The TRIM-NHL protein NHL-2 is a co-factor in the nuclear and somatic RNAi pathways in C. elegans
Figures
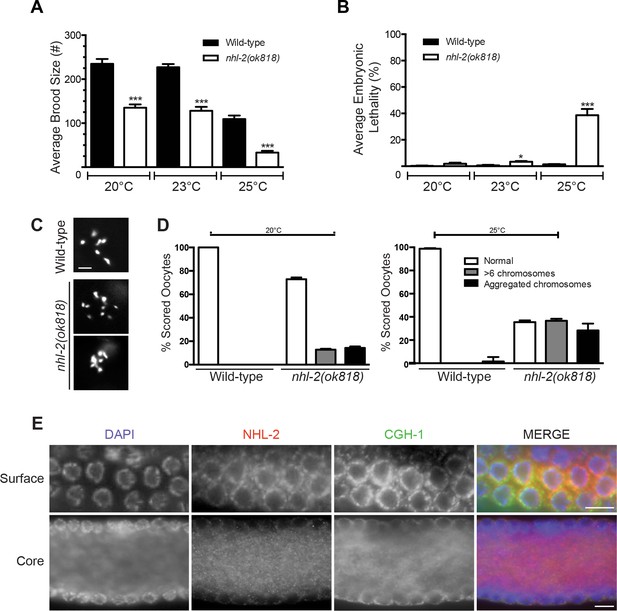
NHL-2 is required for fertility and is enriched in germline granules.
(A) Average brood size and (B) embryonic lethality of nhl-2(ok818) mutants compared to wild-type at 20°C, 23°C and 25°C. Embryonic lethality is significantly different than wild-type animals at higher temperatures but not significantly different than wild-type at 20°C. Brood size is significantly lower across 20°C, 23°C and 25°C when compared to wild-type animals. ***=P values <0.001, *=P values <0.05 error bars represent standard error of mean. (n = 5) (C) Representative images of DAPI stained diakinetic oocytes showing abnormal chromosome morphology in nhl-2(ok818) when compared to wild-type animals. nhl-2(ok818) chromosomes show aggregation or greater than six chromosomal bodies. Scale bar 5 µm, n = 20 (D) Quantification of chromosomal defects in diakinetic oocytes in wild-type and nhl-2(ok818) animals at 20°C and 25°C. (E) NHL-2 co-localizes with CGH-1 in P-granules (top panels, Surface) and gonadal core CGH-1 granules (bottom panels, Core). DAPI (blue), NHL-2 (red), CGH-1 (green). Scale bar 10 µm. Immunostaining with affinity purified NHL-2 antibody was reduced to background in nhl-2(ok818) germlines (Figure 1—figure supplement 2).
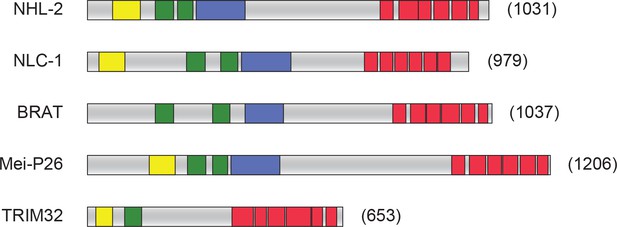
Comparison of representative TRIM-NHL proteins.
(A) Domain organization of selected TRIM-NHL family proteins. The TRIM-NHL family of proteins are conserved among metazoans, however, they often display variable domain organization. Canonical TRIM-NHL proteins have a three zinc-binding domains (a RING finger, one or two B-box motifs) and a coiled-coil domain in the N-terminal half of the proteins, and a series of NHL repeats at the C-terminal end of the protein. Examples shown are from: Drosophila melanogaster (BRAT and Mei-P26), C. elegans (NCL-1 and NHL-2) and Homo sapiens TRIM32. Yellow boxes, RING domain; Green boxes, B-box domains; Blue boxes, coiled-coil domain; Red boxes, NHL repeats.
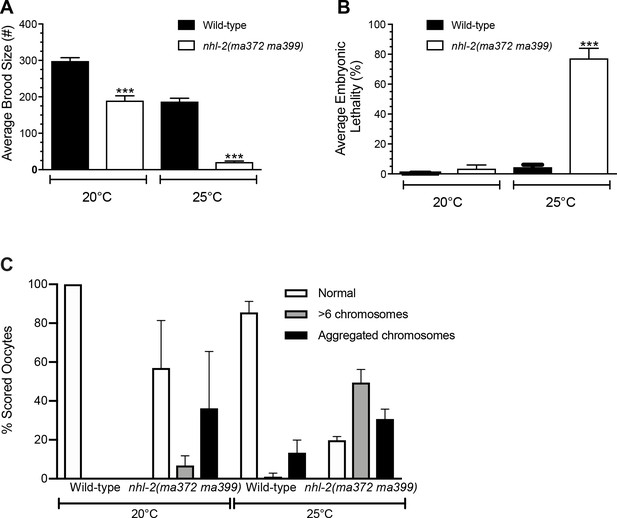
nhl-2(ma372 ma399) mutants display a temperature sensitive reproduction and embryonic lethality defect.
(A) Average brood size and (B) embryonic lethality of nhl-2(ok818) mutants compared to wild-type at 20°C and 25°C. Brood size is significantly lower across 20°C and 25°C when compared to wild-type animals. Embryonic lethality is significantly different than wild-type animals at higher temperatures but not significantly different than wild-type at 20°C. ***=P values <0.001, error bars represent standard error of mean. (n = 5) (C) Quantification of chromosomal defects in diakinetic oocytes in wild-type and nhl-2(ma372 ma399) animals at 20°C and 25°C.
