Drosophila mushroom bodies integrate hunger and satiety signals to control innate food-seeking behavior
Figures
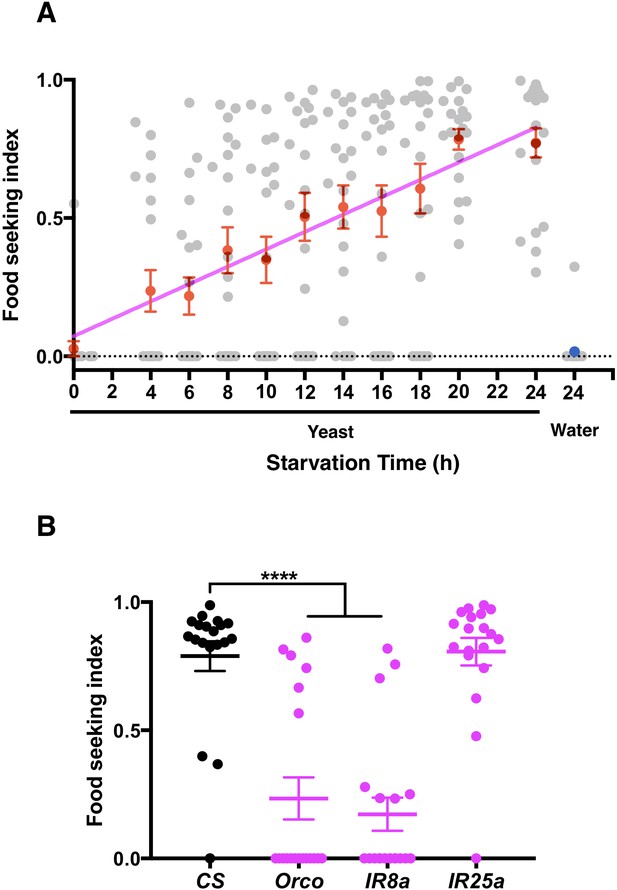
Starvation promotes yeast food-seeking behavior.
(A) The average yeast food-seeking performance (y-axis; see Materials and methods for mode of calculation) increases linearly with the duration of starvation (x-axis). A water-only control for flies starved for 24 hr is also shown. Individual data points and mean ± SEM (n = 20 for each point) are shown. (B) The yeast food-seeking performance of wild-type flies (CS) and flies homozygous for orco2, IR8a1, and IR25a2. The performances of the orco2 and IR8a1 flies were significantly lower than that of the control flies (Kruskal-Wallis, n = 19, p<0.0001). Individual data points and mean ± SEM are shown.
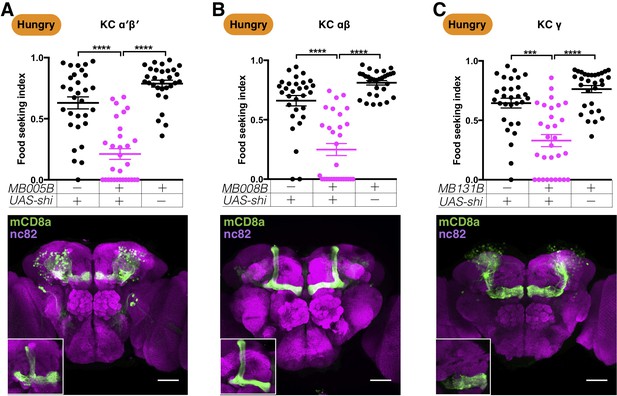
Kenyon cells are required for yeast food-seeking behavior in hungry flies.
(A–C) Male flies starved for 24 hr were assessed for their yeast food-seeking performance. The performance of GAL4;UAS-shits1 flies was statistically different from the controls for (A) MB005B split-GAL4 (α′β′ KCs; Kruskal-Wallis, n = 30, p<0.0001), (B) MB008B split-GAL4 (αβ KCs; Kruskal-Wallis, n = 30, p<0.0001), and (C) MB131B split-GAL4 (γ KCs; Kruskal-Wallis, n = 30, p=0.0003). Individual data points and mean ± SEM are shown. The brain images are full z-projections of confocal stacks showing the expression patterns of the GAL4 lines (green) counter-stained with nc82 antibody (magenta). Insets are z-projections of the MB lobes. Scale bars are 100 µm.
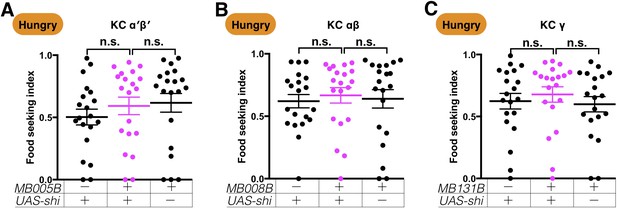
Expression of UAS-shits1 in the KCs does not affect yeast food-seeking behavior in hungry flies at the permissive temperature.
(A) No statistical difference was detected between flies expressing UAS-shits1 in α′β′ KCs and relevant controls (MB005B, Kruskal-Wallis, n = 20, p=0.7165). (B) No statistical difference was detected between flies expressing UAS-shits1 in αβ KCs and relevant controls (MB008B, Kruskal-Wallis, n = 20, p=0.7077). (C) No statistical difference was detected between flies expressing UAS-shits1 in γ KCs and relevant controls (MB131B, Kruskal-Wallis, n = 20, p=0.4803). Individual data points and mean ± SEM are shown.
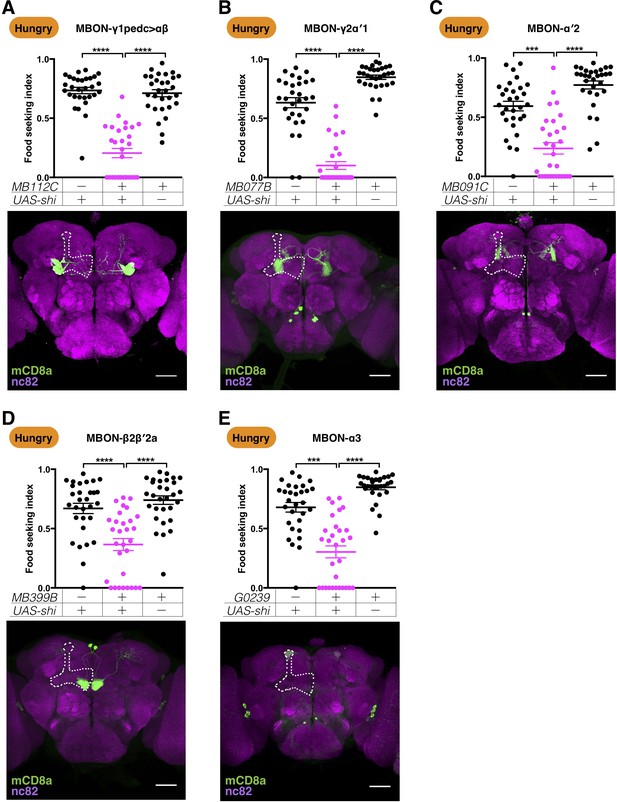
Five MBONs are required for yeast food-seeking behavior in hungry flies.
Male flies starved for 24 hr were assessed for the yeast food-seeking performance. The performance of GAL4;UAS-shits1 flies was statistically lower than the controls for (A) MB112C split-GAL4 (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p<0.0001), (B) MB077B split-GAL4 (MBON-γ2α′1, Kruskal-Wallis, n = 30, p<0.0001), (C) MB091C split-GAL4 (MBON-α′2, Kruskal-Wallis, n = 30, p=0.0004), (D) MB399B split-GAL4 (MBON-β2β′2a, Kruskal-Wallis, n = 30, p<0.0001), and (E) G0239-GAL4 (MBON-α3, Kruskal-Wallis, n = 30, p=0.0002). Individual data points and mean ± SEM are shown. The brain images are full z-projections of confocal stacks showing the expression patterns of the GAL4 lines (green) counter-stained with nc82 antibody (magenta). One side of the MB is outlined by a white dashed line. Scale bars are 100 µm.
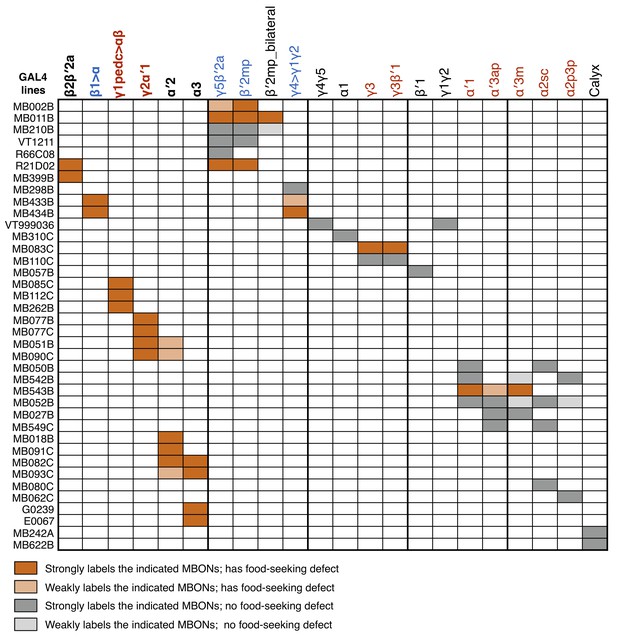
Effects on yeast food-seeking behavior in hungry flies when GAL4 lines labeling different MBONs are used to drive UAS-shits1 expression.
The MBONs labeled by each GAL4 line are indicated. The MBONs potentially involved in hunger-driven food seeking behavior are shown in bold font (the first six MBONs from the left). MBONs with positive or negative valences are in red or blue fonts, respectively (Aso et al., 2014b). All the GAL4;UAS-shits1 flies that showed impaired food seeking at 32°C performed normally at a permissive 23°C.
-
Figure 3—figure supplement 1—source data 1
Source file for the table in Figure 3—figure supplement 1.
- https://doi.org/10.7554/eLife.35264.007
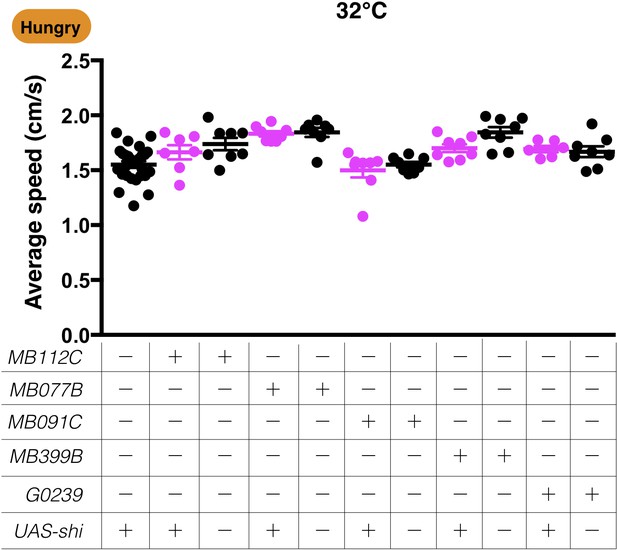
Blocking the MBONs required for yeast food-seeking behavior does not affect the locomotion of flies.
