Lymphatic endothelium stimulates melanoma metastasis and invasion via MMP14-dependent Notch3 and β1-integrin activation
Figures
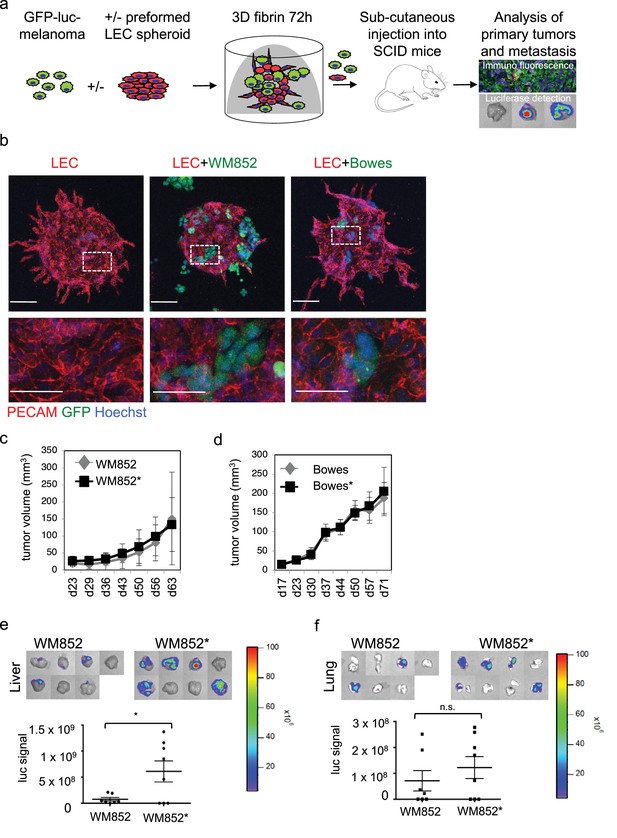
Co-culture of melanoma cells and LECs reveals melanoma invasion into the LEC 3D structures and increases the metastatic potential of WM852 cells in vivo.
(a) Schematic of the experimental pipeline. (b) Confocal images of LEC spheroids (PECAM-1, red) in 3D fibrin matrix (left panel), LEC spheroids co-cultured with WM852 (green, middle panel) or Bowes (green, left panel). The area enclosed in the white square is shown enlarged below each panel. Melanoma cells were stained with GFP (green), and nuclei were counterstained with Hoechst 33342. Maximum intensity Z-projections of confocal stacks are shown. (c,d) Growth rates of the 3D LEC primed WM852* (c) and Bowes* (d) derived tumors (n = 8 for both cell types) compared to control WM852 (n = 7) and Bowes (n = 8) tumors, respectively. (e, f) Distant organ metastasis, detected by bioluminescence imaging of luciferase signal, in liver (e) and lung (f) of SCID mice subcutaneously injected with WM852 alone or co-cultured with LECs (WM852*). Upper panels: representative images of the indicated organs, each box represents an organ from one mouse. Bottom panel: quantification of luciferase signal, each dot represents the luciferase value in one sample. Horizontal line indicates the average, vertical bars represent SEM. *: p<0.05. n.s., non-significant.
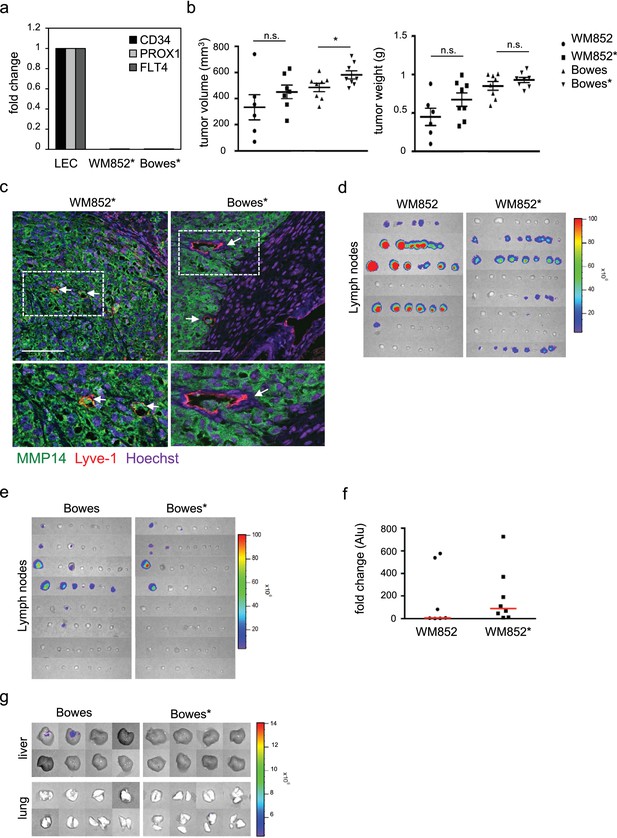
Analysis of mouse xenografts and distant organ metastasis.
(a) mRNA expression of a panel of LEC markers (CD34, PROX1, FLT4) in 3D LEC primed Bowes and WM852 cells (*) before the cells were used for the in vivo xenograft assay. Expression in monotypic-cultured LEC was used as a control and set to one. (b) End point analysis of the volume (left panel) and weight (right panel) of the Bowes/Bowes* and WM852/WM852* tumors. A dot represents one mouse. Error bars indicate s.e.m. *: p<0.05. n.s., non-significant (c) Immunohistochemistry of xenograft sections from the LEC co-cultured WM852* (left panels) or Bowes* (right panels) stained for mouse Lyve-1 (red) to detect mouse lymphatic vessels and human MMP14 (green) as a marker of the human melanoma cells. Nuclei were counterstained with Hoechst 33342. Scale bar = 100 µm. (d, e) Luciferase signal from lymph nodes isolated from mice bearing WM852 (n = 7) and WM852*(n = 8) (d), or Bowes and Bowes* (n = 8) (e); each line shows lymph nodes from one mouse derived tumors. (f) Quantitative-PCR (q-PCR) for human Alu sequences from lung genomic DNA isolated from mice bearing WM852 and WM852* derived tumors. Each dot in the graph represents the luciferase signal intensity obtained from one isolated organ per mouse, and the red line marks the median of the samples. (g) Luciferase signal from liver and lungs isolated from mice bearing Bowes or Bowes* derived tumors (n = 8); each box represents an organ from one mouse.
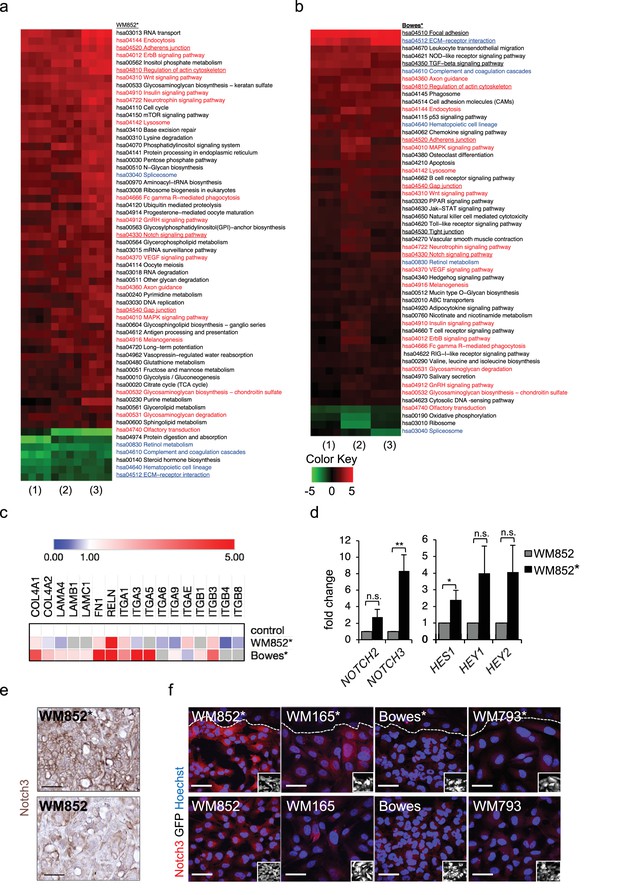
Transcriptomic analysis reveals Notch3 upregulation in the LEC primed WM852 cells.
