Fidaxomicin jams Mycobacterium tuberculosis RNA polymerase motions needed for initiation via RbpA contacts
Figures
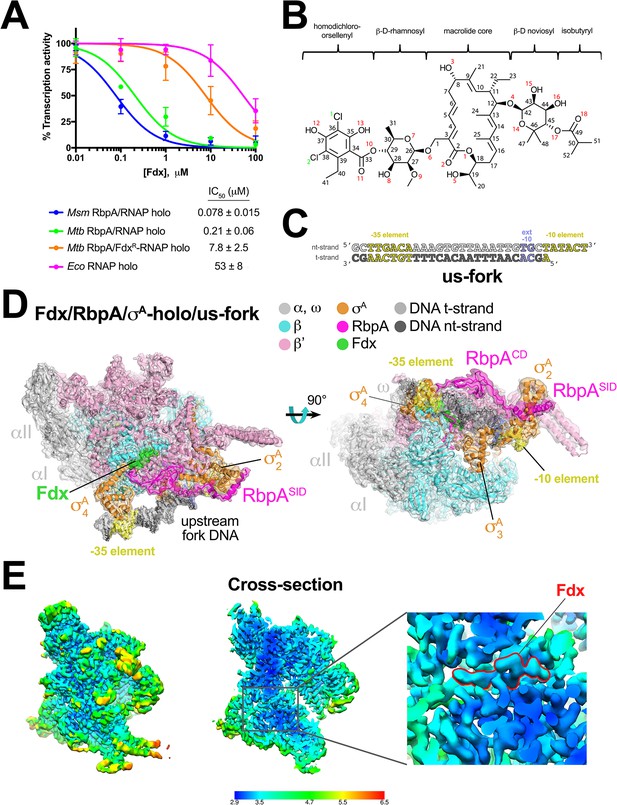
Structure of an Mtb RbpA/TIC with Fdx at 3.4 Å resolution.
(A) Fdx inhibits mycobacterial RbpA/σA-holo transcription greater than 250-fold more effectively than Ecoσ70-holo in in vitro abortive initiation assays. The error bars denote the standard error from a minimum of three experiments (for some points, the error bars are smaller than the width of the point and are not shown). (B) Chemical structure of Fdx (Serra et al., 2017). (C) Synthetic us-fork promoter fragment used for cryo-EM experiments. The DNA sequence is derived from the full con promoter (Gaal et al., 2001). The nontemplate-strand DNA (top strand) is colored light gray; the template-strand DNA (bottom strand), dark grey. The −35 and −10 elements are shaded yellow. The extended −10 (Keilty and Rosenberg, 1987) is colored violet. (D) The 3.4 Å resolution cryo-EM density map of the Fdx/RbpA/σA-holo/us-fork complex is rendered as a transparent surface colored as labeled. Superimposed is the final refined model; proteins are shown as a backbone ribbon, Fdx and the nucleic acids are shown in stick format. (E) Views of the cryo-EM map colored by local resolution based on blocres calculation (Cardone et al., 2013). The left view shows the entire map, while the middle view shows a cross-section of the map sliced at the level of the Fdx binding pocket. The boxed region is magnified on the right. Density for the Fdx molecule is outlined in red.
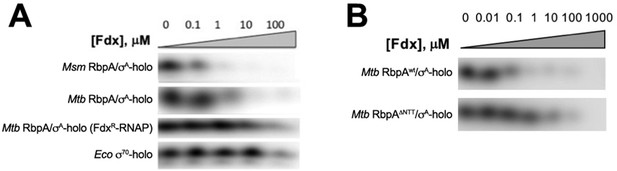
Abortive Transcription assays to determine IC50 for Fdx.
(A, B) Abortive transcription activity of samples (labeled) after incubation with the indicated concentrations of Fdx.
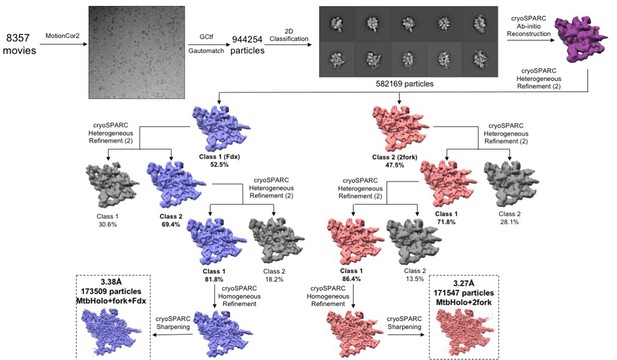
Data processing pipeline for the cryo-EM movies of the Fdx/RbpA/σA-holo/us-fork complexes.
