Functional divergence of paralogous transcription factors supported the evolution of biomineralization in echinoderms
Figures
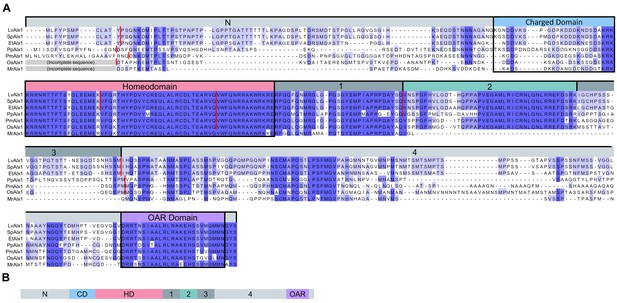
Comparison of the predicted amino acid sequences of Alx1 across the echinoderms.
(A) Clustal Omega alignment of the Alx1 sequences from Lytechinus variegatus (LvAlx1), Strongylocentrotus purpuratus (SpAlx1), Eucidaris tribuloides (EtAlx1), Parastichopus parvimensis (PpAlx1), Patiria miniata (PmAlx1), Ophiothrix spiculata (OsAlx1) and Metacrinus rotundus (MrAlx1). The Alx1 proteins share a highly conserved homeodomain (HD) and a C-terminal OAR domain. The region directly upstream of the homeodomain that contains several aspartic acid and lysine residues was designated as the charged domain (CD). The C-terminal region between the homeodomain and OAR domain was divided into four sub-regions, designated Domains 1–4, based on splicing patterns and evolutionary conservation. The degree of sequence similarity is reflected by the intensity of the violet shading. The red lines indicate known splice junctions. As there are no genomic data available for M. rotundus, the splice junctions are not known for this species. (B) Schematic representation of the domain organization of the echinoderm Alx1 protein.
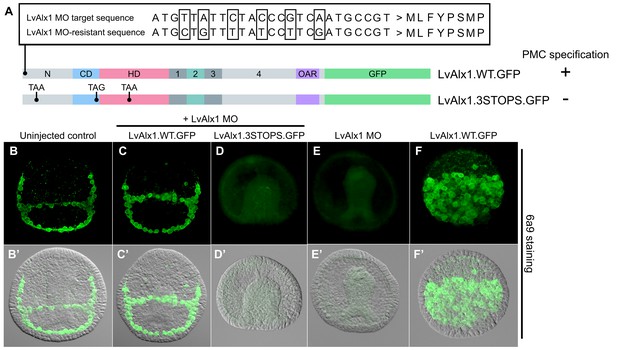
Expression of MO-resistant LvAlx1 rescues PMC specification in LvAlx1 morphants.
(A) Six silent mutations were introduced into the MO target site to generate a MO-resistant Lv-alx1 mRNA (LvAlx1.WT.GFP). Boxed nucleotides indicate changes introduced. A mutant form of LvAlx1.WT.GFP that contained three in-frame premature stop codons was also generated (LvAlx1.3STOPS.GFP). (B–F) Embryos fixed at the mid-late gastrula stage and labelled with a PMC-specific monoclonal antibody (6a9). (B’–F’) DIC/fluorescence composite images of the same embryos. (B, B’) Uninjected control embryo with 6a9-positive PMCs arranged in a ring-like pattern. (C, C’) Embryo co-injected with LvAlx1 MO and LvAlx1.WT.GFP mRNA exhibiting normal PMC specification and patterning. (D, D’) Embryo co-injected with LvAlx1 MO and LvAlx1.3STOPS.GFP. This embryo has no 6a9-positive cells, similar to embryos injected with LvAlx1 MO alone (E, E’). (F, F’) Embryos injected with LvAlx1.WT.GFP mRNA alone showing the induction of supernumerary PMCs.