(a–b) Generally Applicable Gene-set Enrichment (GAGE) for RNA-seq pathway analysis of LEC primed (a) WM852* and (b) Bowes* cells. Samples were compared to their respective cells derived from monotypic cultures. Three biological replicates per sample group and four run replicates were used. In the heatmap, red represents upregulated and green downregulated pathways in WM852* and Bowes*. Pathways enriched in both cell lines are marked with red text if they were similarly upregulated or downregulated, and blue text if they were differentially upregulated or downregulated. Pathways enriched only in one cell line are marked with black text. Pathways with underlined text were used for further analysis. (c) Heatmap depicting average expression fold change of the differentially expressed ECM-receptor interaction pathway genes in the RNA-seq of LEC primed WM852* and Bowes* cells. The WM852 and Bowes cells from monotypic cultures were used as controls, and set to one. Red represents upregulated and blue downregulated genes in WM852* and Bowes*. Adjusted p-values are less than 0.05 for all genes shown. (d) Relative mRNA fold change of the indicated targets in WM852 and WM852*. *:p<0.05; **p<0.001; n.s., non-significant. (e) Representative images of Notch3 immunohistochemistry in the WM852 and WM852* derived xenografts. Scale bar = 50 µm. (f) Representative confocal images of Notch3 staining (red) in different GFP expressing melanoma cell lines (GFP positive cells shown in the inset) cultured in the presence (*, upper panels) or absence (bottom panels) of LECs. Nuclei are counterstained with Hoechst 33342. The dashed line indicates the LEC-melanoma (below the line) border. Scale bar = 50 µm. Full size confocal images are available as a source data 1.
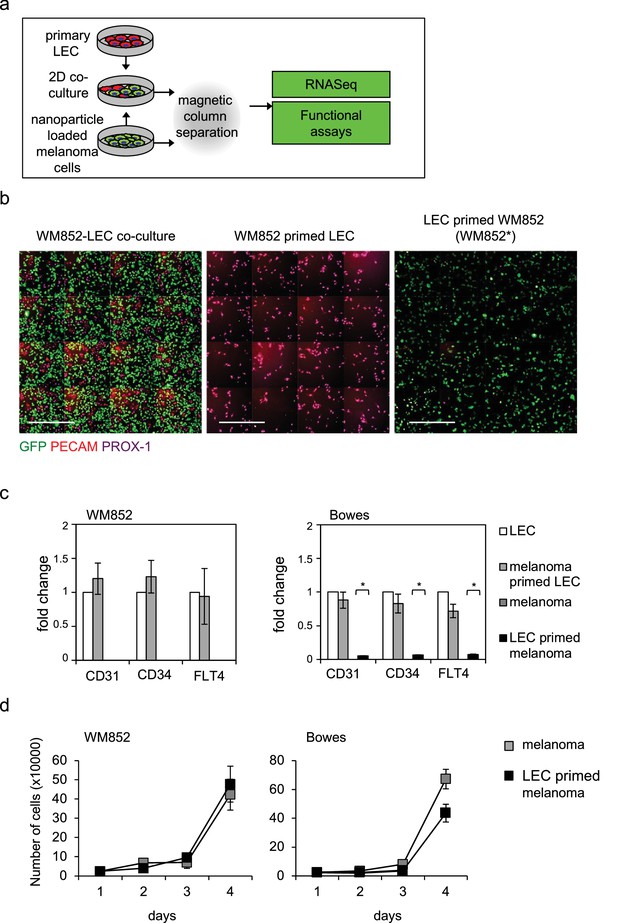
Optimization of magnetic separation of melanoma and LEC cells following co-culture of LEC and melanoma cells.
(a) Schematic of the workflow for cell separation from the melanoma-LEC 2D co-cultures. (b) WM852-LEC co-culture (left), flow through of the magnetic column (middle) and the separated elute (right) were stained for GFP (green), PECAM (red), and PROX-1 (magenta) and visualized by indirect immunofluorescence using high-content imaging. Panels of 16 (4 × 4) images are shown. (c) Relative mRNA fold changes of the indicated targets in LECs and melanoma cells derived from monotypic cultures or co-cultures (*) followed by magnetic separation. Average of three independent experiments, error bars indicate SEM. *: p<0.05. (d) Cell count at different time points following magnetic separation of WM852 (left) and Bowes (right) from monotypic culture or after LEC priming in the co-culture. Graphs show an average of two independent experiments, error bars indicate SD.
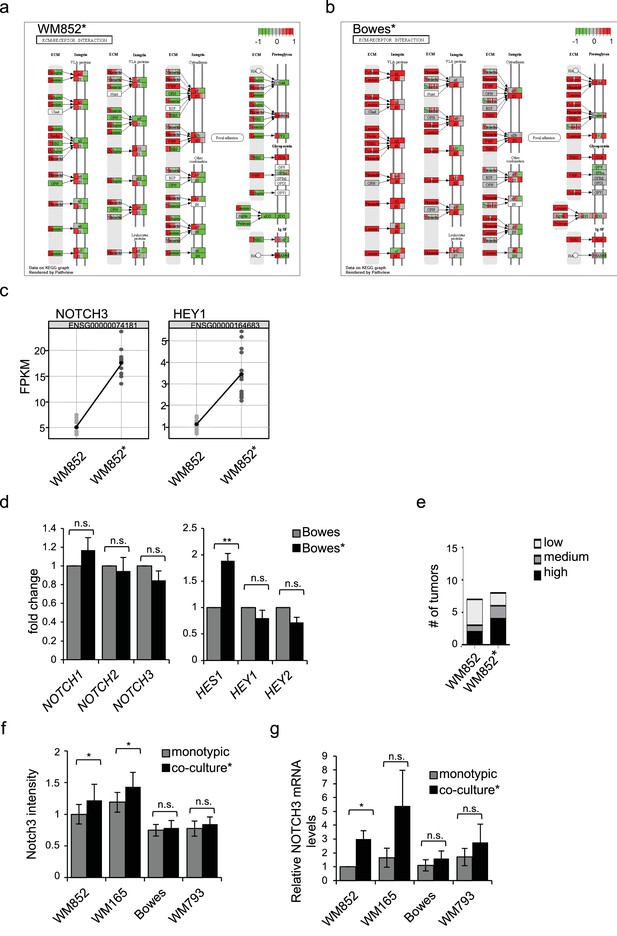
LEC priming induced differential changes in the ECM-receptor interaction pathway and Notch signaling pathway.