Flowchart showing the image processing pipeline for the cryo-EM data of Mtb Fdx/RbpA/σA-holo/us-fork complexes starting with 8357 dose-fractionated movies collected on a 300 keV Titan Krios (FEI) equipped with a K2 Summit direct electron detector (Gatan). Movies were frame aligned and summed using MotionCor2 (Zheng et al., 2017). CTF estimation for each micrograph was calculated with Gctf (Zhang, 2016). A representative micrograph is shown following processing by MotionCor2. Particles were autopicked from each micrograph with Gautomatch and then sorted by 2D classification using RELION (Scheres, 2012) to assess quality. The ten highest populated classes from the 2D classification are shown. After initial cleaning, the dataset contained 582,169 particles. A subset of particles was used to generate an ab-initio map in cryoSPARC (Punjani et al., 2017). Using the low-pass filtered (60 Å) ab-initio map as a template, 3D heterogeneous refinement was performed in cryoSPARC in a binomial-like fashion. Two major classes were observed: Fdx/RbpA/σA-holo/us-fork (blue) and RbpA/σA-holo//(us-fork)2 (red). Multiple rounds of 3D classification were performed; colored classes represent the selected classes while grey classes were thrown out. The final sets of particles for each class were refined using cryoSPARC homogenous refinement and then sharpened for model building.
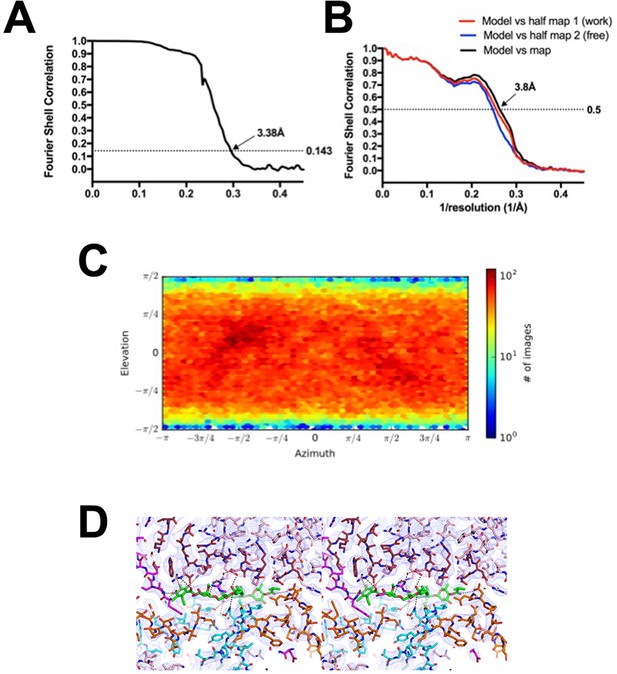
Fdx/RbpA/σA-holo/us-fork class.
(A) Gold-standard FSC (Rosenthal and Henderson, 2003), calculated by comparing the two independently determined half-maps from cryoSPARC. The dotted line represents the 0.143 FSC cutoff which indicates a nominal resolution of 3.38 Å. (B) FSC calculated between the refined structure and the half map used for refinement (work, red), the other half map (free, blue), and the full map (black). (C) Angular distribution calculated in cryoSPARC for particle projections. Heat map shows number of particles for each viewing angle (less = blue, more = red). (D) Stereo view of the Fdx binding pocket with cryo-EM density (blue mesh) superimposed.
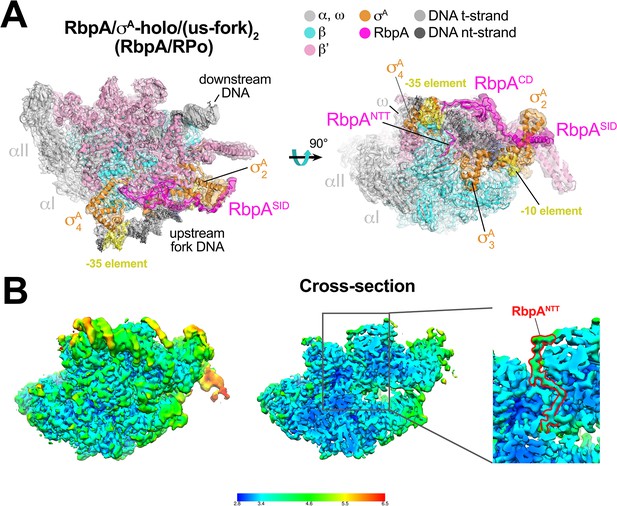
Structure of an Mtb RbpA/RPo mimic at 3.3 Å resolution.
