Current CRISPR gene drive systems are likely to be highly invasive in wild populations
Figures
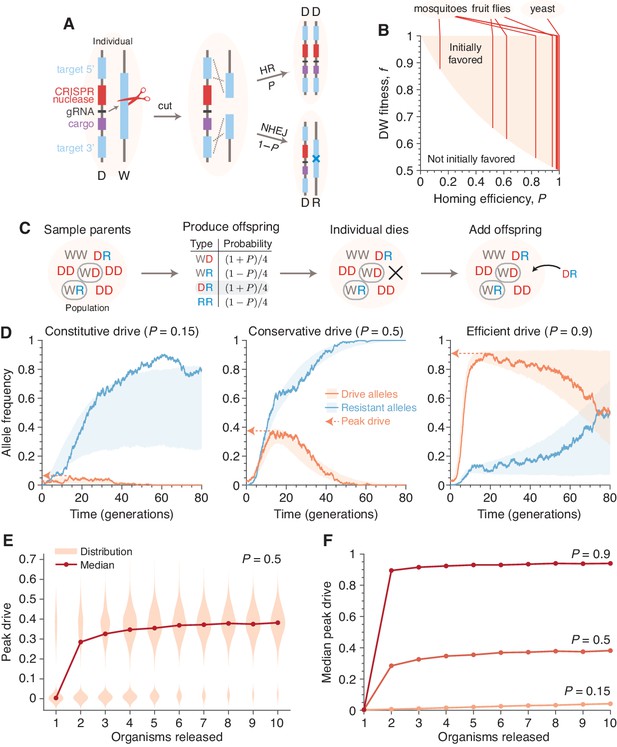
Existing alteration-type CRISPR gene drive systems should invade well-mixed wild populations.
(A) Typical construction and function of alteration-type CRISPR gene drive systems. A drive construct (D), including a CRISPR nuclease, guide RNA (gRNA), and ‘cargo’ sequence, induces cutting at a wild-type allele (W) with homology to sequences flanking the drive construct. Repair by homologous recombination (HR) results in conversion of the wild-type to a drive allele, or repair by nonhomologous end-joining (NHEJ) produces a drive-resistant allele (R). (B) Drives are predicted to invade by deterministic models when the fitness of DW heterozygotes, , and the homing efficiency, , are in the shaded region. Vertical lines indicate empirical efficiencies from Appendix 1—table 1. (C) Diagram of a single step of the gene-drive Moran process. (D) Finite-population simulations of 15 drive individuals released into a wild population of size 500, assuming conservative () or high () homing efficiencies, as well as a low-efficiency, constitutively active system (). Individual sample simulations (solid lines), and 50% confidence intervals (shaded), calculated from simulations. Drive-allele frequencies red and resistant-allele frequencies blue. Peak drive, or maximum frequency reached, is illustrated by dashed lines and arrows. (E) Peak drive distributions and medians with varying numbers of individual organisms released (). (F) Medians of peak drive distributions for varying homing efficiencies (, bottom; , middle; , top). Throughout, we assume neutral resistance () and a 10% dominant drive fitness cost ().
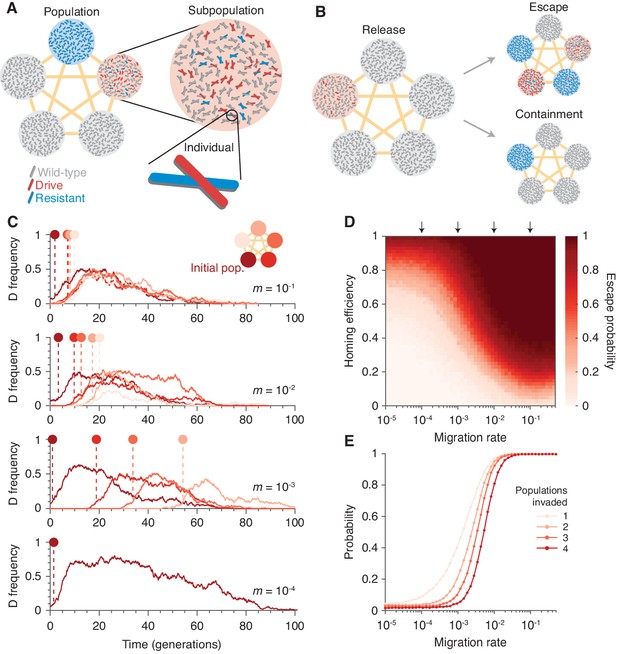
Existing CRISPR gene drive systems should invade linked subpopulations connected by gene flow.