(a–b) ECM-receptor interaction (hsa04512) KEGG pathway map rendered with Pathview from RNA-seq data described in Figure 2 of (a) WM852* and (b) Bowes* cells. Red represents upregulated and green downregulated genes in WM852* and Bowes*. (c) FKPM values of NOTCH3 and HEY1 gene expression in WM852* cells in RNA sequencing analysis in Figure 2A. A dot represents one replicate. (d) mRNA fold change of the indicated targets in Bowes and Bowes*. Average from three independent experiments is shown. **: p<0.001; n.s., non-significant. (e) Quantification of Notch3 staining intensity in xenografts originating from WM852 (n = 7) or WM852* (n = 8). (f) Quantification of Notch3 signal intensity for the indicated cell lines. Average of the intensity was calculated in four images/condition (from two independent experiments. More than 100 cells were always analysed per condition. Error bars indicate SD. *:p<0.05; n.s., non-significant. (g) NOTCH3 mRNA fold change in the indicated melanoma cell lines from monotypic culture or co-cultured with LEC (*). Data averaged from four independent experiments is shown. *:p<0.05; n.s., non-significant.
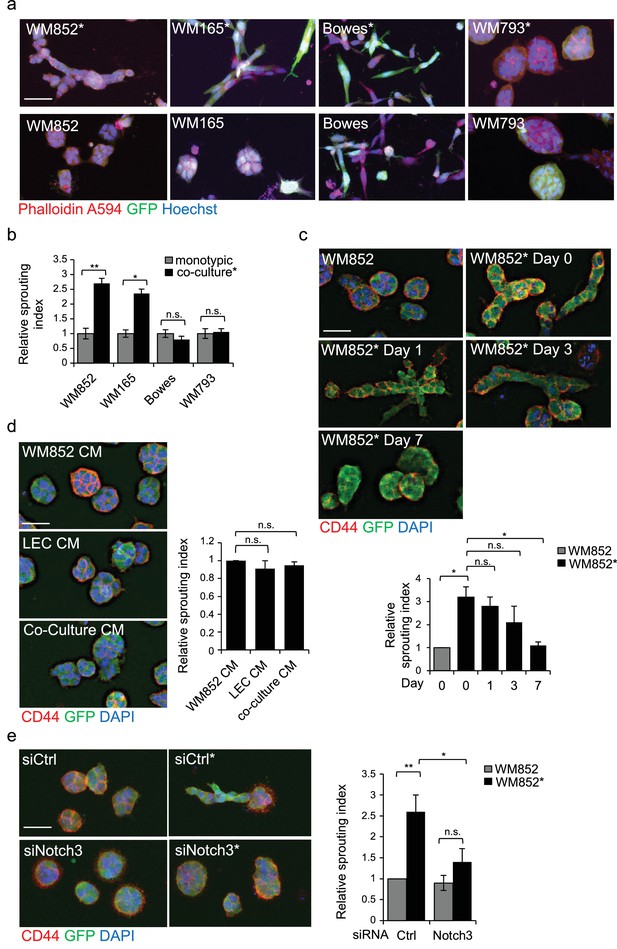
LEC interaction modifies the 3D growth phenotype of the melanoma cells.
(a) Representative confocal images of 3D fibrin assays after magnetic separation of the indicated melanoma cell lines co-cultured with LEC (*, upper panels) or from monotypic culture (bottom panels). GFP expressing (green) melanoma cells were stained with Phalloidin A594 (red), nuclei are counterstained with Hoechst 33342 (blue). Maximum intensity Z-projections of the confocal stacks are shown. Scale bar = 50 µm. (b) Quantification of the sprouting index for the samples in (a). Graphs show the average of at least three images per condition per two independent experiments, error bars indicate the SEM p<0.05; **; p<0.01; n.s., non-significant. (c) Representative images of the 3D fibrin assay of WM852 and WM852* after magnetic separation at the indicated day after separation. The graph represents the average of three images per condition analysed in each of the two independent experiments. Error bars represent SEM. *: p<0.05; n.s., non-significant. (d) Representative images of the 3D fibrin assay of monotypic WM852 treated with conditioned media (CM) from the indicated sources. The graph represents the average of three images per condition. Error bars represent SEM. n.s., non-significant. (e) Representative images of the 3D fibrin assay of WM852 and WM852* treated with the indicated siRNAs for 72 hr prior to magnetic separation and fibrin embedding. The graph represents the average of three images per condition per three independent experiments. Error bars represent SEM. *: p<0.05; **; p<0.01; n.s., non-significant.
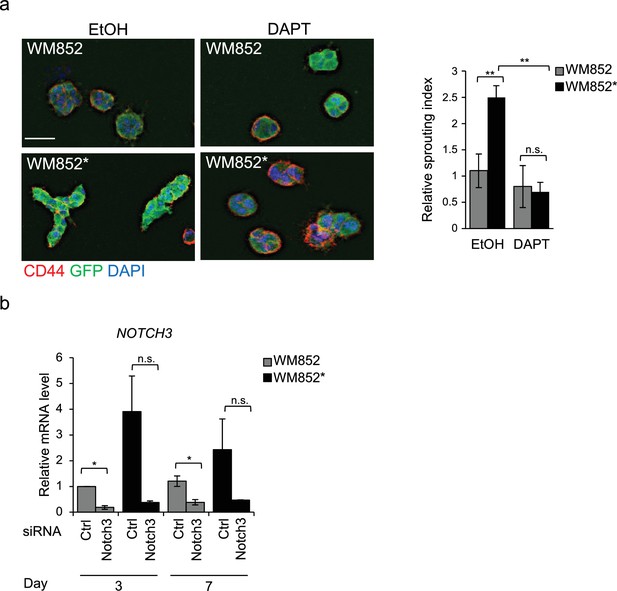
3D growth phenotype of metastatic melanoma cells is Notch3 dependent.
(a) Representative images (left) and quantification (right) of 3D fibrin assay of WM852 and WM852* treated with vehicle (EtOH) or DAPT for 48 hr during co-culture, after magnetic separation and also during the 3D fibrin assay. Relative sprouting index from three independent experiments is shown; **: p<0.01; n.s., non-significant. (b) NOTCH3 mRNA fold change in WM852 and WM852* treated with the indicated siRNAs for 72 hr and subjected to magnetic separation after co-culture and analysed at the indicated time points. *: p<0.05; n.s., non-significant.
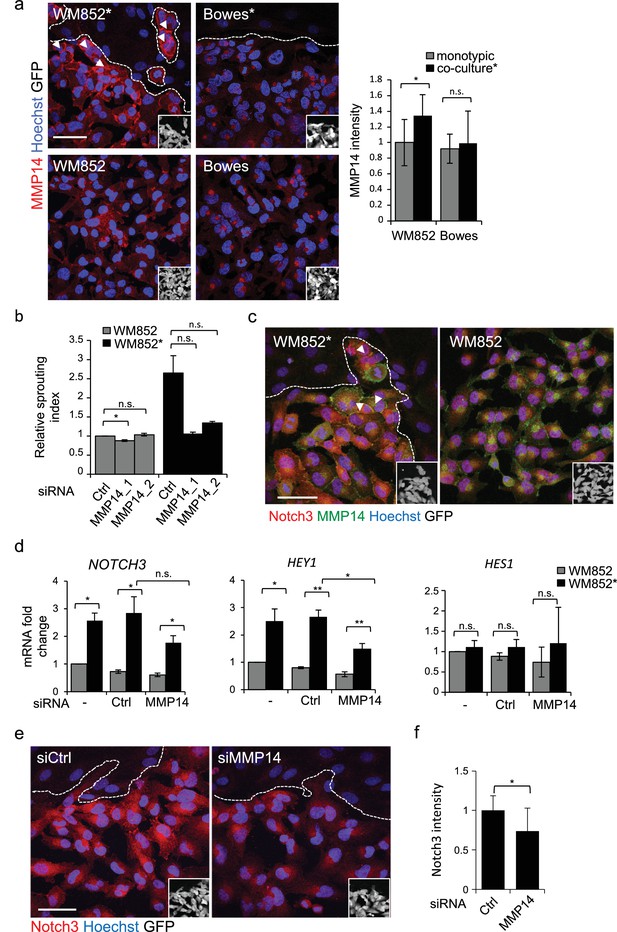
MMP14 is required for the increased sprouting growth of LEC primed melanoma cells in 3D.