The average speed for GAL4;UAS-shits1 flies at a restrictive 32°C is shown. No significant difference was detected (Kruskal-Wallis, 6 ≤ n ≤ 28, p>0.9999). Individual data points and mean ± SEM are shown.
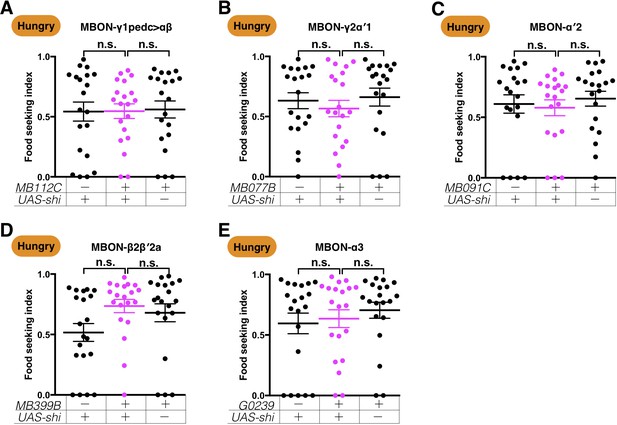
Expression of UAS-shits1 in the five MBONs does not affect yeast food-seeking behavior in hungry flies at the permissive temperature.
No statistical difference was detected between relevant controls and flies expressing UAS-shits1 in (A) MBON-γ1pedc>αβ (MB112C, Kruskal-Wallis, n = 20, p=0.8705), (B) MBON-γ2α′1 (MB077B, Kruskal-Wallis, n = 20, p=0.6004), (C) MBON-α′2 (MB091C, Kruskal-Wallis, n = 20, p=0.5172), (D) MBON-β2β′2a (MB399B, Kruskal-Wallis, n = 20, p=0.0762), and (E) MBON-α3 (G0239, Kruskal-Wallis, n = 20, p=0.7276). Individual data points and mean ± SEM are shown.
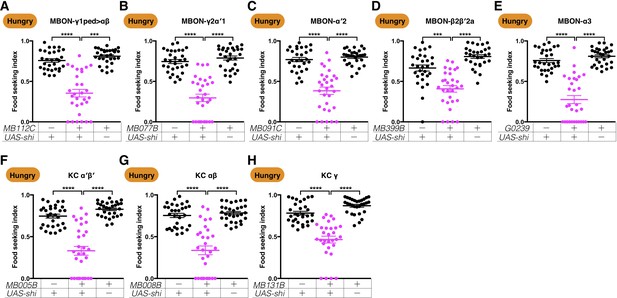
The five MBONs and the KCs are required during the seeking phase in our food-seeking assay.
Male flies starved for 24 hr were tested for their yeast food-seeking performance. In these experiments, a fly was considered as finding the food target whenever it touched the yeast drop. The performance of GAL4;UAS-shits1 flies was significantly lower than the controls for (A) MB112C split-GAL4 (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p<0.0001), (B) MB077B split-GAL4 (MBON-γ2α′1, Kruskal-Wallis, n = 30, p<0.0001), (C) MB091C split-GAL4 (MBON-α′2, Kruskal-Wallis, n = 29–30, p<0.0001), (D) MB399B split-GAL4 (MBON-β2β′2a, Kruskal-Wallis, n = 30, p=0.0003), (E) G0239-GAL4 (MBON-α3, Kruskal-Wallis, n = 30, p<0.0001), (F) MB005B split-GAL4 (α′β′ KCs; Kruskal-Wallis, n = 30, p<0.0001), (G) MB008B split-GAL4 (αβ KCs; Kruskal-Wallis, n = 30, p<0.0001), and (H) MB131B split-GAL4 (γ KCs; Kruskal-Wallis, n = 30, p<0.0001). Individual data points and mean ± SEM are shown.
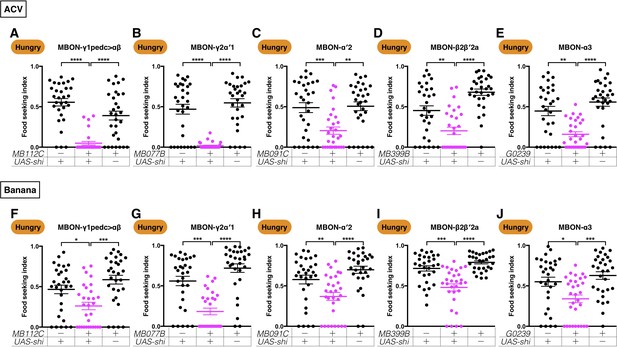
Five MBONs are required for hungry flies to seek ACV and banana odors.
Male flies starved for 24 hr were assessed for their performance in seeking ACV (A–E) or banana odor (F–J) at a restrictive 32°C. The performance of GAL4;UAS-shits1 flies was significantly lower than the controls for (A) MB112C split-GAL4 (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p<0.0001), (B) MB077B split-GAL4 (MBON-γ2α′1, Kruskal-Wallis, n = 30, p<0.0001), (C) MB091C split-GAL4 (MBON-α′2, Kruskal-Wallis, n = 30, p=0.0025), (D) MB399B split-GAL4 (MBON-β2β′2a, Kruskal-Wallis, n = 30, p=0.009), (E) G0239-GAL4 (MBON-α3, Kruskal-Wallis, n = 30, p=0.0016), (F) MB112C split-GAL4 (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p=0.0246), (G) MB077B split-GAL4 (MBON-γ2α′1, Kruskal-Wallis, n = 30, p=0.0004), (H) MB091C split-GAL4 (MBON-α′2, Kruskal-Wallis, n = 30, p=0.0091), (I) MB399B split-GAL4 (MBON-β2β′2a, Kruskal-Wallis, n = 30, p=0.0002), and (J) G0239-GAL4 (MBON-α3, Kruskal-Wallis, n = 30, p=0.0101). Individual data points and mean ± SEM are shown.
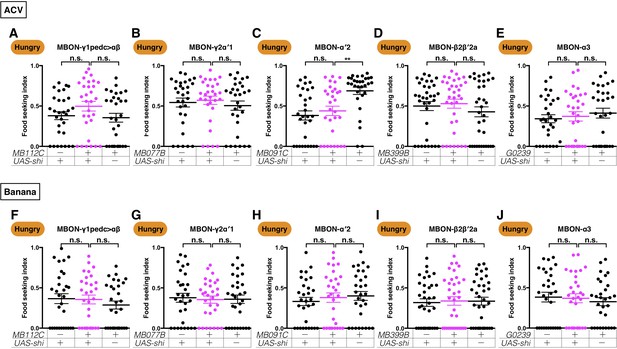
Expression of UAS-shits1 in the five MBONs does not affect ACV and banana odor-seeking behavior in hungry flies at the permissive temperature.
Male flies starved for 24 hr were assessed for their performance in seeking ACV (A–E) or banana odor (F–J) at a permissive 23°C. The performance of GAL4;UAS-shits1 flies was not different from the controls for (A) MB112C split-GAL4 (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p=0.122), (B) MB077B split-GAL4 (MBON-γ2α′1, Kruskal-Wallis, n = 30, p=0.7937), (C) MB091C split-GAL4 (MBON-α′2, Kruskal-Wallis, n = 30, p>0.9999), (D) MB399B split-GAL4 (MBON-β2β′2a, Kruskal-Wallis, n = 30, p=0.6095), (E) G0239-GAL4 (MBON-α3, Kruskal-Wallis, n = 30, p=0.6322), (F) MB112C split-GAL4 (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p=0.6241), (G) MB077B split-GAL4 (MBON-γ2α′1, Kruskal-Wallis, n = 30, p=0.9690), (H) MB091C split-GAL4 (MBON-α′2, Kruskal-Wallis, n = 30, p=0.5855), (I) MB399B split-GAL4 (MBON-β2β′2a, Kruskal-Wallis, n = 30, p=0.9329), and (J) G0239-GAL4 (MBON-α3, Kruskal-Wallis, n = 30, p=0.6649). Individual data points and mean ± SEM are shown.
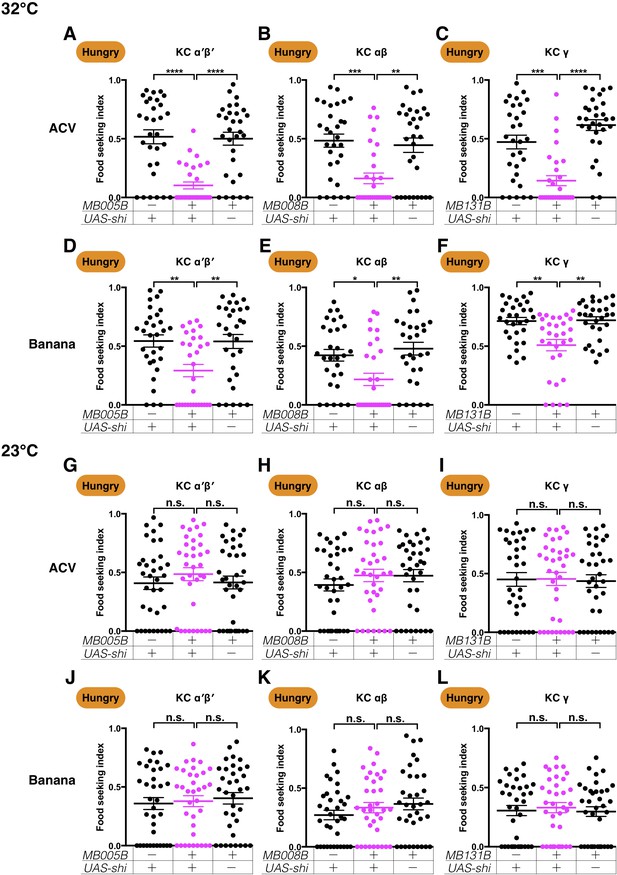
KCs are required for hungry flies to seek ACV and banana odors.
Male flies starved for 24 hr were assessed for their performance in seeking ACV (A–C and G–I) or banana odor (D–F and J–L) at a restrictive 32°C (A–F) or a permissive 23°C (G–L). At 32°C, the performance of GAL4;UAS-shits1 flies was significantly lower than the controls for (A) MB005B split-GAL4 (α′β′ KCs; Kruskal-Wallis, n = 30, p<0.0001), (B) MB008B split-GAL4 (αβ KCs; Kruskal-Wallis, n = 30, p=0.0039), (C) MB131B split-GAL4 (γ KCs; Kruskal-Wallis, n = 30, p=0.0004), (D) MB005B split-GAL4 (α′β′ KCs; Kruskal-Wallis, n = 30, p=0.0079), (E) MB008B split-GAL4 (αβ KCs; Kruskal-Wallis, n = 30, p=0.0286), and (F) MB131B split-GAL4 (γ KCs; Kruskal-Wallis, n = 30, p=0.0034). (G–L) At 23°C, all flies performed normally (Kruskal-Wallis, n = 35, p>0.3). Individual data points and mean ± SEM are shown.