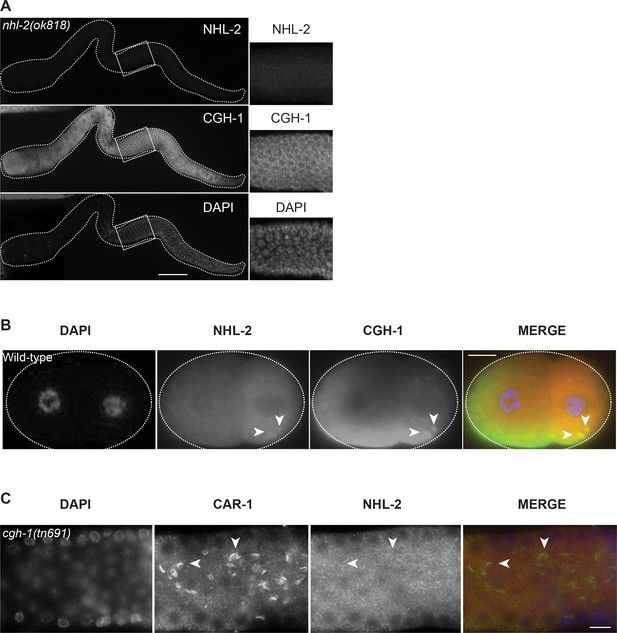
The NHL-2 antibody is specific.
(A) NHL-2 and CGH-1 staining of dissected nhl-2(ok818) hermaphrodite germlines demonstrates that the NHL-2 antibody is specific and that CGH-1 localizes properly in the absence of nhl-2. Scale bar 50 µm. (B) NHL-2 and CGH-1 co-staining of wild-type embryos show that NHL-2 is enriched in the germ cell lineage (P lineage, posterior/right), where it is found in the cytoplasm and in P granules (arrowheads). Scale bar 10 µm. (C) NHL-2 and CAR-1 co-staining of temperature sensitive mutant cgh-1(tn691) grown at the restrictive temperature shows that NHL-2 does not co-localize with CAR-1 in the ‘sheets’ in the gonad core. Arrowheads point to the CAR-1 solid sheets. Scale bar 10 µm.
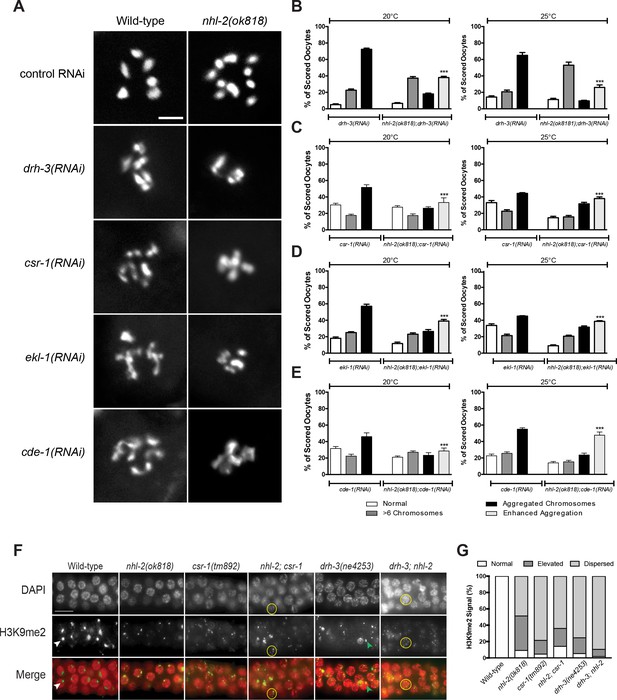
NHL-2 genetically interacts with the CSR-1 22G-RNA pathway.
(A) Representative images of DAPI stained oocyte diakinetic chromosomes showing abnormal chromosome morphology when drh-3, csr-1, ekl-1 and cde-1 were knocked down by RNAi in wild-type and nhl-2(ok818) animals. Scale bar 5 µm, n = 90 oocytes. (B, C, D and E) Chromosomes from each phenotype were binned in one of four categories: normal, oocytes with >6 chromosomal bodies, aggregated chromosomes, and enhanced aggregation (where aggregation exceeded that observed in wild-type animals, ***=P values <0.001). (F) Pachytene region of dissected adult male germlines stained with DAPI (red) and H3K9me2 (green) in wild-type and single and double mutants nhl-2(ok818), csr-1(tm892) and drh-3(ne4253). Wild-type males had one strong signal of H3K9me2 (yellow arrowhead), while single and double mutants had increased levels of nuclei with multiple foci (green arrowhead) or dispersed staining patterns (yellow circles) of H2K9me2 staining. Scale bar 50 µm. Over 109 nuclei were score from a minimum of 11 germline. (G) Quantification of H3K9me2 distribution in germ cells from each phenotype scored as normal, elevated, or dispersed from each phenotype. nhl-2(ok818) mutants display similar proportions of germ cells with abnormal H3K9me2 markings as the double mutants.
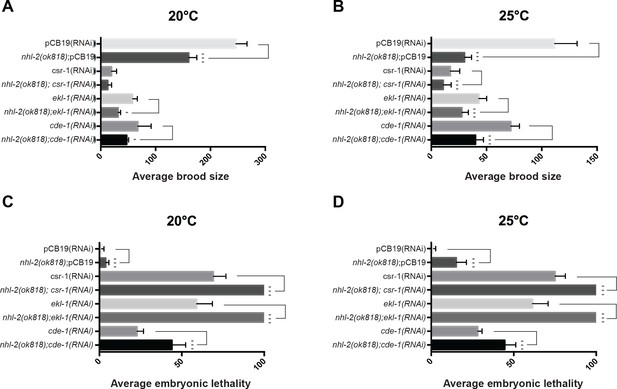
Synthetic brood size reduction and embryonic lethality between nhl-2(ok818) and the CSR-1 pathway.
(A) Knockdown of CSR-1 pathway genes in nhl-2(ok818) mutants results in synthetic reductions in brood size that are significantly different than knockdown in wild-type animals treated with control RNAi, pCB19. (B) Average embryonic lethality of unhatched embryos from the same brood size assay. Knockdown of CSR-1 pathway genes in nhl-2(ok818) mutants results in sign ificantly higher embryonic lethality compared to knockdown in wild-type animals. ***=P values <0.001, *=P values <0.05. Error bars represent standard error of mean. n = 5.