(a) Right panels: representative confocal images of MMP14 (red) expression in WM852 and Bowes co-cultured with LECs (*, upper panels) and from monotypic culture (bottom panels). Nuclei were counterstained with Hoechst 33342. GFP-expressing melanoma cells are shown white in the inset. Arrowheads indicate MMP14 localization to the cell-cell contacts. The dashed line indicates the LEC-melanoma (below the line) border. Scale bar = 50 µm. Left panel: quantification of MMP14 intensity analysed in four images per condition from two independent experiments. More than 100 cells were always analysed per condition. Average is shown, error bars represent SD; *: p<0.05. n.s., non-significant. (b) Quantification of the 3D sprouting index of WM852 and WM852* treated with the indicated siRNAs for 72 hr followed by magnetic separation and the 96 hr fibrin assay. The graph represents the average of three images per condition in each of the two independent experiments, error bars indicate SEM; *: p<0.05. n.s., non-significant. (c) Representative confocal images of MMP14 (green) and Notch3 (red) in WM852 and WM852*. Arrowheads indicate the cell-cell junction where MMP14 and Notch3 co-localize. Nuclei were counterstained with Hoechst 33342. The dashed line indicates the LEC–melanoma (below the line; GFP positive cells (white) in the inset) border. Scale bar = 50 µm. (d) mRNA fold change of the indicated targets in WM852 and WM852* upon treatment with the indicated siRNA for 72 hr and following magnetic separation. Graphs show the average of three independent experiments, error bars indicate SEM, *: p<0.05; **: p<0.01. n.s., non-significant (e) Representative confocal images of Notch3 staining (red) in WM852* treated with the indicated siRNAs for 72 hr. Nuclei were counterstained with Hoechst 33342. The dashed lines indicate the LEC-WM852 (GFP positive cells (white) in the inset) border. Scale bar = 50 µm. (f) Quantification of Notch3 signal intensity of WM852* treated as in (e) and described in (a). Error bars indicate SD; *: p<0.05. Full size confocal images are available as source data 2.
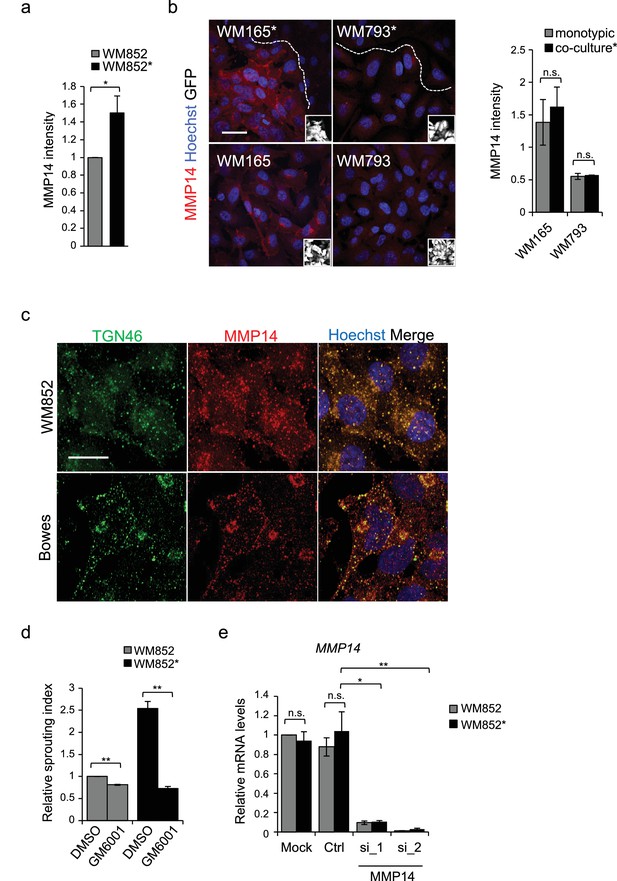
MMP14 levels increase and MMP14 activity is needed for 3D sprouting growth in LEC-primed metastatic melanoma.
(a) Flow cytometry analysis of MMP14 levels in WM852 and WM852*. Mean fluorescence intensity normalized to WM852 is shown, error bars represent SD across three independent experiments; *: p<0.05. (b) Left panels: representative images of MMP14 staining (red) in WM165 and WM793 cells (GFP positive cells (white) in the insets) cultured in the presence (*, upper panels) or absence (bottom panels) of LECs. Nuclei were counterstained with Hoechst 33342. The dashed line indicates the LEC-melanoma (below the line) border. Scale bar = 50 µm. Right panel: quantification of MMP14 signal intensity for the indicated cell lines. Average of the intensity is shown from four images/condition from two independent experiments. More than 100 cells were always analysed per condition. Error bars indicate SD. n.s., non-significant (c) Confocal images of Bowes (upper panels) and WM852 (lower panels) stained with TGN46 (green) and MMP14 (red) antibodies, nuclei were counterstained with Hoechst 33342. Scale bar: 25 µm. (d) Relative sprouting index of WM852 and WM852* treated with the vehicle (DMSO) or panMMP inhibitor GM6001 for 48 hr during co-culture and after magnetic separation as well as during the 3D fibrin assay. Error bars indicate SEM; **: p<0.01. (e) MMP14 mRNA fold change in WM852 and WM852* treated with the indicated siRNAs for 72 hr; *: p<0.05; **: p<0.01. n.s., non-significant. Full size confocal images are available as source data 2.
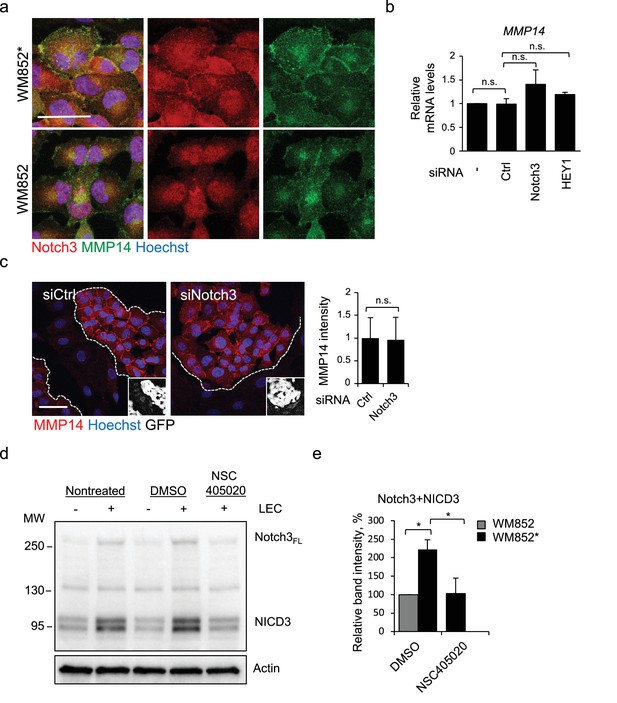
Notch3 does not regulate MMP14 expression, but MMP14 positively regulates Notch3.
