Myotubularin related protein-2 and its phospholipid substrate PIP2 control Piezo2-mediated mechanotransduction in peripheral sensory neurons
Figures
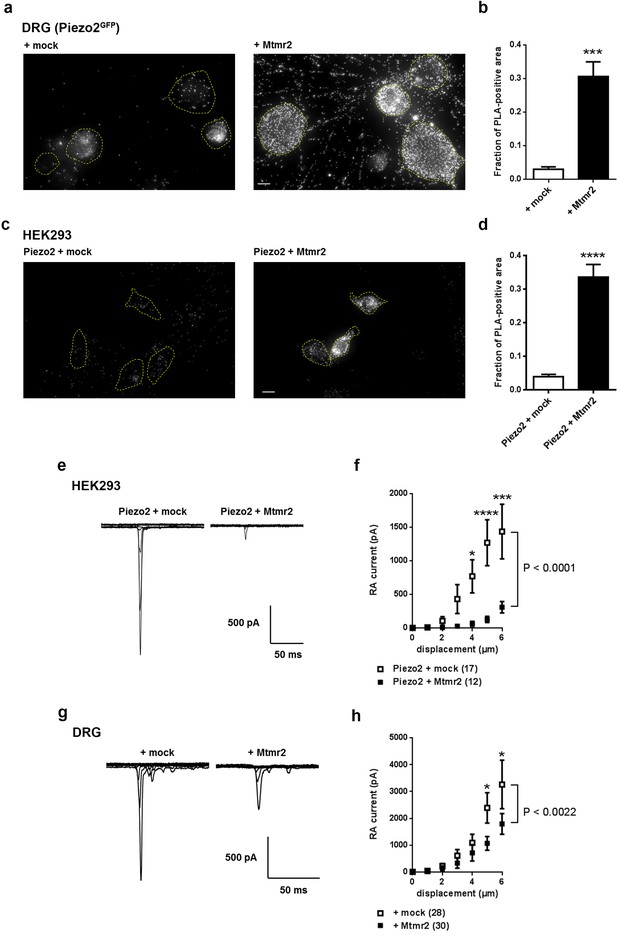
Mtmr2 suppresses Piezo2-mediated RA-MA currents in HEK293 cells and DRG neurons.
(a–d) Representative images (a,c) and quantification (b,d) of a proximity ligation assay (PLA) in cultured DRG neurons (a,b) of Piezo2GFP mice (Woo et al., 2014) and HEK293 cells (c,d). As anti-Mtmr2 antibodies failed to work in neuronal cultures, DRG were transfected with Mtmr2-myc or mock-myc and PLA was performed with antibodies against Piezo2 and myc. Please note the distribution of the PLA signal in soma and neurites of DRG. HEK293 cells were co-transfected with Piezo2-GST-IRES-GFP and Mtmr2-myc or Piezo2-GST-IRES-GFP and mock-myc and PLA was performed with antibodies against GST and myc. Only cells with pronounced GFP signal (due to expression of pmaxGFPVector in DRG and Piezo2-GST-IRES-GFP in HEK293 cells) were considered for the analysis. Cell boundaries are demarcated in yellow. In both cell types, DRG and HEK293 cells, transfection of Mtmr2-myc exhibited significantly stronger PLA signal compared to controls (b,d). Scale bar: 10 µm. Quantification of the total area of PLA signal/total soma area (fraction of PLA-positive area) in DRG cultures (p<0.0001; Mann-Whitney test; + mock: n = 53 neurons; + Mtmr2-myc: n = 53 neurons) (b). The quantification of the intensity of PLA signal in neurites of cultured DRG neurons can be found in Figure 1—figure supplement 1c. Quantification of the total area of PLA signal/total cell area in HEK293 cells (fraction of PLA-positive area) (p<0.0001; Mann-Whitney test; Piezo2-GST + mock: n = 60 cells; Piezo2-GST + Mtmr2-myc: n = 54 cells) (d). Additional controls for PLA in HEK293 cells can be found in Figure 1—figure supplement 1d. (e) Representative traces of RA-MA currents in HEK293 cells upon co-expression of Piezo2 with mock or Mtmr2 and (f) stimulus-current curves. Overexpression of Mtmr2 suppressed Piezo2 current magnitudes compared to mock overexpression (Piezo2 + mock: n = 17 cells; Piezo2 + Mtmr2: n = 12 cells; 2-way ANOVA suggested a significant effect (P<0.0001) of Mtmr2 overexpression on Piezo2 currents; Holm-Sidak’s multiple comparisons test was used to compare both conditions at individual stimulus magnitudes, p-values are indicated by * in the graph). The displacement threshold was increased upon co-expression of Mtmr2 (p=0.0098; Mann-Whitney test; Supplementary file 1). The inactivation time constant of RA-MA currents remained unchanged (Supplementary file 1). (g) Representative traces of RA-MA currents in primary cultures of DRG neurons and (h) stimulus-current curves showed a significant decrease in RA-MA current magnitude upon overexpression of Mtmr2 compared to mock ( + mock: n = 28 neurons; + Mtmr2: n = 30 neurons; 2-way ANOVA suggested a significant effect (P<0.0022) of Mtmr2 overexpression on RA-MA currents; Holm-Sidak’s multiple comparison test was performed to compare both conditions at individual stimulus magnitudes, p-values are indicated by * in the graph). The displacement threshold and inactivation time constant of RA-MA currents were not changed upon overexpression of Mtmr2 in DRG neurons (Supplementary file 1).