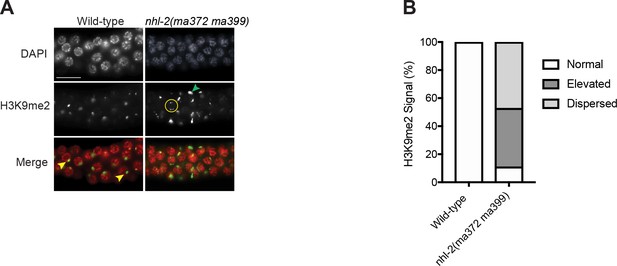
nhl-2(ma372 ma399) mutants show defects in diakinetic chromosome organization and H3K9me2 distribution.
(A) Pachytene region of dissected adult male germlines stained with DAPI (red) and H3K9me2 (green) in wild-type and nhl-2(ma372 ma399) mutants. Wild-type males had one strong signal of H3K9me2 (yellow arrowhead), while single and double mutants had increased levels of nuclei with multiple foci (green arrowhead) or dispersed staining patterns (yellow circles) of H2K9me2 staining. Scale bar 50 µm. Over 109 nuclei were score from a minimum of 11 germline. (C) Quantification of H3K9me2 distribution in germ cells from each phenotype scored as normal, elevated, or dispersed from each phenotype.
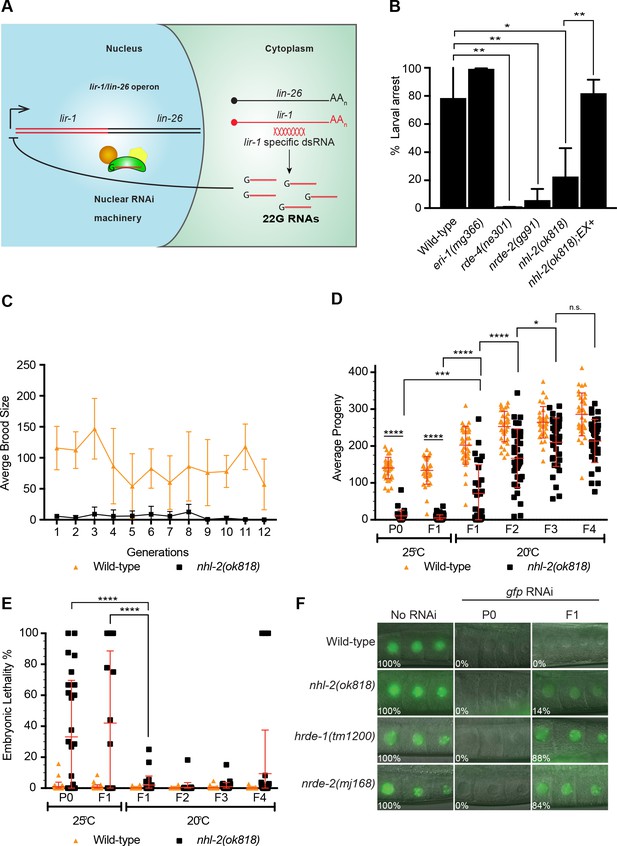
Analysis of nuclear RNAi pathways in nhl-2(ok818) mutants.
(A) The nuclear RNAi pathway silences somatic targets. dsRNA targeting the lir-1 mRNA leads to the generation of 22G small RNAs in the cytoplasm and the nuclear RNAi pathway uses these 22G RNAs to target the endogenous lir-1/let-26 locus for silencing. (B) nhl-2(ok818) worms are resistant to lir-1 RNAi. This phenotype was rescued in transgenic nhl-2(ok818) worms expressing an GFP-NHL-2 extrachromosomal array (Ex+) under control of the putative nhl-2 promoter. Percent larval arrest represents mean of two biological replicates ±SD; **=P values <0.001, *=P values <0.0418, n > 100. (C) nhl-2(ok818) worms display a mortal germline phenotype at 25°C. Error bars indicate mean ±SD, n = 6. (D) Analysis of transgenerational brood size and embryonic lethality (E) of wild-type and nhl-2(ok818) worms. ****=P values <0.0001, ***=P values <0.0004, *=P values <0.0418. Error bars indicate mean ±SD, n = 37. (F) nhl-2(ok818) worms have a minor defect in RNAi inheritance at 20°C. Transgenic worms of the indicated genotypes expressing pie-1::gfp::h2b were fed gfp RNAi for one generation and then grown on OP50 thereafter. Animals were scored for GFP expression using a fluorescence microscope at 63X objective. The percentage of animals expressing GFP is shown, n ≥ 37. Germlines of nhl-2(ok818) worms grown at 25°C were unable to be scored due to the disorganised nature of the proximal gonad.
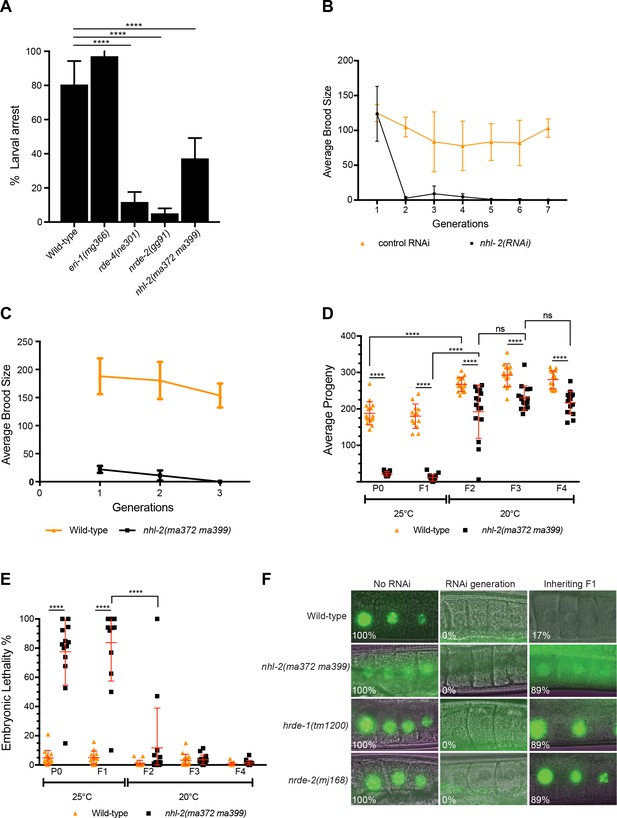
Analysis of nuclear RNAi pathways in nhl-2(ma372 ma399) mutants and nhl-2(RNAi).