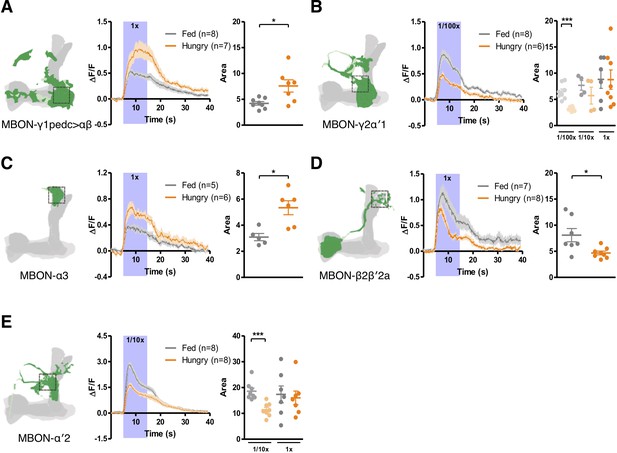
Starvation bi-directionally modulates the responses of MBONs to yeast odor.
Hunger increases (A and C) and decreases (B, D and E) yeast odor-evoked calcium transients (visualized using GCaM6m) in (A) MBON-γ1pedc>αβ (with MB112C-splitGAL4), (B) MBON-γ2α′1 (with MB077B-splitGAL4), (C) MBON-α3 (with G0239-GAL4), (D) MBON-β2β′2a (with MB399B-splitGAL4), and (E) MBON-α′2 (with MB091C-splitGAL4). Schematics indicate where the Ca2+ response was measured. Ca2+ imaging data are mean (solid line) ± SEM (shaded area) normalized curves (see Materials and methods). Wide purple bars indicate the 10 s when yeast odor was presented. Dot plots are quantifications of the area under the curve during the 10 s odor presentation. Individual data points and mean ± SEM are shown. Yeast odor was also tested at 1/10x dilution in (B) and (E) and at 1/100x dilution in (B). Asterisks denote statistical significance; Mann-Whitney test; (A) p=0.0289, (B) p=0.0007, (C) p=0.0173, (D) p=0.0289, (E) p=0.0003.
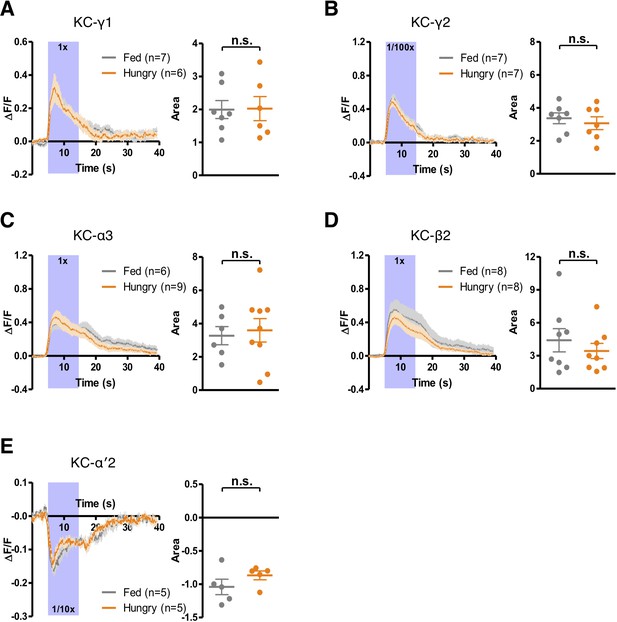
Starvation does not change the responses of KCs to yeast odor.
GCaMP6m was expressed in γ KCs using MB131B split-GAL4 (A and B) or in all KCs using MB010B split-GAL4 (C–E). The Ca2+ signals from different MB lobe compartments—γ1 (A), γ2 (B), α3 (C), β2 (D), and α′2 (E) —were measured when flies were presented with yeast odor. Ca2+ imaging data are mean (solid line) ± SEM (shaded area) normalized curves (see Materials and methods). Wide purple bars indicate the 10 s when yeast odor was presented. Dot plots are quantifications of the area under the curve during the 10 s odor presentation. Individual data points and mean ± SEM are shown. Yeast odor was tested at 1/100x dilution in (B) and 1/10x dilution in (E). No statistical difference was detected between hungry and fed flies in all cases (Mann-Whitney test; p>0.3).
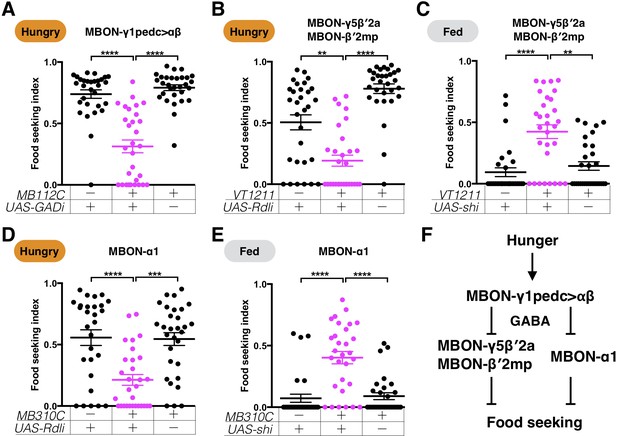
GABAergic MBON-γ1pedc>αβ promotes yeast food-seeking behavior by inhibiting β′2-innervating MBONs and MBON-α1.
Male flies starved for 24 hr (A, B and D) or food-satiated (C and E) were assessed for their yeast food-seeking performance. Individual data points and mean ± SEM are shown. (A) The performance of MB112C;UAS-GAD-RNAi flies was statistically lower than the controls (Kruskal-Wallis, n = 30, p<0.0001). (B) The performance of VT1211-GAL4;UAS-Rdl-RNAi flies was significantly lower than the controls (Kruskal-Wallis, n = 30, p=0.0053). (C) The performance of VT1211-GAL4;UAS-shits1 flies was statistically higher than the controls at a restrictive 32°C (Kruskal-Wallis, n = 30, p=0.0015). (D) The performance of MB310C;UAS-Rdl-RNAi flies was statistically lower than the controls (Kruskal-Wallis, n = 28–30, p=0.0004). (E) The performance of MB310C;UAS-shits1 flies was higher than the controls at a restrictive 32°C (Kruskal-Wallis, n = 30, p<0.0021). (F) A model showing the relationship between MBON-γ1pedc>αβ, MBON-γ5β′2a, MBON-β′2mp, and MBON-α1.
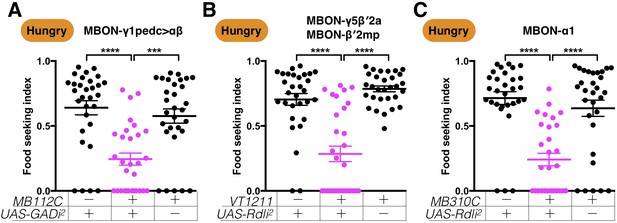
A second set of RNAi lines confirms that GABAergic MBON-γ1pedc>αβ promotes yeast food-seeking behavior by inhibiting β′2-innervating MBONs and MBON-α1.
Male flies starved for 24 hr were assessed for their yeast food-seeking performance. (A) The performance of MB112C;UAS-GAD-RNAi2 flies was statistically lower than the controls (Kruskal-Wallis, n = 30, p=0.0001). (B) The performance of VT1211-GAL4;UAS-Rdl-RNAi2 flies was significantly lower than the controls (Kruskal-Wallis, n = 30, p<0.0001). (C) The performance of MB310C;UAS-Rdl-RNAi2 flies was statistically lower than the controls (Kruskal-Wallis, n = 30, p<0.0001). Individual data points and mean ± SEM are shown.
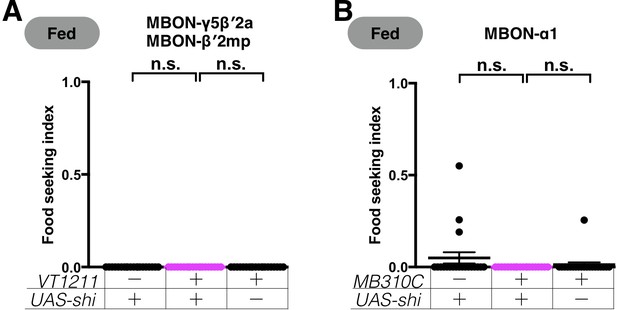
Expression of UAS-shits1 in the MBONs does not affect yeast food-seeking performance at the permissive temperature.
(A) No statistical difference was detected between the controls and flies expressing UAS-shits1 in MBON-γ5β′2a and MBON-β′2mp (VT1211, Kruskal-Wallis, n = 20, p>0.9999). (B) No statistical difference was detected between flies expressing UAS-shits1 in MBON-α1 and relevant controls (MB310C, Kruskal-Wallis, n = 20, p=0.1545). Individual data points and mean ± SEM are shown.
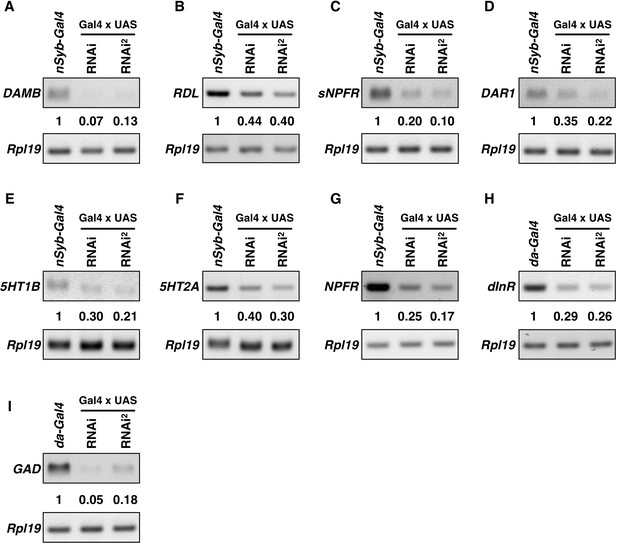
Knockdown efficiency of the RNAi lines.