(A) The 3.3 Å resolution cryo-EM density map of the RbpA/σA-holo/(us-fork)2 complex (RbpA/RPo mimic) is rendered as a transparent surface colored as labeled. Superimposed is the final refined model; proteins are shown as a backbone ribbon, nucleic acids are shown in stick format. (B) Views of the Mtb RbpA/RPo mimic cryo-EM map colored by local resolution based on blocres calculation (Cardone et al., 2013). The left view shows the entire map, while the middle view shows a cross-section of the map sliced at the level of the RbpANTT. The boxed region is magnified on the right. Density for the RbpANTT is outlined in red.
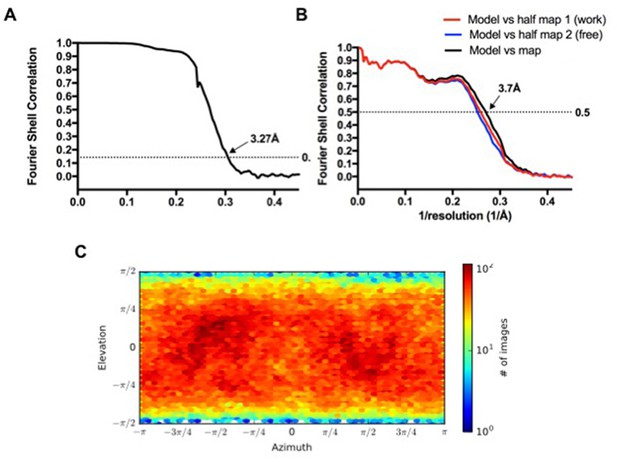
RbpA/σA-holo/(us-fork)2 class.
(A) Gold-standard FSC (Rosenthal and Henderson, 2003), calculated by comparing the two independently determined half-maps from cryoSPARC. The dotted line represents the 0.143 FSC cutoff which indicates a nominal resolution of 3.27 Å. (B) FSC calculated between the refined structure and the half map used for refinement (work, red), the other half map (free, blue), and the full map (black). (C) Angular distribution calculated in cryoSPARC for particle projections. Heat map shows number of particles for each viewing angle (less = blue, more = red).
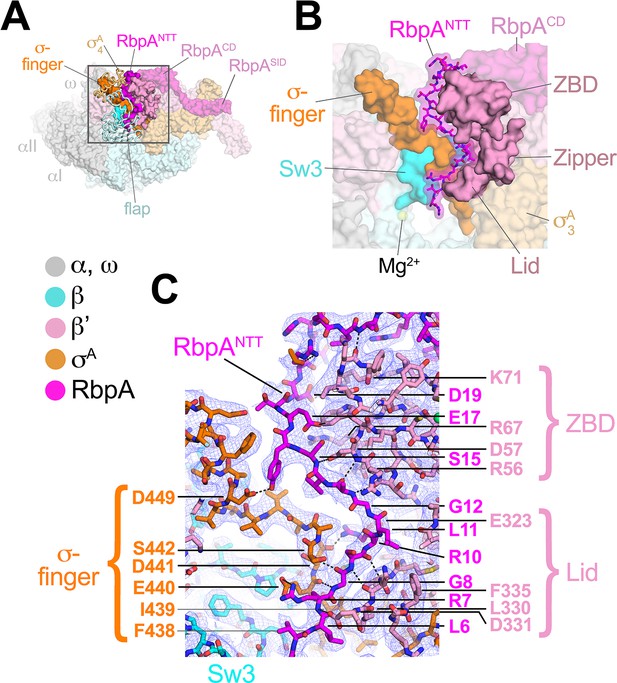
The RbpANTT interacts with conserved structural elements in the RNAP active site cleft.