(A) nhl-2(ma372 ma399) mutants are defective in somatic nuclear RNAi. Percent larval arrest represents mean of two biological replicates ±SD; ***=P values <0.0004, n > 100. (B) RNAi of nhl-2 leads to germline mortality. Average brood sizes were determined over multiple generations for worms grown at 25°C and fed RNAi against nhl-2 or the control (pCB19). (C) nhl-2(ma372 ma399) mutants have a mortal germline at 25°C. Error bars indicate mean ±SD, n = 6. (D) Analysis of transgenerational brood size and embryonic lethality (E) of wild-type and nhl-2(ma372 ma399). ****=P values <0.0001. Error bars indicate mean ±SD. Progeny of 13–15 parent worms per genotype/generation were scored. (F) nhl-2(ma372 ma399) worms have a defect in RNAi inheritance at 20°C. Transgenic worms of the indicated genotypes expressing pie-1::gfp::h2b were fed gfp RNAi for one generation and then grown on OP50 thereafter. Animals were scored for GFP expression using a fluorescence microscope at 63X objective. The percentage of animals expressing GFP is shown, n ≥ 37. Germlines of nhl-2(ma372 ma399) worms grown at 25°C were unable to be scored due to the disorganised nature of the proximal gonad.
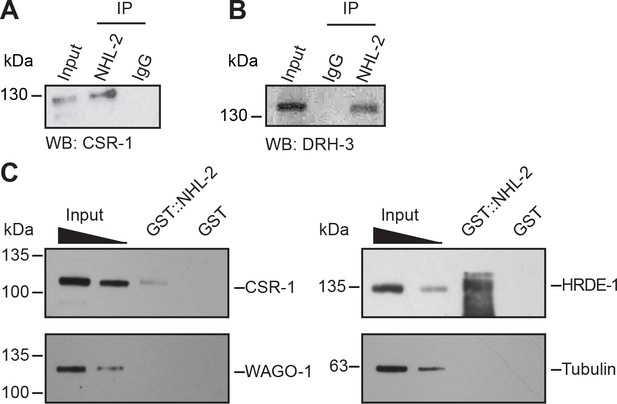
NHL-2 interacts physically with the components of the 22G-RNA pathway.
(A) Western blot using anti-CSR-1 showing no variance between CSR-1 levels in wild-type animals (WT) and nhl-2(ok818) mutants when equal amounts of protein are loaded. Tubulin is shown as a loading control (bottom). CSR-1 co-immunoprecipitates with NHL-2. NHL-2 was immunoprecipitated from one-day-old adult hermaphrodite extracts and CSR-1 was detected by Western blot. (B) DRH-3 co-immunoprecipitates with NHL-2. NHL-2 was immunoprecipitated from one-day-old adult hermaphrodite extracts and DRH-3 was detected by Western blot. (C) GST-pulldown of NHL-2. GST-NHL-2 was able to pulldown CSR-1 and HRDE-1, but not WAGO-1. SDS-PAGE gel of purified GST-NHL-2 is shown in Figure 4—figure supplement 1. (D) Western blot using anti-CSR-1 showing no variance between CSR-1 levels in wild-type animals (WT) and nhl-2(ok818) mutants when equal amounts of protein are loaded. Tubulin is shown as a loading control (bottom).
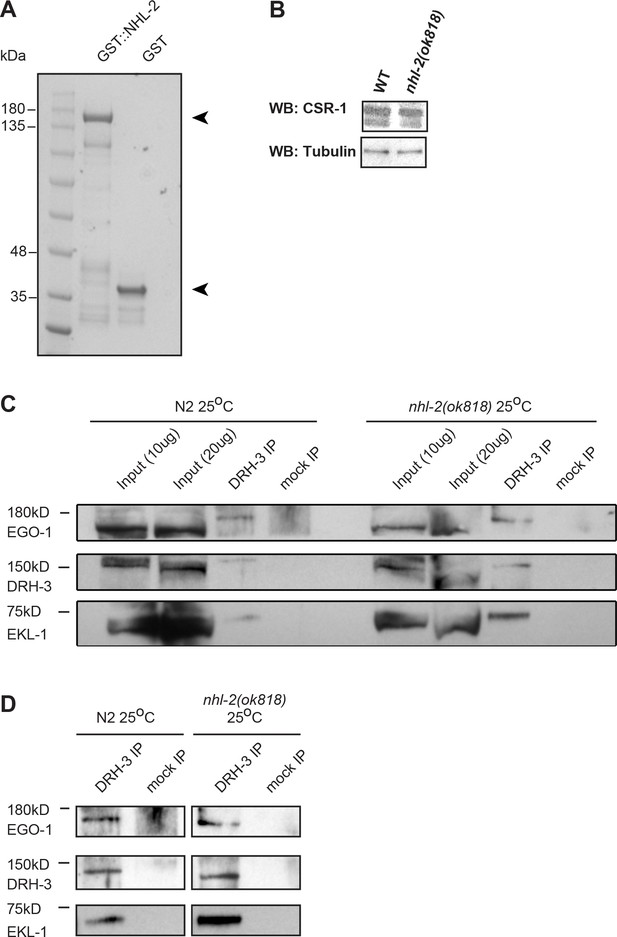
The RdRP complex is intact in nhl-2(ok818) mutants.
