iTAP, a novel iRhom interactor, controls TNF secretion by policing the stability of iRhom/TACE
Figures
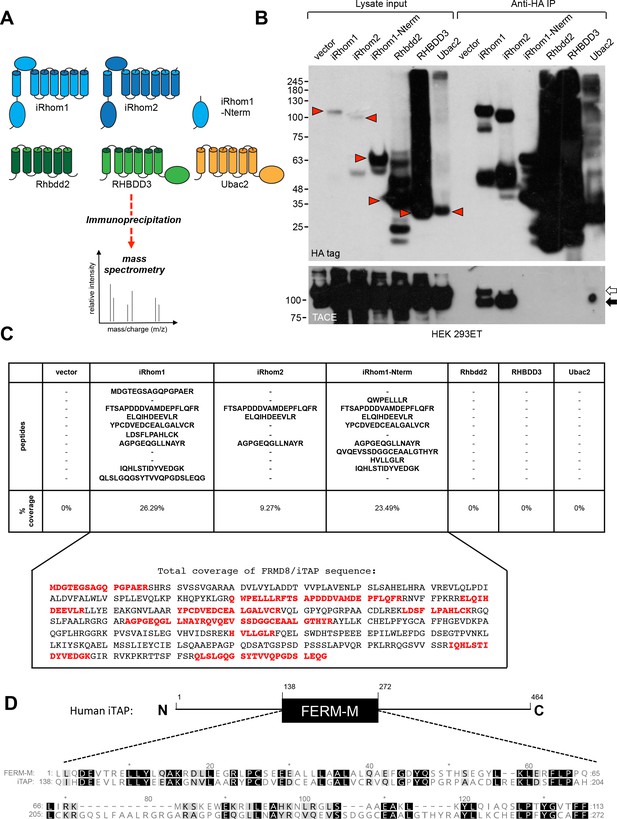
Identification of iTAP as a novel iRhom-interacting protein.
(A). Schematic diagram showing the stable HEK 293ET cell lines expressing iRhom proteins or related rhomboid pseudoproteases as controls, which were subjected to immunoprecipitation followed by mass spectrometry. (B). An example immunoprecipitation indicating that only immunoprecipitates from cell lines expressing WT iRhom1 or iRhom2 contain the binding of the positive control protein, TACE. Here and throughout, immature and mature species of TACE are indicated with white and black arrows respectively. The red arrowheads show the full-length forms of the individual rhomboid-like proteins. (C). Peptides identified by mass spec that were assigned to FRMD8/iTAP. These peptides were found in immunoprecipitates from iRhom1, iRhom2 or the N-terminus of iRhom1 but not in the other samples. The peptides are shown from a representative experiment. All of the peptides found in the iRhom immunoprecipitates were mapped onto the human FRMD8 amino acid sequence (indicated in red). (D). Schematic diagram illustrating the domain structure of human FRMD8/iTAP. Beneath is shown a CLUSTALW alignment of the pfam FERM-M consensus (pfam00373) alongside FRMD8/iTAP. Identical residues are shaded in black and similar residues in grey.
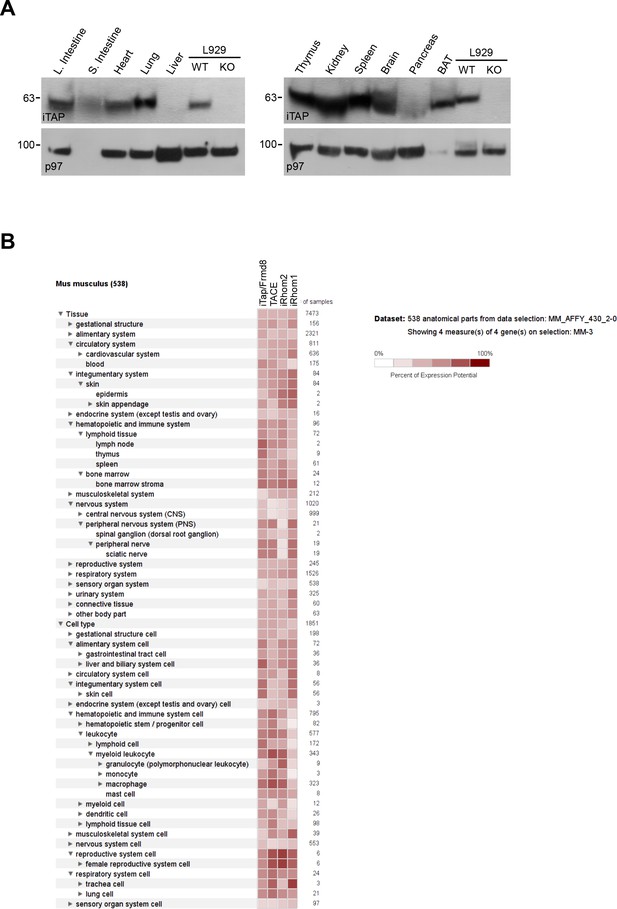
iTAP is broadly expressed in a variety of tissues important for TACE biology.
(A). Lysates were prepared from a panel of tissues from WT C57 BL/6J mice and iTAP was detected by western blot. Lysates from WT versus iTAP KO L929 mouse fibroblasts were used as a control. (B). The extent of co-expression between iTAP/Frmd8, TACE/ADAM17, iRhom2 or iRhom1 in mouse anatomical parts and cell types was investigated by interrogating gene expression data from the GeneChip Mouse Genome 430 2.0 Array via the Genevestigator platform (Hruz et al., 2008).
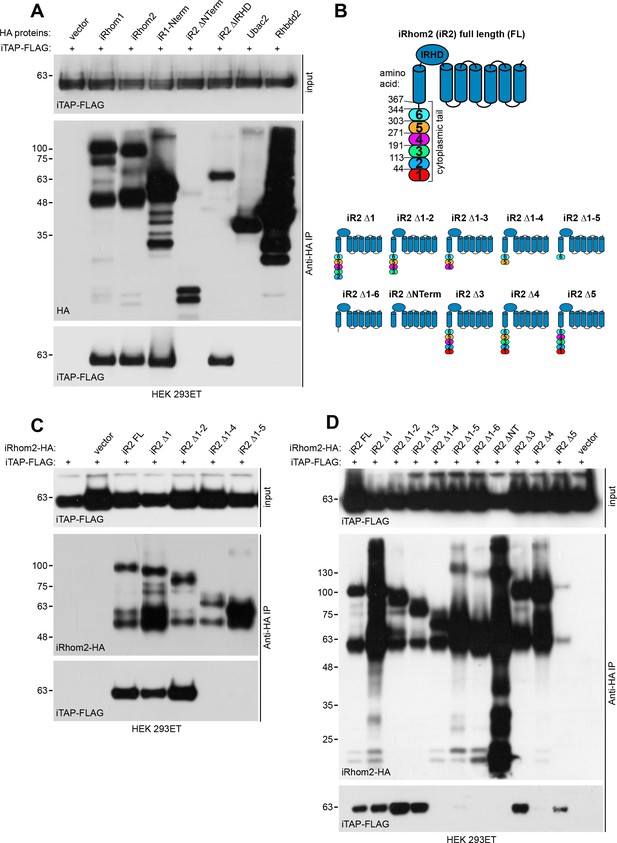
Validation of iTAP binding to iRhoms.
(A). iTAP binds specifically to iRhom −1 and −2. Human iTAP-FLAG was transfected into HEK 293ET cells alongside empty vector or the indicated HA-tagged iRhoms, their deletion mutants or the rhomboid pseudoproteases Ubac2 or Rhbdd2. HA proteins were immunoprecipitated and FLAG binding was assessed by western blot. (B). Schematic diagram indicating truncation mutants of the iRhom2 cytoplasmic tail that were generated to map the iTAP binding region. The cytoplasmic tail of iRhom2 was divided into six arbitrary portions. (C,D). iTAP binds to iRhom2 within an area defined by region 4 of iRhom2 (aa 192-271). HEK 293ET cells were transfected with iTAP-FLAG and iRhom2-HA full length (FL) or the indicated iRhom2-HA deletion constructs shown in (B). Anti-HA immunoprecipitates were assessed for the binding of iTAP-FLAG by western blotting.