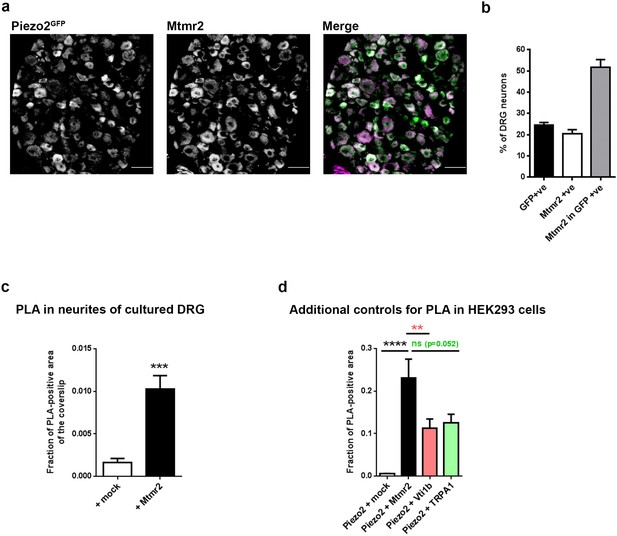
Mtmr2 is expressed in mouse DRG and also in close vicinity to Piezo2.
(a) Representative immunohistochemistry and (b) quantification of Mtmr2-positive neurons in cryosections of DRGs obtained from Piezo2GFP mice (Woo et al., 2014). 20.37 ± 2.01% of DRG neurons exhibit positive Mtmr2 immunolabel and 24.53 ± 1.21% of DRG neurons were positive for GFP indicating the presence of Piezo2. Among Piezo2-positive cells, 51.61 ± 3.71% cells were found to be Mtmr2-positive. n > 2000 neurons, N = 3 independent animals. (c) Quantification of the intensity of PLA signal in neurites of cultured DRG neurons (p<0.0001 compared to mock, Mann Whitney test; + mock: n = 94 regions from six coverslips; + Mtmr2-myc: n = 115 regions from six coverslips. (d) Quantification of PLA signal upon co-expression of Piezo2 with mock, Mtmr2, Vti1b (Vesicle transport through interaction with t-SNAREs homolog 1B) or TRPA1 in HEK293 cells. PLA was performed using antibodies against GST and myc to detect Piezo2-GST-IRES-GFP and myc-tagged proteins, respectively. PLA signal (fraction of PLA-positive area) upon co-transfection of mock, Vti1b and TRPA1 served as negative controls and was indeed much smaller than the PLA signal upon co-transfection of Piezo2 and Mtmr2 (p<0.01, in case of TRPA1 p=0.052 compared to Piezo2 + Mtmr2, Kruskal-Wallis test followed by Dunn´s multiple comparison test; Piezo2-GST + mock: n = 60 cells; Piezo2-GST + Mtmr2-myc: n = 35 cells; Piezo2-GST + Vti1b-myc: n = 54 cells; Piezo2-GST + TRPA1-myc: n = 83 cells). Only cells with pronounced GFP signal (due to expression of Piezo2-GST-IRES-GFP) were considered for the analysis. All experiments were performed in parallel.
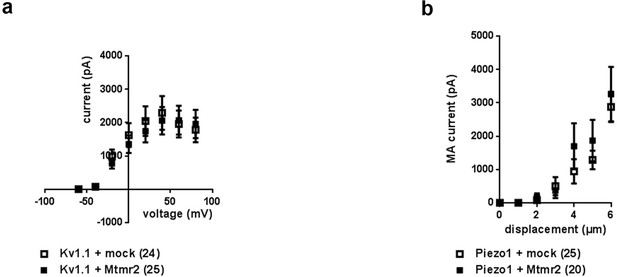
Mtmr2 overexpression does not influence Kv1.1- or Piezo1-mediated currents.
(a) Voltage-current curves upon overexpression of Kv1.1 with Mtmr2 in HEK293 cells showed no significant difference compared to overexpression with mock (Kv1.1 + mock: n = 24 cells; Kv1.1+ Mtmr2: n = 25 cells; ns; 2-way ANOVA). (b) Stimulus-current curves of Piezo1-mediated MA currents upon co-overexpression of Piezo1 (Piezo1-753-myc-IRES-GFP [Coste et al., 2015]) with Mtmr2 compared to mock. No significant difference was observed among conditions (Piezo1 + mock: n = 25 cells; Piezo1 + Mtmr2: n = 20 cells; ns; 2-way ANOVA). The displacement threshold and inactivation time constant of Piezo1-MA currents remained unchanged upon overexpression with Mtmr2 (Supplementary file 1).
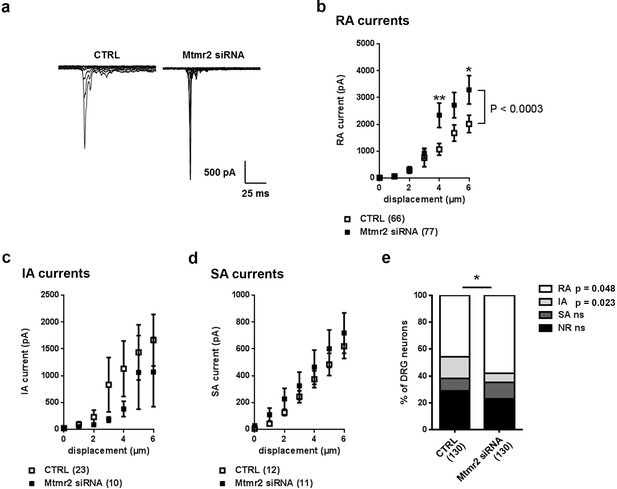
Mtmr2 knockdown potentiates Piezo2 activity in peripheral sensory neurons.
