Architecture of the human mTORC2 core complex
Figures
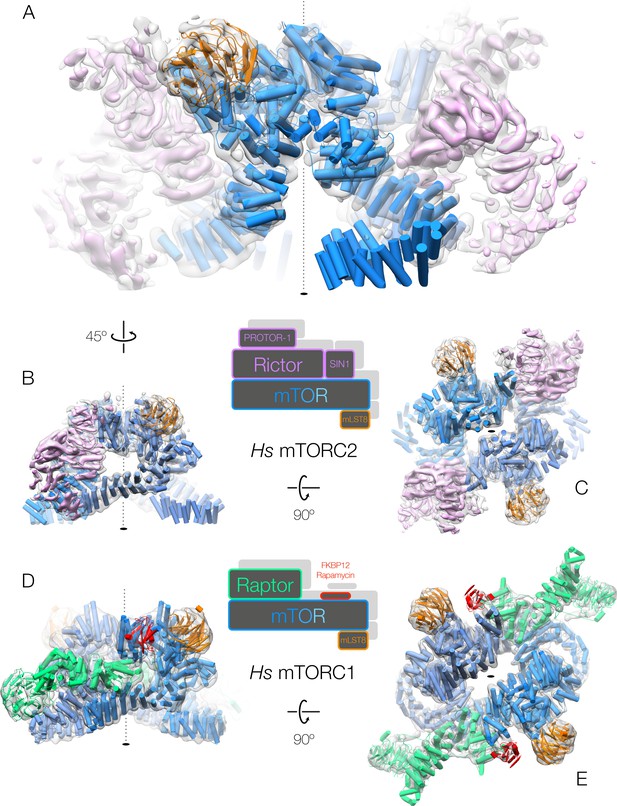
Both human mTOR complexes resolved at intermediate resolution.
(A–C) The architecture of human mTORC2. The structure is shown rotated as indicated by the arrows between the panels. The accessory factor density from focused refinement is shown within the dimeric, C2-symmetric mTORC2 density in pink. (D–E) The architecture of human mTORC1 (Aylett et al., 2016). The structure is shown rotated as indicated by the arrows between the panels. All complexes are shown with cryo-EM density as a grey transparent surface and the fitted structures in cartoon representation, coloured according to the primary structure schematics shown between the corresponding panels.
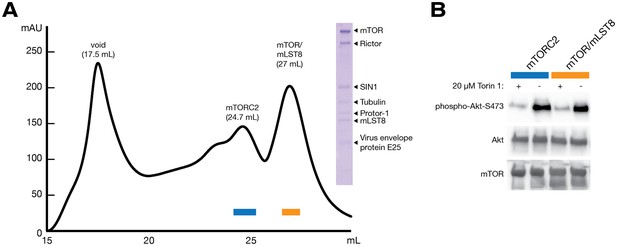
Purification and characterisation of mTORC2.
(A) Size exclusion chromatography elution profile of mTORC2 from a tandem Superose 6 Increase 10/300 GL column. Peaks representing the void, mTORC2 and excess mTOR-mLST8 are labeled. SDS-PAGE analysis of the final mTORC2 sample is shown in inset. Tubulin and virus envelope protein E25 were detected by mass spectrometry as contaminants. Fractions used for the in vitro kinase activity assay are indicated with a blue and orange box. (B) In vitro kinase activity assay of recombinant mTORC2. Western blots showing the phosphorylation state of Akt-1 in the presence and absence of mTOR inhibitor Torin1.