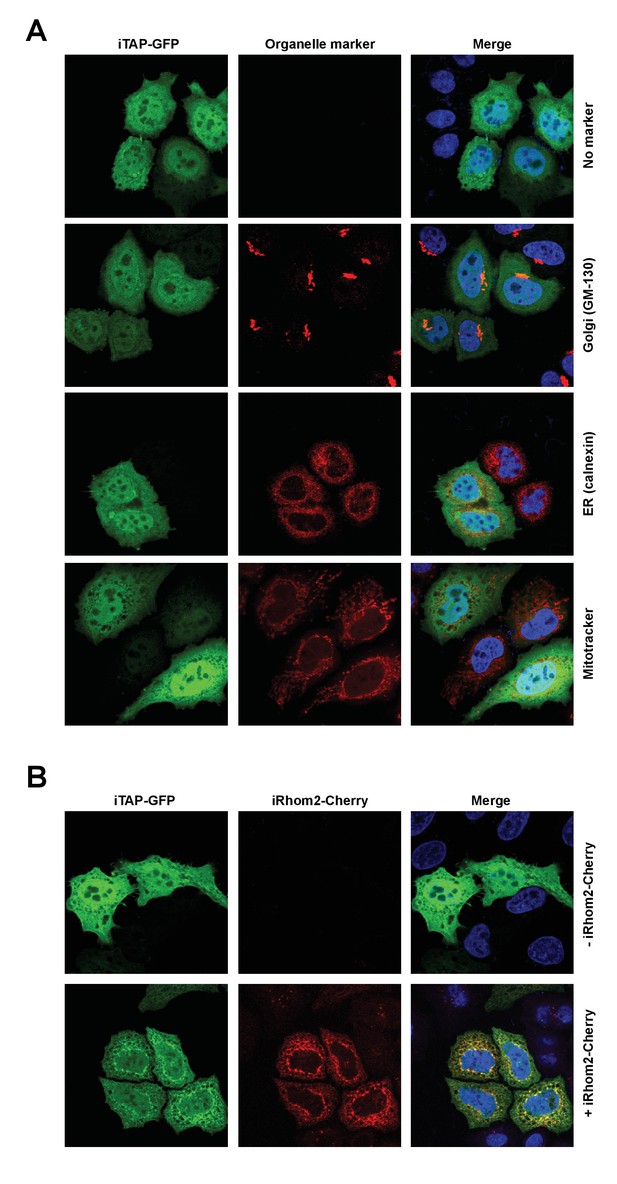
Cellular localization of iTAP.
(A). HeLa cells were transfected with iTAP-GFP. After 24 hr, cells were fixed, permeabilised and immunostained, as indicated, with specific antibodies against Golgi (GM-130) or ER (Calnexin) markers. The fluorescent dye Mitotracker-Red was used to visualise mitochondria. DNA was stained with Hoechst (blue). (B). HeLa cells were transfected with iTAP-GFP alone, or in the presence of iRhom2-Cherry, as indicated. After 24 hr, cells were fixed and stained with Hoechst to visualise DNA (blue). Images were acquired by confocal microscopy.
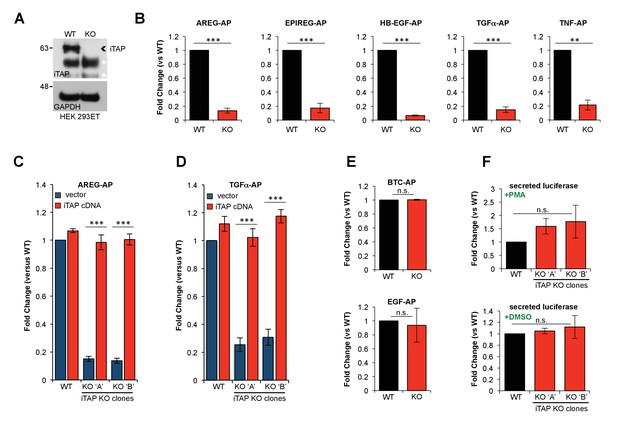
KO of iTAP diminishes TACE proteolytic activity.
(A). Anti-iTAP immunoprecipitates from WT versus iTAP KO HEK 293ET cells were analyzed by immunoblotting. A GAPDH blot is the loading control for the inputs. Non-specific bands are indicated by white asterisks. (B). PMA-stimulated TACE shedding is impaired in iTAP KO cells. The TACE substrates [Amphiregulin (AREG), Epiregulin (EPIREG), Heparin Binding-Epidermal Growth Factor (HB-EGF), Transforming Growth Factor-α (TGFα) and Tumor Necrosis Factor (TNF)] fused to alkaline phosphatase (AP) were transfected into HEK 293ET WT or iTAP KO cells. TACE activity was assessed based on AP activity secreted into the supernatant of the cells as described in materials and methods. (C,D). Expression of iTAP rescues the impaired shedding in iTAP KO cells. WT or iTAP KO HEK 293ET cells stably expressing empty vector or human iTAP were transfected with AREG-AP or TGFα-AP, then challenged in PMA shedding assays as described above. (E). The shedding impairment is specific to TACE. ADAM10 AP-fused substrates [Betacellulin (BTC) and Epidermal Growth Factor (EGF)] were transfected into the WT vs iTAP KO HEK 293ET cells. The cells were treated with the ADAM10 stimulant Ionomycin (IO) and AP activity was measured in the medium. (F). Global secretion is not impaired in iTAP KO cells. WT or iTAP KO cells were transfected with secreted luciferase and luciferase-associated luminescence was measured in the supernatant of PMA-stimulated cells (upper graph) or vehicle (DMSO, lower graph). Here and throughout: KO ‘A’ and KO ‘B’ denotes independent iTAP KO HEK 293ET clones. PMA (1 μM) or IO (2.5 μM) incubations took place for 1 hr following serum starvation. Shedding or secretion values are expressed as fold change relative to WT cells. Data are presented as mean ± standard deviation and represent at least three independent experiments. *=p ≤ 0.05, **=p ≤ 0.01, ***=p ≤ 0.001 and n.s. = non significant.
-
Figure 3—source data 1
PMA-stimulated TACE shedding is impaired in iTAP KO cells.
Alkaline phosphatase assays with TACE substrates.
- https://doi.org/10.7554/eLife.35032.011
-
Figure 3—source data 2
Expression of iTAP rescues the impaired shedding in iTAP KO cells.
Alkaline phosphatase assays with TACE substrates ± iTAP.
- https://doi.org/10.7554/eLife.35032.012
-
Figure 3—source data 3
The shedding impairment is specific to TACE.
Alkaline phosphatase assays with ADAM10 substrates.
- https://doi.org/10.7554/eLife.35032.013
-
Figure 3—source data 4
Global secretion is not impaired in iTAP KO cells.
Secreted luciferase.