(A) An overview of the RbpA/RPo structure is shown as a color-coded molecular surface (color-coding denoted in the key) except the β flap and σA4 domain are shown as backbone worms, revealing the RbpANTT (magenta) underneath. The DNA fragments are not shown. The boxed region is magnified in panel (B). (B) Magnified view of the boxed region from panel (A). The RbpANTT is shown in stick format with a transparent molecular surface. Conserved RNAP structural elements that interact with the RbpANTT are highlighted (βSw3, β'ZBD, β'Zipper, β'Lid, and σ-finger). (C) Further magnified view showing the cryo-EM density (blue mesh) with the superimposed model. Conserved residues of the RbpANTT are labeled, along with conserved residues of the β'ZBD, β'Lid, and σ-finger that interact with the RbpANTT.
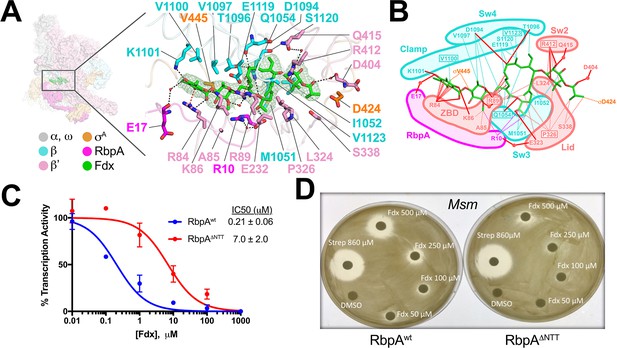
Structural basis for Fdx inhibition of Mtb transcription and the role of the RbpANTT.
(A) (left) Overview of the Fdx/RbpA/σA-holo/us-fork structure, shown as a molecular surface (the DNA is not shown). The boxed region is magnified on the right. (right) Magnified view of the Fdx binding pocket at the same orientation as the boxed region on the left. Proteins are shown as α-carbon backbone worms. Residues that interact with Fdx are shown in stick format. Fdx is shown in stick format with green carbon atoms. Water molecules are shown as small pink spheres. Hydrogen-bonds are indicated by dashed gray lines. Cation-π interactions (between β'R84 and the aromatic ring of the Fdx homodichloroorsellinic acid moiety and β'R89 and the conjugated double-bond system centered between C4 and C5 of the macrolide core) are represented by red dashed lines. (B) Schematic summary of the Fdx contacts with σA-holo and RbpA. Fdx is shown in stick format with green carbon atoms. Thin dashed lines represent van der Waals contacts (≤4.5 Å), thick red lines represent hydrogen bonds (<4 Å). The thin dashed red lines denote cation-π interactions. (C) The RbpANTT is required for optimal inhibition of Mtb transcription by Fdx in in vitro abortive initiation assays. The error bars denote the standard error from a minimum of three experiments (for some points, the error bars are smaller than the width of the point and are not shown). (D) Zone of inhibition assays with Msm cells show that loss of the RbpA-NTT (RbpAΔNTT) leads to loss of Fdx sensitivity in vivo.
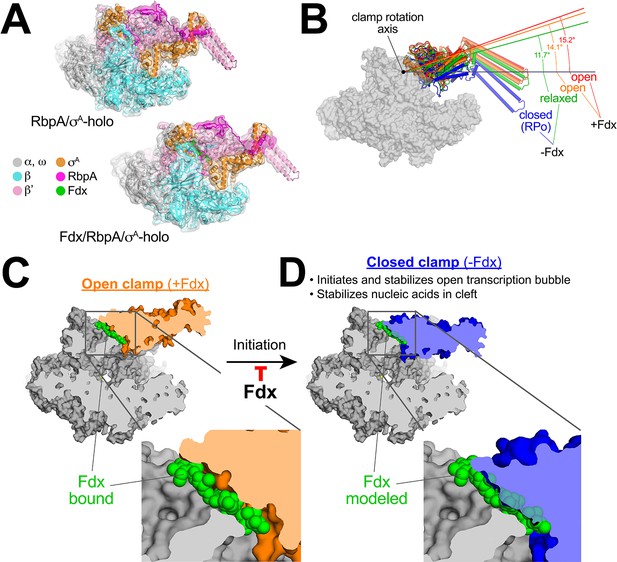
Mechanism of Fdx inhibition of bacterial RNAP.