(a) Representative enlargement with separated channels of the images in Figure 4c. Merge, Notch3 (red) and MMP14 (green) stainings are shown. (b) MMP14 mRNA fold change in WM852* treated with the indicated siRNAs for 72 hr. Data averaged from two independent experiments is shown. n.s., non-significant. (c) Right panels: representative confocal images of MMP14 (red) in WM852* after the cells were treated with indicated siRNAs for 72 hr. Nuclei were counterstained with Hoechst 33342. The dashed lines indicate LEC-melanoma (GFP expressing cells (white in the inset) border. Scale bar = 50 µm. Left panel: quantification of the MMP14 intensity analysed in four images per condition from two independent experiments. More than 100 cells were always analysed per condition. Average is shown, error bars represent SD. n.s., non-significant (d) Immunoblot using anti-Notch3 (upper panels) and anti-actin (lower panels) antibodies of WM852 and WM852* (+LEC) treated as indicated and subjected to magnetic separation prior to sample preparation. Notch3FL: full length Notch3; NICD3: Notch3 intracellular domain. Panels represent the bands from the same membrane and same exposure time. (e) Relative Notch3 and cleaved NICD3 band intensity normalized to actin of two independent experiments performed as in (d). Average is shown, error bars represent SD; *: p<0.05. Full size confocal images are available as source data 2.
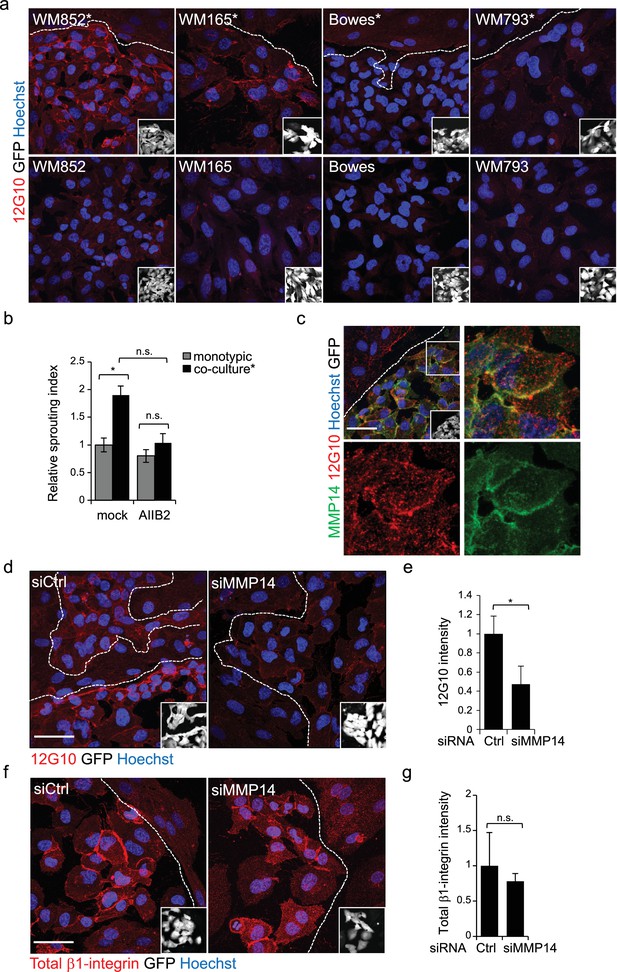
Change in the 3D growth phenotype of the LEC primed melanoma cells is β1-integrin dependent.
(a) Representative confocal images of active β1-integrin (12G10) staining (red) in the indicated melanoma cell lines (GFP positive cells (white) in the inset) in the presence (*, upper panels) or absence (bottom panels) of LECs. Nuclei were counterstained with Hoechst 33342. The dashed line indicates the LEC-melanoma border. Scale bar = 50 µm. (b) Quantification of 3D sprouting index in WM852 and WM852* mock treated or treated with β1-integrin blocking antibody (AIIB2) during the 96 hr fibrin growth assay. Graph shows the average of at least three images per condition per two independent experiments, error bars indicate the SEM; *: p<0.05. n.s., non-significant. (c) Representative confocal image of active β1-integrin (12G10, red) and MMP14 (green) staining of WM852* (white cells in the inset). Nuclei were counterstained with Hoechst 33342. The dashed lines indicate the border between LEC and WM852 (white, GFP positive WM852 cells in the inset). The right and bottom panels show an enlargement of the area enclosed within the white square as a merge, Notch3 (red) and MMP14 (green) in separated channels. Scale bar = 50 µm. (d,f) Representative confocal images of WM852* treated with the indicated siRNAs for 72 hr and stained for active β1-integrin with 12G10 (d, red), or total β1-integrin with P5D2 (f, red) antibodies. Nuclei were counterstained with Hoechst 33342. The dashed lines indicate the LEC-WM852 borders (white, GFP positive WM852 cells in the inset) border. Scale bar = 50 µm. Quantification of the average 12G10 (e) and total β1-integrin (g) signal intensity in WM852* (white) cells. Four images/condition were quantified from two independent experiments. More than 100 cells were always analysed per condition; error bars indicate SD. *: p<0.05. n.s., non-significant. Full size confocal images are available as a source data 3.
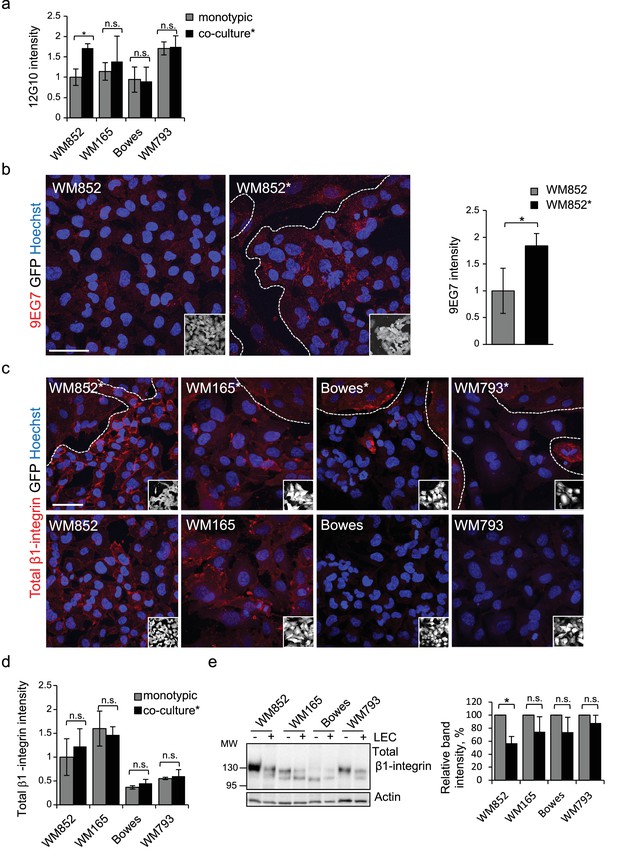
Quantification of active and total β1-integrin in monotypic and LEC-primed melanoma cells.