- https://doi.org/10.7554/eLife.35032.014
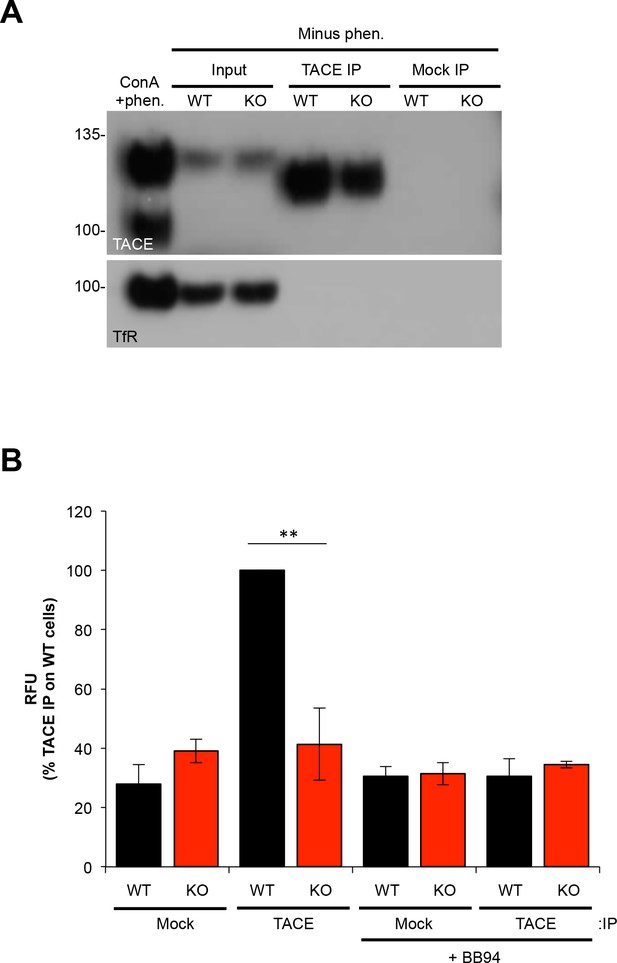
Assessment of proteolytic activity in TACE IPs from WT versus iTAP KO cells.
TACE immunoprecipitates (IPs) were used in a TACE substrate cleavage assay. (A). The specificity of the TACE antibody in immunoprecipitations is demonstrated by probing IPs with TACE and transferrin Receptor (TfR) antibodies. To avoid blocking TACE activity, the IPs and subsequent assays were performed in the absence of 1,10-phenathroline (phen.). As post-lysis TACE cleaves off the cytoplasmic tail containing the epitope recognized by the western blotting antibody, only the immature form of TACE is detected by immunoblotting (see Materials and methods for details). A ConA enriched lysate, prepared in the presence of 1,10-phenathroline, is shown on the left-hand side as a positive control for TACE detection. (B). Total TACE immunoprecipitates from WT or iTAP KO HEK 293ET cells were subjected to a TACE peptide-substrate fluorogenic cleavage assay. Data are presented as mean ± standard error of 3 independent experiments. **=p ≤ 0.01.
-
Figure 3—figure supplement 1—source data 1
iTAP ablation impairs the performance of TACE in activity assays.
- https://doi.org/10.7554/eLife.35032.010
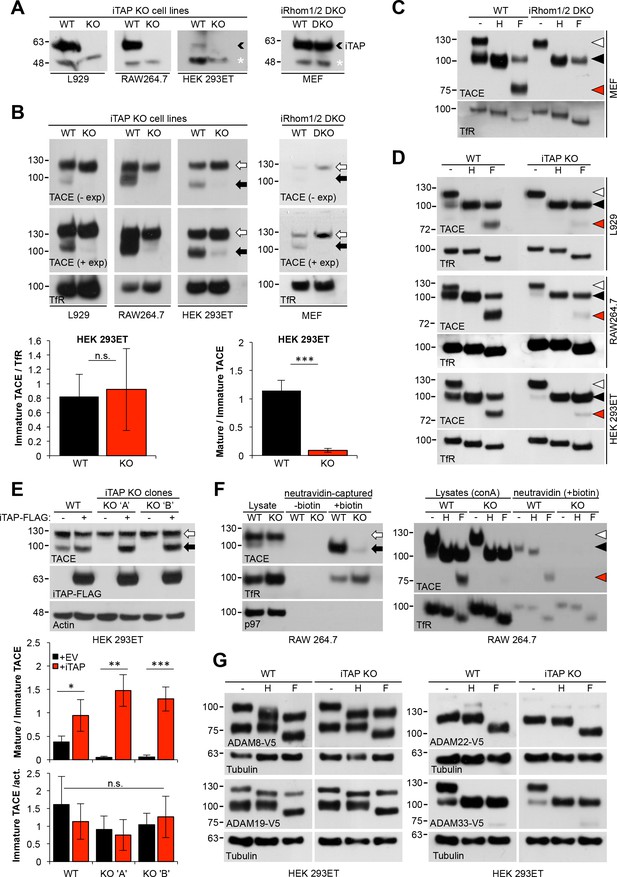
Mature TACE is specifically depleted in iTAP KO cells.
(A). iTAP was knocked out in L929, RAW 264.7 and HEK 293ET cells using CRISPR. Lysates were immunoblotted with anti-iTAP antibodies. A small black arrowhead indicates iTAP protein whereas a non-specific band (white asterisk) serves as a loading control. (B). Glycoproteins from lysates isolated from the cells in (A) were enriched using concanavalin A-sepharose (conA) and TACE levels were assessed by western blot. Here and throughout, the immature form of TACE is indicated by a white arrow, whereas, the mature form is denoted by a black arrow. iRhom double KO MEFs were used as a reference and the transferrin receptor (TfR) as a loading control. Lower panels: densitometry in HEK 293ET. Left hand panel: Levels of immature TACE normalized to TfR. Right hand panel: levels of mature TACE as a relative proportion of immature TACE in WT and iTAP KO HEK 293ET. (C,D). Validation of mature and immature TACE detection in panels of WT versus iRhom2 DKO (C) or iTAP KO (D) cells, by deglycosylation. ConA enriched lysates from the cell lines in (A) were treated with endoglycosidase H (Endo-H; H; which cleaves ER-resident glycans only) and PNGase F (F; which cleaves both ER and post-ER glycans). Here and throughout: the immature TACE is indicated with white arrowheads; the black arrowhead denotes both glycosylated mature TACE and deglycosylated immature TACE respectively (which have similar electrophoretic mobility), whereas red arrowheads denote the fully deglycosylated, mature, TACE polypeptide. (E). iTAP expression restores the presence of mature TACE in iTAP KO cells. Lysates from WT or iTAP KO HEK 293ET stably expressing empty vector (-, EV) or human iTAP (+) were screened for mature TACE. Actin was used as a loading control. Middle and lower panels: densitometric analysis indicates that iTAP expression increases the levels of mature TACE but does not affect the levels of immature TACE. Middle panel: levels of mature TACE as a relative proportion of immature TACE in WT and KO upon iTAP or EV expression in WT and iTAP KO HEK 293ET clones. Lower panel: Levels of immature TACE after normalization to actin. (F). iTAP KO cells lack mature cell surface TACE. Left hand panel: RAW 264.7 WT or iTAP KO were surface-biotinylated in vivo and lysates were enriched for biotinylated proteins with neutravidin resin. Probing for TfR was used as a cell surface positive control protein whereas anti-p97 probing demonstrates that intracellular proteins were not labeled. Right hand panel: Cell surface biotinylated proteins were deglycosylated using Endo-H (H) or PNGase F (F). ConA enriched lysates were run as mobility controls, for immature and mature TACE. Blots were probed for TACE and for TfR as a control protein. (G). Loss of iTAP has no impact on the mature species of other ADAM metalloproteases. HEK 293ET WT or KO cells were transfected with the indicated panel of V5-tagged ADAMs. The lysates were deglycosylated as described above and Tubulin serves as a loading control. Throughout: Data are presented as mean ± standard deviation and represent three independent experiments. *=p ≤ 0.05, **=p ≤ 0.01, ***=p ≤ 0.001 and n.s. = non significant.
-
Figure 4—source data 1
iTAP KO cells are depleted in mature TACE levels.
Densitometric analyses of mature/immature TACE levels.
- https://doi.org/10.7554/eLife.35032.016
-
Figure 4—source data 2
iTAP expression restores the presence of mature TACE in iTAP KO cells.
Densitometric analyses of mature/immature TACE levels.