(a) Representative traces of RA-MA currents in primary cultures of DRG neurons and (b) stimulus-current curves for RA-MA currents upon nucleofection of Mtmr2 siRNA showed a significant increase in RA-MA current magnitude compared to nucleofection with AllStar Negative Control siRNA (CTRL: n = 66 neurons; Mtmr2 siRNA: n = 77 neurons; 2-way ANOVA suggested a pronounced effect (P<0.0003) of Mtmr2 knockdown on RA-MA currents; Holm-Sidak’s multiple comparisons test was used to compare both conditions at individual stimulus magnitudes, p-values are indicated by * in the graph). The displacement threshold and inactivation time constant of RA-MA currents remained unchanged upon knockdown of Mtmr2 (Supplementary file 1). Of note, MA current properties cannot be compared between different experiments or treatments of DRG cultures, for example Figure 2b cannot be compared to overexpression of Mtmr2 (Figure 1). Cultures were differently nucleofected (siRNA vs. plasmids) and recorded on different days in vitro (DIV) according to established protocols (please see Materials and methods for details). Hence matching controls were performed for each set of data. (c) Stimulus-current curves show IA-MA currents were unaffected by knockdown of Mtmr2 in DRG neurons (CTRL: n = 23 neurons; Mtmr2 siRNA: n = 10 neurons; ns; 2-way ANOVA). (d) Stimulus-current curves show SA-MA currents were unchanged upon Mtmr2 knockdown (n = 11–12 neurons per condition; ns; 2-way ANOVA). Of note, the displacement thresholds and inactivation time constants of IA-MA and SA-MA currents remained unchanged upon Mtmr2 siRNA nucleofection (Supplementary file 1). (e) Stacked histograms show the number of cells exhibiting different MA currents upon knockdown of Mtmr2 in cultured DRG. The proportions of cells exhibiting RA and IA currents were significantly changed in cultures transfected with Mtmr2 siRNA. RA:IA:SA:NR (% of total; rounded to whole numbers): CTRL: 46:16:9:29; Mtmr2 siRNA: 58:7:12:23; p<0.044 overall and for the proportion of RA/total (p=0.048) and IA/total (p=0.023), respectively; Chi-square test;≥130 neurons were analyzed per condition). NR (non-responsive), refers to cells which showed no MA current.
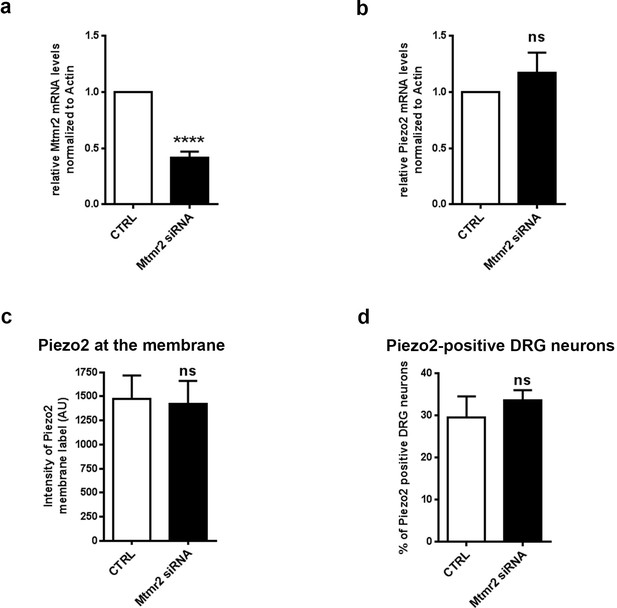
Mtmr2 knockdown in cultured DRG neither affects Piezo2 mRNA nor Piezo2 membrane levels or overall expression.
(a) Quantification of Mtmr2 mRNA upon siRNA transfection in DRG neurons showed a significant decrease in Mtmr2 mRNA (Actin as reference: 0.42 ± 0.05; p<0.0001; one sample t-test; N = 8 independent DRG cultures). Of note, our qPCR results indicate successful siRNA-mediated knockdown of Mtmr2 across the whole coverslip, which also includes non-transfected neurons and glia cells. Therefore our data do not report on the transfection efficiency and extent of Mtmr2 knockdown in individual neurons. (b) Quantification of Piezo2 mRNA upon Mtmr2 knockdown in DRG neurons confirmed that Piezo2 mRNA levels are similar to controls (Actin (Actb) as reference: 1.17 ± 0.18; ns; one sample t-test; N = 8 independent DRG cultures). (c) Quantification of Piezo2 membrane expression in DRG cultures derived from Piezo2GFP mice (Woo et al., 2014) upon Mtmr2 knockdown. No difference was observed among conditions (CTRL: 1470 ± 246.7 AU; Mtmr2 siRNA: 1424 ± 237.5 AU; n = 22 neurons per condition from N = 2 independent DRG cultures; ns; Mann-Whitney test). AU, arbitrary units. (d) Percentage of Piezo2-positive cells in DRG cultures derived from Piezo2GFP mice treated with CTRL or Mtmr2 siRNA. No difference was observed among analyzed conditions (CTRL: 29.59 ± 4.97%; Mtmr2 siRNA: 33.59 ± 2.45%; ns; Mann-Whitney test; n > 1000 neurons per condition; N = 3 independent DRG cultures).
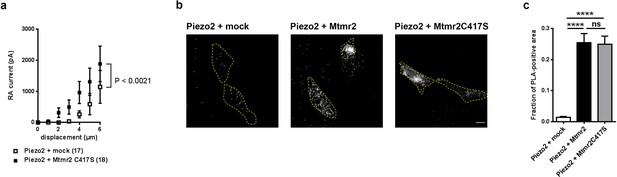
Catalytic activity of Mtmr2 is necessary to suppress Piezo2-mediated RA-MA currents.
(a) Stimulus-current curves upon co-expression of Piezo2 with the catalytically inactive Mtmr2 C417S mutant in HEK293 cells compared to mock controls. Mtmr2 C417S overexpression slightly, but not significantly increased Piezo2 RA-MA currents especially at lower stimulus magnitudes (Piezo2 + mock: n = 17 cells; Piezo2 + Mtmr2 C417S: n = 18 cells). 2-way ANOVA reported a significant (P<0.0021) overall effect of Mtmr2 C417S overexpression on RA-MA currents, however a Holm-Sidak’s multiple comparisons test showed no significant difference between currents at individual stimulus magnitudes.). Of note, the displacement thresholds and inactivation time constant were unaffected upon overexpression with Mtmr2 C417S compared to mock (Supplementary file 1). (b–c) Representative images (b) and quantification (c) of PLA signal using antibodies against GST and myc to detect Piezo2-GST-IRES-GFP and Mtmr2 C417S-myc, respectively. PLA signal upon co-transfection of Piezo2-GST + Mtmr2 C417S-myc was indistinguishable from Piezo2-GST + Mtmr2 myc and significantly stronger than Piezo2-GST + mock. Cell boundaries are demarcated in yellow. Only cells with pronounced GFP signal (due to expression of Piezo2-GST-IRES-GFP) were considered for the analysis. Scale bar: 10 µm. Quantification of the number of the total area of PLA signal/total cell area (fraction of PLA-positive area) (c); p<0.0001 compared to Piezo2-GST + mock, Kruskal-Wallis test followed by Dunn’s Multiple Comparison Test; Piezo2-GST + mock: n = 75 cells; Piezo2-GST + Mtmr2-myc: n = 70 cells; Piezo2-GST + Mtmr2 C417S-myc: n = 70 cells.