(A) SDS-PAGE gel of purified GST-NHL-2 and GST used in GST Pull-down experiments. Arrowheads indicate the positions of GST-NHL-2 and GST. (B) Western blot using anti-CSR-1 showing no variance between CSR-1 levels in wild-type animals (WT) and nhl-2(ok818) mutants when equal amounts of protein are loaded. Tubulin is shown as a loading control (bottom). (C) An IP of DRH-3 also precipitates the RdRP complex components EKL-1 and EGO-1 in wild-type and nhl-2(ok818) mutants at 25°C. Western blots are probed with anti-EGO-1 (top), anti-DRH-3 (center), anti-EKL-1 (bottom). Sizes are marked, and a mock IP (beads only) is shown as a negative control. All antibodies are as used in (Claycomb et al., 2009; Gu et al., 2009). (D) A darker exposure of the DRH-3 and mock IP lanes.
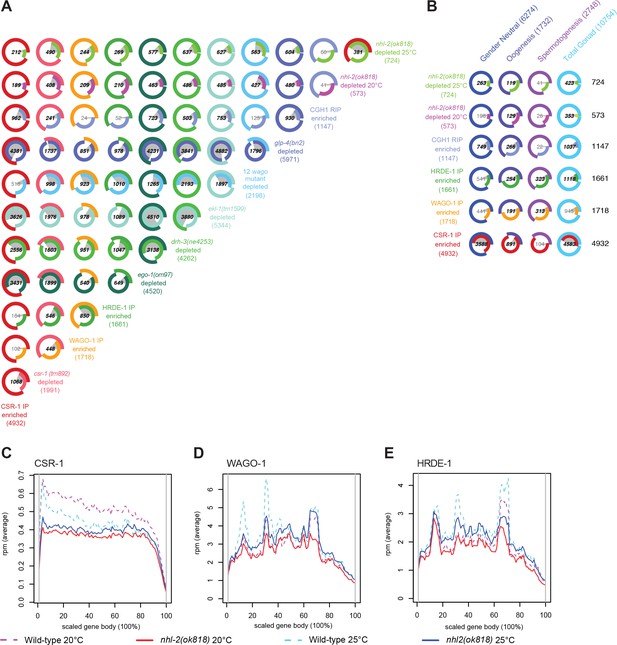
Comparison of small RNAs in nhl-2(ok818) mutants to other small RNA and mRNA profiles.
(A) Venn-pie diagrams show comparisons between sets of genes that are the targets of 22G-RNAs. For mutant strains, genes are two-fold or greater depleted of 22G-RNAs in mutants relative to appropriate wild-type controls, with a minimum of ten rpm. For Argonaute IPs, genes are two-fold or greater enriched for 22G-RNAs in the IP relative to a total small RNA input sample, with a minimum of ten rpm. Numbers shown in bold demonstrate statistically significant overlap. (B) Venn-pie diagrams show comparisons between 22G-RNA gene targets (as in (A), left/rows), and mRNA transcriptome data (bottom/columns) from Total gonads (the union of entire male and female gonad transcriptomes), male gonads (Spermatogenesis), female gonads (Oogenesis), and gender neutral (the overlap between genes expressed in both male and female gonads) genes (Ortiz et al., 2014). Numbers shown in bold demonstrate statistically significant overlap. (C–E) Distribution of 22G-RNA reads in nhl-2(ok818) mutants along the scaled gene bodies are shown for CSR-1 target genes (C), WAGO-1 target genes (D), and HRDE-1 target genes (E) in nhl-2(ok818) and wild-type animals.
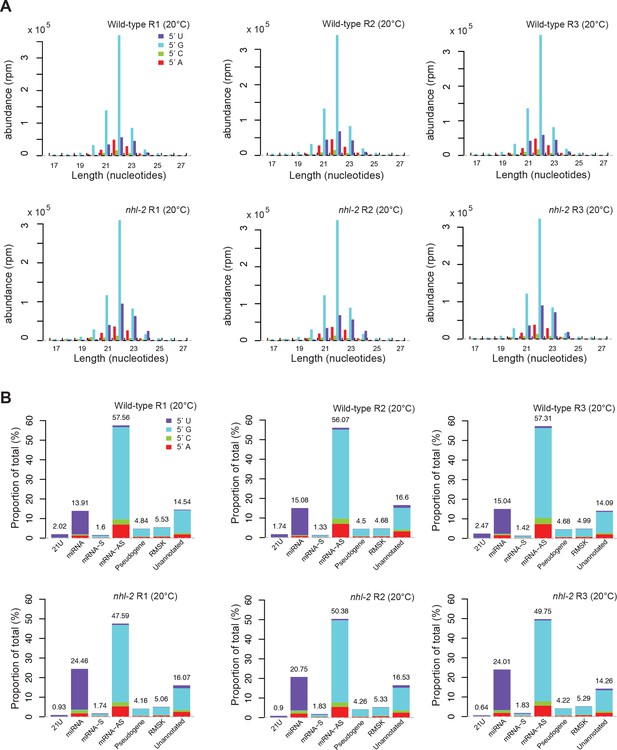
Analysis of small RNA populations in nhl-2(ok818) mutants and wild-type animals at 20°C.
(A) Length and first nucleotide distribution of small RNAs in wild-type and nhl-2(ok818) mutants at 20°C. Small RNA abundance is measured in reads per million (rpm) for all small RNAs that are mapped to the genome or exon-exon junctions. (B) Proportions of reads which were aligned to 21U-RNAs (piRNAs), miRNAs, sense (S) mRNAs, antisense (AS) mRNAs, pseudogenes, RepeatMasker annotated repeat elements (RMSK), and the unannotated portion of the genome, with the first nucleotide distribution of small RNAs in each category (Note that all classes except for miRNAs and piRNAs include 22G-RNAs and 26G-RNAs).