(A) Cryo-EM density maps and superimposed refined models for Mtb RbpA/σA-holo (5.2 Å resolution) and Mtb Fdx/RbpA/σA-holo (6.5 Å resolution). (B) RNAP clamp conformational changes for four cryo-EM structures determined in this work. The RbpA/RPo (Figure 2A) structure was used as a reference to superimpose the other structures via α-carbon atoms of the structural core module (Supplementary file 2), revealing a common core RNAP structure (shown as a gray molecular surface) but with large shifts in the clamp modules. The clamp modules are shown as backbone cartoons with cylindrical helices and color-coded (blue, closed clamp of RPo; green, relaxed clamp of RbpA/σAholo; orange, open clamp of Fdx/RbpA/σA-holo/us-fork; red, open clamp of Fdx/RbpA/σA-holo). The clamp conformational changes can be characterized as rigid body rotations about a rotation axis perpendicular to the page (denoted). The angles of clamp opening for the different structures are shown (relative to the blue closed RPo clamp, 0° opening). (C) The core RNAP from the 3.4 Å resolution Fdx/RbpA/σA-holo/us-fork structure is shown as a gray molecular surface but with the open clamp colored orange. The structure is sliced at the level of the Fdx binding pocket (the bound Fdx is shown in green). The boxed region is magnified below, showing the tight fit of the Fdx molecule in a narrow gap between the clamp and the rest of the RNAP. (D) The core RNAP from the 3.3 Å resolution RbpA/RPo structure is shown as a gray molecular surface but with the closed clamp colored blue. The structure is sliced at the level of the (empty) Fdx binding pocket. Fdx, modeled from the structure shown in (C), is shown in green. The boxed region is magnified below. Fdx cannot bind to RNAP with a closed clamp because clamp closure pinches off the Fdx binding site. Clamp closure is required for initiation and stabilization of the transcription bubble (Feklistov et al., 2017) and also for stable binding of nucleic acids in the RNAP cleft.
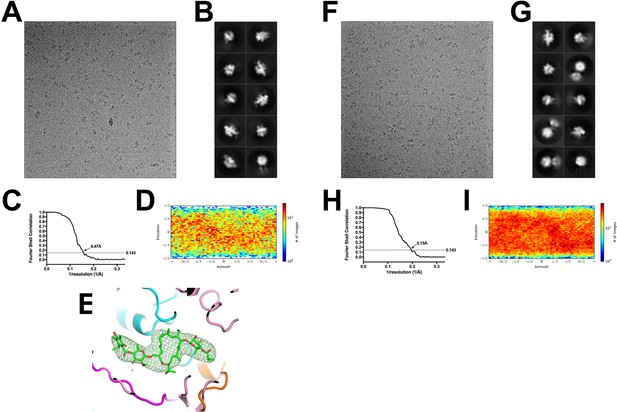
Cryo-EM of the Mtb RbpA/σA-holo and Fdx/RbpA/σA-holo complexes.
(A) Representative micrograph of the Fdx/RbpA/σA-holo complex in vitreous ice. (B) The ten highest populated classes from cryoSPARC 2D classification. (C) Gold-standard FSC of the Mtb Holo/Fdx complex. The gold-standard FSC was calculated by comparing the two independently determined half-maps from cryoSPARC. The dotted line represents the 0.143 FSC cutoff which indicates a nominal resolution of 6.47 Å. (D) Angular distribution calculated in cryoSPARC for Mtb Fdx/RbpA/σA-holo particle projections. Heat map shows number of particles for each viewing angle (less = blue, more = red). (E) Difference map showing Fdx is bound in 6.5 Å resolution structure. (F) Representative micrograph of the Mtb RbpA//σA-holo complex in vitreous ice. (G) The ten highest populated classes from cryoSPARC 2D classification. (H) Gold-standard FSC of the Mtb Holo complex. The gold-standard FSC was calculated by comparing the two independently determined half-maps from cryoSPARC. The dotted line represents the 0.143 FSC cutoff which indicates a nominal resolution of 5.15 Å. (I) Angular distribution calculated in cryoSPARC for Mtb Holo particle projections. Heat map shows number of particles for each viewing angle (less = blue, more = red).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | Eco BL21(DE3) | EMD-Millipore (Novagen; Darmstadt, Germany) | ||