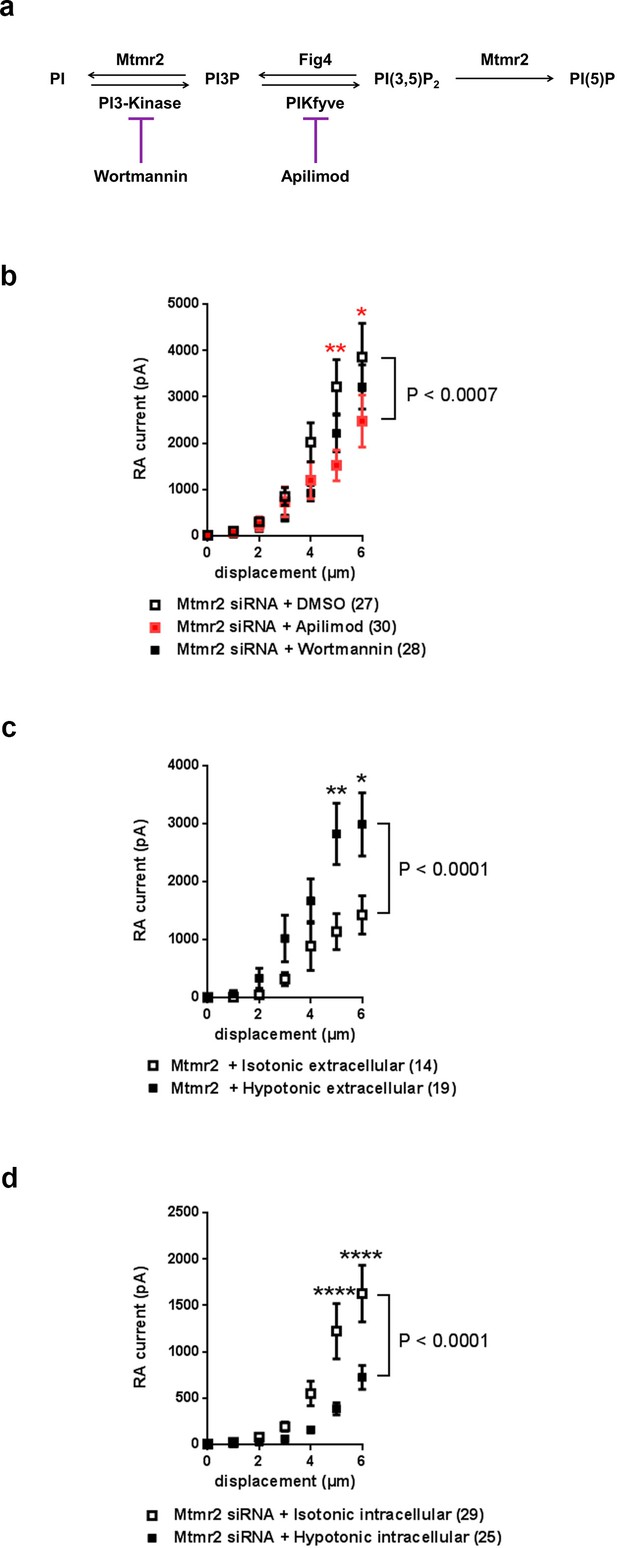
Mtmr2 modulates Piezo2-mediated RA-MA currents mainly via PI(3,5)P2.
(a) Scheme illustrating the major steps of PI(3,5)P2 synthesis and turnover including commonly used inhibitors and their targets. Wortmannin is an inhibitor of the phosphatidylinositol 3-kinase (PI3-Kinase) while Apilimod inhibits phosphatidylinositol 3-phosphate 5-kinase (PIKfyve). The Fig4 gene encodes a polyphosphoinositide phosphatase. (b) Stimulus-current curves after addition of Wortmannin, Apilimod or vehicle (DMSO) to Mtmr2 siRNA-treated neurons (Mtmr2 siRNA + DMSO: n = 27 neurons; Mtmr2 siRNA + Wortmannin: n = 28 neurons; Mtmr2 siRNA + Apilimod: n = 30 neurons). 2-way ANOVA suggested a significant (P<0.0007) overall effect on RA-MA currents. Holm-Sidak’s multiple comparisons test was performed to compare both conditions to DMSO at individual stimulus magnitudes. While no significant difference between Wortmannin and DMSO at individual stimulus magnitudes was observed, Apilimod application showed a significant reduction of currents compared to DMSO, p-values are indicated by * in the graph. Similarly, only Apilimod treatment increased the displacement threshold (p=0.0055 compared to DMSO-treated neurons, Kruskal-Wallis test followed by Dunn´s multiple comparisons test; Supplementary file 1). The inactivation time constants were unaltered by either treatment (Supplementary file 1). (c) Hypotonic extracellular solution counteracted the inhibition of Piezo2 RA-MA currents caused by Mtmr2 overexpression. Stimulus-current curves for Piezo2 RA-MA currents upon extracellular hypotonic stress application to DRG neurons overexpressing Mtmr2 (Mtmr2 + Isotonic extracellular solution: n = 14 neurons; Mtmr2 + Hypotonic extracellular solution: n = 19 neurons; 2-way ANOVA suggested that extracellular hypotonic stress had a significant (P<0.0001) effect on RA-MA currents. Holm-Sidak’s multiple comparisons test was performed to compare both conditions at individual stimulus magnitudes, p-values are indicated by * in the graph. The displacement threshold of RA-MA currents and inactivation time constant of RA-MA currents were unchanged (Supplementary file 1). (d) Hypotonic intracellular solution counteracted the potentiation of Piezo2 RA-MA currents caused by Mtmr2 knockdown. Stimulus-current curves for Piezo2 RA-MA currents upon intracellular hypotonic stress application to DRG neurons treated with Mtmr2 siRNA (Mtmr2 siRNA + Isotonic intracellular solution: n = 29 neurons; Mtmr2 siRNA + Hypotonic intracellular solution: n = 25 neurons; 2-way ANOVA suggested that intracellular hypotonic stress had a significant (P<0.0001) effect on RA-MA currents. Holm-Sidak’s multiple comparisons test was performed to compare both conditions at individual stimulus magnitudes, p-values are indicated by * in the graph. The displacement threshold of RA-MA currents was increased upon intracellular hypotonic stress (p=0.0131; Mann-Whitney test; Supplementary file 1). The inactivation time constant of RA-MA currents was unchanged (Supplementary file 1).