RT-PCR results for DAMB, Rdl, sNPFR, DAR1, 5HT1B, 5HT2A, NPFR, dInR, GAD, and Rpl19 are shown. Pan-neuronal nSyb-GAL4 (A–G) or da-GAL4 (H–I) were used to drive two independent RNAi lines (Key resources table). Total RNA was extracted from adult brains (A–G), or—because flies with knockdown of dInR and GAD using nSyb-GAL4 failed to survive to adulthood—third-instar larvae (H–I). Rpl19 was used as an internal control. Intensity of the PCR bands was normalized to Rpl19, and the quantification of intensities relative to the nSyb-GAL4-only controls are shown.
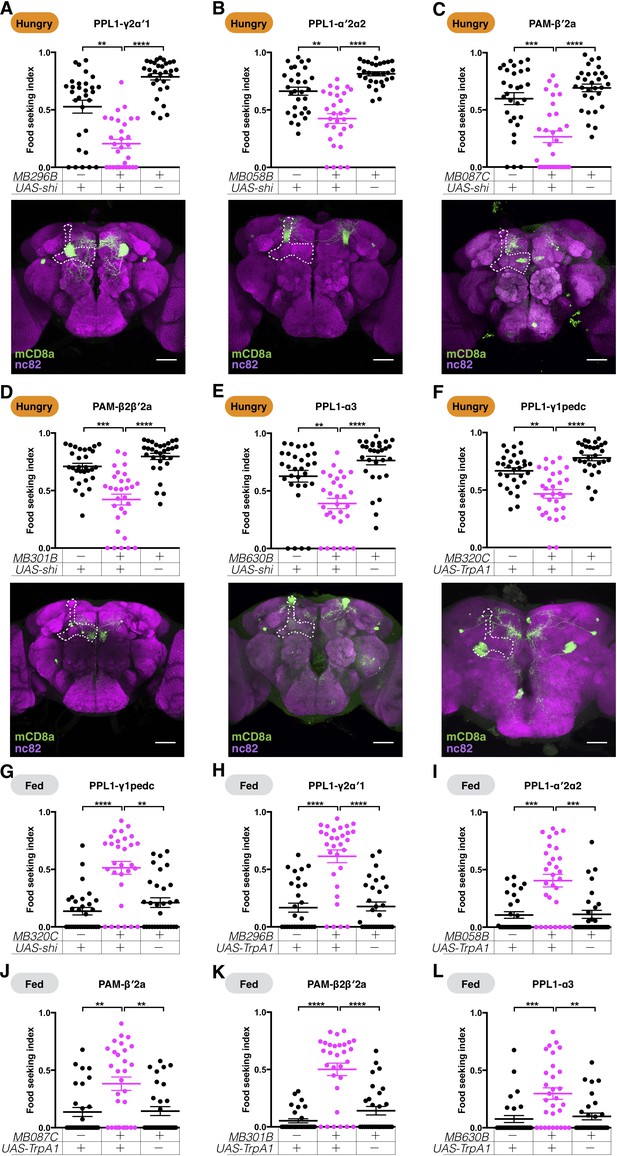
DANs mediate hunger-control of yeast food-seeking behavior.
(A–E) Male flies starved for 24 hr were assessed for their yeast food-seeking performance. At a restrictive 32°C, the performance was significantly different between the controls and flies expressing UAS-shits1 in (A) PPL1-γ2α′1 (MB296B, Kruskal-Wallis, n = 30, p=0.0018), (B) PPL1-α′2α2 (MB058B, Kruskal-Wallis, n = 30, p=0.0012), (C) PAM-β′2a (MB087C, Kruskal-Wallis, n = 30, p=0.0002), (D) PAM-β2β′2a (MB301B, Kruskal-Wallis, n = 30, p=0.0001), and (E) PPL1-α3 (MB630B, Kruskal-Wallis, n = 30, p=0.0023) DANs. (F) The performance of MB320C;UAS-TrpA1 male flies starved for 24 hr was lower than the controls (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p=0.003). (G) The performance of male MB320C;UAS- shits1 fed flies was statistically better than the controls (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p=0.001). (H–L) Food-satiated male flies were tested for their yeast food-seeking performance. At 32°C, the performance was statistically different between the controls and flies expressing UAS-TrpA1 in (H) PPL1-γ2α′1 (MB296B, Kruskal-Wallis, n = 30, p<0.0001), (I) PPL1-α′2α2 (MB058B, n = 30, p=0.0004), (J) PAM-β′2a (MB087C, Kruskal-Wallis, n = 30, p=0.0056), (K) PAM-β2β′2a (MB301B, Kruskal-Wallis, n = 30, p<0.0001), and (L) PPL1-α3 (MB630B, Kruskal-Wallis, n = 30, p=0.0049) DANs. Individual data points and mean ± SEM are shown. The brain images in (A–F) are full z-projections of confocal stacks showing the expression patterns of the GAL4 lines (green) counter-stained with nc82 antibody (magenta). One side of the MB is outlined by a white dashed line. Scale bars are 100 µm.
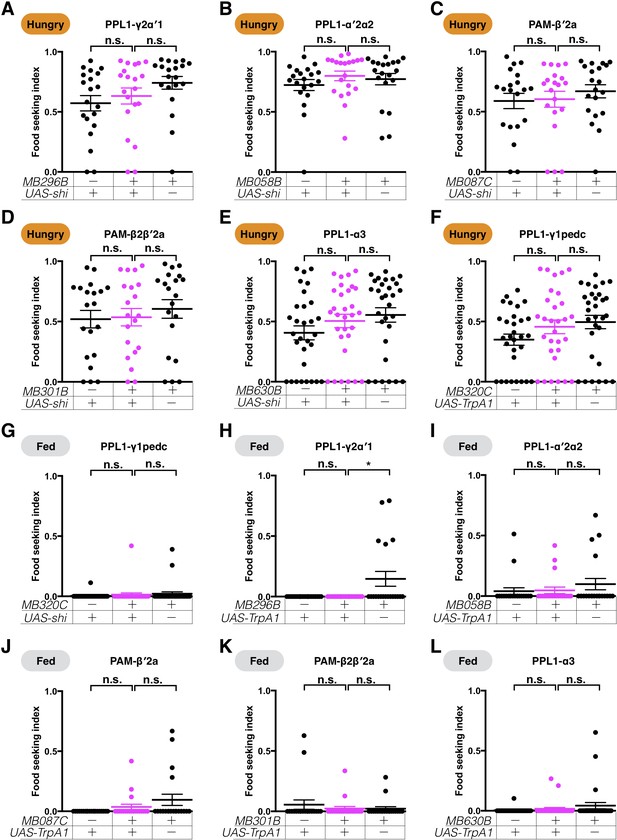
Expression of UAS-shits1 or UAS-TrpA1 in the DANs does not affect yeast food-seeking performance at the permissive temperature.
No statistical difference was detected between the controls and flies expressing UAS-shits1 in (A) PPL1-γ2α′1 (MB296B, Kruskal-Wallis, n = 20, p=0.0737), (B) PPL1-α′2α2 (MB058B, Kruskal-Wallis, n = 20, p=0.1662), (C) PAM-β′2a (MB087C, Kruskal-Wallis, n = 20, p=0.6230), (D) PAM-β2β′2a (301B, Kruskal-Wallis, n = 20, p=0.5019), and (E) PPL1-α3 (MB630B, Kruskal-Wallis, n = 30, p=0.1456) DANs. (F) No statistical difference was detected between flies expressing UAS-TrpA1 in PPL1-γ1pedc DANs and relevant controls (MB320C, Kruskal-Wallis, n = 30, p=0.0794). (G) No statistical difference was detected between flies expressing UAS-shits1 in PPL1-γ1pedc DANs and relevant controls (MB320C, Kruskal-Wallis, n = 30, p=0.7715). No statistical difference was detected between the controls and flies expressing UAS-TrpA1 in (H) PPL1-γ2α′1 (MB296B, Kruskal-Wallis, n = 20, p=0.6225), (I) PPL1-α′2α2 (MB058B, Kruskal-Wallis, n = 20, p=0.199), (J) PAM-β′2a (MB087C, Kruskal-Wallis, n = 20, p=0.9928), (K) PAM-β2β′2a (301B, Kruskal-Wallis, n = 20, p=0.4434), and (L) PPL1-α3 (MB630B, Kruskal-Wallis, n = 30, p=0.5547) DANs. Satiety states (fed or hungry) are indicated in each figure. Individual data points and mean ± SEM are shown.
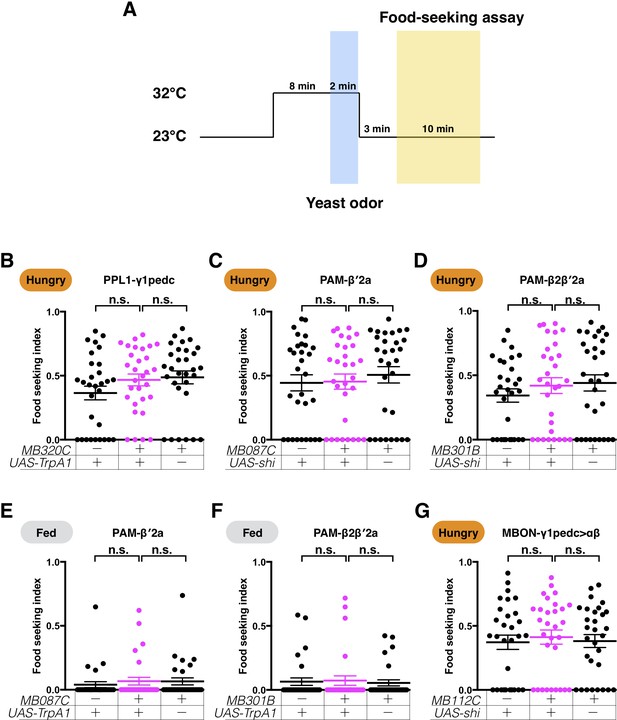
Pre-conditioning flies by pairing yeast odor with the activation or silencing of DANs and MBONs does not affect their yeast food-seeking performance.
(A) A schematic of the behavioral protocol. (B–G) Under this protocol, no difference in yeast food-seeking performance was detected between the controls and (B) MB320C;UAS-TrpA1 (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p=0.1979), (C) MB087C;UAS-shits1 (PAM-β′2a, Kruskal-Wallis, n = 30, p=0.612), (D) MB301B;UAS-shits1 (PAM-β2β′2a, Kruskal-Wallis, n = 30, p=0.375), (E) MB087C;UAS-TrpA1 (PAM-β′2a, Kruskal-Wallis, n = 30, p=0.6082), (F) MB301B;UAS-TrpA1 (PAM-β2β′2a, Kruskal-Wallis, n = 30, p=0.9636), and (G) MB112C;UAS-shits1 (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p=0.8135) flies. Satiety states (fed or hungry) are indicated in each figure. Individual data points and mean ± SEM are shown.