- https://doi.org/10.7554/eLife.35032.017
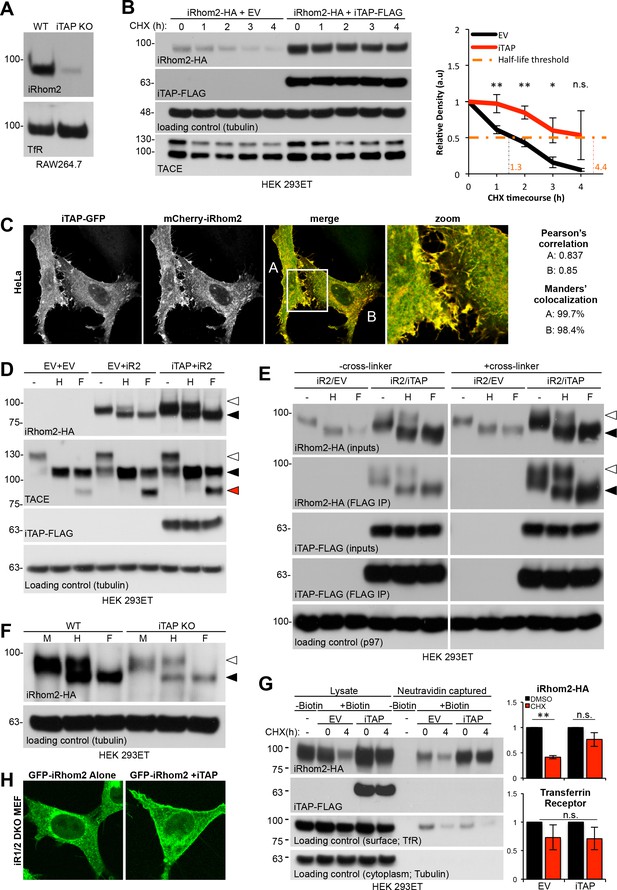
iTAP is required to promote iRhom stability at the cell surface.
(A). iRhom2 is depleted in iTAP KO cells. Lysates from WT vs iTAP KO RAW 264.7 were probed for endogenous iRhom2. The transferrin receptor (TfR) is a loading control. (B). iTAP expression enhances the stability of iRhom2. Stable iRhom2-HA-expressing HEK 293ET were transiently transfected with empty vector (EV) or iTAP-FLAG. 48 hr post-transfection, the cells were treated with 100 µg/mL Cycloheximide (CHX) for the indicated durations. The stability of iRhom2 was assessed by HA blotting. The graph to the right of the panel indicates the relative density of iRhom2-HA bands from cells expressing EV (black) versus iTAP (red). The half-life of iRhom2 under both conditions is calculated. (C). iTAP and iRhom2 co-localize. HeLa cells were transfected with iTAP-GFP and mCherry-iRhom2. Two areas, A and B, were selected for the calculation of the Pearson’s correlation and Manders’ colocalization co-efficients, respectively. (D). iTAP expression enhances the post-ER form of iRhom2. HEK 293ET expressing stably iRhom2-HA minus or plus stably expressed iTAP-FLAG were deglycosylated with Endo-H or PNGase F. (E). iRhom2-HA stably expressing HEK 293ET cells were transiently transfected with EV or iTAP-FLAG. Cells were treated ± the thiol-reducible cell-permeable crosslinker, DSP, and then anti-FLAG immunoprecipitations were performed from the lysates. Prior to SDS-PAGE and immunoblotting, lysates and co-immunoprecipitates were denatured in the presence of DTT to break the DSP-mediated covalent cross-links. Samples containing iRhom2-HA were deglycosylated as described before. (F). ER exit of iRhom2 is not impaired in iTAP KO cells. WT or iTAP KO HEK 293ET were transiently transfected with iRhom2-HA. Their lysates were deglycosylated as described. Endo-H–sensitive (black arrowhead) and –insensitive (white arrowhead) bands are noted. (G). iTAP expression stabilizes iRhom2 on the cell surface. The same cell lines as in (B), (E), were subject to a cell surface biotinylation protocol and the cell surface levels of iRhom2 in response to CHX treatment were evaluated. The graphs on the right hand side show densitometric analysis of the surface fractions of iRhom2-HA (upper graph) or TfR (lower graph) (H). iRhom1/2 DKO MEFs stably expressing mouse eGFP-iRhom2 either alone or together with mouse iTAP-mCherry were imaged as live cells. The eGFP-iRhom2 signal is shown.
-
Figure 5—source data 1
iTAP expression enhances the stability and half-life of iRhom2.
Densitometric analysis on iRhom2-HA in a CHX course.
- https://doi.org/10.7554/eLife.35032.020
-
Figure 5—source data 2
iTAP expression stabilizes iRhom2 on the cell surface.
Densitometric analysis of the cell surface fractions of iRhom2-HA or TfR.
- https://doi.org/10.7554/eLife.35032.021
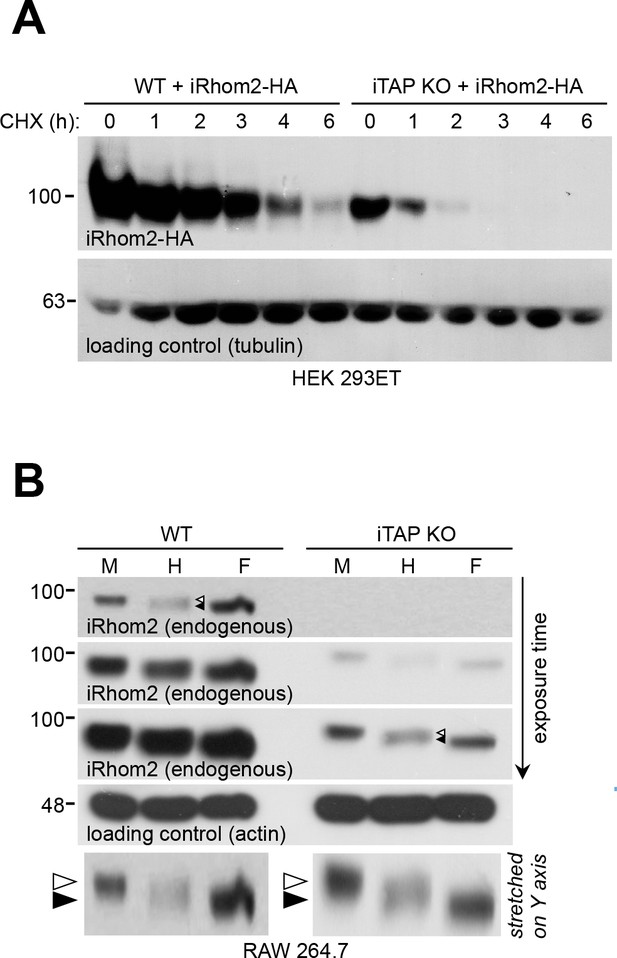
iTAP ablation increases iRhom2 degradation but doesn’t affect its ER exit.
(A). Absence of iTAP destabilizes iRhom2. WT or iTAP KO HEK 293ET cells were transiently transfected with iRhom2-HA. 48 hr post transfection, the cells were treated with 100 µg/mL CHX for the indicated durations. The stability of iRhom2 was then assessed by HA blotting. Tubulin was used as loading control. (B). Deglycosylation of endogenous iRhom2 in lysates from WT versus iTAP KO RAW 264.7 cells. The lysates were run on 4–12% gradient gels. The arrowheads in the Endo-H lanes denote a doublet containing Endo-H sensitive (upper arrowhead) and insensitive (lower arrowhead) species of iRhom2 detected in both WT and iTAP KO cells. The two panels at the bottom are cropped from the upper (WT) versus third from top (KO) iRhom2 exposures respectively. The two, cropped images were grouped together, then artificially stretched, to the same degree, along the Y axis to accentuate the difference in iRhom2 mobility in response to Endo-H versus PNGase F.