(a) Quantification of active β1-integrin (12G10) intensity in the indicated melanoma cell lines from monotypic cultures or after co-culture with LEC (*). Average of the intensity was calculated in four images/condition from two independent experiments. More than 100 cells were always analysed per condition. Error bars indicate SD, *: p<0.05; n.s., non-significant. (b) Left and middle panels: representative confocal images of WM852 and WM852* stained with 9EG7 antibody for active β1-integrin (red). Nuclei were counterstained with Hoechst 33342 (blue). Dashed line indicates LEC-melanoma border. Scale bar = 50 µm. Right panel: quantification of 9EG7 intensity in the indicated cell types performed as in (a). (c) Representative confocal images of β1-integrin staining (red) in the different GFP expressing melanoma cell lines (GFP positive cells white in the inset) cultured in the presence (*, upper panels) or absence (bottom panels) of LECs. Nuclei were counterstained with Hoechst 33342. The dashed line indicates the LEC-melanoma (below the line) border. Scale bar = 50 µm. (d) Quantification of total β1-integrin intensity in the indicated melanoma cell lines from monotypic cultures or co-cultured with LEC (*) as described in (a). n.s., non-significant (e) Left panels: Immunoblot of indicated cell lines derived from monotypic or LEC co-cultures and separated by magnetic separation prior to sample preparation and stained with the indicated antibodies. Right panel: Percentages of β1-integrin intensities normalized to actin from two independent experiments. Error bars indicate SD; *: p<0.05; n.s., non-significant. Full size confocal images are available as a source data 4.
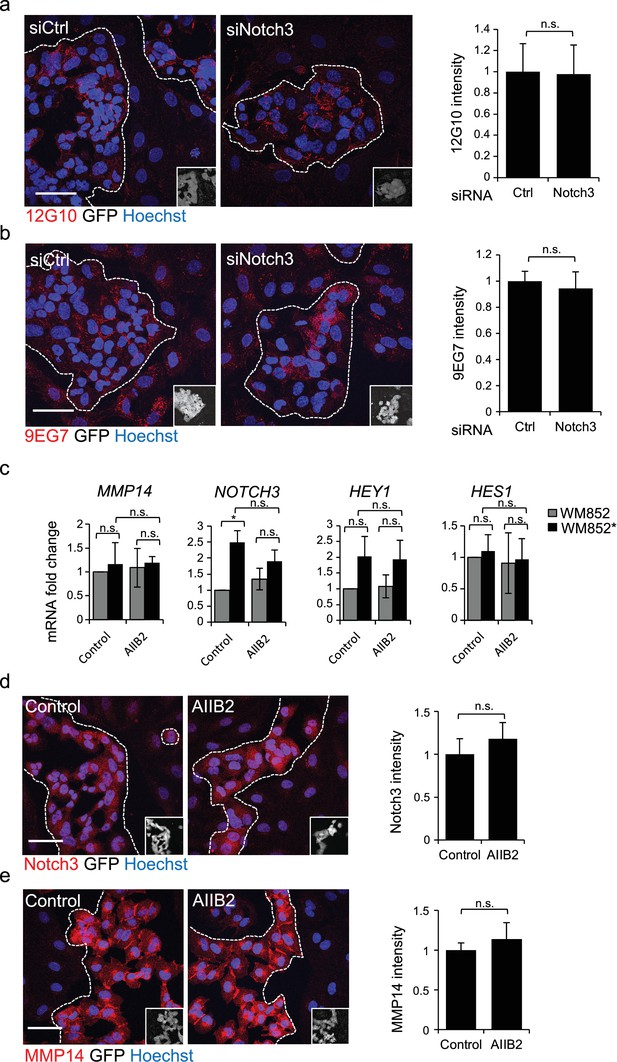
β1-integrin activity does not affect MMP14 and Notch3 expression.
(a, b) Left and middle panels: representative confocal images of active β1-integrin expression using 12G10 (a) and 9EG7 antibodies (red) in WM852* cells treated with indicated siRNAs for 72 hr. Nuclei were counterstained with Hoechst 33342. GFP-expressing melanoma cells are shown white in the inset. The dashed line indicates the LEC-melanoma border. Scale bar = 50 µm. Right panels: Quantification of the average of 12G10 (a) and 9EG7 (b) signal intensities. Four images/condition were quantified from two independent experiments. More than 100 cells were always analysed per condition; error bars indicate SD. n.s., non-significant. (c) mRNA fold change of the indicated targets in WM852 and WM852* following treatment with AIIB2 β1-integrin blocking antibody for 48 hr and magnetic separation. Graphs show the average of two independent experiments, error bars indicate SD; *: p<0.05; n.s., non-significant. (d,e) Left and middle panels: Representative confocal images of control and AIIB2 β1-integrin blocking antibody treated WM852* cells stained with Notch3 (d, red), or MMP14 (e, red) antibodies. Nuclei were counterstained with Hoechst 33342. The dashed lines indicate the LEC-WM852 border (white, GFP positive WM852 cells in the inset) border. Scale bar = 50 µm. Right panels: Quantification of the average Notch3 (d) and MMP14 (e) signal intensity in WM852* (white) cells as described in (a,b); error bars indicate SD. n.s., non-significant. Full size confocal images are available as a source data 4.
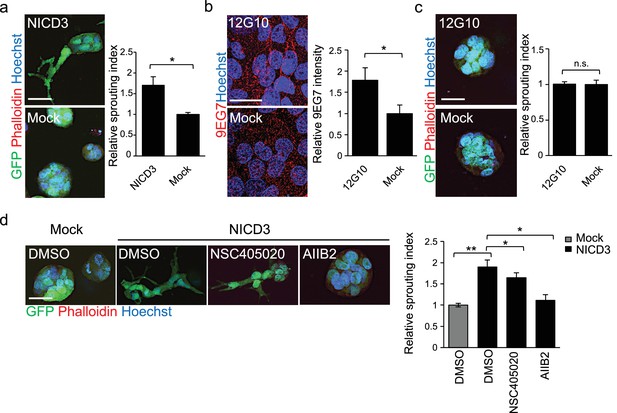
NICD3 overexpression provokes a β1 integrin-dependent 3D sprouting in WM793 cells.
(a) Representative images (left) and quantification (right) of the 3D fibrin assay of WM793 upon transfection of either a NICD3 expressing vector or a control vector. Scale bar = 50 µm. Four images/condition were quantified from two independent experiments. Graph shows the average of at least three images per condition per two independent experiments, error bars indicate the SEM. *p<0.05. (b) Representative images (left) and quantification (right) as in of mock and 12G10 antibody treated WM793 cells. Active β1-integrin was detected with 9EG7 antibody (red). Nuclei were counterstained with Hoechst 33342. Four images/condition were quantified from two independent experiments. More than 100 cells were always analysed per condition; error bars indicate SD. Scale bar 50 µm. *p<0.05. (c) Representative images (left) and quantification (right) of 3D fibrin assay as in (a) of WM793 treated with 12G10 antibody for four days or mock treated. Scale bar = 50 µm. n.s., non-significant. (d) Representative images (left) and quantification (right) of 3D fibrin assay as in (a) of WM793 upon transfection of either a NICD3 expressing vector or a control vector (mock) and treated with either DMSO or the indicated inhibitors. Scale bar = 50 µm, *p<0.05; **p<0.01.