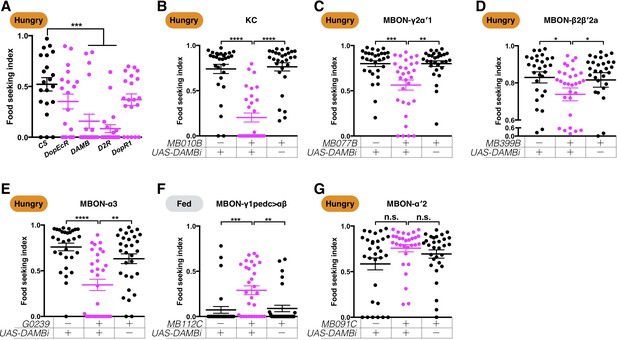
The dopamine receptor DAMB is required pre- and post-synaptically to regulate yeast food-seeking behavior.
(A) The yeast food-seeking performance in 24-hr-starved male wild-type flies (CS, n = 21) and flies homozygous for DopEcR (n = 19), DAMB (n = 20), D2R (n = 20), and DopR1 (n = 20) was assessed. The performance of the DAMB and D2R flies was significantly lower than for the wild-type flies (Kruskal-Wallis, p=0.0024 for DAMB; p=0.0003 for D2R). (B–G) Male flies starved for 24 hr (B–E and G) or food-satiated (F) were assessed for their yeast food-seeking performance. The performance of GAL4;UAS-DAMB-RNAi flies was statistically different from the controls for (B) MB010B (all KCs, Kruskal-Wallis, n = 29–30, p<0.0001), (C) MB077B (MBON-γ2α′1, Kruskal-Wallis, n = 30, p=0.0018), (D) MB399B (MBON-β2β′2a, Kruskal-Wallis, n = 29–30, p=0.0285), (E) G0239 (MBON-α3, Kruskal-Wallis, n = 30, p=0.007). (F) MB112C (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p=0.0028), but not for (G) MB091C (MBON-α′2, Kruskal-Wallis, n = 30, p=0.5757). Satiety states (fed or hungry) are indicated in each figure. Individual data points and mean ± SEM are shown.
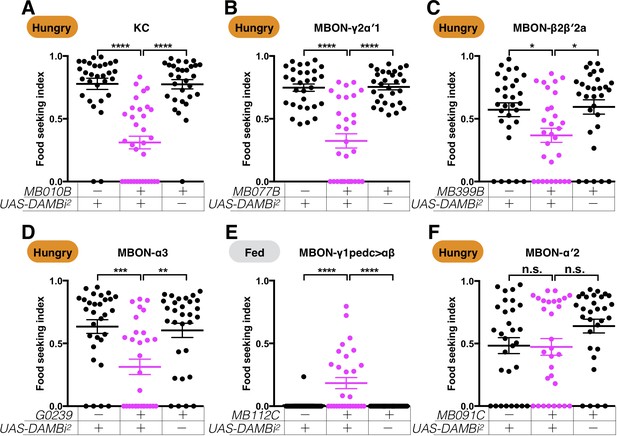
A second set of RNAi lines confirms that the dopamine receptor DAMB is required both pre- and post-synaptically to regulate yeast food-seeking behavior.
Male flies starved for 24 hr (A–D and F) or food-satiated (E) were assessed for their yeast food-seeking performance. The performance of GAL4;UAS-DAMB-RNAi2 flies was statistically different from the controls for (A) MB010B (all KCs, Kruskal-Wallis, n = 30–33, p<0.0001), (B) MB077B (MBON-γ2α′1, Kruskal-Wallis, n = 30, p<0.0001), (C) MB399B (MBON-β2β′2a, Kruskal-Wallis, n = 30, p=0.0488), (D) G0239 (MBON-α3, Kruskal-Wallis, n = 30, p=0.0036), and (E) MB112C (MBON-γ1pedc>αβ, Kruskal-Wallis, n = 30, p<0.0001), but not for (G) MB091C (MBON-α′2, Kruskal-Wallis, n = 30, p=0.1624). Satiety states (fed or hungry) are indicated in each figure. Individual data points and mean ± SEM are shown.
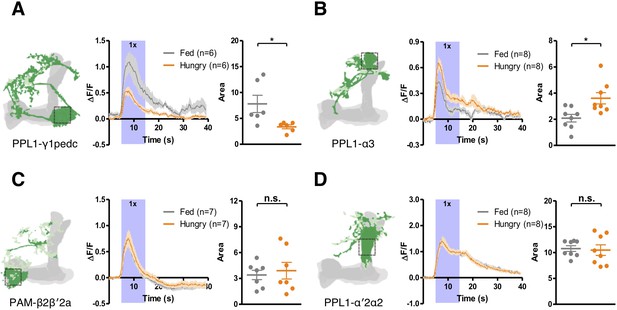
Starvation modulates yeast odor-evoked responses in some DANs.
GCaMP6m was expressed in PPL1-γ1pedc DANs using MB320C split-GAL4 (A), PPL1-α3 DANs using G0239-GAL4 (B), PAM-β2β′2a DANs using MB301B split-GAL4 (C), and PPL1-α′2α2 DANs using MB058B split-GAL4 (D). The Ca2+ signals were measured when flies were presented with yeast odor. Schematics indicate where the Ca2+ response was measured. Ca2+ imaging data are mean (solid line) ± SEM (shaded area) normalized curves (see Materials and methods). Wide purple bars indicate the 10 s when yeast odor was presented. Dot plots are quantifications of the area under the curve during the 10 s odor presentation. Individual data points and mean ± SEM are shown. Statistical differences were detected in (A) PPL1-γ1pedc (Mann-Whitney test, n = 6, p=0.026) and (B) PPL1-α3 (Mann-Whitney test, n = 8, p=0.014) DANs, but not in (C) PAM-β2β′2a (Mann-Whitney test, n = 7–8, p=0.7789) or (D) PPL1-α′2α2 (Mann-Whitney test, n = 8, p=0.7984) DANs.
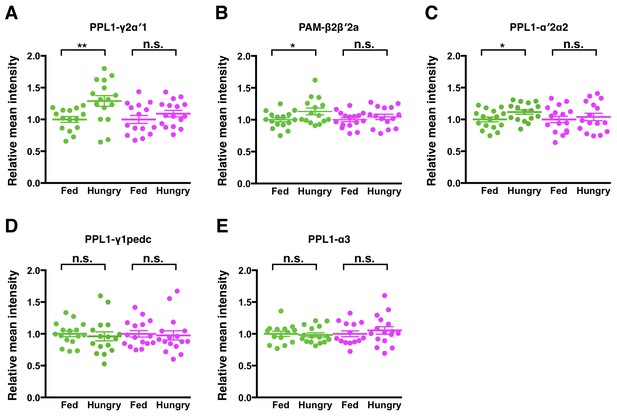
Starvation may increase the density or size of active zones in some DANs.
Transgenes UAS-DSyd-1-GFP and UAS-DenMark were expressed in PPL1-γ2α′1 DANs using MB296B split-GAL4 (A), PAM-β2β′2a DANs using MB301B split-GAL4 (B), PPL1-α′2α2 DANs using MB058B split-GAL4 (C), PPL1-γ1pedc DANs using MB320C split-GAL4 (D), and PPL1-α3 DANs using G0239-GAL4 (E). Mean intensity of DSyd-1-GFP (green dots) was significantly different between hungry and fed flies for (A) PPL1-γ2α′1 (unpaired t-test, n = 15–16, p=0.0056), (B) PAM-β2β′2a (unpaired t-test, n = 15–16, p=0.0406), and (C) PPL1-α′2α2 (unpaired t-test, n = 16, p=0.0314) DANs, but not for (D) PPL1-γ1pedc (unpaired t-test, n = 16, p=0.6393) or (E) PPL1-α3 (unpaired t-test, n = 14–16, p=0.7481) DANs. Mean intensity of DenMark (magenta dots) did not differ between hungry and fed flies in all cases (A–E) (unpaired t-test, n = 15–16, p>0.25). Data are normalized to the mean intensities of the fed flies. Individual data points and mean ± SEM are shown.
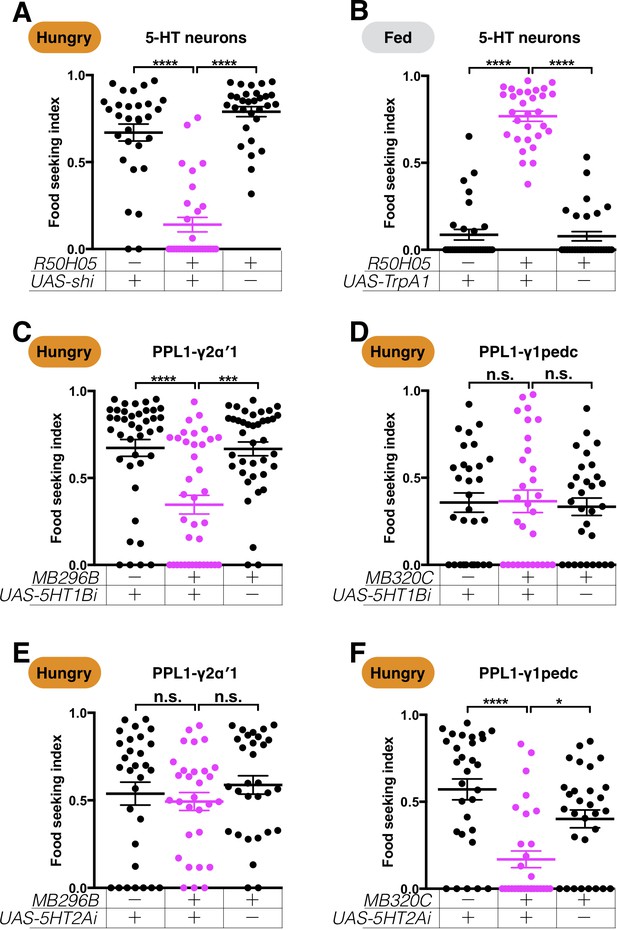
Serotonin regulates PPL1-γ2α′1 and PPL1-γ1pedc DANs via different receptors.
The yeast food-seeking performance of 24-hr-starved (A and C–F) and food-satiated (B) male flies was assessed. (A) The performance of R50H05-GAL4;UAS-shits1 flies was statistically worse than for the control flies at a restrictive 32°C (5-HT neurons, Kruskal-Wallis, n = 30, p<0.0001). (B) The performance of R50H05-GAL4;UAS-TrpA1 flies was statistically better than for the control flies at a restrictive 32°C (5-HT neurons, Kruskal-Wallis, n = 30, p<0.0001). (C) The performance of MB296B;UAS-5HT1B-RNAi flies was statistically worse than for the control flies (PPL1-γ2α′1, Kruskal-Wallis, n = 39–40, p=0.0003). (D) The performance of MB320C;UAS-5HT1B-RNAi flies was not statistically different from that of control flies (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p>0.9999). (E) The performance of MB296B;UAS-5HT2A-RNAi flies was not statistically different from that of control flies (PPL1-γ2α′1 Kruskal-Wallis, n = 30, p=0.5116). (F) The performance of MB320C;UAS-5HT2A-RNAi flies was statistically worse than that of control flies (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p=0.0263). Individual data points and mean ± SEM are shown.