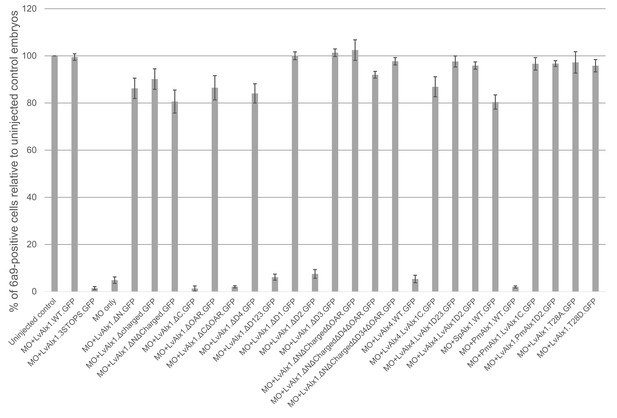
Quantitative analysis of MO rescue experiments.
Each vertical bar indicates the mean percentage of 6a9-positive cells relative to uninjected control embryos, calculated from 80 to 100 embryos. Vertical lines represent standard errors. An average of 6a9-positive cells of control embryos of a given batch was first calculated. Subsequently, for each individual experimental embryo, a percentage of 6a9-positive cells relative to the average control value was calculated and the mean of those values was determined.
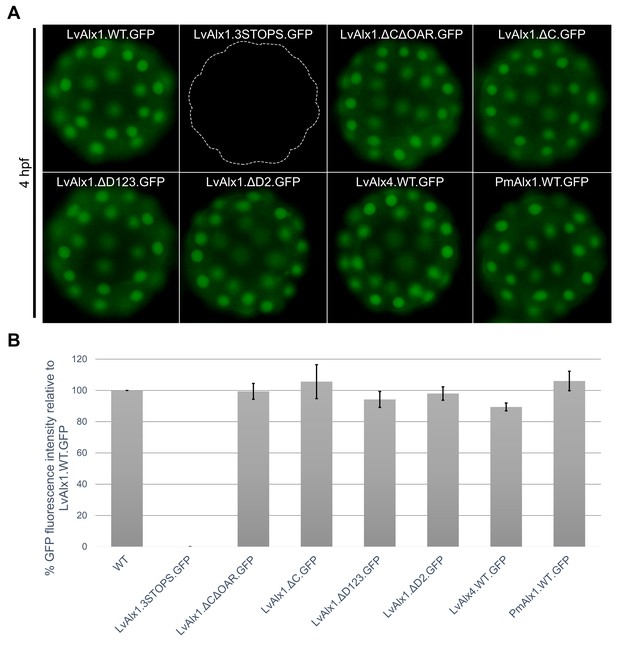
LvAlx1 with premature stop codons, LvAlx1 C region mutants, LvAlx4 and PmAlx1 are expressed at levels similar to that of LvAlx1.WT.GFP.
(A) Fluorescent images showing expression of LvAlx1.GFP proteins in living embryos at 4 hpf. The dotted line indicates the outline of embryo injected with LvAlx1.3STOPS.GFP, which cannot be translated into a GFP fusion protein. (B) Quantification of the GFP fluorescence intensities. Each vertical bar indicates the average percentage of normalized integrated density value relative to embryos injected with LvAlx1.WT.GFP, calculated from 15 to 40 embryos for each construct. Vertical lines represent standard errors. For each embryo, the average integrated density value of GFP fluorescence was normalized with the integrated density value of Texas Red dextran to account for differences in total embryo area and injection volume. An average integrated density value was first calculated for LvAlx1.WT.GFP of a given batch. Subsequently, for each individual experimental embryo, a percentage GFP fluorescence intensity relative to the average control value was calculated and the mean of those values was determined.
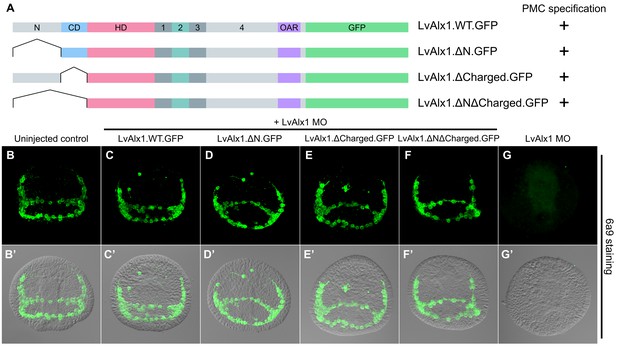
The N-terminus is dispensable for LvAlx1 function in PMC specification and patterning.