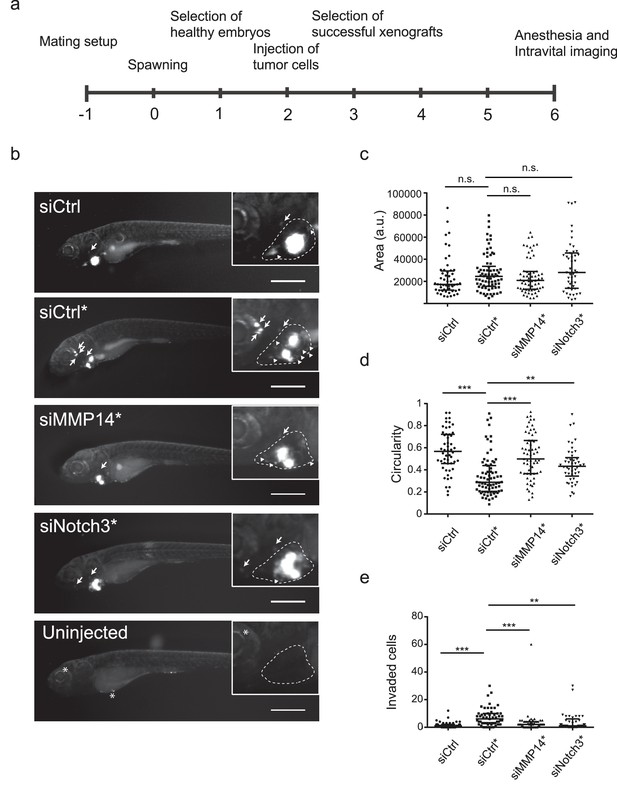
In vivo invasion and dissemination of LEC-primed melanoma cells are dependent on MMP14 and Notch3.
(a) Time line of zebrafish xenograft experiments. (b) Intravital fluorescence microscopy images of six dpf zebrafish embryos taken four days post injection (4 dpi). Fluorescence in GFP channel is shown. Scale bar, 500 µm. Inset shows magnification of the primary tumor. Tumor cells invading outside pericardial space are marked with an arrow, invading cells in pericardial cavity with a triangle and unspecific fluorescence in eye and yolk sac with an asterisk (*). The outline of pericardial cavity is depicted with dashed line. (c) Quantification of area of primary tumors at 4 dpi. siCtrl, n = 52; siCtrl*, n = 74; siMMP14*, n = 61; siNotch3*, n = 46. (d) Quantification of circularity of primary tumors at 4 dpi. siCTRL, n = 52; siCtrl*, n = 74; siMMP14*, n = 61; siNotch3*, n = 46. (e) Quantification of melanoma cells invaded outside pericardial cavity. siCtrl, n = 43; siCtrl*, n = 55; siMMP14*, n = 44; siNotch3*, n = 37. (c–e) Non-parametric Kruskal-Wallis test with Dunn´s multiple comparison test was used, and in addition to individual data points, the median and interquartile range has been plotted. n.s., non significant (p>0.05); **p<0.01; ***p<0.001.

(a) Flow cytometry analysis of active b1-integrin levels in WM852 and WM852* using 12G10 (upper) and 9EG7 (lower) antibodies.
(b) Confocal images of WM852* replated after separation on coverslips for 24h (left panels), WM852 +LEC co-culture (not subjected to magnetic separation; middle panels) and monotypic WM852 (right panels). Cells were stained with the indicated antibodies (red), nuclei were counterstained with Hoechst 33342. Scale bar: 20mM. (c) Representative confocal z-stack projections of 3D fibrin assay after magnetic separation of WM852 cells co-cultured with LECs (*,right panel) or from monotypic culture (left panel). The GFP expressing (green) melanoma cells were stained with 12G10 antibody (red), nuclei were counterstained with Hoechst 33342 (blue). Scale bar = 50 µm.
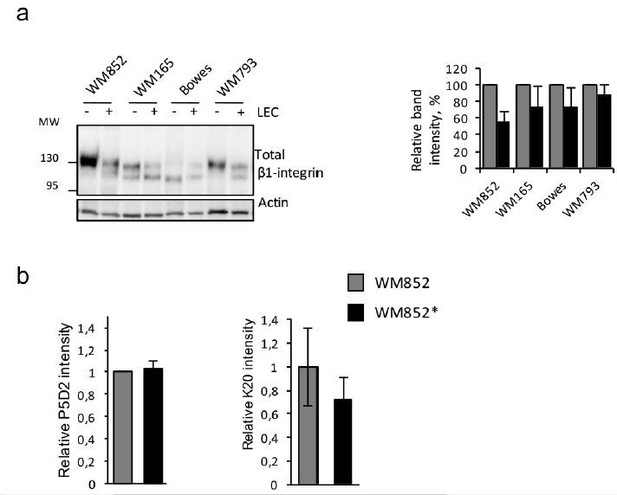
(a) Left panel: immunoblot of indicated cells lines cultured with or without LECs and separated by magnetic separation and stained with indicated antibodies.
Right panel: quantification of relative β1-integrin band intensities from two independent experiments as shown in left panel. The signal intensity was normalized to actin. The average is shown, error bars represent SD. (b) Flow cytometry analysis of total β1-integrin levels in WM852 and WM852* using P5D2 (left) and K20 antibodies. Mean fluorescence intensity normalized to WM852 is shown, error bars represent SD.
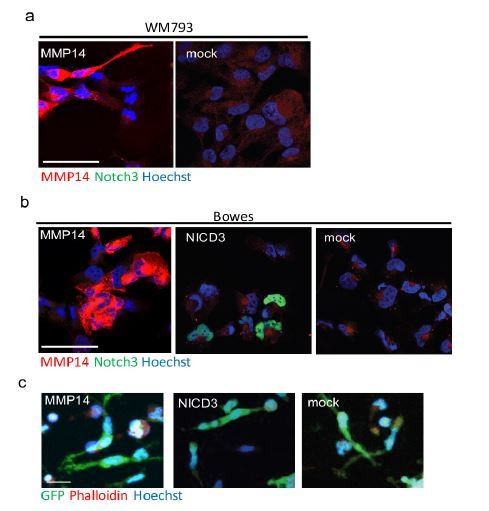
(a) Confocal images of WM793 transiently transfected with MMP14 expressing plasmid (left) or a control vector (right) stained with Notch3 (green) and MMP14 (red) antibodies, nuclei were counterstained with Hoechst 33342.
(b) Confocal imaged of Bowes transiently transfected with MMP14 (left), NICD3 (middle) expressing plasmid, or a control vector (right) stained with Notch3 (green) and MMP14 (red) antibodies, nuclei were counterstained with Hoechst 33342. (c) Representative confocal images of 3D fibrin assays of Bowes cells treated as in (b). GFP expressing (green) melanoma cells were stained with Phalloidin A594 (red), nuclei are counterstained with Hoechst 33342 (blue). Maximum intensity Z-projections of the confocal stacks are shown. Scale bars= 50 µm.