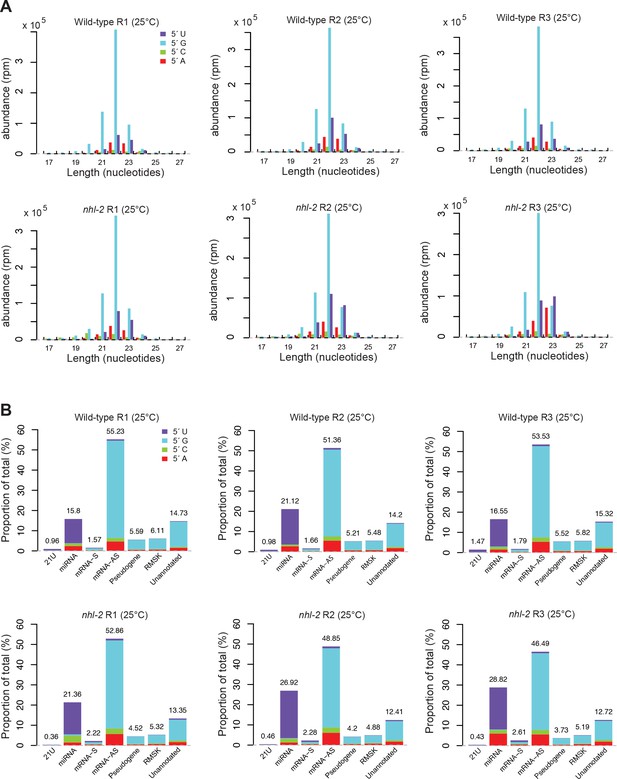
Analysis of small RNA populations in nhl-2(ok818) mutants and wild-type animals at 25°C.
(A) Length and first nucleotide distribution of small RNAs in wild-type and nhl-2(ok818) mutants at 25°C. Small RNA abundance is measured in reads per million (rpm) for all small RNAs that are mapped to the genome or exon-exon junctions. (B) Proportions of reads which were aligned to 21U-RNAs (piRNAs), miRNAs, sense (S) mRNAs, antisense (AS) mRNAs, pseudogenes, RepeatMasker annotated repeat elements, and the unannotated portion of the genome, with the first nucleotide distribution of small RNAs in each category (Note that all classes except for miRNAs and piRNAs include 22G-RNAs and 26G-RNAs).
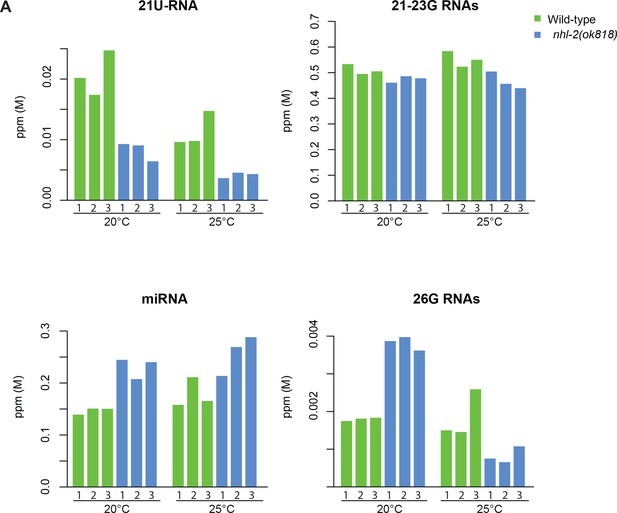
Comparison of abundance (in ppm) for different categories of small RNAs in wild-type (green) and nhl-2(ok818) mutants (blue) at 20°C vs. 25°C.
Biological replicates are indicated as 1, 2 and 3.
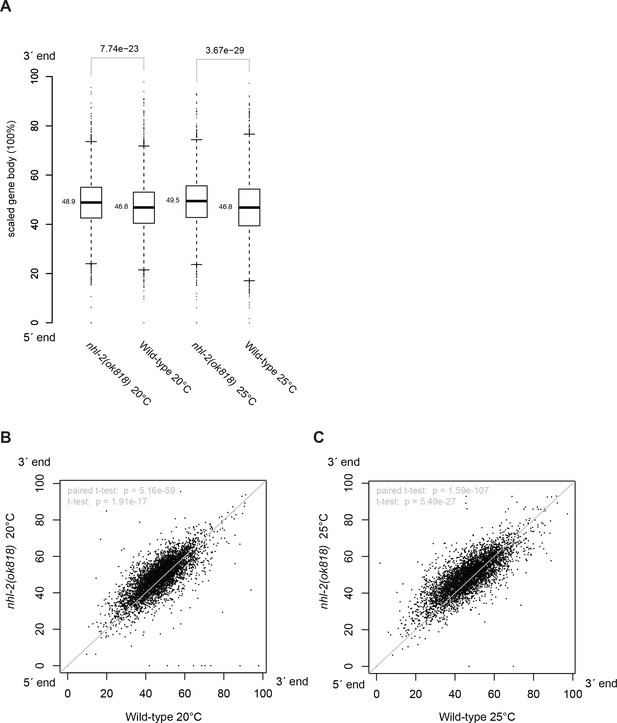
The distribution of 22G-RNAs is reduced at the 5´-end of CSR-1 targets in nhl-2(ok818) relative wild-type control samples.
Along the scaled gene body, the coordinates of the centroids of the areas under the 22G-RNA density curves (Figure 5C–E) were calculated and compared. The centroid positions for 4,392 CSR-1 target genes between mutants and wild-type controls were compared in boxplot with Wilcoxon rank sum test p-values (A) and scatter plots (B,C) with t-test p-values.