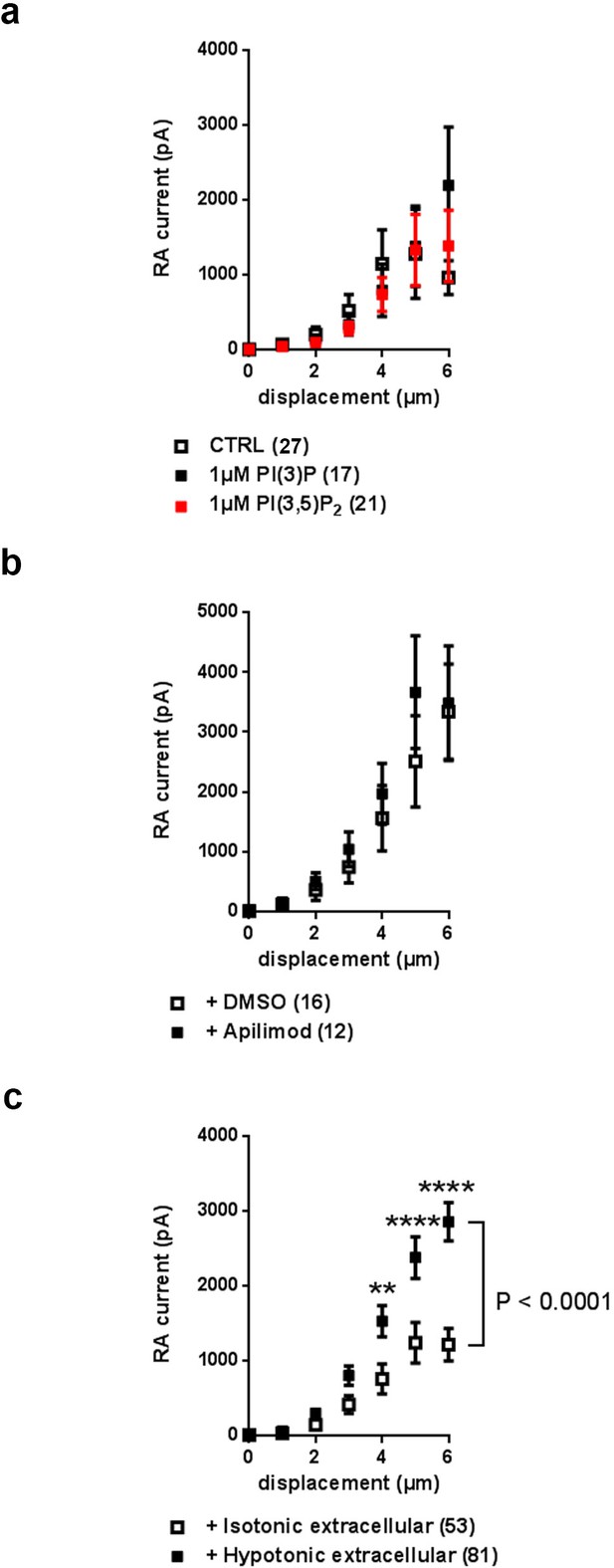
Effect on Piezo2 RA-MA currents upon application of PIPs or Apilimod in cultured DRG.
(a) Stimulus-current curves of RA-MA currents are not altered upon addition of 1 µM PI(3)P or 1 µM PI(3,5)P2 (CTRL: n = 27 neurons; 1 µM PI(3)P: n = 17 neurons; 1 µM PI(3,5)P2: n = 21 neurons; ns; 2-way ANOVA). The displacement threshold and inactivation time constant of RA-MA currents also remained unchanged upon application of PIPs (Supplementary file 1). (b) Stimulus-current curves of RA-MA currents upon application of Apilimod or DMSO (vehicle) to wild type, untreated DRG neurons ( + DMSO: n = 16 neurons; + Apilimod: n = 12 neurons; ns; 2-way ANOVA). The displacement threshold and inactivation time constant (τ) of RA-MA currents remained unchanged upon treatment with Apilimod (Supplementary file 1). (c) Stimulus-current curves demonstrate Piezo2 RA-MA potentiation upon application of extracellular hypotonic stress to DRG cultures compared to isotonic conditions (Isotonic extracellular: n = 53 neurons; Hypotonic extracellular: n = 81 neurons). 2-way ANOVA suggested that hypotonic stress had a significant (P<0.0001) effect on RA-MA currents. Holm-Sidak’s multiple comparisons test was performed to compare conditions at individual stimulus magnitudes, p-values are indicated by * in the graph. The displacement threshold of RA-MA currents was significantly decreased by hypotonic stress (p=0.0021; Mann-Whitney test; Supplementary file 1). Given that Jia and colleagues (Jia et al., 2016) reported slower inactivation of MA currents upon prolonged hypotonic stress, we wanted to ensure that measured currents in our paradigm of acute hypotonic stress were of the RA-type. In fact, this was the case as judged by comparable inactivation time constants (Supplementary file 1). Of note, MA current properties shown here in wild type, untreated DRG neurons cannot be compared to nucleofected neurons in Figure 4. Nucleofection with siRNA or plasmids (i) alters neuronal and Piezo2 activity and (ii) required recordings to be performed on different days in vitro (2 DIV and 3 DIV, respectively) according to established protocols. Please see Materials and methods for more details on variability in DRG cultures. For this reason each dataset consists of experiments and respective matching controls measured in parallel (i) in the same mouse cohort and (ii), where possible, on each experimental day.