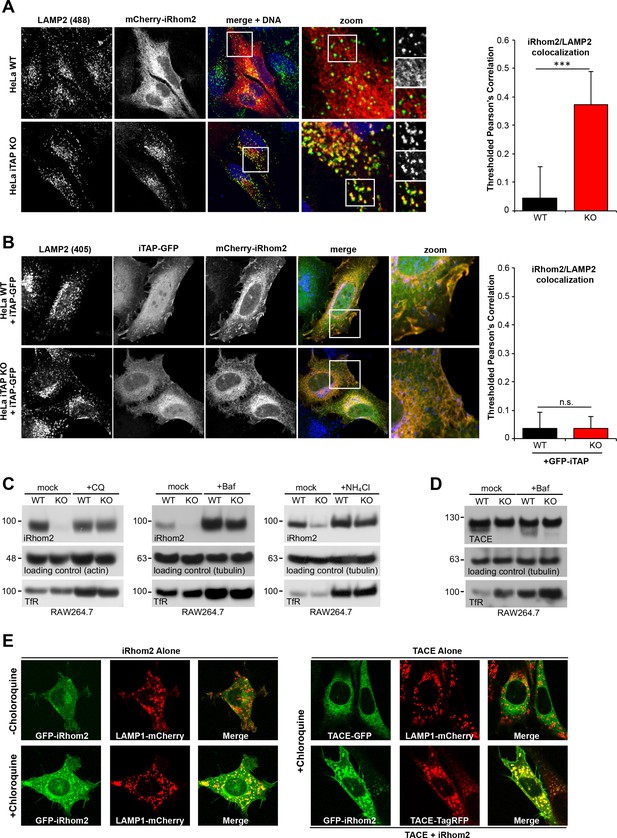
iTAP is required to prevent the trafficking of the sheddase complex to the lysosomes, where iRhom2/TACE are degraded.
(A). WT or iTAP KO HeLa cells were transfected with mCherry-iRhom2. Fixed cells were immunostained for the lysosomal marker LAMP2 and stained with DAPI. iRhom2/LAMP2 co-localization was quantified (right-hand graph). (B). The phenotype observed in (A) is reverted upon the co-expression of iTAP-GFP, resulting in no co-localization of mCherry-iRhom2 with endogenous LAMP2. (C). WT or iTAP KO RAW 264.7 cells were treated with 50 µM Chloroquine (CQ) for 48 hr, 100 µM Bafilomycin (Baf) for 16 hr, or 10 mM ammonium chloride (NH4Cl) for 48 hr and endogenous iRhom2 levels were detected by western blotting. Actin or tubulin were used as loading controls and the Transferrin Receptor (TfR) acts as a control for the inhibition of lysosomal hydrolases. (D). Lysates from cells treated with Baf as described above, were conA-enriched and probed for TACE on a western blot. Tubulin and TfR are controls for loading and lysosomal inhibition, respectively. (E). In the left-hand panel, eGFP-iRhom2 was stably expressed in WT MEFs (expressing endogenous iTAP) in the presence of lysosomal marker LAMP1-mCherry, and the subcellular localization of both proteins was imaged in live cells using confocal microscopy in the absence (upper row) or presence (lower row) of 10 µM Chloroquine. The results indicate that iRhom2 alone is trafficked into the lysosomes. In the right-hand panel, WT MEFs stably expressing TACE-GFP and LAMP1-mCherry in the presence of 10 µM Chloroquine were imaged similarly. The results indicate that TACE alone does not localize in lysosomes (upper row). However, lysosomal trafficking of TACE-TagRFP is induced by the presence of co-expressed eGFP-miRhom2 (lower row).
-
Figure 6—source data 1
Quantification of mCherry-iRhom2/LAMP2 colocalization analyses.
- https://doi.org/10.7554/eLife.35032.024
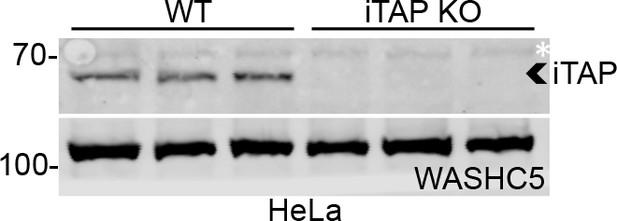
Validation of iTAP KO in HeLa cells.
Lysates (three technical replicates) from WT vs iTAP KO HeLa cells were immunoblotted with antibodies specific to iTAP (upper panel) or WASHC5 as a loading control.
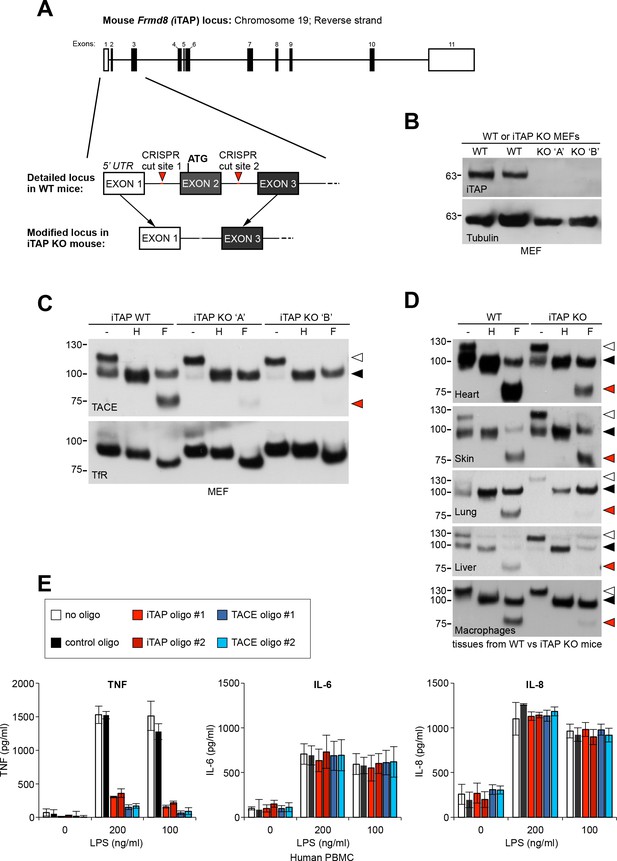
iTAP is essential for TACE maturation and function in primary cells and tissues from human and mouse.
(A). Schematic representation of the CRISPR targeting strategy to delete mouse Frmd8 (iTAP) gene using two guide RNAs flanking the first coding exon (exon 2). In the upper schematic of the Frmd8 locus, open boxes indicate non-coding exons whereas filled boxes indicate coding exons (B). Mouse embryonic fibroblasts (MEFs) were isolated from WT versus two independent iTAP KO E14.5 embryo littermates. The loss of iTAP at the protein level is shown by immunoblotting. (C). Mature TACE is diminished in iTAP KO MEFs. ConA-enriched lysates from MEFs isolated from WT versus iTAP KO embryos were deglycosylated as described previously. The transferrin receptor (TfR) is used as a loading control. (D). Mature TACE is depleted or diminished in TACE-relevant tissues from iTAP KO mice. ConA-enriched lysates from WT vs iTAP KO mouse tissues and bone marrow-derived macrophages, were deglycosylated as described previously. TACE was detected by western blot. The immature and mature species of TACE are indicated with white arrowheads and black arrowheads respectively, whereas red arrowheads denote the fully deglycosylated mature polypeptide. The experiment was performed twice with lysates isolated from tissues from two individual KO mice. (E). iTAP is essential for TACE physiological regulation in human primary cells. Isolated primary human peripheral blood mononuclear cells (PBMC) were differentiated into monocytes, then electroporated with the indicated siRNAs. Cells were then stimulated with the indicated concentrations of lipopolysaccharide (LPS). After 18 hr, the concentration of the cytokines TNF, IL-6 and IL-8 secreted into the supernatants, was measured by ELISA. The experiment was done three independent times and data from one representative experiment is shown. Data presented as mean ± standard error from triplicate measurements.