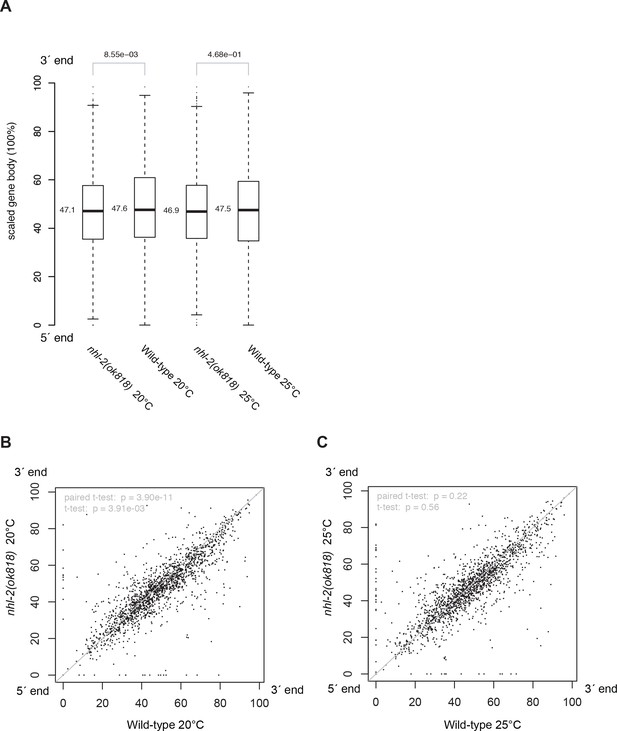
The distribution of 22G-RNAs is reduced slightly at the 3´-end of WAGO-1 targets in nhl-2(ok818) relative wild-type control samples.
Along the scaled gene body, the coordinates of the centroids of the areas under the 22G-RNA density curves (Figure 5C–E) were calculated and compared. The centroid positions for 1718 WAGO-1 target genes between mutants and wild-type controls were compared in boxplot with Wilcoxon rank sum test p-values (A) and scatter plots (B, C) with t-test p-values.
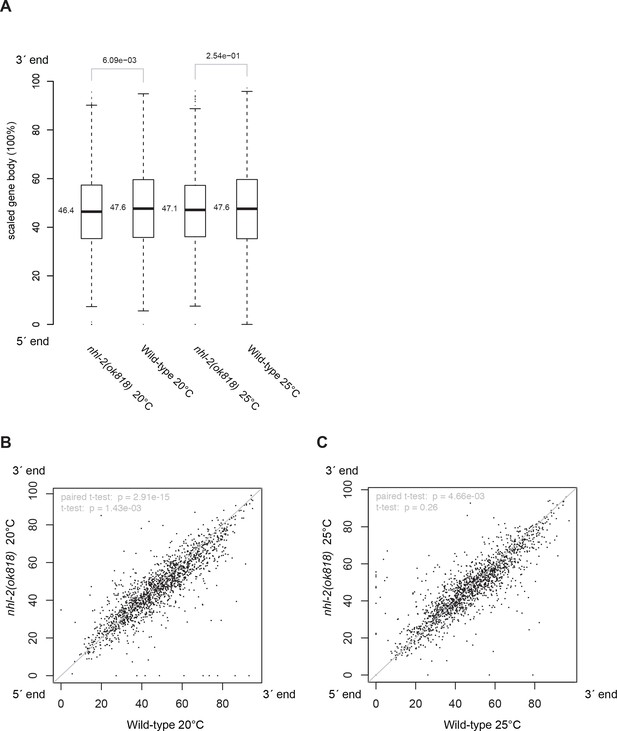
The distribution of 22G-RNAs is reduced slightly at the 3´-end of HRDE-1 targets in nhl-2(ok818) relative wild-type control samples.
Along the scaled gene body, the coordinates of the centroids of the areas under the 22G-RNA density curves (Figure 5C–E) were calculated and compared. The centroid positions for 1661 HRDE-1 target genes between mutants and wild-type controls were compared in boxplot with Wilcoxon rank sum test p-values (A) and scatter plots (B, C) with t-test p-values.
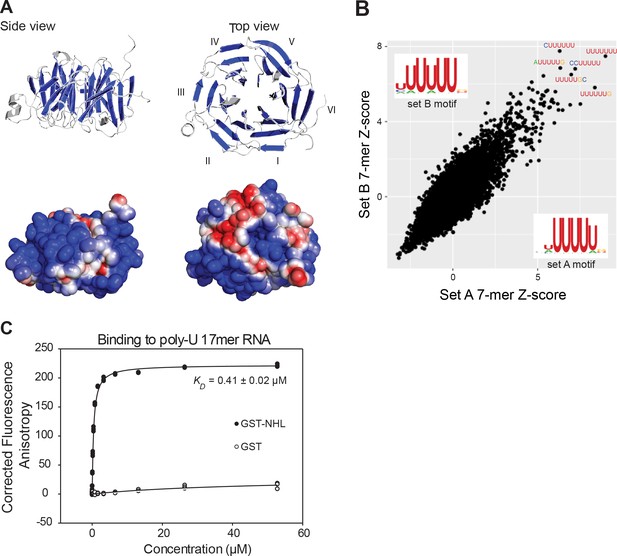
The NHL domain of NHL-2 binds RNA.
(A) Side and top view of the homology-based structure prediction of the NHL-2 NHL domain and its six β-propellers marked blue. Electrostatic surface representation of the NHL-2 NHL domain showing negative regions in blue and positive in red is shown below. The six blades of the β propeller of the NHL domain are numbered (I–VI). (B) Identification of RNA-binding motif of purified GST-NHL-2-NHL domain by RNAcompete. The top five high-scoring 7-mers, and the scatter plots, displaying Z scores and motifs for the two halves of the RNA pool (set A and set B) are shown. Sequence logos were derived by aligning the top ten high-scoring 7-mers. (C) Quantitative assessment of RNA-binding of GST-NHL-2 NHL domain and GST. The fluorescence anisotropy of reactions contained 1 nM fluorescently-labeled 17mer poly-U single-stranded RNA and increasing concentrations of GST-NHL-2 NHL domain (closed circles) or GST alone (open circles).