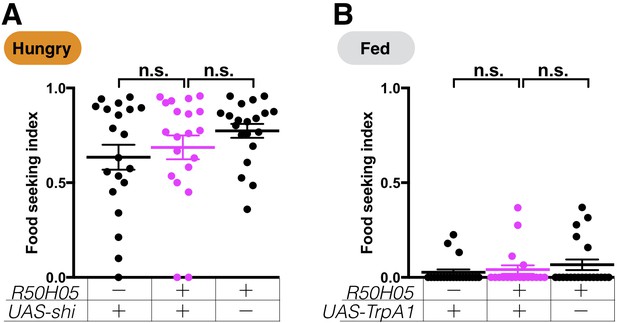
Expression of UAS-shits1 or UAS-TrpA1 in serotoninergic neurons does not affect yeast food-seeking performance at the permissive temperature.
(A) No statistical difference was detected between flies expressing UAS-shits1 in serotoninergic neurons and the relevant controls (R50H05, Kruskal-Wallis, n = 20, p=0.5377). (B) No statistical difference was detected between flies expressing UAS-TrpA1 in serotoninergic neurons and the relevant controls (R50H05, Kruskal-Wallis, n = 20, p=0.6367). Individual data points and mean ± SEM are shown. Satiety states (fed or hungry) are indicated in each figure.
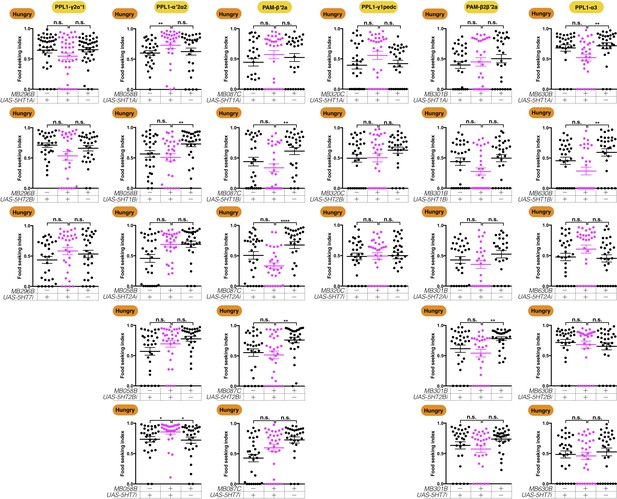
RNAi knockdown of serotonin receptors in the yeast-seeking DANs.
The yeast food-seeking performance of 24-hr-starved male flies with RNAi knockdown of different serotonin receptors (5HT1A, 5HT1B, 5HT2A, 5HT2B, and 5HT7) in different DANs is shown. Data are individual data points and mean ± SEM. The names of the GAL4 lines (MB296B, MB058B, MB087C, MB320C, MB301B, and MB630B) and the neurons they label (PPL1-γ2α′1, PPL1-α′2α2, PAM-β′2a, PPL1-γ1pedc, PAM-β2β′2a, and PPL1-α3 DANs) are indicated. For all combinations shown here, there is no significant difference in yeast-seeking performance between RNAi knockdown flies and at least one of their relevant controls (Kruskal-Wallis, n = 25–30, p>0.05), except for flies with knockdown of 5HT7 in PPL1-α′2α2 DANs that exhibited slightly enhanced yeast food-seeking performance (Kruskal-Wallis, n = 30, p=0.0099). The latter finding suggests that PPL1-α′2α2 DANs may be regulated by tonic inhibitory serotonin inputs.
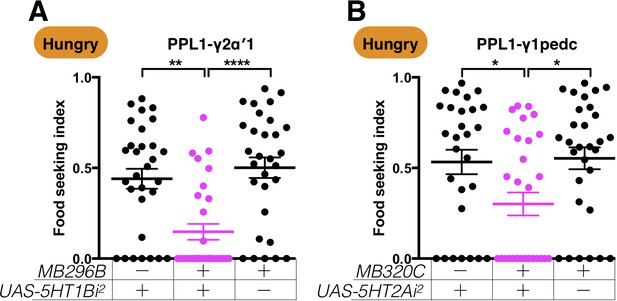
A second set of RNAi lines confirms the importance of 5HT1B and 5HT2A receptors in regulating yeast food-seeking behavior.
(A) The yeast food-seeking performance of MB296B;UAS-5HT1B-RNAi2 flies was statistically worse than that of the control flies (PPL1-γ2α′1, Kruskal-Wallis, n = 30, p=0.0012). (B) The yeast food-seeking performance of MB320C;UAS-5HT2A-RNAi2 flies was statistically worse than that of the control flies (PPL1-γ1pedc, Kruskal-Wallis, n = 28–30, p=0.0388). Individual data points and mean ± SEM are shown.
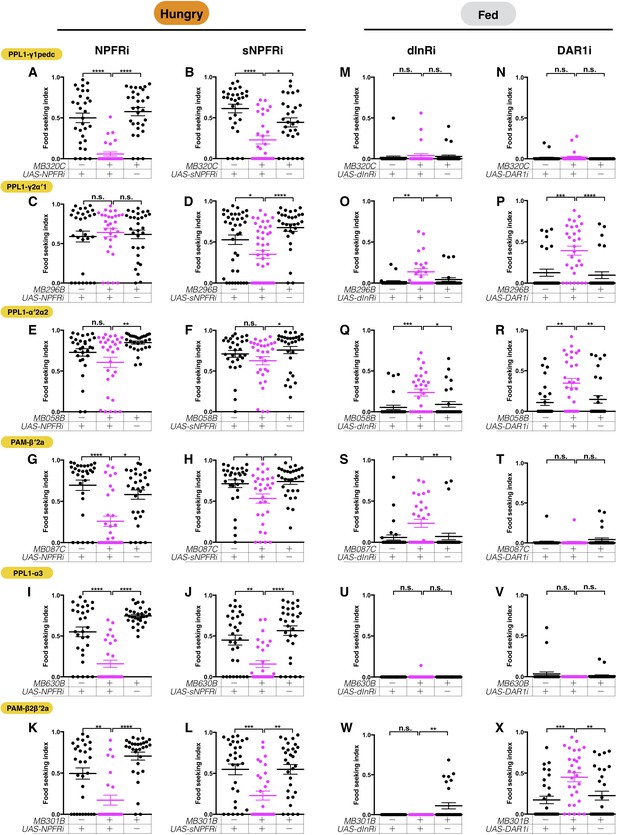
The six assessed DANs are regulated by combinations of hunger and satiety signals.
The yeast food-seeking performance of 24-hr-starved (A–L) and food-satiated (M–X) male flies was assessed. (A) The performance of MB320C;UAS-NPFR-RNAi flies was impaired (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p<0.0001). (B) The performance of MB320C;UAS-sNPFR-RNAi flies was impaired (PPL1-γ1pedc, Kruskal-Wallis, n = 29–30, p=0.0438). (C) The performance of MB296B;UAS-NPFR-RNAi flies was normal (PPL1-γ2α′1, Kruskal-Wallis, n = 30, p>0.9999). (D) The performance of MB296B;UAS-sNPFR-RNAi flies was impaired (PPL1-γ2α′1, Kruskal-Wallis, n = 30–44, p=0.0145). (E) The performance of MB058B;UAS-NPFR-RNAi flies was normal (PPL1-α′2α2, n = 30, p>0.9999). (F) The performance of MB058B;UAS-sNPFR-RNAi flies was normal (PPL1-α′2α2, Kruskal-Wallis, n = 30, p=0.4948). (G) The performance of MB087C;UAS-NPFR-RNAi flies was impaired (PAM-β′2a, Kruskal-Wallis, n = 30, p=0.0155). (H) The performance of MB087C;UAS-sNPFR-RNAi flies was impaired (PAM-β′2a, Kruskal-Wallis, n = 30, p=0.0162). (I) The performance of MB630B;UAS-NPFR-RNAi flies was impaired (PPL1-α3, Kruskal-Wallis, n = 30, p<0.0001). (J) The performance of MB630B;UAS-sNPFR-RNAi flies was impaired (PPL1-α3, Kruskal-Wallis, n = 30, p=0.0038). (K) The performance of MB301B;UAS-NPFR-RNAi flies was impaired (PAM-β2β′2a, Kruskal-Wallis, n = 25–30, p=0.0087). (L) The performance of MB301B;UAS-sNPFR-RNAi flies was impaired (PAM-β2β′2a, Kruskal-Wallis, n = 30, p=0.0013). (M) The performance of MB320C;UAS-dInR-RNAi flies was normal (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p>0.9999). (N) The performance of MB320C0;UAS-DAR1-RNAi flies was normal (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p>0.9999). (O) The performance of MB296B;UAS-dInR-RNAi flies was enhanced (PPL1-γ2α′1, Kruskal-Wallis, n = 24–25, p=0.0199). (P) The performance of MB296B;UAS-DAR1-RNAi flies was enhanced (PPL1-γ2α′1, Kruskal-Wallis, n = 30, p=0.0002). (Q) The performance of MB058B;UAS-dInR-RNAi flies was enhanced (PPL1-α′2α2, Kruskal-Wallis, n = 29–30, p=0.0126). (R) The performance of MB058B;UAS-DAR1-RNAi flies was enhanced (PPL1-α′2α2, Kruskal-Wallis, n = 30, p=0.009). (S) The performance of MB087C;UAS-dInR-RNAi flies was enhanced (PAM-β′2a, Kruskal-Wallis, n = 30, p=0.0192). (T) The performance of MB087C;UAS-DAR1-RNAi flies was normal (PAM-β′2a, Kruskal-Wallis, n = 30, p=0.3645). (U) The performance of MB630B;UAS-dInR-RNAi flies was normal (PPL1-α3, Kruskal-Wallis, n = 30, p=0.6620). (V) The performance of MB630B;UAS-DAR1-RNAi flies was normal (PPL1-α3, Kruskal-Wallis, n = 30, p=0.6699). (W) The performance of MB301B;UAS-dInR-RNAi flies was normal (PAM-β2β′2a, Kruskal-Wallis, n = 25–30, p>0.9999). (X) The performance of MB301B;UAS-DAR1-RNAi flies was enhanced (PAM-β2β′2a, Kruskal-Wallis, n = 30, p=0.0071). Individual data points and mean ± SEM are shown.