(A) Constructs generated with different N-terminal deletions and truncations. (B–G) Embryos fixed at the mid-late gastrula stage and labelled with 6a9 antibody. (B’–G’) DIC/fluorescence composite images of the same embryos. (B, B’) Uninjected control embryo. (C–F’) Embryos co-injected with LvAlx1 MO and LvAlx1 mRNA with different N-terminal mutations exhibit normal PMC specification and patterning, similar to the positive control embryo co-injected with LvAlx1 MO and LvAlx1.WT.GFP mRNA (C, C’). Total deletion of the region upstream of the homeodomain (LvAlx1.ΔNΔCharged.GFP) also does not affect PMC specification or patterning (F, F’). (G, G’) Embryo injected with LvAlx1 MO alone (negative control).
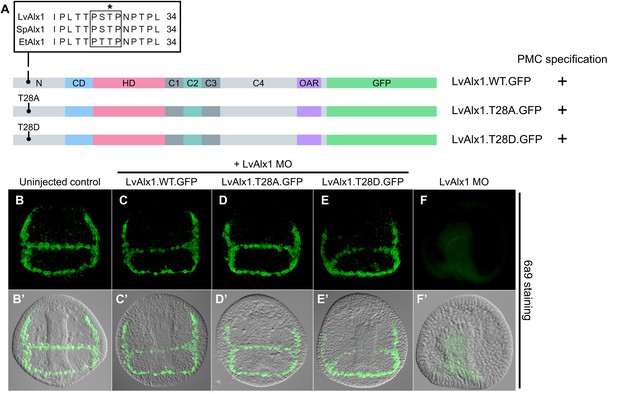
The putative MAPK phosphorylation site T28 is not essential for PMC specification or patterning.
(A) Alignment of part of the N-terminal region of Alx1 from L. variegatus (LvAlx1), S. purpuratus (SpAlx1) and E. tribuloides (EtAlx1). Boxed region indicates protein sequences that conform to the consensus MAPK phosphorylation site, PXS/TP. Asterisk indicates the threonine residue that was mutated to either alanine (T28A) or aspartic acid (T28D). (B–G) Embryos fixed at the mid-late gastrula stage and labelled with 6a9 antibody. (B’–G’) DIC/fluorescence composite images of the same embryos. (B, B’) Uninjected control embryo. (D–E’) Embryos co-injected with LvAlx1 MO and the phosphorylation-null mutant (LvAlx1.T28A.GFP) or the phosphomimetic mutant (LvAlx1.T28D.GFP) show normal numbers of 6a9-positive cells, indicating rescue of PMC specification in LvAlx1 morphants, similar to those injected with LvAlx1 MO and LvAlx1.WT.GFP (C, C’). (F, F’) Negative control embryo injected with LvAlx1 MO alone.
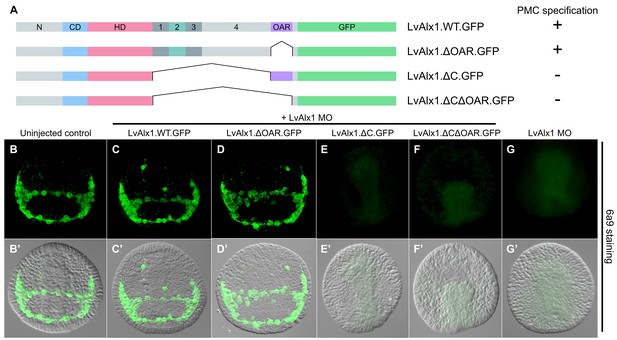
The C-terminal region contains motifs essential for Alx1 function.
(A) Constructs generated with different C-terminal deletions and truncations. (B–G) Embryos fixed at the mid-late gastrula stage and labelled with 6a9 antibody. (B’–G’) DIC/fluorescence composite images of the same embryos. (B, B’) Uninjected control embryo. (C, C’) Positive control embryo co-injected with LvAlx1 MO and LvAlx1.WT.GFP. (D, D’) Deletion of the highly conserved OAR domain does not affect rescue of PMC specification or patterning. (E, E’ and F, F’) Embryos co-injected with LvAlx1 MO and LvAlx1 mRNA lacking C-terminal sequences (LvAlx1.ΔC.GFP and LvAlx1.ΔC.ΔOAR.GFP) contain no 6a9-positive cells, similar to embryos injected with LvAlx1 MO alone (G, G’).