(A) Diagram of well-mixed subpopulations (circles) linked by gene flow (edges). Individuals represented by chromosomes with wild-type (gray), drive (red), or resistant (blue) haplotypes. (B) Few drive homozygotes are released in one subpopulation. The drive escapes if it invades another subpopulation before going extinct. Otherwise it is contained. (C) Typical simulations for varying migration rates (, top, to , bottom), following introduction into a single subpopulation. Lines represent drive frequencies in each subpopulation. Circles correspond to the time the drive invades a subpopulation. Population color is by invasion order, not predetermined. (D) Escape probability as a function of homing efficiency, , and migration rate, . Arrows indicate migration rates from B. Each pixel is calculated from simulations. (E) Probability of invading 1, 2, 3, or 4 additional populations (aside from the originating population, which is typically invaded), assuming a homing efficiency of . Each data point is calculated from simulations. Throughout, we consider five subpopulations connected in a complete graph, each consisting of 100 individuals. Initially, 15 drive homozygotes are introduced into one subpopulation. Resistance is neutral () and the drive confers a dominant cost ().
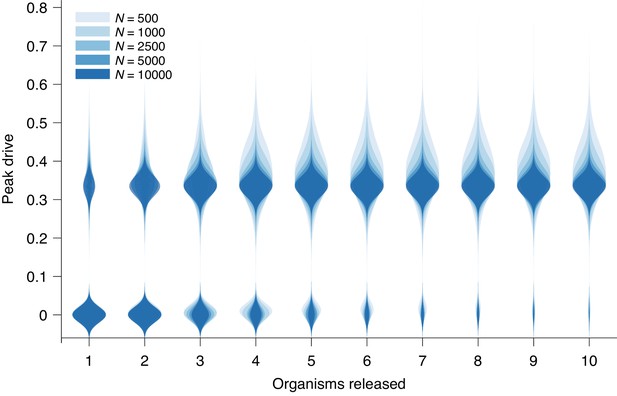
Peak drive distributions for variable release and population sizes.
Parameters are chosen to correspond to Figure 1E: , and neutral resistance. Population sizes are, from light to dark, . Note that corresponds exactly to Figure 1E. Each distribution corresponds to simulations.
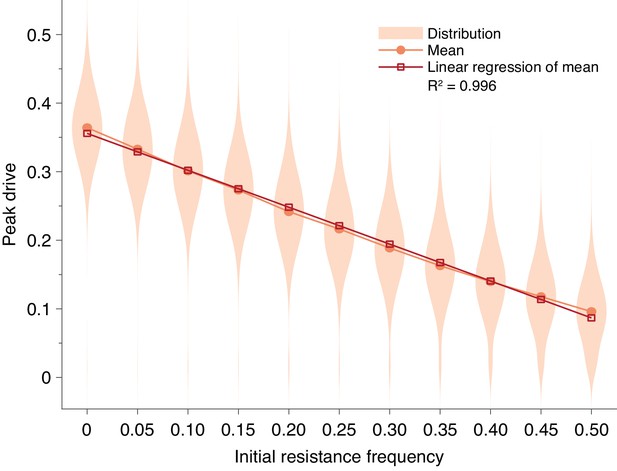
Pre-existing drive-resistant allele frequency linearly decreases peak drive.
Distributions (violin plots), means (orange, circles) and linear regression of the mean values (red, squares). Parameters are chosen to correspond to Figure 1E: , , neutral resistance, . Each distribution corresponds to simulations.