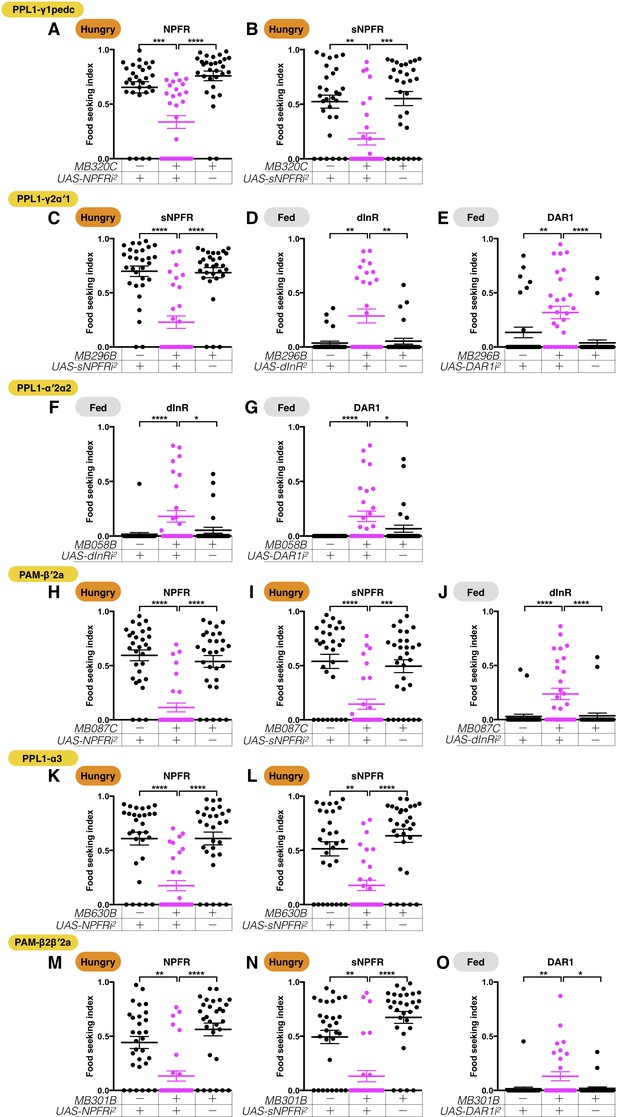
A second set of RNAi lines confirms that the six assessed DANs are regulated by combinations of hunger and satiety signals.
The yeast food-seeking performance of 24-hr-starved or food-satiated male flies was assessed. (A) The performance of MB320C;UAS-NPFR-RNAi2 flies was impaired (PPL1-γ1pedc, Kruskal-Wallis, n = 30, p=0.0007). (B) The performance of MB320C;UAS-sNPFR-RNAi2 flies was impaired (PPL1-γ1pedc, Kruskal-Wallis, n = 29–30, p=0.0016). (C) The performance of MB296B;UAS-sNPFR-RNAi2 flies was impaired (PPL1-γ2α′1, Kruskal-Wallis, n = 30, p<0.0001). (D) The performance of MB296B;UAS-dInR-RNAi2 flies was enhanced (PPL1-γ2α′1, Kruskal-Wallis, n = 30–44, p=0.0057). (E) The performance of MB296B;UAS-DAR1-RNAi2 flies was enhanced (PPL1-γ2α′1, Kruskal-Wallis, n = 30, p=0.0071). (F) The performance of MB058B;UAS-dInR-RNAi2 flies was enhanced (PPL1-α′2α2, n = 30, p=0.0243). (G) The performance of MB058B;UAS-DAR1-RNAi2 flies was enhanced (PPL1-α′2α2, Kruskal-Wallis, n = 30, p=0.0158). (H) The performance of MB087C;UAS-NPFR-RNAi2 flies was impaired (PAM-β′2a, Kruskal-Wallis, n = 30, p<0.0001). (I) The performance of MB087C;UAS-sNPFR-RNAi2 flies was impaired (PAM-β′2a, Kruskal-Wallis, n = 30, p=0.001). (J) The performance of MB087C;UAS-dInR-RNAi2 flies was enhanced (PAM-β′2a, Kruskal-Wallis, n = 30, p<0.0001). (K) The performance of MB630B;UAS-NPFR-RNAi2 flies was impaired (PPL1-α3, Kruskal-Wallis, n = 30, p<0.0001). (L) The performance of MB630B;UAS-sNPFR-RNAi2 flies was impaired (PPL1-α3, Kruskal-Wallis, n = 30, p=0.002). (M) The performance of MB301B;UAS-NPFR-RNAi2 flies was impaired (PAM-β2β′2a, Kruskal-Wallis, n = 30, p=0.0019). (N) The performance of MB301B;UAS-sNPFR-RNAi2 flies was impaired (PAM-β2β′2a, Kruskal-Wallis, n = 30, p=0.0037). (O) The performance of MB301B;UAS-DAR1-RNAi2 flies was enhanced (PAM-β2β′2a, Kruskal-Wallis, n = 30, p=0.021). Individual data points and mean ± SEM are shown. Satiety states (fed or hungry) are indicated in each figure.
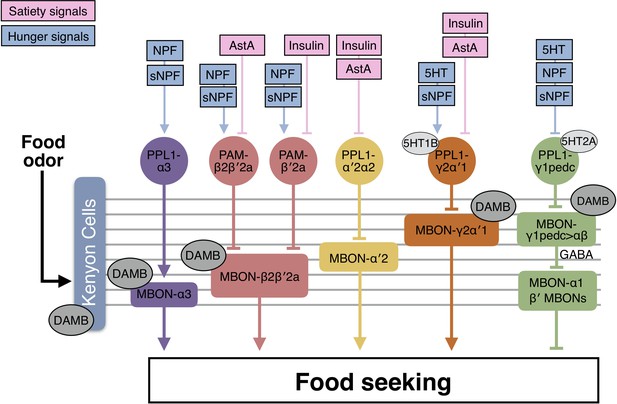
A model for the neural mechanics of the MB circuit in controlling food-seeking behavior.
During food seeking, odors activate the KCs and in turn activate MBON-α3, MBON-β2β′2a, MBON-α′2, MBON-γ2α′1 and MBON-γ1pedc>αβ. GABAergic MBON-γ1pedc>αβ inhibits the downstream neurons that suppress food-seeking behavior, including β′2-innervating MBONs and MBON-α1. KC-to-MBON connectivity is regulated by the corresponding DANs. The DANs are regulated by combinations of hunger and satiety signals. When flies are food-satiated, satiety signals like insulin and AstA suppress PPL1-γ2α′1, PPL1-α′2α2, PAM-β′2a, and PAM-β2β′2a DANs. When flies are starved, hunger signals including serotonin (5HT), NPF, and sNPF activate PPL1-α3, PAM-β2β′2a, PAM-β′2a, and PPL1-γ2α′1 DANs, whereas they suppress PPL1-γ1pedc DANs. Dopamine signals pre- and post-synaptically mediated by the DAMB receptor fine-tune the KC-to-MBON connectivity and modulate the collective output of the MBONs driven by food odor. Therefore, hunger state tunes the odor-driven output of the MBONs to regulate food-seeking behavior.
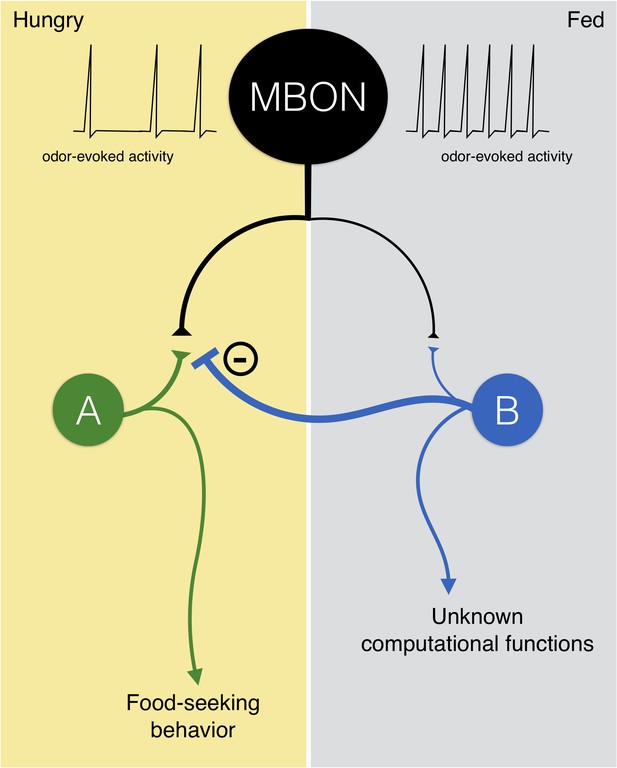
A hypothetical model for the MBONs that show reduced responses to yeast odor in hungry flies.
The MBONs have strong connections with neuron A and weak connections with neuron B. When flies are starved, weak activities of the MBONs evoked by yeast odor are sufficient to activate neuron A but not neuron B. The activation of neuron A drives food-seeking behavior. When flies are fed, strong yeast odor-evoked activity in the MBONs activates neuron B, which in turns inhibits neuron A. The outputs from neuron B might be important for other computational functions, but the flies do not exhibit food-seeking behavior. Importantly, in this model the output from MBONs is still required for food-seeking behavior in hungry flies, so silencing these MBONs with UAS-shits1 leads to impaired food-seeking behavior.