-
Figure 7—source data 1
iTAP is essential for TNF secretion in primary macrophages.
TNF ELISA.
- https://doi.org/10.7554/eLife.35032.027
-
Figure 7—source data 2
iTAP is not essential for IL-6 secretion.
IL-6 ELISA.
- https://doi.org/10.7554/eLife.35032.028
-
Figure 7—source data 3
iTAP is not essential for IL-8 secretion.
IL-8 ELISA.
- https://doi.org/10.7554/eLife.35032.029
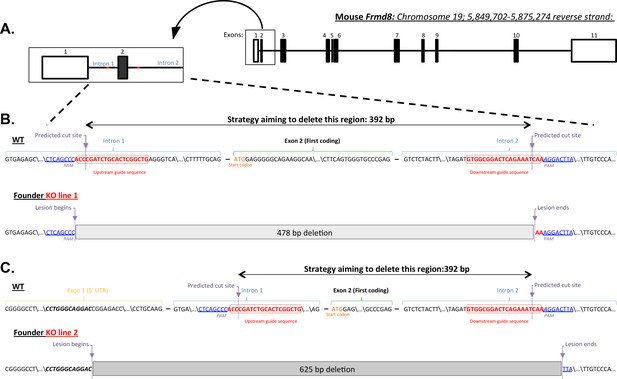
Mouse Frmd8/iTAP gene targeting via CRISPR.
(A). Schematic diagram showing the Frmd8 locus. Non-coding exons are denoted by open rectangles, whereas coding exons are shaded black. An enlarged area showing Frmd8 exons 1 and 2 is included to indicate the regions targeted by the CRISPR strategy. (B,C). Detailed maps of the WT and mutant loci of two different Frmd8 KO founder lines that were generated. A pair of gRNAs (indicated in red) was selected to engineer the deletion of the first coding exon of mouse Frmd8 as described in materials and methods. The PAM sequence and the theoretical Cas9 cut sites 3–4 bp from the PAM are indicated. The resulting founder animals were genotyped from tail biopsies and the identity of the lesions revealed by DNA sequencing. Two lines of animals (1 and 2) were established following germline transmission of the mutant alleles from individual founders. (B). KO line 1 contains a 478 bp deletion that removes exon 2 along with parts of introns 1 and 2. (C). KO line 2 contains a larger deletion of 625 bp that starts within the non-coding exon 1, deletes all of intron 1, exon 2 and part of intron 2, as shown. In both cases, the precise nucleotide sequence on the 5’ and 3’ boundaries of the mutant lines is indicated and aligned with the WT genomic sequence. The loss of iTAP at the protein level in both founder lines was confirmed by western blotting.
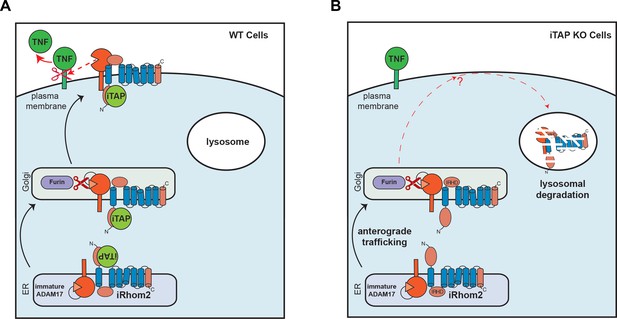
Schematic model showing regulation of the cell surface stability of the sheddase complex by iTAP.
(A). In WT cells the iRhom2/TACE sheddase complex successfully transits from the ER to the Golgi apparatus, where TACE undergoes maturation (prodomain removal). The sheddase complex then traffics to the cell surface, where TACE cleaves it substrates (e.g. TNF, EGFR ligands), enabling their release for signaling. iTAP, which loads onto the sheddase complex in the ER, remains associated with the sheddase complex and ensures the stability of the complex on the cell surface, promoting the cleavage of TACE substrates. (B). By contrast, in iTAP KO cells, the sheddase complex is aberrantly sorted to the lysosome, where iRhom2 and mature TACE are degraded. As a result, no TACE substrates are released for signaling. The dotted arrows indicate a putative itinerary taken by the sheddase complex in iTAP KO cells. The sheddase complex may be destabilized on the cell surface: aberrantly targeted for endocytosis and shunted to the lysosome. Alternatively, the sheddase complex may be endocytosed from the cell surface at the normal rate, but loss of iTAP may result in a defect in recycling the complex back to the cell surface, favouring delivery to the lysosome.
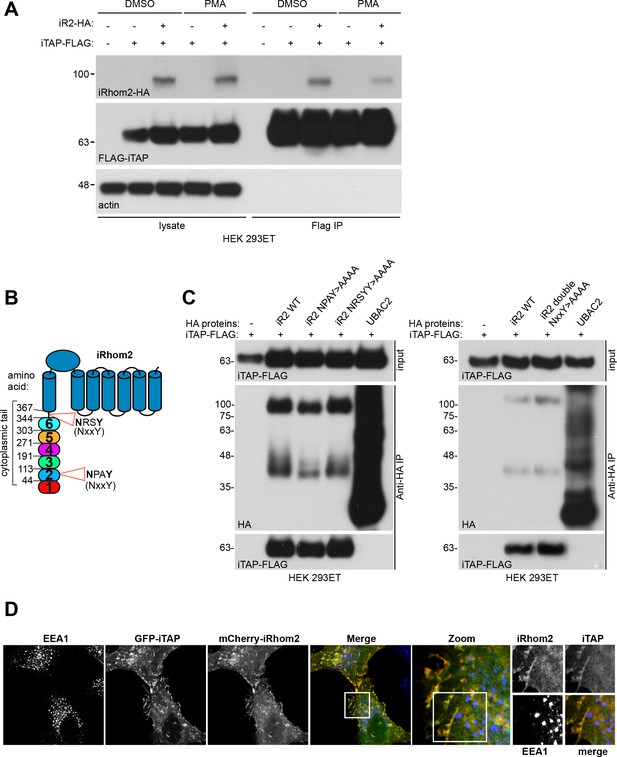
iTAP does not bind to features commonly recognized by FERM domain proteins.
(A). iTAP appears not to have a high affinity for actin. HEK 293ET cells transiently expressing vector or iRhom2-HA and iTAP-FLAG were starved overnight then stimulated with or without PMA (1 µM) for 30 min. PMA was shown to alter the steady state of iRhom interactions with clients (Cavadas et al., 2017) and hence is used to assess potentially differential binding upon stimulated conditions. An anti-FLAG IP was performed on the cell lysates and binding was assessed by western blotting (B). Schematic representation of the location of the NxxY motifs within the iRhom2 sequence. NxxY is a common consensus binding motif for FERM proteins (e.g. sorting nexins) that participate in vesicular sorting functions. (C). iTAP binding is independent of NxxY motifs. HEK 293ET cells were transiently transfected with iTAP-FLAG and vector, iRhom2-HA, single (left hand panel) or double (right hand panel) NxxY >AAAA mutants of iRhom2-HA, or Ubac2-HA as a negative control. Cell lysates were subjected to anti-HA co-precipitation. iTAP binding to iRhom was assessed by western blotting. (D). iTAP does not co-localize with early endosomes. HeLa cells co-expressing iTAP-GFP and mCherry-iRhom2 were immunostained for EEA1, a marker of early endosomes. The white box indicates a magnified area that is separated into all three channels in the right hand images.
Tables
iTAP interacts with endogenous iRhoms.
