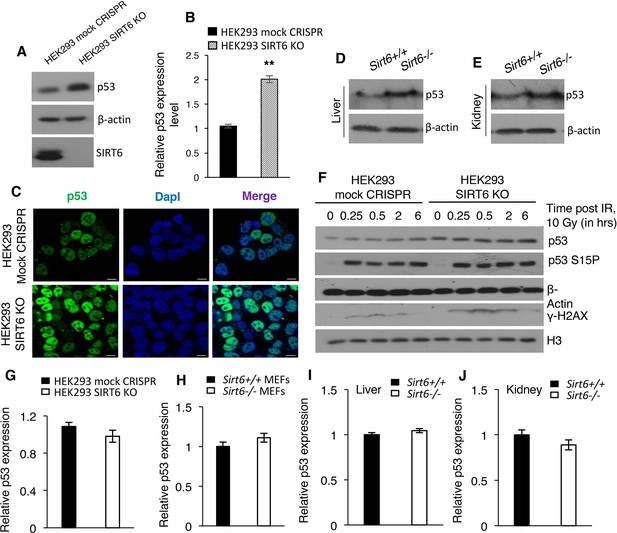
SIRT6 negatively regulates the stability of p53.
(A) Western blotting analysis of p53 protein expression in SIRT6 KO and control HEK293 cells. (B) Quantification of data presented in (A). Data represent mean ± SEM, n = 3. **p<0.01 calculated using Student’s t-test. (C) Immunofluorescence staining to confirm enhanced p53 expression in HEK293 mock CRISPR and SIRT6 KO cells. Scale bar, 10 µm. (D) Western blotting analysis of p53 protein expression in liver of wild-type (WT) and Sirt6-/- mice. (E) Western blotting analysis of p53 protein expression in kidneys of wild-type (WT) and Sirt6-/- mice. (F) Western blotting analysis of p53 and phosphorylation of p53 at serine 15 in HEK293 mock CRISPR and SIRT6 KO cells in response to 10 Gy of γ-irradiation. (G) qPCR analysis of p53 expression in HEK293 mock CRISPR and SIRT6 KO cells (with respect to Gapdh controls). Data represent mean ± SEM, n = 3. (H) qPCR analysis of p53 expression in Sirt6+/+ and Sirt6-/- MEFs (with respect to Gapdh controls). Data represent mean ± SEM, n = 3. (I) qPCR analysis of p53 expression in liver of Sirt6+/+ and Sirt6-/- mice (with respect to Gapdh controls). Data represent mean ± SEM, n = 3. (J) qPCR analysis of p53 expression in kidneys of Sirt6+/+ and Sirt6-/- mice (with respect to Gapdh controls). Data represent mean ± SEM, n = 3.
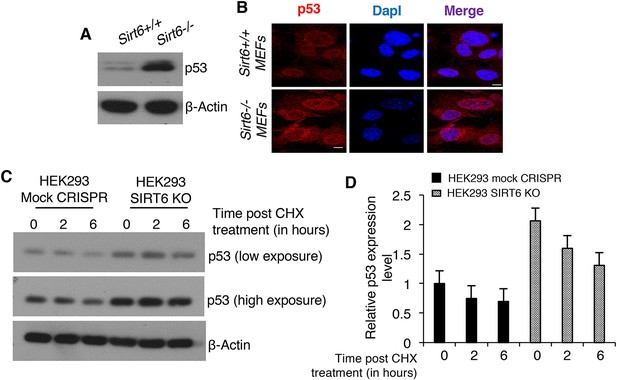
Analyses of the stability of p53 in the presence and absence of SIRT6.
(A) Western blotting analysis of p53 protein expression in Sirt6+/+ and Sirt6-/- MEFs. (B) Immunofluorescence staining to confirm enhanced p53 protein expression in WT and Sirt6-/- MEFs. Scale bar, 10 µm. (C) Western blotting analysis of p53 levels in HEK293 mock CRISPR and SIRT6 KO cells after 6 hr of treatment with 150 µg/ml cycloheximide (CHX) to block protein synthesis. (D) Quantification of data presented in (C). Data represent mean ± SEM, n = 3.
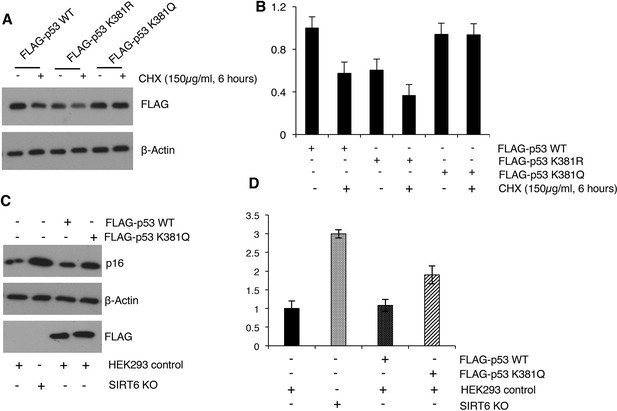
Acetyl mimic mutant p53 K381Q confers stability to p53 and imparts senescence-like properties to cells.
(A) FLAG-tagged p53 wild-type (WT), lysine 381 to arginine (K381R, non-acetylatable) mutant, and lysine 381 to glutamine (K381Q, acetyl-mimic) mutants were ectopically expressed in HEK293 cells. Then western blotting analysis of p53 levels (using FLAG antibodies) was performed in cells after 6 hr of treatment with 150 µg/ml cycloheximide (CHX) to block protein synthesis. (B) Quantification of data presented in (A). Data represent mean ± SEM, n = 3. (C) FLAG-tagged p53 wild-type (WT) and lysine 381 to glutamine (K381Q, acetyl-mimic) mutant were ectopically expressed in HEK293 cells, keeping SIRT6 KO cells as control and whole cell lysate was collected post 48 hr of transfection. Then western blotting analysis was performed to determine p16 expression levels in the cells, with respect to corresponding β-actin controls. (D) Quantification of data presented in (C). Data represent mean ± SEM, n = 3.
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32127.019