Tables
| Reagent type (species) or resource | Designation | Sourcor reference | Identifiers | Additional Information |
|---|---|---|---|---|
| Danio Rerio casper strain (roy-/-; mitfa-/-) | PMID: 18371439 | |||
| Mus Musculus C.B-17/IcrHanTMHSD | Harlan, Indianapolis, IN, USA | |||
| Human primary juvenile foreskin lymphatic endothelial cells | Promocell, Heidelberg, Germany | |||
| Adult Dermal lymphatic endothelial cells | Lonza,Basel, Switzerland | |||
| WM852 | Wistar Institute Philadelphia | RRID:CVCL_6804 | ||
| WM165 | Wistar Institute Philadelphia | RRID:CVCL_L033 | ||
| Bowes | D.B. Rifkin, Rockfeller University | RRID:CVCL_3317 | ||
| WM793 | Wistar Institute Philadelphia | RRID:CVCL_8787 | ||
| HEK293FT | Biomedicum Functional Genomic Unit, University of Helsinki | |||
| siRNA | Notch3 siRNA Notch3 siRNA | Dharmacon, Lafayette, CO Ambion,Waltham, MA | L-011093-00-0005 4392420 | |
| siRNA | MMP14 siRNA | Qiagen,Hilden, Germany | SI03648841; SI00071176 | |
| Transfected construct | NICD3pCLE | AddGene,Cambridge, MA;PMID: 16508304 | Plasmid #26894 | |
| Transfected construct | MMP14 Sport6 | GenomeBiology UnitUniversity of Helsinki | NA | |
| Transfected construct | PcDNA3 | Invitrogen,Carlsbad, CA | NA | |
| Antibody | Pecam | DAKO,Santa Clara, CA | M0823 | IFA 3D 1:500 |
| Antibody | VE-cadherin | BD Pharmingen, San Jose, CA | 553927 | IFA 1:1000 |
| Antibody | GFP | Prof. Jason Mercer, UCL, London | NA | IFA 3D 1:1000 |
| Antibody | Notch3 | Santa Cruz Biotechnology,Dallas, TX | sc-5593 | IFA 1:50 WB 1:250 |
| Antibody | MMP14 EP1264Y | Abcam,Cambridge, UK | 51074 | IFA 1:100 |
| Antibody | MMP14 (LEM clone) | Chemicom,Waltham, MA | MAB3328 | IFA 1:300 FACS 1:100 |
| Antibody | active β1-integrin 12G10 | Abcam | 30394 | IFA 1:300 FACS 1:100 |
| Antibody | active β1-integrin 9EG7 | BD Pharmingen | 553715 | IFA 1:100 FACS 1:100 |
| Antibody | Total β1-integrin | Abcam | 52971 | WB 1:1000 |
| Antibody | TGN46 | Sigma,St. Louis, MO | T7576 | IFA 1:500 |
| Antibody | LYVE1 | Reliatech,Wolfenbüttel, Germany | 103-PA50AG | IHC 1:200 |
| Antibody | Alexa594-phalloidin | Thermo Fisher,Waltham, MA | 21833 | IFA 3D 1:200 |
| Sequence-based reagents | ||||
| CD31 RTqPCR primers (for, rev) | AACAGTGTTGACATGAAGAGCC, TGTAAAACAGCACGTCATCCTT | |||
| CD34 RTqPCR primers (for, rev) | TGGGCATCACTGGCTATTTC, CCACGTGTTGTCTTGCTGAA | |||
| FLT4 RTqPCR primers (for, rev) | GACAGCTACAAATACGAGCATCTG, CTGTCTTGCAGTCGAGCAGAA | |||
| NOTCH1 RTqPCR primers (for, rev) | GAGGCGTGGCAGACTATCATGC, CTTGTACTCCGTCAGCGTGA | |||
| NOTCH2 RTqPCR primers (for, rev) | CCTGGGCTATACTGGGAGCTACTG ,ACACCCTGATAGCCTGGGACAC | |||
| NOTCH4 RTqPCR primers (for, rev) | AATCCCACTGCCTCCAGACT, TTGTGGCAAAGGGAAGAGAC | |||
| HES1 RTqPCR primers (for, rev) | TCAACACGACACCGGATAAA, TCAGCTGGCTCAGACTTTCA | |||
| HEY1 RTqPCR primers (for, rev) | GTTCGGCTCTAGGTTCCATGT, CGTCGGCGCTTCTCAATTATTC | |||
| HEY2 RTqPCR primers (for, rev) | TTGAGAAGACTTGTGCCAACTG, GTGCGTCAAAGTAGCCTTTACC | |||
| MMP14 RTqPCR primers (for, rev) | GCAGAAGTTTTACGGCTTGCAA, CCTTCGAACATTGGCCTTGAT | |||
| ACT RTqPCR primers (for, rev) | TCACCCACACTGTGCCATCTACGA, CAGCGGAACCGCTCATTGCCAATGG | |||
| GAPDH RTqPCR primers (for, rev) | TCACCACCATGGAGAAGGCT, GCCATCCACAGTCTTCTGGG | |||
| Commercial assay or kit | NucleoSpin RNA II kit | Macherey Nagel,Düren, Germany | 740955 | |
| Commercial assay or kit | SYBR Green PCR mix | Fermentas, Waltham, MA | 4415440 | |
| Commercial assay or kit | QuantiTect Primer Assay NOTCH3 | Qiagen | QT00003374 | |
| Commercial assay or kit | dextran coated magnetic nanoparticles extran coated magnetic nanoparticles | fluidMAG-DX, Chemicell, Berlin, Germany | 4104–5 | |
| Commercial assay or kit | MidiMACS separator | Miltenyi Biotec,Bergisch Gladbach, Germany | 130-042-302 | |
| Commercial assay or kit | LS column | Miltenyi Biotec | 130-042-401 | |
| Chemical compound, drug | Lipofectamine RNAiMax | Invitrogen | 13778150 | |
| Chemical compound, drug | DAPT | Sigma | D5942 | Used at 10 μM |
| Chemical compound, drug | GM6001 | Tocris Biosciences, Bristol, UK | 2983/10 | Used at 10 μM |
| Chemical compound, drug | NSC 405020 | Selleckchem,Munich, Germany | S8072 | Used at 50 μM |
| Chemical compound, drug | AIIB2 | DSHB hybridoma, from Johanna Ivaska | RRID:AB_528306 | Used 1:10 |
| Software, algorithm | Bioimage XD (http://www.bioimagexd.net/) | PMID: 22743773 | NA | |
| Software, algorithm | CellProfiler | PMID: 17076895 | RRID:SCR_007358 | |
| Software, algorithm | Adobe Photoshop | RRID:SCR_014199 | ||
| Software, algorithm | ImageJ | RRID:SCR_003070 | ||
| Software, algorithm | Pathview https://pathview.uncc.edu | RRID: SCR_002732 | ||
| Software, algorithm | Morpheus https://software.broadinstitute.org/morpheus/ | RRID: SCR_014975 | ||
Additional files
-
Soure data 1
Full size confocal images of Figure 2.
- https://doi.org/10.7554/eLife.32490.018
-
Soure data 2
Full size confocal images of Figure 4a,c,e; Figure 4—figure supplement 1b,c; Figure 4—figure supplement 2a,c.
- https://doi.org/10.7554/eLife.32490.019
-
Soure data 3
Full size confocal images of Figure 5a,c,d,f.
- https://doi.org/10.7554/eLife.32490.020
-
Soure data 4
Full size confocal images of Figure 5—figure supplement 1b,c; Figure 5—figure supplement 2a,b,d,e.
- https://doi.org/10.7554/eLife.32490.021
-
Supplementary file 1
related to Figure 2 and containing more than two fold significantly altered genes.
- https://doi.org/10.7554/eLife.32490.022
-
Supplementary file 2
related to Figure 2 and containing all genes significantly up- and downregulated listed for both cell lines in the selected pathways.
- https://doi.org/10.7554/eLife.32490.023
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32490.024