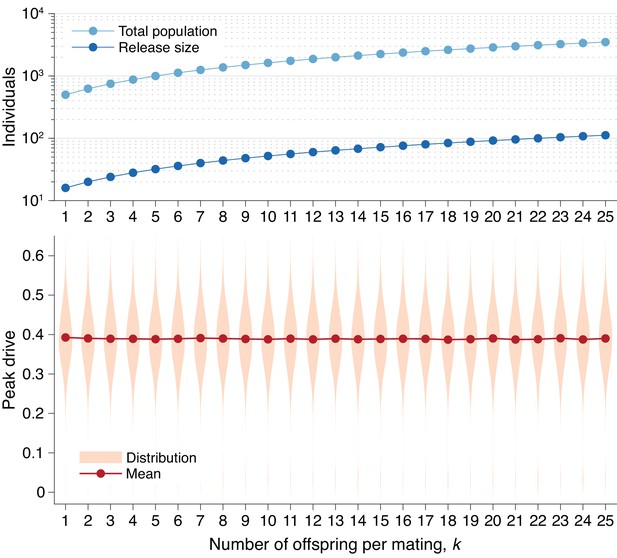
Peak drive distributions for varying numbers of offspring per mating with effective population and release sizes held constant.
(top) Population and release sizes used in the simulations below. For the case , we use our usual population size of with an initial release of drive homozygotes. According to Equation (5), the effective total population and release sizes in this case are and . For other values of , we use values of and which maintain constant effective population and release sizes: and . These values are plotted: (light blue) and (dark blue). (bottom) Peak drive distributions assuming values of and as in the above plot. All employ , , and neutral resistance. Each distribution includes 5000 simulations.
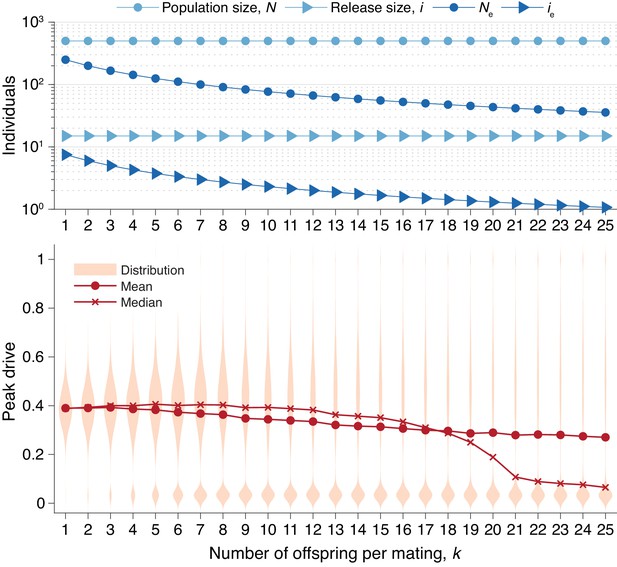
Peak drive distributions for varying numbers of offspring per mating with census population and actual release sizes held constant.
(top) Population and release sizes used in the simulations below. Actual population size, (light blue, circles) and actual release size, (light blue, triangles). Note that and are constant. Effective values calculated via Equation (5): population size, (dark blue, circles) and release size, (dark blue, triangles). (bottom) Peak drive distributions for simulations using indicated values of and population and release sizes as depicted above. Compare with Figure 5 which holds the effective population and release sizes constant, whereas here we hold the census population and release sizes constant. All simulations employ , , and neutral resistance. Each distribution includes 5000 simulations.
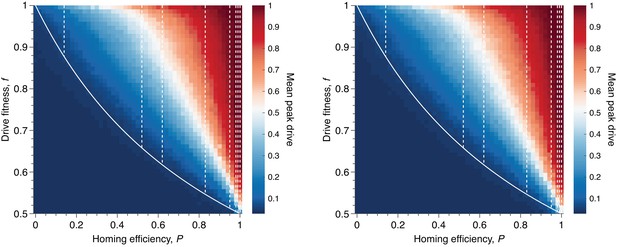
Mean peak drive for varying homing efficiency, , and drive-individual fitness values, (i.e., individuals with genotypes WD, DD, and DR), assuming that fitness affects birth rate (left) or death rate (right).