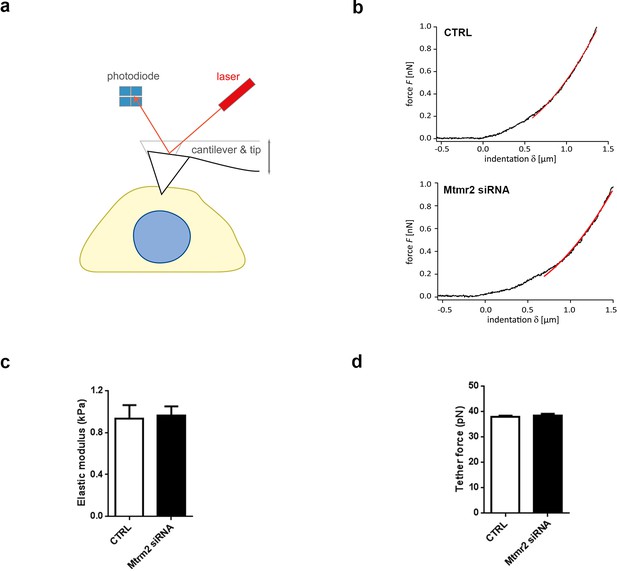
Mtmr2 knockdown does not obviously alter mechanical properties of cultured DRG neurons.
(a) Sketch of the AFM set-up used to measure the mechanical properties of cultured DRG neurons transfected with CTRL or Mtmr2 siRNA. (b) Representative force-indentation curves for CTRL and Mtmr2 siRNA treated DRG neurons. (c) Quantification of effective Young’ modulus (Eeff) of DRG neurons shows no significant difference between CTRL and Mtmr2 siRNA treatment (CTRL: 0.93 ± 0.13 kPa, n = 44 neurons from N = 4 independent cultures; Mtmr2 siRNA: 0.96 ± 0.09 kPa, n = 48 neurons from N = 4 independent cultures; ns; Mann-Whitney test). (d) Quantification of tether force also showed no difference between conditions (37.74 ± 0.55 pN, n = 53 from N = 4 independent cultures; Mtmr2 siRNA: 38.30 ± 0.76 pN, n = 54 from N = 4 independent cultures; ns; Mann-Whitney test).
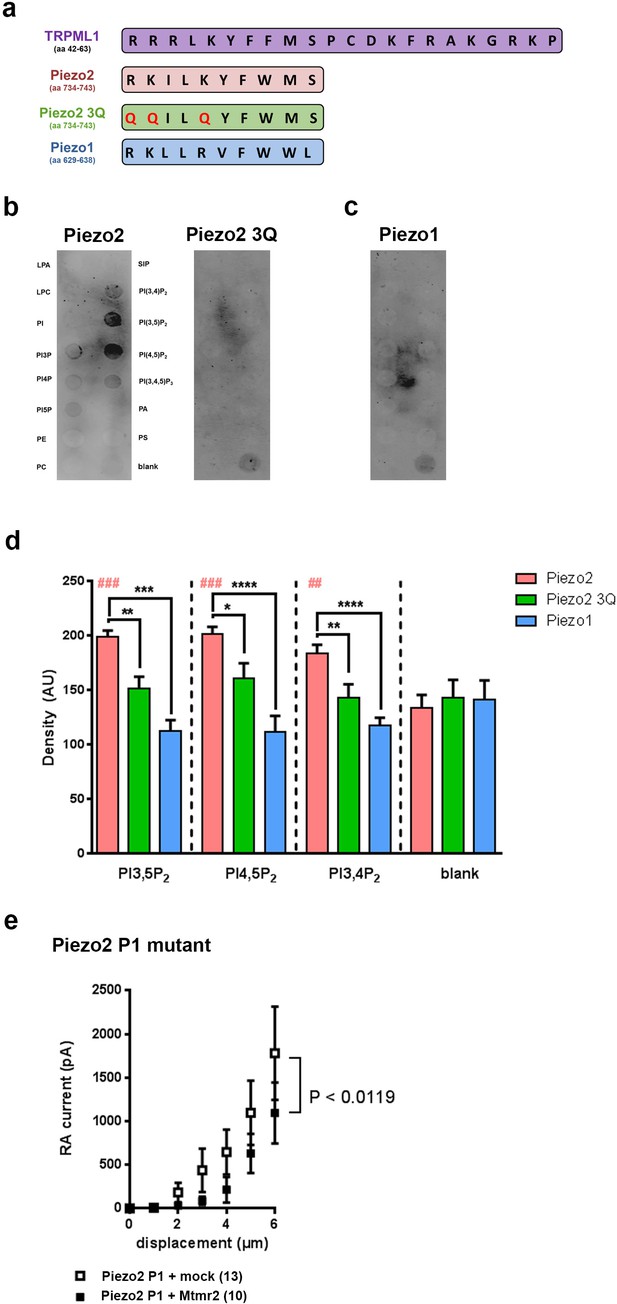
Murine Piezo2, but not Piezo1, harbors a PIP2 binding motif.