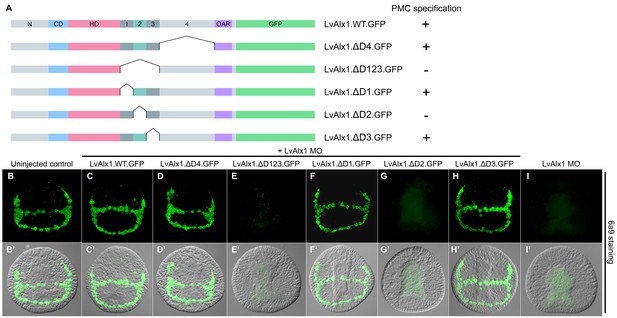
Domain 2 is essential for Alx1 function.
(A) Constructs with deletions within the C-terminal region. (B–I) Embryos fixed at the mid-late gastrula stage and labelled with 6a9 antibody. (B’–I’) DIC/fluorescence composite images of the same embryos. (B, B’) Uninjected control embryo. (C, C’) Positive control embryo co-injected with LvAlx1 MO and LvAlx1.WT.GFP. (D, D’) Deletion of the relatively large Domain 4 does not affect rescue of PMC specification and patterning. (E, E’) Deletion of Domains 1–3 together abolishes rescue, as does deletion of Domain 2 alone (G, G’). (F, F’ and H, H’) Deletion of Domains 1 or 3 individually does not affect rescue. (I, I’) Embryo injected with LvAlx1 MO alone (negative control).
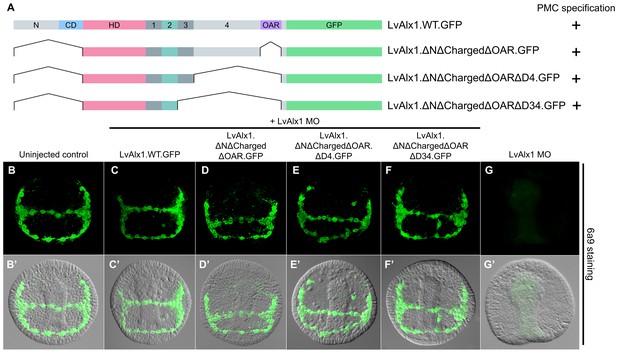
Determination of a minimal construct sufficient to rescue LvAlx1 morphants.
(A) Constructs tested. All mutant constructs contained the homeodomain and Domain 2, both of which were required for PMC specification. (B–G) Embryos fixed at the mid-late gastrula stage and labelled with 6a9 antibody. (B’–G’) DIC/fluorescence composite images of the same embryos. (B, B’) Uninjected control embryo. (C, C’) Positive control embryo co-injected with LvAlx1 MO and LvAlx1.WT.GFP (D–F’) LvAlx1 mRNA that lacked the N-terminus, charged domain, OAR domain and Domains 3 and 4 (LvAlx1.ΔNΔChargedΔOARΔD34.GFP) was sufficient to rescue LvAlx1 morphants. (G, G’) Embryo injected with LvAlx1 MO alone (negative control).
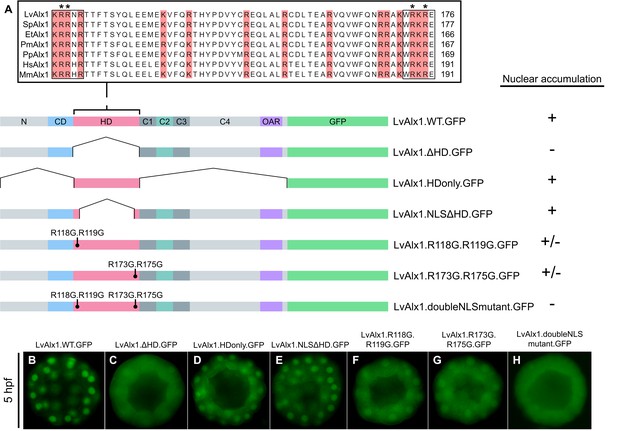
Basic residues flanking the LvAlx1 homeodomain function as nuclear localization signals.