The left panel corresponds to our standard model, shown in Figure 1C, while the right panel represents a modification: parents are chosen uniformly, and individuals die with probability proportional to the inverse of their fitness. The solid white line shows the boundary from Figure 1B indicating whether the drive is predicted to invade by deterministic models. The drive is only expected to invade based on deterministic models if the fitness/homing efficiency pair lie above the boundary. The dashed white lines indicate the empirically measured homing efficiencies from Appendix 1—table 1 and Figure 1B. Each point in the grid () depicts an average of 100 simulations. Parameters used include a population size of 500, with an initial release of 15 drive homozygotes to ensure that trajectories establish. Neutral resistance is assumed throughout with no standing genetic variation.
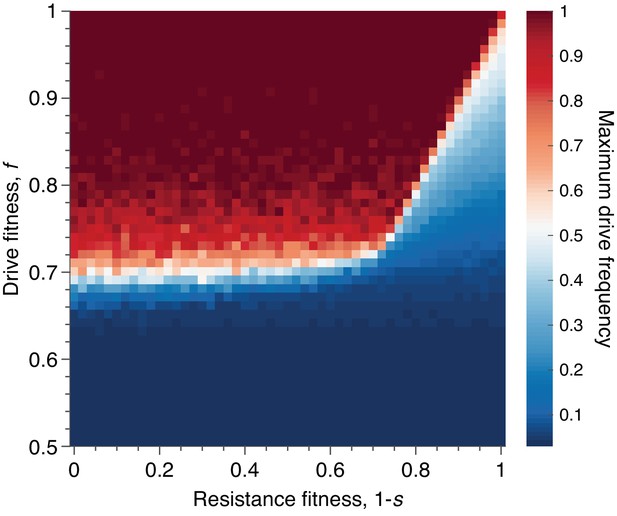
Mean peak drive for varying drive-individual fitness values, , and resistant-individual (RR) fitness values, , where is the cost associated with resistance.
Each point in the grid () depicts an average of 100 simulations. Parameters used include homing efficiency , population size of 500, with an initial release of 15 drive homozygotes to ensure that trajectories establish. Throughout we assume no standing genetic variation (i.e., the initial frequency of the resistant allele is 0).
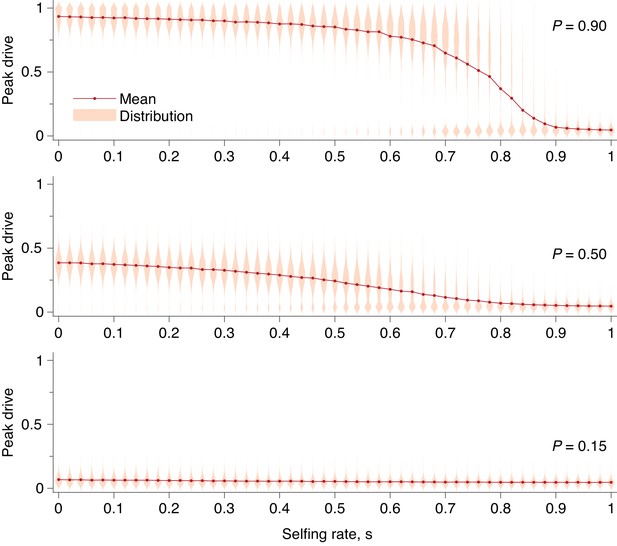
Peak drive distributions and means for varying selfing rates in our partial selfing model.
(top) Effective drive, . (middle) conservative drive, , and (bottom) constitutive drive, . Each distribution comprises simulations. Parameters used include a population size of 500 with an initial release of 15 drive homozygotes. Neutral resistance is assumed throughout with no standing genetic variation, and the offspring number per mating is .