(a) Schematic view of the PI(3,5)P2 binding region of TRPML1 identified elsewhere (Dong et al., 2010) and the region in murine Piezo2 that exhibits pronounced sequence similarity to the PI(3,5)P2 binding region of TRPML1. The indicated sequences were used to generate peptides for Piezo2, the Piezo2 3Q mutant and Piezo1. All peptides were tagged with a FLAG-epitope to allow for detection with anti-Flag antibodies on immunoblots. (b–d) Representative peptide-lipid binding assays followed by immunoblotting after incubation with indicated peptides (b, c) and densitometric quantification (d). The arrangement of lipids on the lipid-strip is indicated; blank, no lipid was spotted. The Piezo2 peptide strongly binds to PI(3,5)P2 and PI(4,5)P2, and weakly to PI(3,4)P2. Neither the Piezo2 3Q mutant peptide nor the Piezo1 peptide exhibited significant binding to any of the lipids tested. One-way ANOVA followed by Dunnett’s multiple comparisons test was used to compare spot signal densities for each peptide to the respective blank, p-values are indicated by # in the graph (d). In addition, one-way ANOVA followed by Holm-Sidak’s multiple comparisons test was used to compare spot signal densities across the three peptides, p-values are indicated by * in the graph (d). The graph only presents data for those lipids, to which the Piezo2 peptide exhibited significant binding. Please see Figure 5—figure supplement 1 for summarized data on lipids tested with the Piezo2 peptide. Experiments using the Piezo2 peptides were independently repeated 6 times, of which four times were conducted in parallel with experiments using the Piezo1 peptide. AU, arbitrary units. (e) Stimulus-current curves upon co-expression of the Piezo2 P1 mutant with Mtmr2 in HEK293 cells compared to mock controls. Mtmr2 only slightly attenuated MA currents (Piezo2 P1 mutant + mock: n = 13 cells; Piezo2 P1 mutant + Mtmr2: n = 10 cells). 2-way ANOVA reported a significant (P<0.0119) overall effect of Mtmr2 overexpression on RA-MA currents of the Piezo2 P1 mutant, however a Holm-Sidak’s multiple comparisons test showed no significant difference between currents at individual stimulus magnitudes. Of note, the displacement threshold and inactivation time constant were also unaffected upon overexpression of Mtmr2 compared to mock (Supplementary file 1).
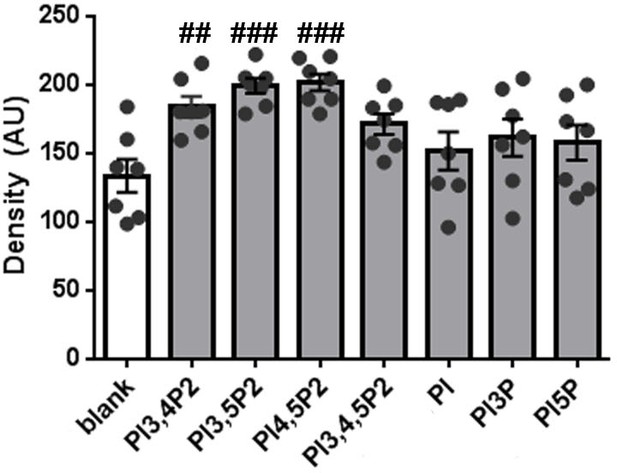
Quantification of peptide-lipid binding assays using the Piezo2 peptide.
Piezo2 peptide densitometry data were analyzed using one-way ANOVA followed by Dunnett’s multiple comparisons test to compare spot signal densities for different lipids to the blank, p-values are indicated by # in the graph. AU, arbitrary units.
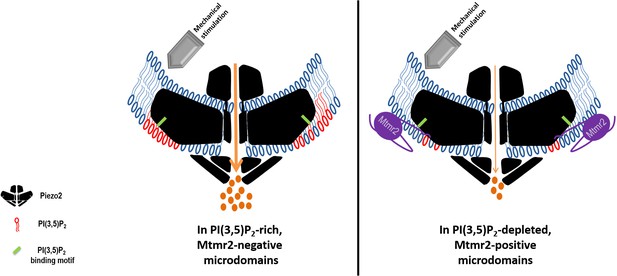
Working model: Local control of Piezo2 function by interdependent actions of Mtmr2 and PI(3,5)P2.
Mtmr2 controls the abundance of PI(3,5)P2 by dephosphorylation (please see Figure 4a). Mtmr2 and Piezo2 expression as well as PI(3,5)P2 might be compartmentalized in membrane microdomains. Piezo2 localization in Mtmr2-negative microdomains would facilitate its access to local PI(3,5)P2 and consequently potentiate Piezo2 RA-MA currents (left side). On the other hand, high Mtmr2 levels and its localization in the proximity of Piezo2 would augment PI(3,5)P2 turnover, thereby decreasing local PI(3,5)P2 availability and suppressing Piezo2 RA-MA currents (right side). One could further speculate that Mtmr2, via binding Piezo2, might recruit Piezo2 to membrane microdomains depleted of PI(3,5)P2. This would provide an active mechanism to inhibit Piezo2 RA-MA currents in membrane compartments – may they be at the plasma membrane or intracellular membranes. Ultimately, Mtmr2 and PI(3,5)P2 may contribute to dynamically tuning touch sensitivity of an organism in response to diverse conditions modulating Mtmr2 and PI(3,5)P2 levels (e.g. osmotic stress as indicated by results shown in Figure 4).The following questions await further clarification: (i) How are Mtmr2/PI(3,5)P2 regulated during (patho)physiological conditions in the somatosensory system, (ii) does the modulation of RA-MA currents require additional yet to be identified effector proteins, and (iii) are other PIPs also involved, for example PI(4,5)P2 shown to modulate Piezo2 function (Borbiro et al., 2015)? As the structure of Piezo2 has not been resolved yet, Piezo2 is depicted after the recently solved structure of Piezo1 (Ge et al., 2015; Guo and MacKinnon, 2017; Saotome et al., 2018). If this structure holds also true for Piezo2, the PI(3,5)P2 binding domain (depicted in green) would roughly be localized within the first third of the N-terminal blade.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (mouse) | B6/J mice | RRID: IMSR_JAX:000664 | bred in the animal facility of the MPIem Goettingen | |