(A) Top: Clustal Omega alignment of Alx1 homeodomains from L. variegatus (LvAlx1), S. purpuratus (SpAlx1), E. tribuloides (EtAlx1), P. miniata (PmAlx1), P. parvamensis (PpAlx1), H. sapiens (HsAlx1), and M. musculus (MmAlx1). Boxed sequences indicate the two NLS-like motifs flanking the homeodomain. Asterisks show the NLS residues targeted for mutations. Bottom: LvAlx1 constructs used in nuclear localization studies. (B–H) GFP fluorescence of embryos injected with the different NLS mutants. (B) LvAlx1.WT.GFP shows robust nuclear accumulation throughout development. (C) Removal of the homeodomain (LvAlx1.ΔHD.GFP) causes a diffuse protein distribution. (D) The homeodomain alone (LvAlx1.HDonly.GFP) is sufficient to drive nuclear accumulation. (E) Retaining only the two NLS motifs from the homeodomain (LvAlx1.NLSΔHD.GFP) allows the protein to accumulate in the nucleus. (F and G) Mutation of either of the NLS motifs result in weaker nuclear accumulation. (H) Mutation of both NLS motifs result in completely diffused protein distribution.
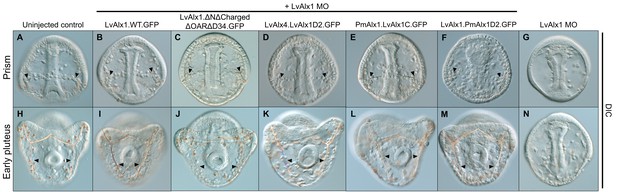
Rescued embryos show normal skeleton development at late embryonic stages.
Embryos were visualized using DIC optics. (A–F) Prism stage embryos (ventral views). Arrowheads indicate spicule primordia. (H–M) Early pluteus stage embryos (posterior views). Arrowheads indicate elongated skeletal rods. (G, N) LvAlx1 morphants at the prism (G) and early plutues (N) stages showing complete lack of skeletal elements. .
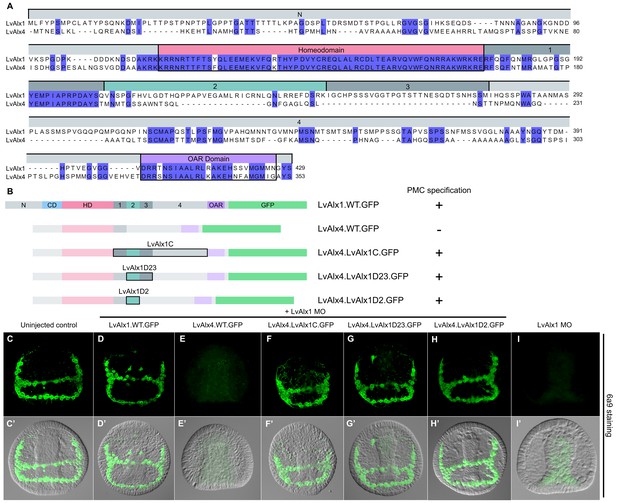
LvAlx1 and LvAlx4 are not functionally redundant; however, insertion of LvAlx1 Domain 2 into LvAlx4 is sufficient to confer skeletogenic function.
(A) Clustal Omega alignment of the predicted amino acid sequences of LvAlx4 and LvAlx1. (C–I) Embryos fixed at the mid-late gastrula stage and labelled with 6a9 antibody. (C’–I’) DIC/fluorescence composite images of the same embryos. (C, C’) Uninjected control embryo. (D, D’) Positive control embryo co-injected with LvAlx1 MO and LvAlx1.WT.GFP. (E, E’) Embryo co-injected with LvAlx1 MO and LvAlx4 mRNA showing no 6a9-positive cells. (F, F’, G, G’ and H, H’) Chimeric forms of LvAlx4 containing the entire C-terminal region of LvAlx1, Domains 2 and 3, or Domain 2 alone, are able to rescue PMC specification and patterning. (I, I’) Embryo injected with LvAlx1 MO alone (negative control).