Videos
Single fly food-seeking assay
https://doi.org/10.7554/eLife.35264.035Tables
| Reagent type | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Fly line | E0067-Gal4 | (Pai et al., 2013) | NA | Gift from Ann-Shyn Chiang |
| Fly line | G0239-Gal4 | (Pai et al., 2013) | Flybase: FBti0132502 | |
| Fly line | MB002B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68305 | |
| Fly line | MB005B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68306 | |
| Fly line | MB008B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68291 | |
| Fly line | MB011B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68294 | |
| Fly line | MB018B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68296 | |
| Fly line | MB027B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68301 | |
| Fly line | MB050B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68365 | |
| Fly line | MB051B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68275 | |
| Fly line | MB052B-SplitGal4 | (Aso et al., 2014a) | NA | Gift from Yoshinori Aso |
| Fly line | MB057B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68277 | |
| Fly line | MB062C-SplitGal4 | (Aso et al., 2014a) | NA | Gift from Yoshinori Aso |
| Fly line | MB077B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68283 | |
| Fly line | MB077C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68284 | |
| Fly line | MB080C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68285 | |
| Fly line | MB082C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68286 | |
| Fly line | MB083C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68287 | |
| Fly line | MB085C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68288 | |
| Fly line | MB090C-SplitGal4 | (Aso et al., 2014a) | NA | Gift from Yoshinori Aso |
| Fly line | MB091C-SplitGal4 | (Aso et al., 2014a) | NA | Gift from Yoshinori Aso |
| Fly line | MB093C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68289 | |
| Fly line | MB110C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68262 | |
| Fly line | MB112C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68263 | |
| Fly line | MB131B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68265 | |
| Fly line | MB210B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68272 | |
| Fly line | MB242A-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68307 | |
| Fly line | MB262B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68254 | |
| Fly line | MB298B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68309 | |
| Fly line | MB310C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68313 | |
| Fly line | MB399B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68369 | |
| Fly line | MB433B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68324 | |
| Fly line | MB434B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68325 | |
| Fly line | MB542B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68372 | |
| Fly line | MB543B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68335 | |
| Fly line | MB549C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68373 | |
| Fly line | MB622B-SplitGal4 | (Aso et al., 2014a) | NA | Gift from Yoshinori Aso |
| Fly line | R21D02-Gal4 | (Owald et al., 2015) | RRID:BDSC_48939 | |
| Fly line | R50H05-Gal4 | (Albin et al., 2015) | RRID:BDSC_38764 | |
| Fly line | R66C08-Gal4 | (Owald et al., 2015) | RRID:BDSC_49412 | |
| Fly line | VT1211-Gal4 | (Owald et al., 2015) | VDRC: 202324 | |
| Fly line | VT999036-Gal4 | (Aso et al., 2014a) | NA | Gift from Yoshinori Aso |
| Fly line | MB320C-SplitGAL4 | (Aso et al., 2014a) | RRID:BDSC_68253 | |
| Fly line | MB058B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68278 | |
| Fly line | MB087C-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68366 | |
| Fly line | MB296B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68308 | |
| Fly line | MB301B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68311 | |
| Fly line | MB630B-SplitGal4 | (Aso et al., 2014a) | RRID:BDSC_68334 | |
| Fly line | *UAS-DAMBi | (Pimentel et al., 2016) | VDRC: 105324; RRID:FlyBase_FBst0478846 | |
| Fly line | *UAS-GADi | (Barnstedt et al., 2016) | VDRC: 32344; RRID:FlyBase_FBst0459538 | |
| Fly line | *UAS-Rdli | (Cheung and Scott, 2017) | VDRC: 41103; RRID:FlyBase_FBst0463935 | |
| Fly line | UAS-5HT1Ai | (Lee et al., 2011) | VDRC: 106094; RRID:FlyBase_FBst0472248 | |
| Fly line | *UAS-5HT1Bi | (Mohammad et al., 2016) | VDRC: 109929; RRID:FlyBase_FBst0476393 | |
| Fly line | *UAS-5HT2Ai | (Mohammad et al., 2016) | VDRC: 102105; RRID:FlyBase_FBst0475582 | |
| Fly line | UAS-5HT2Bi | (Mohammad et al., 2016) | VDRC: 102356; RRID:FlyBase_FBst0478281 | |
| Fly line | UAS-5HT7i | (Dietzl et al., 2007) | VDRC: 104804; RRID:FlyBase_FBst0472736 | |
| Fly line | *UAS-NPFRi | (Ni et al., 2009) | RRID:BDSC_25939 | |
| Fly line | *UAS-sNPFRi | (Hong et al., 2012) | VDRC: 38925; RRID:FlyBase_FBst0462758 | |
| Fly line | *UAS-dInRi | (Loh et al., 2017) | RRID:BDSC_35251 | |
| Fly line | *UAS-DAR1i | (Yamagata et al., 2016) | NA | Gift from Hiromu Tanimoto |
| Fly line | *UAS-DAMBi2 | (Hattori et al., 2017) | RRID:BDSC_51423 | |
| Fly line | *UAS-GADi2 | (Koganezawa et al., 2016) | RRID:BDSC_51794 | |
| Fly line | *UAS-Rdli2 | (Koganezawa et al., 2016) | RRID:BDSC_52903 | |
| Fly line | *UAS-5HT1Bi2 | (Kaneko et al., 2017) | RRID:BDSC_33418 | |
| Fly line | *UAS-5HT2A2 | (Ni et al., 2009) | RRID:BDSC_56870 | |
| Fly line | *UAS-NPFRi2 | (Dietzl et al., 2007) | VDRC: 107663; RRID:FlyBase_FBst0481454 | |
| Fly line | *UAS-dInRi2 | (Ni et al., 2009) | RRID:BDSC_51518 | |
| Fly line | *UAS-sNPFRi2 | (Hu et al., 2017) | RRID:BDSC_27507 | |
| Fly line | *UAS-DAR1i2 | (Yamagata et al., 2016) | NA | Gift from Hiromu Tanimoto |
| Fly line | Orco2 | (Larsson et al., 2004) | RRID:BDSC_23130 | |
| Fly line | IR25a2 | (Abuin et al., 2011) | RRID:BDSC_41737 | |
| Fly line | IR8a1 | (Abuin et al., 2011) | RRID:BDSC_41744 | |
| Fly line | DAMB | (Selcho et al., 2009) | Flybase: FBab0047678 | |
| Fly line | DopR1 (dumb2) | (Qin et al., 2012) | Exelixis: f02676; RRID:FlyBase_FBst1017920 | |
| Fly line | D2Rf06521 | (Marella et al., 2012) | Exelixis: f06521; RRID:FlyBase_FBst1020637 | |
| Fly line | DopEcRc02142 | (Inagaki et al., 2012) | Exelixis: c02142; RRID:FlyBase_FBst1006135 | |
| Fly line | UAS-TrpA1 | (Hamada et al., 2008) | Flybase: FBtp0040248 | Gift from Scott Waddell |
| Fly line | UAS-shits1 | (Kitamoto, 2001) | Flybase: FBtp0013545 | Gift from Scott Waddell |
| Fly line | UAS-mCD8-GFP | (Lee and Luo, 1999) | Flybase: FBtp0002652 | Gift from Scott Waddell |
| Fly line | UAS-GCaMP6m | (Chen et al., 2013) | RRID:BDSC_42748 | |
| Fly line | UAS-DenMark,UAS-Dsyd-1::GFP | (Owald et al., 2015) | NA | Gift from Scott Waddell |
| Fly line | nSyb-GAL4 | (Pauli et al., 2008) | Flybase: FBtp0041245 | Gift from Scott Waddell |
| Fly line | da-GAL4 | (Wang et al., 2017) | RRID:BDSC_55851 | |
| Antibody | Mouse anti-brp (nc82) | Developmental Studys Hybridoma Bank (DSHB), IA, USA | RRID:AB_2314866 | |
| Antibody | Chicken anti-GFP | Abcam, UK | RRID:AB_300798 | 1:5000 |
| Antibody | Rabbit anti-Dsred | Clontech, CA, USA | RRID:AB_10013483 | 1:500 |
| Antibody | Rat anti-mCD8α | Thermo Fisher Scientific, MA, USA | RRID:AB_10392843 | 1:100 |
| Antibody | Goat anti-rabbit (Cy3) | Jackson ImmunoResearch, PA, USA | RRID:AB_2338006 | 1:400 |
| Antibody | Goat anti-mouse (Cy3) | Jackson ImmunoResearch, PA, USA | RRID:AB_2338692 | 1:400 |
| Antibody | Donkey anti-chicken (Alexa 488) | Jackson ImmunoResearch, PA, USA | RRID:AB_2340375 | 1:400 |
| Antibody | Goat anti-rat (Alexa 488) | Thermo Fisher Scientific, MA, USA | RRID:AB_141373 | 1:400 |
| Food odor | Dry yeast | Ferminpan red, Italy | NA | |
| Food odor | Apple cider vinegar | Alce Nero, Italy | NA | |
| Food odor | Banana powder | Gen Asia Biotech, Taiwan | NA | |
| Chemical | Sucrose | Merck, Germany | Cat# 107687 | |
| Chemical | Agar | BD, NJ, USA | Cat# 214530 | |
| Chemical | Formaldehyde | Sigma, MO, USA | Cat# F8775 | |
| Chemical | PBS | Sigma, MO, USA | Cat# P4417 | |
| Chemical | Gold Antifade reagent | Thermo Fisher Scientific, MA, USA | Cat# S36937 | |
| Chemical | paraffin wax | Sigma, MO, USA | Cat# 327304 | |
| Serum | Normal goat serum | Jackson ImmunoResearch, PA, USA | RRID:AB_2336990 | |
| Kits | TRIzol RNA Isolation reagents | Thermo Fisher Scientific, MA, USA | Cat# 15596026 | |
| Kits | SuperScript IV First-Strand Synthesis System for RT-PCR Kit | Thermo Fisher Scientific, MA, USA | Cat# 18091050 | |
| Software | Prism 7 | GraphPad, CA, USA | RRID:SCR_002798 | |
| Software | MATLAB 2017a | MathWorks, MA, USA | RRID:SCR_001622 | |
| Software | Fiji/ImageJ | Fiji | RRID:SCR_002285 |
-
*Asterisks indicate the RNAi lines whose knockdown efficiencies are shown in Figure 6—figure supplement 3
Primers used for RT-PCR
https://doi.org/10.7554/eLife.35264.036| Primer list | Primer sequence (5′ → 3′) |
|---|---|
| sNPFR-F | CCAACTGGAGCCTAACGTCG |
| sNPFR-R | AACTGGTTGTGAATGATCCCG |
| 5HT1B-F | TTGGTTGCATCTCTGGCAGTG |
| 5HT1B-R | CCGGTCCCAATATCCATCCATT |
| 5HT2A-F | TTCACACTGCGACACTTCAAT |
| 5HT2A-R | GGGGTGTAGGATGTGCTGT |
| InR-F | CCGCAAGCAGTGAAGAAGC |
| InR-R | CGTCGTCTCCACTTCGTCAAA |
| DAR1-F | CCCGTATTCTTTGGCATTATCGG |
| DAR1-R | GGCCAGGTTGATTATCAGCAGA |
| RDL-F | CACAGGCAACTATTCGCGTTT |
| RDL-R | GCGATTGAGCCAAAATGATACC |
| GAD-F | CACCAACGACCGGAACGAG |
| GAD-R | TGGGGATGTCCCGTCTTAACT |
| DAMB-F | CATCTCCGAGGATGTCTACTTCT |
| DAMB-R | CCATCGCAGGACTCAAGGTG |
| NPFR-F | ATCAGCATGAATCAGACGGAGC |
| NPFR-R | GATGCCGGTCGTCCAGATA |
| Rpl19-F | TCTCTAAAGCTCCAGAAGAGGC |
| Rpl19-R | CGATCTCGTTGATTTCATTGGGA |
Additional files
-
Source code 1
MATLAB program used to generate the moving trajectory of a recorded fly.
- https://doi.org/10.7554/eLife.35264.037
-
Source code 2
MATLAB program used to analyze the moving speed of a recorded fly based on the trajectory generated by Source code 1.
- https://doi.org/10.7554/eLife.35264.038
-
Transparent reporting form
- https://doi.org/10.7554/eLife.35264.039