| Strain (mouse) | Piezo2GFP | kind gift of Ardem Patapoutian | bred in the animal facility of the MPIem Goettingen | |
| Cell line (human) | HEK293 | purchased from ATCC | RRID: CVCL_0045 | Cells were not tested for mycoplasma contamination; cells were authenticated by ATCC upon purchase |
| Antibody | Rabbit anti-Mtmr2 (1:100) | Biotechne, #NBP1-33724 | RRID: AB_2147841 | |
| Chicken anti-GFP (1:500) | Thermo Fisher Scientific, #A10262 | RRID: AB_2534023 | ||
| Rabbit anti-GST (1:500) | Santa Cruz, #sc-459 | |||
| Mouse anti-myc (1:750, 1:500) | Santa Cruz, #sc-47694 | RRID: AB_627266 | ||
| Mouse anti-FLAG (1:100) | Sigma Aldrich, #F1804 | RRID: AB_262044 | ||
| Rabbit anti-Piezo2 (1:200) | Novus Biologicals, #NBP1-78624 | RRID: AB_11005294 | ||
| Recombinant DNA reagent | pCMVSport6 Piezo2-GST IRES GFP | kind gift of Ardem Patapoutian | mouse Piezo2 | |
| pCMV6-Entry Mtmr2-myc-DDK | Origene, #MR215223 | mouse Mtmr2 | ||
| Mtmr2C417S-myc-DDK | mouse Mtmr2 C417S | Mutation generated using Q5 Site-Directed Mutagenesis kit (New England BioLabs) | ||
| pCMV Sport6 Piezo1-753-myc-IRES GFP | kind gift of Ardem Patapoutian | mouse Piezo1 | Myc tag was inserted at amino acid 753 as described inCoste et al., 2015. | |
| pGEM-Teasy Kv1.1-HA | mouse Kv1.1 | Custom-made and sequence-verified | ||
| pCMVSport6 | kind gift of ArdemPatapoutian | |||
| pCDNA3.1-myc-His | Invitrogen, #V80020 | |||
| pCNDA3-GST | kind gift of Ardem Patapoutian | |||
| pCMVSport6 Piezo2 P1 mutant-GST IRES GFP | mouse Piezo2 P1 mutant | Mutation generated using Q5 Site-Directed Mutagenesis kit (New England BioLabs) | ||
| pCDNA3.1-myc-His TRPA1 | kind gift of Ardem Patapoutian | mouse TRPA1 | ||
| pCMV6-Vti1b-myc-DDK | Origene | |||
| Sequence-based reagent | Mtmr2 forward primer for qPCR | MPIem DNA Core Facility | TGTACCCCACCATTGAAGAAA | |
| Mtmr2 reverse primer for qPCR | MPIem DNA Core Facility | TAAGAGCCCCTGCAAGAATG | ||
| Piezo2 forward primer for qPCR | MPIem DNA Core Facility | AGGCAGCACATAGGATGGAT | ||
| Piezo2 reverse primer for qPCR | MPIem DNA Core Facility | GCAGGGTCGCTTCAGTGTA | ||
| Actb forward primer for qPCR | MPIem DNA Core Facility | GATCAAGATCATTGCTCCTCCTG | ||
| Actb reverse primer for qPCR | MPIem DNA Core Facility | CAGCTCAGTAACAGTCCGCC | ||
| Gapdh forward primer for qPCR | MPIem DNA Core Facility | CAATGAATACGGCTACAGCAAC | ||
| Gapdh reverse primer for qPCR | MPIem DNA Core Facility | TTACTCCTTGGAGGCCATGT | ||
| Piezo2 mutagenesis forward primer | MPIem DNA Core Facility | GTCTTCTGGTGGCTCGTGGTCATTTATACCATGTTGG | ||
| Piezo2 mutagenesis reverse primer | MPIem DNA Core Facility | ACGCAGCAGCTTCCTCCACCACTCGTAGTGCAC | ||
| Mtmr2 mutagenesis forward primer | MPIem DNA Core Facility | GTGGTACACTCCAGTGATGGATG | ||
| Mtmr2mutagenesis reverse primer | MPIem DNA Core Facility | CACAGACGTCTTCCCAGA | ||
| Peptide, recombinant protein | Piezo2-FLAG tagged | Custom-made by GenScript | EWWRKILKYFWMSVVIDYKDDDDKQNN | |
| Piezo2 3Q-FLAG tagged | Custom-made by GenScript | EWWQQILQYFWMSVVIDYKDDDDKQNN | ||
| Piezo1-FLAG tagged | Custom-made by GenScript | TLWRKLLRVFWWLVDYKDDDDKQNN | ||
| Chemical compound, drug | Wortmannin | Sigma Aldrich | ||
| Apilimod | Bertin Pharma | |||
| PI(3,5)P2 | Echelon | |||
| PI(3)P | Echelon | |||
| Software, algorithm | Fitmaster | HEKA Electronik GmbH | ||
| Patchmaster | HEKA Electronik GmbH | |||
| ImageJ | NIH (Schindelin et al., 2015) | RRID: SCR_003070 | ||
| GraphPad Prism 6.01 | GraphPad Software | RRID: SCR_015807 |
Additional files
-
Supplementary file 1
Summary of properties of MA currents elicited under various conditions in HEK293 cells and DRG neurons
The table shows the displacement threshold and inactivation time constant (τ) values for all electrophysiological data presented in this study (please see Materials and methods for details on the calculation of each value). Values are represented as mean ± SEM and cell numbers are indicated by ‘n’. Data were not significant (ns) unless otherwise mentioned. Please note: for SA-MA currents, it was not possible to fit the current traces of all cells with a mono or bi-exponential fit (please see Materials and methods for details), hence the cell numbers measured for the inactivation time constant (τ) are lower than actual cell numbers measured and reported in Figure 2.
- https://doi.org/10.7554/eLife.32346.015
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32346.016