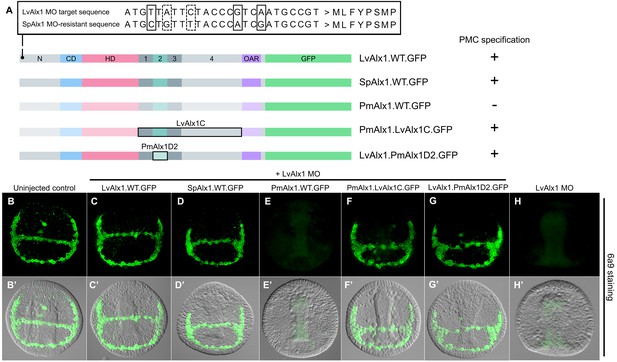
Alx1 proteins from closely related echinoderm species, but not from distantly related species, are functionally interchangeable with LvAlx1.
(A) Three silent mutations (solid boxes) were introduced into the S. purpuratus alx1 translational start site to generate a form of Sp-alx1 mRNA (SpAlx1.WT.GFP) that was resistant to the LvAlx1 MO. Dashed boxes indicate natural nucleotide differences between Lv-alx1 and Sp-alx1. The P. miniata alx1 translational start site is very different from that of Lv-alx1 and hence no mutations were necessary. (B–G) Embryos fixed at the mid-late gastrula stage and labelled with 6a9 antibody. (B’–G’) DIC/fluorescence composite images of the same embryos. (B, B’) Uninjected control embryo. (C, C’) Positive control embryo co-injected with LvAlx1 MO and LvAlx1.WT.GFP. (D, D’) Alx1 from a closely related species, the sea urchin S. purpuratus (SpAlx1), rescues PMC specification and patterning in LvAlx1 morphants. (E, E’) Alx1 from a more distantly related species, the sea star P. miniata (PmAlx1), is not interchangeable with LvAlx1. (F, F’) A chimeric form of PmAlx1 containing the C-terminal region of LvAlx1 between the homeodomain and OAR domain (PmAlx1.LvAlx1C.GFP) rescues PMC specification and patterning in LvAlx1 morphants. (G, G’). A chimeric form of LvAlx1 which had the endogenous Domain 2 replaced with Domain 2 of PmAlx1 (LvAlx1.PmAlx1D2.GFP) rescues PMC specification. (H, H’) Embryo injected with LvAlx1 MO alone (negative control).
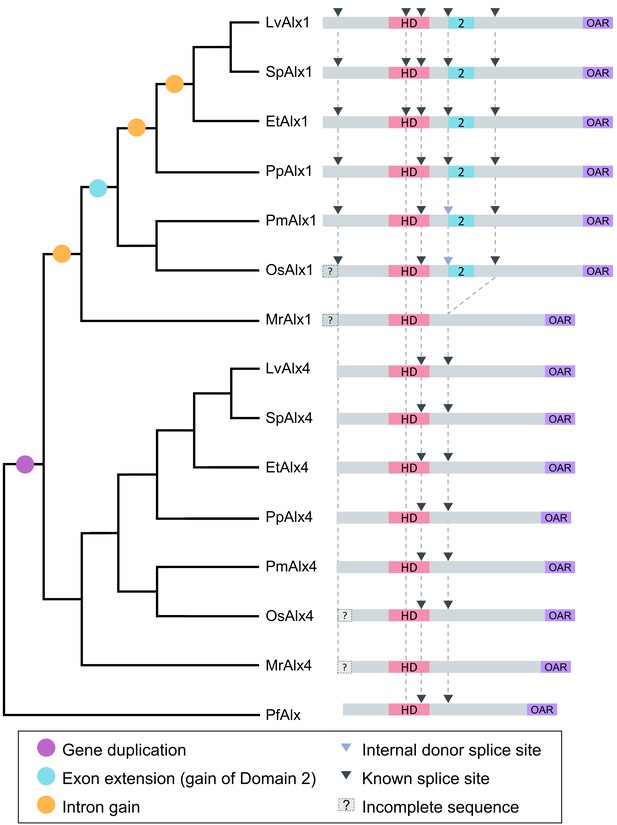
Evolution of echinoderm alx gene structure.