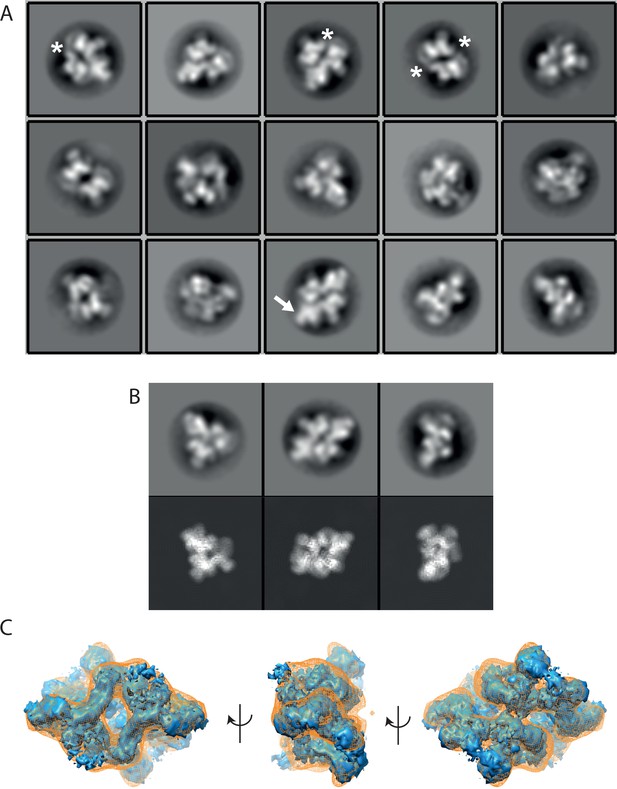
Analysis of negatively stained mTORC2 without applying chemical fixation.
(A) Most populated class averages (out of 80 class averages) of a 2D classification of negatively stained non-cross-linked mTORC2. In some class averages a peripheral density (arrow) is underrepresented (star). (B) Comparison of representative class averages (upper panel) with reprojections of the cryo-EM reconstruction with matching projection angles (lower panel). (C) Reconstruction of the non-cross-linked mTORC2 (orange mesh) from negatively stained grids superimposed on a down sampled cryo-EM density of the cross-linked mTORC2.
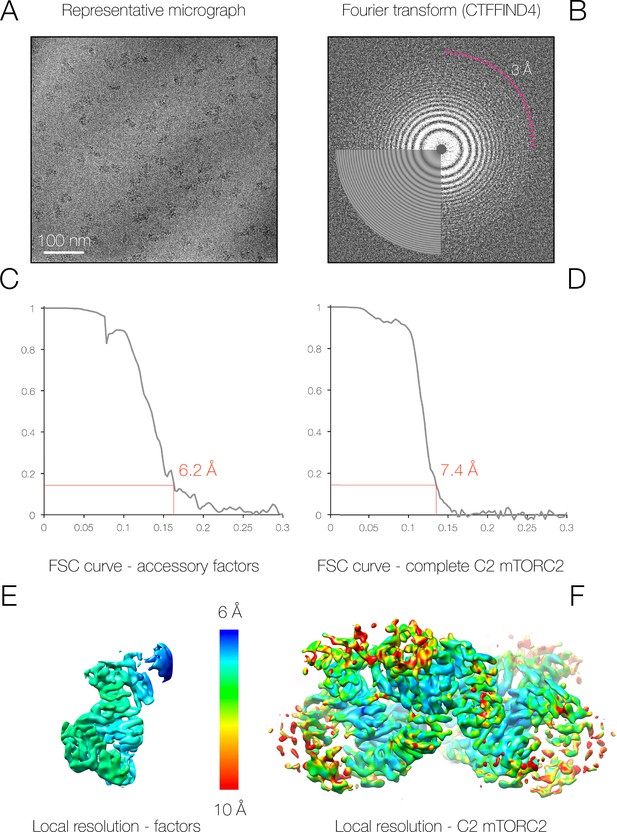
Sample micrograph and resolution statistics for mTORC2.
(A–B) Sample micrograph and its power spectrum. Thon rings visible to 3 Å are indicated. (C–D) Fourier shell correlation plots for the full mTORC2 dimeric complex and the mTORC2 accessory protein region, calculated between independently refined half sets (gold-standard) (Scheres, 2012). (E–F) The surface of each mTORC2 density is shown coloured according to local resolution (6 Å in blue through to 10 Å in red) (Kucukelbir et al., 2014).