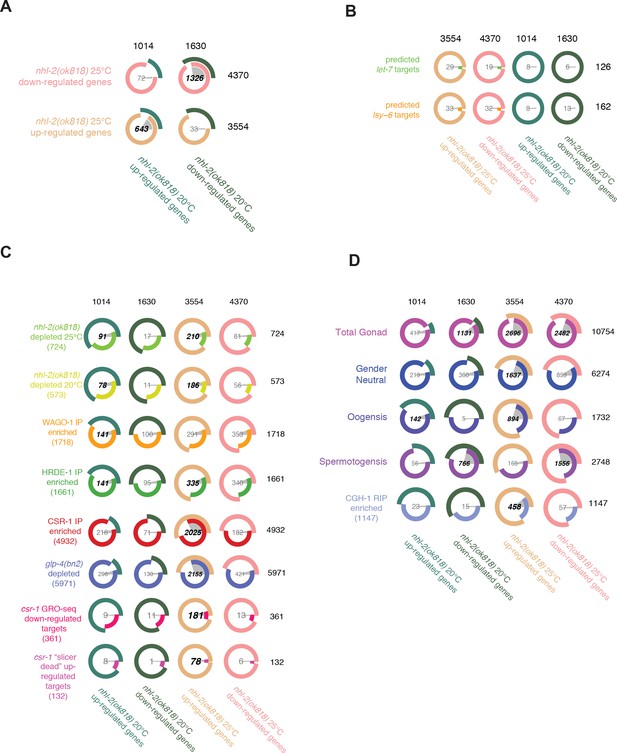
Analysis of the 22G-RNAs targeting the nhl-2(ok818) mRNA transcriptome.
(A) Venn-pie diagrams indicate the number of genes enriched in or depleted of 22G-RNAs in nhl-2(ok818) mutants relative to wild-type worms at 20°C and 25°C, as determined using EdgeR. Numbers in bold demonstrate statistically significant overlap. Each row corresponds to a gene set, with its label on the right. Each Venn-pie diagram indicates the overlap between the gene set of its row and the gene set whose label is in its column. (B) Venn-pie diagrams show comparisons between the predicted targets of the miRNAs let-7 and lsy-6 (determined using TargetScan6), and genes that are mis-regulated in nhl-2(ok818) mutants. (C) Venn-pie diagrams show comparisons between the genes depleted of 22G-RNAs in nhl-2(ok818) mutants and the genes enriched in 22G-RNAs in CSR-1, WAGO-1, or HRDE-1 IP samples with nhl-2(ok818) mRNA-seq data. (D) Venn-pie diagrams show comparisons between nhl-2(ok818) mRNA-seq data and genes depleted of 22G-RNAs in glp-4(bn2) mutants (which have very few germ cells) or germline expressed gene mRNA-seq data (Ortiz et al., 2014) (as in Figure 4B).
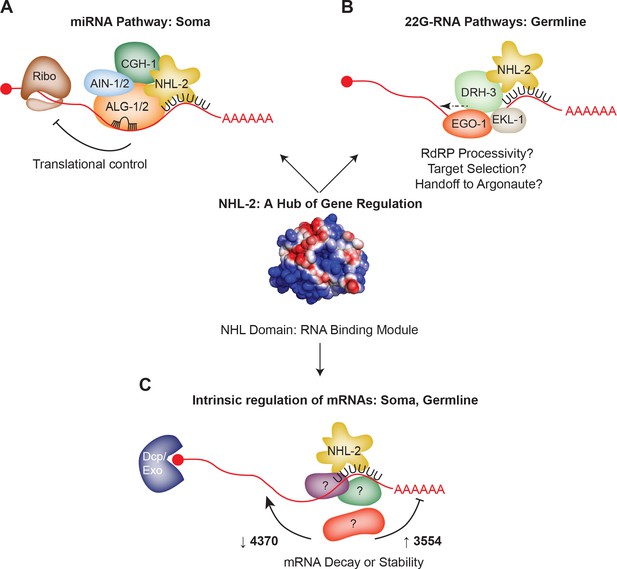
Model: NHL-2 acts as a hub of gene regulatory activity.
(A) NHL-2 interacts with the miRNA pathway in the soma, where it may regulate the translation of let-7 and lsy-6 targets. The NHL RNA binding domain of NHL-2 may help to reinforce target selection of miRISC in this pathway. (B) In the germline, NHL-2 interacts with the 22G-RNA pathway, specifically via genetic and physical interactions with the RdRP helicase DRH-3. In this capacity, NHL-2 could impact the processivity or reloading of the RdRP complex on mRNA templates, and/or could aid in target mRNA selection via its RNA binding capacity. Owing to its genetic and physical interaction with CSR-1, NHL-2 could also be involved in the handoff of 22G-RNAs from the RdRP complex to Argonaute. If NHL-2 mediates gene regulation via 22G-RNAs, it is likely to be at the level of translation. (C) The intrinsic RNA binding capacity of NHL-2 could enable it to regulate a large number of transcripts in the soma and germline, both positively and negatively. This activity could be influenced by additional binding partners, and we speculate that it may occur in P bodies (soma) and P granules (germline) based on the localization pattern of NHL-2.
Additional files
-
Supplementary file 1
Small RNA sequencing statistics of four samples in nhl-2(ok818) mutants and wild-type animals at 25°C or 20°C, Related to Figures 4 and 6.
- https://doi.org/10.7554/eLife.35478.023
-
Supplementary file 2
Genes altered in small RNA and mRNA sequencing datasets generated and investigated in this paper, Related to Figures 4 and 6.
- https://doi.org/10.7554/eLife.35478.024
-
Supplementary file 3
Master file of small RNA and mRNA-seq data used in this study.
- https://doi.org/10.7554/eLife.35478.025
-
Supplementary file 4
Source code file for the custom Shell and Perl (5.10.0) scripts used for Small RNA analysis.
- https://doi.org/10.7554/eLife.35478.026
-
Transparent reporting form
- https://doi.org/10.7554/eLife.35478.027