Lysates from HEK 293ET cells expressing iTAP-FLAG versus cells containing empty vector or expressing a panel of control proteins (TNF-FLAG, STING-FLAG, SREBP2-FLAG) were immunoprecipitated for FLAG and subjected to mass spectrometry. Peptides assigned to iRhom1 or iRhom2 are found specifically in iTAP precipitates.
| iRhom1 | Vector | iTAP | TNF | STING | SREBP2 |
|---|---|---|---|---|---|
| Peptide counts | 0 | 27 | 0 | 0 | 0 |
| Sequence coverage [%] | 0 | 37.7 | 0 | 0 | 0 |
| iRhom2 | Vector | iTAP | TNF | STING | SREBP2 |
|---|---|---|---|---|---|
| Peptide counts | 3 | 44 | 2 | 2 | 2 |
| Sequence coverage [%] | 2.9 | 45.4 | 1.8 | 2.2 | 2 |
Mendelian ratios of embryos isolated at embryonic day 14.5 (E14.5), or pups at (P1) post-partum obtained from crosses obtained between iTAP heterozygous mice.
https://doi.org/10.7554/eLife.35032.032| Developmental stage | iTAP +/+ | iTAP +/- | iTAP -/- | Total no. of animals |
|---|---|---|---|---|
| E14.5 % Ratio | 10 32.2 1 | 17 54.8 1,7 | 4 12.9 0,4 | 31 |
| P1 | 54 | 116 | 50 | 220 |
| % Ratio | 24.55 1 | 52.73 2.14 | 22.73 0.93 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Homo sapiens) | FRMD8; iTAP, human | n/a | ENSG00000126391 | |
| Gene (Mus musculus) | Frmd8; iTAP, mouse | n/a | ENSMUSG00000024816 | |
| Strain, strain background (Mm; C57BL/6) | wild type; WT mice | Jackson Laboratories | RRID:IMSR_JAX:000664 | |
| Genetic reagent (Mm,C57BL/6) | iTAP KO mice | this paper | n/a | Generated by CRISPR (more details in M and M section and Figure 7—figure supplement 1) |
| Cell line (Hs) | HEK 293ET | 10.1038/sj.emboj.7601743 | RRID:CVCL_6996 | |
| Cell line (Hs) | HeLa | ATCC | ATCC CCL-2; RRID:CVCL_0030 | |
| Cell line (Mm) | RAW 264.7 | Sigma | 91062702; RRID:CVCL_0493 | |
| Cell line (Mm) | L929 | ATCC | ATCC CCL-1; RRID:CVCL_0462 | |
| Cell line (Hs) | HEK 293ET iTAP KO | this paper | n/a | Generated by CRISPR (more details in M and M section) |
| Cell line (Mm) | RAW 264.7 iTAP KO | this paper | n/a | Generated by CRISPR (more details in M and M section) |
| Cell line (Mm) | L929 iTAP KO | this paper | n/a | Generated by CRISPR (more details in M and M section) |
| Cell line (Mm) | MEF iTAP KO | this paper | n/a | Isolated from iTAP KO mouse embryos |
| Cell line (Mm) | DKO MEF | 10.1038/embor.2013.128. | n/a | Isolated from iRhom1/2 DKO mouse embryos |
| Biological sample (Mm) | tissues from WT and KO mice | other | n/a | Isolated with standard techniques from the mice described here |
| Antibody | anti-TACE; TACE Ab 318 | https://doi.org/10.1016 /j.jim.2011.06.015 | n/a | |
| Antibody | anti-TACE for Immunoprecipitation | R and D systems | R and D 9301 | |
| Antibody | anti-TACE | abcam | Ab39162; RRID:AB_722565 | Ab39163 |
| Antibody | anti-P97 | Thermo-Fisher | MA1-21412; RRID:AB_557663 | |
| Antibody | anti-TACE | R and D systems | 9301; RRID:AB_2223551 | |
| Antibody | anti-TACE | 10.1016/j.jim.2011.06.015 | Ab318 | monoclonal anti-TACE antibody (Trad et al., 2011) was used for the detection of deglycosylated TACE when no conA was used |
| Antibody | anti-tubulin | IGC antibody facility | Clone YL1/2; RRID:AB_793541 | |
| Antibody | anti-HA-HRP | Roche | 3F10; RRID:AB_2314622 | |
| Antibody | anti-HA | Biolegend | 901501; RRID:AB_291259 | |
| Antibody | anti-V5-HRP | Life Technologies | R961-25 | |
| Antibody | anti-Frmd8 | Abnova | 157H00083786-B01P; RRID:AB_1573641 | |
| Antibody | anti-Transferrin Receptor | Life Technologies | 13–6800; RRID:AB_2533029 | |
| Antibody | anti-Flag-HRP | Sigma | A8592; RRID:AB_439702 | |
| Antibody | anti-GAPDH | Cell signalling Technology | 2118; RRID:AB_561053 | |
| Antibody | anti-actin | Abcam | Ab8227; RRID:AB_2305186 | |
| Antibody | anti-iRhom2 | 10.1126/science.1214400 | n/a | Anti-iRhom2 polyclonal antibodies specific to the mouse iRhom2 N- terminus (amino acids 1–373) or raised against the iRhom homology domain, were previously described (Adrain et al., 2012) |
| Antibody | Anti-HA magnetic beads | Pierce | 88836, Pierce | |
| Antibody | Mouse anti-GFP | IGC antibody Facility | clone 19F7 | Used as IP negative control in Figure 3—figure supplement 1 |
| Antibody | Anti-Flag M2 affinity gel | Sigma | A220 | |
| Antibody | MagnaBind Goat Anti- Rabbit IgG Beads | Thermo Scientific | 21356 | |
| Antibody | MagnaBind Goat Anti- Mouse IgG Beads | Thermo Scientific | 21354 | |
| Antibody | mouse anti-LAMP2 | DSHB | Clone H4B4; RRID:AB_2134755 | |
| Antibody | rabbit anti-EEA1 | Cell signalling Technology | 3288; RRID:AB_2096811 | |
| Antibody | rabbit anti-Strumpellin/ WASHC5 | Santa Cruz | SC87442; RRID:AB_2234159 | |
| Transfected construct (Hs) | h iRhom1-HA (plasmid) | this paper | n/a | |
| Transfected construct (Hs) | h iRhom1 Nterm HA (plasmid) | this paper | n/a | |
| Transfected construct (Mm) | Rhbdd2-HA (plasmid) | this paper | n/a | |
| Transfected construct (Hs) | RHBDD3-HA (plasmid) | this paper | n/a | |
| Transfected construct (Mm) | Ubac2-HA (plasmid) | this paper | n/a | |
| Recombinant DNA reagent | pM6P.blast-GFP (plasmid) | Felix Randow | n/a | |
| Recombinant DNA reagent | pLEX.blast (plasmid) | this paper | n/a | Details in M and M |
| Transfected construct (Mm) | iRhom2-Cherry (plasmid) | doi:10.1038/ni.3510 | n/a | |
| Transfected construct (Hs) | ADAM33-V5 (plasmid) | 10.1038/embor.2013.128. | n/a | |
| Transfected construct (Hs) | ADAM19-V5 (plasmid) | 10.1038/embor.2013.128. | n/a | |
| Transfected construct (Hs) | ADAM22-V5 (plasmid) | 10.1038/embor.2013.128. | n/a | |
| Transfected construct (Hs) | ADAM8-V5 (plasmid) | 10.1038/embor.2013.128. | n/a | |
| Transfected construct | TNF-FLAG (plasmid) | this paper | n/a | |
| Transfected construct (Hs) | STING-FLAG (plasmid) | doi:10.1038/ni.3510 | n/a | |
| Transfected construct (Hs) | LAMP1-mCherry (plasmid) | this paper | n/a | Details in M and M |
| Transfected construct (Mm) | GFP-iRhom2 (plasmid) | this paper | n/a | Details in M and M |
| Transfected construct (Mm) | mCherry-iRhom2 (plasmid) | this paper | n/a | Details in M and M |