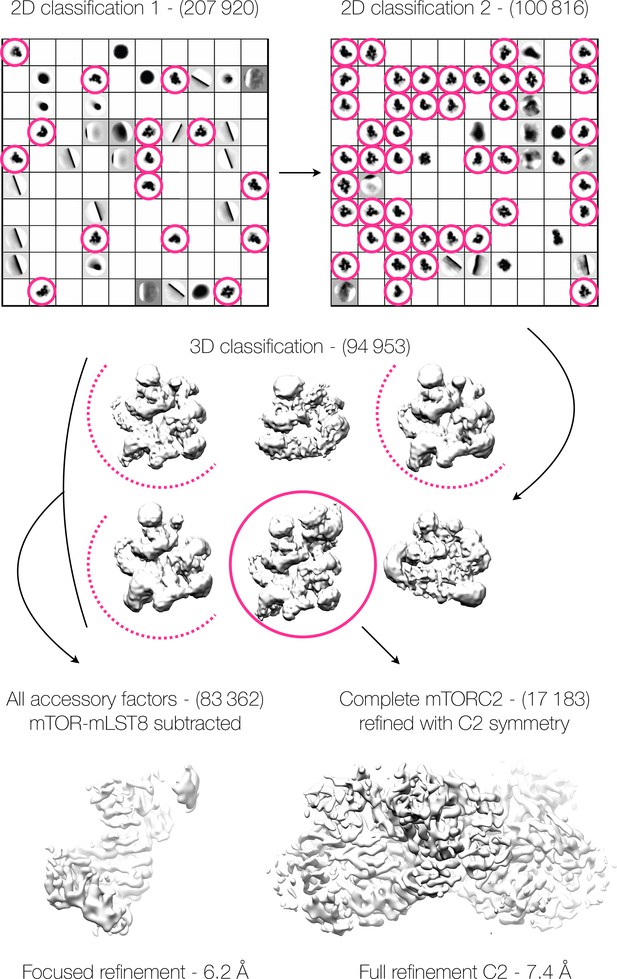
Cryo-EM classification schematic.
The reconstructions from each round of classification, the number of particles involved in each step, and the retention of particles between stages of the classification are shown as a flow diagram. Magenta circles indicate the classes from which particles were retained at each stage, dotted half-circles indicate single-sided particles retained after subtraction of the opposing density.
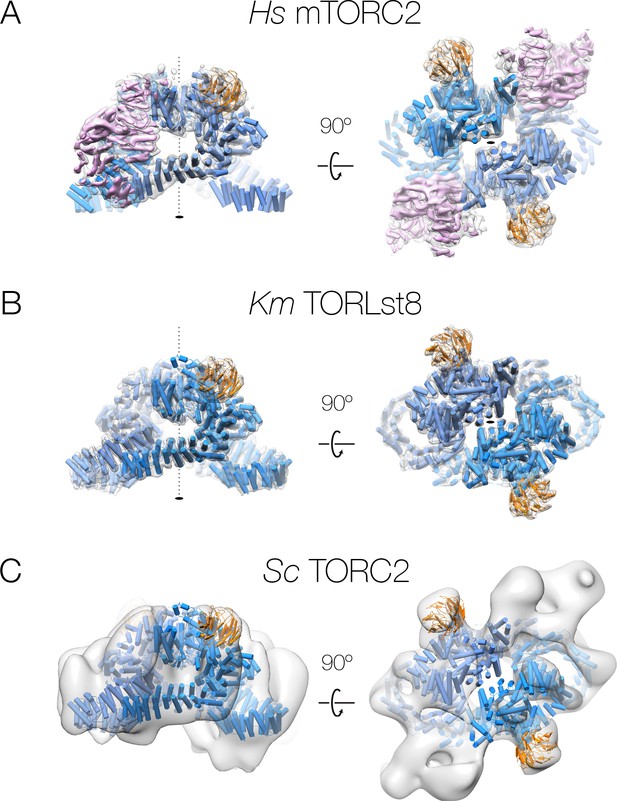
Comparison of human and yeast TOR complexes:.
(A) human mTORC2, (B) K.m. TOR-Lst8, (C) S.c. TORC2. Each structure is shown rotated as indicated by the arrows. The accessory factor density from focused refinement is shown within the C2-symmetric mTORC2 density. All complexes are shown with the corresponding cryo-EM density as a grey transparent surface, the fitted structures in cartoon representation, and are coloured according to the colour scheme from Figure 1.