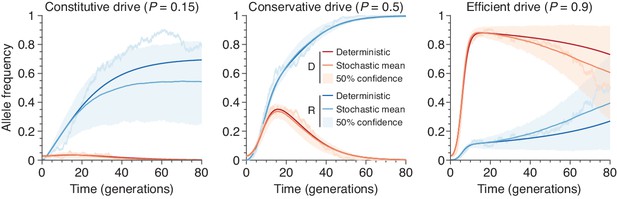
Finite-population simulations of 15 drive individuals released into a wild population of size 500, assuming low () or high () homing efficiencies, as well as a low-efficiency, constitutively active system ().
Deterministic results (dark lines) and means of simulations (medium lines), individual sample simulations (light lines), and 50% confidence intervals (shaded). Drive frequencies red and resistant-allele frequencies blue.
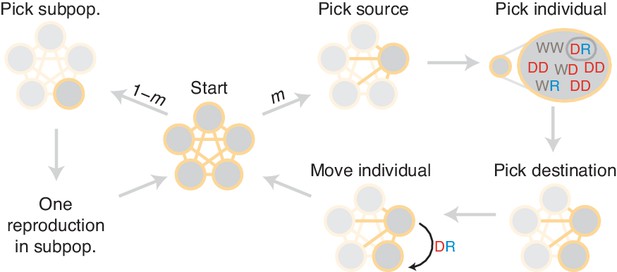
Diagram of simulation scheme.
In each time step, a migration occurs with probability , or a mating happens with probability . If a migration occurs, a source population is chosen randomly proportional to its size; an individual is chosen uniformly at random, then a destination is chosen uniformly at random, and the individual is moved. If a mating occurs, the dynamics proceed as in the well-mixed case for a particular subpopulation (Figure 1C).
Tables
Empirical homing efficiencies for all CRISPR gene drive systems published to date.
Details can be found in the Appendix.
| Organism | Ref. | System name | Efficiency |
|---|---|---|---|
| Yeast | (DiCarlo et al., 2015) | ade2::sgRNA | |
| ade2::sgRNA + URA3 | |||
| sgRNA + ABD1 | |||
| cas9 + sgRNA | |||
| ADE2 + sgRNA + cas9 | |||
| Fruit flies | (Gantz and Bier, 2015) | -MCR | |
| (Champer et al., 2017) | nanos | ||
| vasa | |||
| additional nanos | 40–62% | ||
| additional vasa | 37–53% | ||
| Mosquitoes | (Gantz et al., 2015) | AsMCRkh2 (male) | |
| AsMCRkh2 (female) | |||
| (Hammond et al., 2016) | AGAP011377 | ||
| AGAP005958 | |||
| AGAP007280 |
Gantz et al., An. stephensi transgenic male lines.
(left) Phenotypes of G progeny. (right) Phenotypes of G progeny.
| G crosses | Reference | ||
|---|---|---|---|
| GWT, larval | Table S3 | ||
| GWT, larval | Table S4 | ||
| GWT, adult | Table S5 | ||
| GWT, adult | Table S6 | ||
| Total | — | ||
| G crosses | Reference | ||
| Cross 6, larval | Table S7 | ||
| Cross 8, larval | Table S8 | ||
| Cross 6, adult | Table S10 | ||
| Cross 8, adult | Table S11 | ||
| Total | — |
Gantz et al., An. stephensi transgenic male lines.
(left) Phenotypes of G larvae. (right) Phenotypes of G adults.
| G larvae | Reference | ||
|---|---|---|---|
| Cross | Table S7 | ||
| Cross 2 | Table S7 | ||
| Cross 3 | Table S8 | ||
| Cross 4 | Table S8 | ||
| Total | — | ||
| G adults | Reference | ||
| Cross 1 | Table S10 | ||
| Cross 2 | Table S10 | ||
| Cross 3 | Table S11 | ||
| Cross 4 | Table S11 | ||
| Total | — |
Additional files
-
Transparent reporting form
- https://doi.org/10.7554/eLife.33423.014