Left side: molecular phylogeny of echinoderm Alx1 and Alx4 proteins (based on Koga et al., 2016), with a hemichordate as an outgroup. Branch lengths are arbitrary. Right side: intron-exon organization of echinoderm alx1 and alx4 genes. Lv: Lytechinus variegatus (euechinoid sea urchin); Sp: Strongylocentrotus purpuratus (euechinoid sea urchin); Et: Eucidaris tribuloides (cidaroid sea urchin); Pp: Parastichopus parvimensis (sea cucumber); Pm: Patiria miniata (sea star); Os: Ophiothrix spiculata (brittle star); Mr: Metacrinus rotundus (sea lily); Pf: Ptychodera flava (hemichordate). Splice sites for Mr-alx1 and Mr-alx4 are not known. Dotted lines are for alignment purposes only.
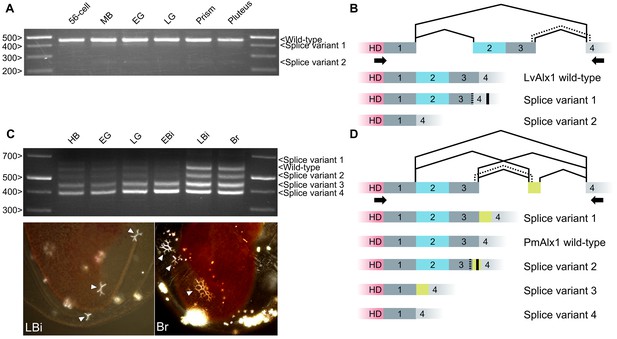
Inclusion of Domain 2 is regulated by alternative splicing.
(A) Agarose gel showing different LvAlx1 splice forms. Wild-type LvAlx1 is the predominant splice form throughout embryonic development. MB: mesenchyme blastula; EG: early gastrula; LG: late gastrula. (B) Sequencing results show the inclusion of the exon containing Domains 2 and 3 in LvAlx1 splice variant 1. Cryptic splice sites (dotted line) in the Domain 3 and Domain 4-containing exons are utilized, however, resulting in a frameshift and premature stop codon (solid line). In LvAlx1 splice variant 2, the Domain (2 + 3) exon is skipped. The reading frame of Domain 4 is not altered. (C) Top: agarose gel showing different PmAlx1 splice forms. There are five PmAlx1 splice variants across the developmental stages tested. HB: hatched blastula; EG: early gastrula; LG: late gastrula; EBi: early bipinnaria larva; LBi: late bipinnaria larva; Br: brachiolaria larva. Bottom: images of living, late bipinnaria and brachiolaria larvae, viewed with partially crossed polarizers to highlight skeletal elements (arrowheads). (D) Sequencing results show the inclusion of an alternative exon (shown in purple) between Domains 3 and 4 in PmAlx1 splice variant 1. In PmAlx1 splice variant 2, the same alternative exon is included, but cryptic splice sites (dotted line) in Domain 3- and Domain 4-containing exons are utilized, causing a frameshift and resulting in a premature stop (solid line). In PmAlx1 splice variant 3, an internal splice site near the 5’ end of Domain 2 is utilized and Domains 2 and 3 are skipped, while the alternative exon is included. In PmAlx1 splice variant 4, a cryptic splice site in Domain 2 is utilized and Domains 2 and 3 and the alternative exon are skipped. Arrows indicate the positions of PCR primers flanking the regions of interest.
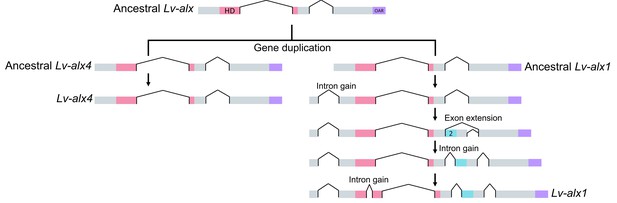
Model representing the evolution of L. variegatus alx genes.
Following gene duplication, the ancestral Lv-alx1 underwent rapid evolution through multiple intron gains and more importantly, acquired Domain 2 through exonization of previously non-coding sequences. By contrast, Lv-alx4 retained an intron-exon structure very similar to that of the ancestral Lv-alx4.
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.32728.018