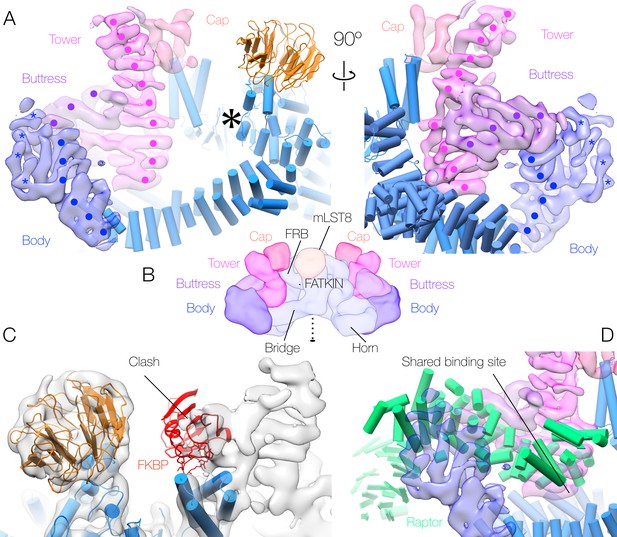
Human mTORC2 accessory factor density and binding sites.
(A) Accessory factor density bound to the mTOR dimer. The regions of α-solenoid are indicated by surface colour, and the visible helical hairpins denoted by points (‘•’ - well-ordered regions believed to correspond to Rictor) or 5-pointed stars (‘*’ - poorly ordered regions that cannot be definitively assigned to Rictor or Protor-1). mTOR-mLST8 is shown in cartoon representation, and the mTOR active site is indicated by a large 6-pointed asterisk ‘*’. (B) Schematic overview of the 3-dimensional layout of the mTORC2 complex in the same colour scheme. Well-ordered α-solenoidal repeat density, almost all of which will correspond to Rictor, makes up the tower and buttress. The cap and body each consist of less well-ordered density; the body comprises density connecting to the buttress, and is believed to contain both regions of Rictor and peripheral density belonging to Protor-1 (see Supplement 1), whereas the cap contains density continuing from the tower, which is believed to consist of regions of Rictor and possibly SIN1. (C) Superimposition of the mTORC2 density upon the mTORC1 model, indicating the clash between FKBP and the density corresponding to the cap (black arrow). (D) Comparison between the binding site (black arrow) of Raptor within mTORC1 (cartoon representation) and the corresponding binding site for the mTORC2 accessory factors (surface representation). All panels are coloured according to the scheme in Figure 1.
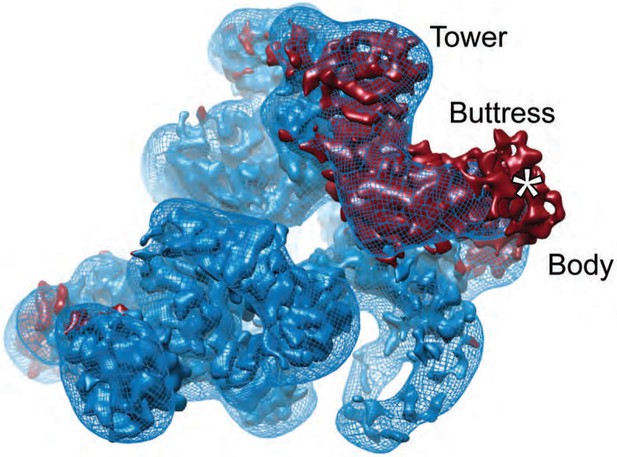
Assignment of density to Rictor/SIN-1.
Negatively stained EM reconstruction of mTORC2-ΔProtor-1 (blue mesh) superimposed on the cryo-EM reconstruction of mTORC2 (blue surface) with the accessory factor density colored red. The mTORC2-ΔProtor-1 accessory factor density is very similar in size and shape to that in the mTORC2 cryo-EM reconstruction indicating that the ‘tower’ and ‘buttress’ structure are part of Rictor as is most of the ‘body’. Density in the mTORC2 ‘body’ absent in the mTORC2-ΔProtor-1 reconstruction (*) might correspond to partial density for Protor-1. We note that this region corresponds to the part of S.c. TORC2 tentatively identified as Bit2/61 (Gaubitz et al., 2015).
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.33101.011