| Transfected construct (Mm) | TACE-GFP (plasmid) | this paper | n/a | Details in M and M |
| Transfected construct (Mm) | TACE-TagRFP (plasmid) | this paper | n/a | Details in M and M |
| Transfected construct (Mm) | mouse iTAP-mCherry (plasmid) | this paper | n/a | Details in M and M |
| Transfected construct (Hs) | human iTAP-GFP (plasmid) | this paper | n/a | Details in M and M |
| Transfected construct (Mm) | iRhom2 NPAY > AAAA (plasmid) | this paper | n/a | Quick change-based mutagenesis on iRhom2-HA plasmid described in M and M |
| Transfected construct (Mm) | iRhom2 NRSY > AAAA (plasmid) | this paper | n/a | Quick change-based mutagenesis on iRhom2-HA plasmid described in M and M |
| Transfected construct (Mm) | iRhom2 Double NxxY > AAAA (plasmid) | this paper | n/a | Quick change-based mutagenesis on iRhom2-HA plasmid described in M and M |
| Transfected construct (Hs) | SREBP2-FLAG (plasmid) | DOI: 10.1083/jcb.201305076 | n/a | |
| Recombinant DNA reagent | EGF-AP; EGF (plasmid) | doi: 10.1083/jcb.200307137 | n/a | |
| Recombinant DNA reagent | Betacellulin-AP; BTC-AP; BTC (plasmid) | doi: 10.1083/jcb.200307137 | n/a | |
| Recombinant DNA reagent | TNF-AP; TNF (plasmid) | doi: 10.1083/jcb.200307137 | n/a | |
| Recombinant DNA reagent | TGFα-AP; TGFα (plasmid) | doi: 10.1083/jcb.200307137 | n/a | |
| Recombinant DNA reagent | HB-EGF-AP; HB-EGF (plasmid) | doi: 10.1083/jcb.200307137 | n/a | |
| Recombinant DNA reagent | Epiregulin-AP; EPIREG-AP; EPIREG (plasmid) | doi: 10.1083/jcb.200307137 | n/a | |
| Recombinant DNA reagent | Amphiregulin-AP; AREG-AP; AREG (plasmid) | doi: 10.1083/jcb.200307137 | n/a | |
| Recombinant DNA reagent | secreted luciferase | 10.1038/embor.2013.128. | n/a | |
| Recombinant DNA reagent | pLenti-Crispr version2 (plasmid) | Addgene | 52961 | |
| Recombinant DNA reagent | pLentiCas9-Blast(plasmid) | Addgene | 52962 | |
| Recombinant DNA reagent | pLentiGuide-Puro(plasmid) | Addgene | 52963 | |
| Recombinant DNA reagent | pgRNAbasic(plasmid) | doi: 10.1242/dev.133074 | n/a | |
| Recombinant DNA reagent | pT7-Cas9(plasmid) | doi: 10.1242/dev.133074 | n/a | |
| Sequence-based reagent (siRNA; Hs) | iTAP oligo #1 | Santa Cruz | sc-96500 | |
| Sequence-based reagent (siRNA; Hs) | iTAP oligo #2 | GE Dharmacon | M-018955-01-0005 | |
| Sequence-based reagent (siRNA; Hs) | TACE oligo #1 | Santa Cruz | sc-36604 | |
| Sequence-based reagent (siRNA; Hs) | TACE oligo #2 | GE Dharmacon | M-003453-01-0005 | |
| Sequence-based reagent (CRISPR Guide sequence; Hs; cell lines) | gRNA targeting exon 1 | this paper | n/a | 5’-GCCCCGCTGAGCGATCCCAC-3’ |
| Sequence-based reagent (CRISPR Guide sequence; Hs; cell lines) | gRNA targeting exon 4 | this paper | n/a | 5’-ACGTGTTCTTCCCAAAGCGG-3’ |
| Sequence-based reagent (CRISPR Guide sequence; Hs; cell lines) | gRNA targeting exon 2 | this paper | n/a | 5'-TGACGTGCTGGTATACCTAG-3' |
| Sequence-based reagent (CRISPR Guide sequence; Hs; cell lines) | gRNA targeting exon 6 | this paper | n/a | 5’-GGCACTTGAGGAGATAGGCG-3’ |
| Sequence-based reagent (CRISPR Guide sequence; Mm; cell lines) | gRNA targeting first exon | this paper | n/a | 5’-TTCGGTGGGACCGCTCCGCA-3’ |
| Sequence-based reagent (CRISPR Guide sequence; Mm; cell lines) | gRNA targeting second exon | this paper | n/a | 5’-GCACTACTGTATCATCCGCC-3’ |
| Commercial assay or kit | fluorogenic TACE substrate peptide | ENZO Life Sciences | BML-P235-0001 | |
| Commercial assay or kit | 1-step PNPP Substrate | Thermo Fisher | PIER37621 | |
| Commercial assay or kit | Fugene 6 | Promega | 2691-SC | |
| Commercial assay or kit | Endoglycosidase; Endo-Hf | NEB | 174P0703 | |
| Commercial assay or kit | PNGase F | NEB | 174P0704 | |
| Commercial assay or kit | Concanavalin A Agarose; conA | G-biosciences | 786–216 | |
| Commercial assay or kit | MEGAshortscript T7 Kit | Thermo Fisher | AM1354 | |
| Commercial assay or kit | MEGAclear kit | Thermo Fisher | AM1908 | |
| Commercial assay or kit | Gibson Assembly Master Mix | New England Biolabs | 174E2611 | |
| Commercial assay or kit | KOD Hotstart DNA polymerase | Novagen | 71086–5 | |
| Commercial assay or kit | TOPO TA cloning kit for sequencing | Invitrogen | 450030 | |
| Commercial assay or kit | mMESSAGE mMACHINE T7 Ultra Kit | Thermo Fisher | AM1345 | |
| Commercial assay or kit | human TNF ELISA kit | R and D Systems | DY210 | |
| Commercial assay or kit | human IL-6 ELISA kit | R and D systems | DY-206 | |
| Commercial assay or kit | human IL-8 ELISA kit | R and D systems | DY-208 | |
| Chemical compound, drug | Polyethylenimine; PEI | Sigma | 408727 | |
| Chemical compound, drug | 1,10-phenanthroline | Sigma | 131377 | |
| Chemical compound, drug | Bafilomycin A1 | Santa cruz | 201550 | |
| Chemical compound, drug | Chloroquine | Sigma | C6628 | |
| Chemical compound, drug | Ammonium chloride | Acros | 10676052 | |
| Chemical compound, drug | Ionomycin | Cayman | CAYM11932-5 | |
| Chemical compound, drug, (E.coli 055:B5) | Lipopolysaccharide; LPS | Santa cruz | (sc-221855A) | |
| Chemical compound, drug | Phorbol 12-myristate 13-acetate; PMA | Sigma | P1585 | |
| Chemical compound, drug | Cycloheximide; CHX | Santa cruz | sc-3508 | |
| Chemical compound, drug | Dithiobis (succinimidyl propionate); DSP | Pierce | 10731945 | |
| Chemical compound, drug | Sulfo-NHS-LC-Biotin | Thermo Scientific, | 21335 | |
| Other | Lenti-X Concentrator | Clontech | Clonetech: 631231 | |
| Other | NeutrAvidin agarose; Neutravidin resin | Thermo Fisher | 11885835 | |
| Other | NuPAGE Novex 4–12% Bit- Tris Protein Gels 1.0 mm | Novex; Life Technologies | NP0322BOX | |
| Other | Opti-MEM I Reduced Serum Medium, GlutaMAX | Life Technologies | 51985–026 | |
| Software, algorithm | Fiji | PMID: 22743772 | RRID:SCR_002285 | |
| Software, algorithm | Illustrator | Adobe | Adobe Creative Suite | |
| Software, algorithm | GraphPad Prism | GraphPad Software, Inc. | ||
| Software, algorithm | Volocity | PerkinElmer | ||
| Software, algorithm | Geneious | Biomatters Ltd. | Javaversion 1.8. 0_71-b15 | |
| Software, algorithm | perkin elmer 2030 workstation | PerkinElmer |