| Strain, strain background (Mycobacterium smegmatis mc2155) | MGM6029: Msm mc2155 rpoC:rpoC- ppx-10his hyg | PMID: 28067618 | ||
| Strain, strain background (Mycobacterium smegmatis mc2155) | MGM6234: Msm mc2155ΔrbpA attB:: rbpA(28-114) kan | PMID: 28067618 | ||
| Recombinant DNA reagent | pAC22 | PMID: 24713321 | pET28a derivative. Encodes M. bovis RNAP. β contains a S450Y substitution (RifR) and a short N-terminal insertion at codon 2 (LEGCIL); β′ has C-terminal His8 tag; β and β′ are fused with a short linker (LARHGGSGA) | |
| Recombinant DNA reagent | pACYCDuet- 1_Ec_rpoZ | PMID: 23389035 | ||
| Recombinant DNA reagent | pET21a-Ecoσ70 | PMID: 24218560 | ||
| Recombinant DNA reagent | pet21C-MsmRbpA | PMID: 28067618 | ||
| Recombinant DNA reagent | pet21C-MtbRbpA | PMID: 28067618 | ||
| Recombinant DNA reagent | pET28a | EMD-Millipore (Novagen) | ||
| Recombinant DNA reagent | pET-SUMO MsmσA | PMID: 28067618 | ||
| Recombinant DNA reagent | pET-SUMO MtbσA | PMID: 28067618 | ||
| Recombinant DNA reagent | pET-SUMO MsmRbpAΔNTT | PMID: 28067618 | ||
| Recombinant DNA reagent | pET-SUMO MtbRbpAΔNTT | PMID: 28067618 | ||
| Recombinant DNA reagent | pGEMABC | PMID: 23389035 | Addgene 45398 | |
| Recombinant DNA reagent | pMP55 | this paper | pAC22 derivative encoding Mtb RNAP with β S450Y. Derived from pAC22 by P69R substitution, removal of the N-terminal β insertion, and substitutions of increased predicted-strength RBSs for the rpoA, rpoZ, and rpoB::C RBSs. | |
| Recombinant DNA reagent | pMP57 | this paper | pMP55 with β Q1054H. Fdx resistant. | |
| Recombinant DNA reagent | pMP61 | this paper | pMP55 with wild-type S450 in place of β Y450. | |
| Recombinant DNA reagent | pMP62 | this paper | pMP61 with β S450L. | |
| Recombinant DNA reagent | pRARE2 | EMD-Millipore (Novagen) | ||
| Recombinant DNA reagent | pUC57-AP3 | PMID: 25510492 | ||
| Chemical compound, drug | Fidaxomicin | VWR International, Inc. (Radnor, PA) | 2832–1 | |
| Chemical compound, drug | 3-([3-cholamidopropyl]dimethylammonio)−2 -hydroxy-1-propanesulfonate (CHAPSO) | Sigma-Aldrich (St. Louis, MO) | C4695 | |
| Software, algorithm | Blocres | PMID: 23954653 | ||
| Software, algorithm | Chimera | PMID: 15264254 | ||
| Software, algorithm | Coot | PMID: 15572765 | ||
| Software, algorithm | CryoSPARC | PMID: 28165473 | ||
| Software, algorithm | EMAN2 | PMID: 16859925 | ||
| Software, algorithm | Gautomatch | http://www.mrc-lmb.cam.ac.uk/kzhang/Gautomatch | ||
| Software, algorithm | Gctf | PMID: 26592709 | ||
| Software, algorithm | Leginon | PMID: 20817100 | ||
| Software, algorithm | Molprobity | PMID: 20057044 | ||
| Software, algorithm | MotionCor2 | PMID: 28250466 | ||
| Software, algorithm | Phenix | PMID: 20124702 | ||
| Software, algorithm | PyMOL | Schrödinger, LLC (New York, NY) | http://www.pymol.org | |
| Software, algorithm | RELION | PMID: 23000701 | ||
| Software, algorithm | Serial EM | PMID: 16182563 | ||
| Software, algorithm | Unblur | PMID: 26023829 | ||
| Other | C-flat CF-1.2/1.3 400 mesh gold grids | Electron Microscopy Sciences (Hatfield, PA) | CF413-100-Au |
Additional files
-
Supplementary file 1
Model statistics.
- https://doi.org/10.7554/eLife.34823.013
-
Supplementary file 2
Superimposition of cryo-EM structures.
- https://doi.org/10.7554/eLife.34823.014
-
Transparent reporting form
- https://doi.org/10.7554/eLife.34823.015





